Abstract
Cardiac magnetic resonance imaging (CMR) has emerged as a valuable diagnostic tool for cardiac diseases. However, a significant drawback of CMR is its slow imaging speed, resulting in low patient throughput and compromised clinical diagnostic quality. The limited temporal resolution also causes patient discomfort and introduces artifacts in the images, further diminishing their overall quality and diagnostic value. There has been growing interest in deep learning-based CMR imaging algorithms that can reconstruct high-quality images from highly under-sampled k-space data. However, the development of deep learning methods requires large training datasets, which have so far not been made publicly available for CMR. To address this gap, we released a dataset that includes multi-contrast, multi-view, multi-slice and multi-coil CMR imaging data from 300 subjects. Imaging studies include cardiac cine and mapping sequences. The ‘CMRxRecon’ dataset contains raw k-space data and auto-calibration lines. Our aim is to facilitate the advancement of state-of-the-art CMR image reconstruction by introducing standardized evaluation criteria and making the dataset freely accessible to the research community.
Similar content being viewed by others
Background & Summary
Cardiac magnetic resonance (CMR) imaging has emerged as a valuable technique for diagnosing cardiovascular diseases, thanks to its superior soft tissue contrast and non-invasive nature. CMR imaging can provide anatomical, functional, and tissue characteristics information of the heart, and has been reported in many population studies1,2,3,4,5. Cine magnetic resonance imaging (MRI) is currently considered the gold standard for non-invasive evaluation of cardiac functionality. Cardiac T1 and T2 quantitative mapping are widely used in evaluating intracellular disturbances of cardiomyocytes5. Myocardial T1 characterization is valuable for detecting and assessing various cardiomyopathies, while T2 changes have been observed in edematous regions in patients with infarction, hemorrhage, graft rejection or myocarditis.
However, a significant drawback of CMR is its slow imaging speed, resulting in low patient throughput and compromised clinical diagnostic quality. The limited temporal resolution also causes patient discomfort and introduces artifacts in the images, further diminishing their overall quality and diagnostic value. Typically, cine is obtained using an electrocardiography (ECG)-gated segmented gradient sequence, as acquiring the full k-space data within a timeframe short enough to resolve cardiac motion accurately is not feasible. Thus, the entire k-space is segmented and read out over multiple cardiac cycles. However, for patients with impaired breath-hold capacity or cardiac arrhythmia, image degradation will occur due to the long acquisition time and motions, which can influence further diagnosis. Accelerated cine imaging effectively addresses these limitations by utilizing a reduced amount of k-space data, while still maintaining high reconstruction performance. Sufficient acceleration also enables “real-time” imaging, substantially reducing artifacts associated with respiratory motion and arrhythmia. Similar to cine imaging, accelerated T1 and T2 mapping shortens the acquisition window, leading to significant reduction of artifacts associated with respiratory motion and arrhythmia6. Consequently, there has been a growing interest in accelerated CMR image reconstruction from highly under-sampled k-space data.
So far, many artificial intelligence (AI)-based image reconstruction algorithms have shown great potentials in improving imaging performance7,8,9,10,11,12,13,14,15,16,17. However, deep-learning-based methods require large quantities of raw k-space data for model training. The field of CMR imaging still lacks standardized and high-quality datasets that are publicly available. In addition, due to the absence of large public datasets, there is no common gold standard on which they can be properly compared. To date, NYU Langone Health has released ‘fastMRI’ dataset, containing multi-channel brain15, knee16 and prostate17 MRI raw data. However, these datasets are inadequate for the 3D + 1D (time domain) scenario in cardiac imaging. To the best of our knowledge, previous available cardiac raw datasets mainly include OCMR18 and Harvard CMR Dataverse19. The former provides fully sampled as well as prospectively under-sampled cine data, while the latter released cine data with radial sampling trajectories. However, these datasets have limitations in terms of insufficient anatomical views (e.g., 2-chamber, 3-chamber), imaging contrasts (e.g., T1 and T2 mapping), and dataset size, which motivated the release of this ‘CMRxRecon’ dataset. The goal of establishing the ‘CMRxRecon’ dataset is to provide a benchmark dataset that enables the broad research community to promote advances in high quality CMR imaging.
In this paper, we describe the first release of CMR raw k-space data that includes multi-contrast, multi-view and multi- channel cardiac imaging from 300 subjects. Imaging studies include cine and mapping sequences. In addition, we released processed CMR images in The Neuroimaging Informatics Technology Initiative (NIFTI) format and the corresponding scripts of state-of-the-art parallel imaging.
Methods
Subject characteristics
The study was approved by the institutional review board of Fudan University (approval number: FE20017). The data were allowed to be made publicly available as part of the written consent process. All participants were aware of the nature of the study and agreed to make their materials publicly available in anonymized form. Inclusion criteria were defined as: 1) adults without pathologically confirmed diagnosis of cardiovascular disease, and 2) availability of an MRI examination with all imaging sequences. A total of 300 healthy volunteers (160 females and 140 males) were recruited between June 2022 and March 2023 with written informed consent. The mean age of the subjects was 26 ± 5 years.
Image acquisition
Data were acquired on a 3 T scanner (MAGNETOM Vida, Siemens Healthineers, Germany), with a dedicated cardiac coil made up of 32 channels. Participants were placed in a supine position on the table before scans. Electrodes were attached and Electrocardiogram (ECG) signal were recorded during the scan. The ‘Dot’ engine was used for cardiac scout imaging. Figure 1 shows representative CMR images of cardiac cine and mapping released in the dataset.
We followed the recommendation of CMR imaging reported in the previous publication5. The ‘TrueFISP’ readout was used for 2D cardiac cine acquisitions. The collected images included short-axis (SAX), two-chamber (2CH), three-chamber (3CH) and four-chamber (4CH) long-axis (LAX) views. Cardiac cine was acquired through a retrospective ECG-gated segmented approach, wherein k-space is segmented in the phase encoding direction across multiple cardiac cycles. The selection of breath holds was automatically optimized according to the acquisition size, heart rate and slices. Typically, 5~14 slices were acquired for SAX view, while a single slice was acquired for each of the other views. The cardiac cycle was segmented into 12~25 phases with a temporal resolution ~50 ms according to the heart rate. Typical scan parameters were: spatial resolution of 1.5 × 1.5 mm2, slice thickness of 8.0 mm, repetition time (TR) of 3.6 ms, and echo-time (TE) of 1.6 ms. The parallel imaging acceleration factor was R = 3. Signal were acquired with breath-hold. The representative imaging parameters are summarized in Table 1.
T1 mapping was conducted using a modified look-locker inversion recovery (MOLLI) sequence, which acquired 9 images with different T1 weightings (using the 4-(1)-3-(1)-2 scheme, with one heart beat rest). T1 mapping was performed in SAX view only, with typical field-of-view (FOV) of 340 × 340 mm2, spatial resolution of 1.5 × 1.5 mm2, slice number of 5~6, slice thickness of 5.0 mm, TR of 2.7 ms, TE of 1.1 ms, partial Fourier of 6/8, and parallel imaging acceleration factor of R = 2. The inversion time varied among subjects according to the real-time heart rate. Signals were collected at the end of the diastole with ECG triggering.
T2 mapping was performed using T2-prepared (T2prep)-FLASH sequence with three T2 weightings in SAX view, with identical geometrical parameters as used in T1 mapping. Typical imaging parameters were FOV of 340 × 340 mm2, spatial resolution of 1.5 × 1.5 mm2, slice number of 5~6, slice thickness of 5.0 mm, TR of 3.0 ms, TE of 1.3 ms, T2 preparation time of 0/35/55 ms, partial Fourier of 6/8, and parallel imaging acceleration factor of R = 2. Signals were collected at the end of the diastole with ECG triggering.
Image processing
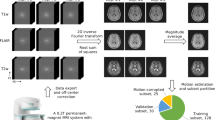
The general workflow to produce the ‘CMRxRecon’ dataset is illustrated in Fig. 2. The raw data with the filename extension ‘.dat’ was exported directly from the scanner using the Siemens “TWIX” tool. The k-space data was then extracted using the “mapVBVD” toolbox (https://github.com/pehses/mapVBVD), which was written in MATLAB. The k-space data were anonymized via conversion to the raw data format. We only included imaging parameters in the raw data while removing all items related to subject identity, e.g., subject name, personal ID, hospital ID, data of exam and date of birth. Those images with poor quality were removed based on visual assessment by experienced radiologists. After these processing steps, the resulting k-space were transformed to ‘.mat’ format (MATLAB 2018a). Each k-space data we provided included 24 calibration lines, which were stored in the same file as the original data. When calculating the undersampling factor, we did not include the calibration lines. The released dataset includes 120 training data, 60 validation data and 120 test data. The training and validation datasets can be used to train reconstruction models and to determine hyperparameter values, while the test dataset is used to compare the results across different approaches. It is worth noting that there is no difference in the data processing procedures for the training, validation, and test data. Therefore, researchers can use these datasets in any combination for their studies. However, if researchers wish to compare their results with the evaluation results published for the validation and test data in the MICCAI CMRxRecon 2023 Challenge20, they can directly use the validation and test data provided in this study.
Data Records
Data description
All the data contains raw k-space data and the auto-calibration lines (ACS, 24 lines). In addition, we provided metadata of the dataset in ‘csv’ format, including FOV, acquisition matrix, slice number, slice thickness, coil number, temporal phases, TR, TE, flip angle and oversampling factor. This dataset is being made public in the Synapse repository21.
Data format
The directory structure of the released dataset can be seen in Supplementary File. The CMRxRecon dataset contains two types of k-space data, i.e., raw k-space data and ACS data, both of which were complex-valued single precision multi-coil data. Detailed descriptions of the data types of cardiac cine and mapping are summarized in Table 2.
Technical Validation
Evaluation on data quality
For data quality, a technician has rated the original images on a 5-point scale. We only retained images with a rating of 5 to construct the dataset. The overall signal-to-noise ratio (SNR) of the dataset is shown in Fig. 3. The SNR of the reconstructed images of both cine and mapping were measured by calculating the mean cardiac signal divided by the standard deviation of the noise. The average SNRs of all channels is 27.344 ± 9.709 for Cine SAX, 28.979 ± 8.919 for Cine LAX, 27.686 ± 8.564 for T1 mapping, and 26.212 ± 9.031 for T2 mapping, respectively. All image qualities are sufficient for cardiac region segmentation and quantitative analysis. The python scripts for image quality evaluation metrics were provided in github (https://github.com/CmrxRecon/CMRxRecon-SciData/tree/main/Evaluation).
The histogram of SNR from cine, T1 mapping and T2 mapping in the reconstructed images. Different frames of T1 and T2 are shown, in which “t1-t9” following “T1” represents images at different inversion times, and “t1-t3” following “T2” represents images at different echo times. The mean and standard deviation of the SNR are labeled underneath. The noise ROI is selected from the corner (as shown in red box), while the signal ROI is selected as the whole cardiac region.
Benchmark reconstruction results
To further evaluate the feasibility of using the provided k-space data for undersampling image reconstruction tasks, we used the GRAPPA22 and ESPIRiT23 methods as benchmarks for examples. Representative results of the ‘CMRxRecon’ dataset with the abovementioned reconstruction algorithms are displayed in Figs. 4–7. The following criteria were used for reconstruction results assessments: normalized mean square error (NMSE), peak SNR (PSNR) and structural similarity index measure (SSIM). Quantitative assessments of the results using this benchmark algorithms are summarized in Tables 3–5. We provided scripts to reconstruct the released data using the above state-of-the-art algorithms in the public GitHub repository: https://github.com/CmrxRecon/CMRxRecon-SciData/tree/main/ReconCode.
Usage Notes
The dataset is public and can be downloaded from Synapse repository through this link (https://doi.org/10.7303/syn52965326.1). Registered Synapse users are able to access the data without the need to get approved. To process the provided k-space data, it is recommended to use the tools we provided in the GitHub repository. In addition to the dataset, we also provided a platform for the evaluation of reconstruction performance (https://www.synapse.org/#!Synapse:syn51471091/wiki/622170).
Code availability
We provide the script to facilitate the use of the released data at https://github.com/CmrxRecon/CMRxRecon-SciData. A brief description of the provided package is as follows:
a) ‘ReconCode’: contains parallel imaging reconstruction code;
b) ‘DemoData’: contain one example data;
c) ‘Evaluation’: contains image quality evaluation code.
References
Nakamura, M. & Sadoshima, J. Mechanisms of physiological and pathological cardiac hypertrophy. Nature Reviews Cardiology 15(7), 387–407 (2018).
Jaffe, A. S., Babuin, L. & Apple, F. S. Biomarkers in acute cardiac disease: the present and the future. Journal of the American college of cardiology 48(1), 1–11 (2006).
Bai, W. et al. A population-based phenome-wide association study of cardiac and aortic structure and function. Nature medicine 26(10), 1654–1662 (2020).
Bai, W. et al. Longitudinal changes of cardiac and aortic imaging phenotypes following COVID-19 in the UK biobank cohort. medRxiv, 2021.11. 04.21265918 (2021).
Wang, C. et al. Recommendation for cardiac magnetic resonance imaging-based phenotypic study: imaging part. Phenomics 1, 151–170 (2021).
Wang, C. et al. Black blood myocardial T2 mapping. Magnetic resonance in medicine. 81(1), 153–166 (2019).
Lyu, J. et al. Region-focused multi-view transformer-based generative adversarial network for cardiac cine MRI reconstruction. Medical Image Analysis 85, 102760 (2023).
Qin, C. et al. Convolutional recurrent neural networks for dynamic MR image reconstruction. IEEE transactions on medical imaging 38(1), 280–290 (2018).
Qin, C. et al. Complementary time‐frequency domain networks for dynamic parallel MR image reconstruction. Magnetic Resonance in Medicine 86(6), 3274–3291 (2021).
Lyu, J., Tong, X. & Wang, C. Parallel Imaging With a Combination of SENSE and Generative Adversarial Networks (GAN). Quantitative Imaging in Medicine and Surgery. 10(12), 2260–2273 (2020).
Lyu, J. et al. DuDoCAF: Dual-Domain Cross-Attention Fusion with Recurrent Transformer for Fast Multi-contrast MR Imaging. International Conference on Medical Image Computing and Computer-Assisted Intervention. 474–484 (Springer, Cham, 2022).
Wang, S. et al. The Extreme Cardiac MRI Analysis Challenge under Respiratory Motion (CMRxMotion). arXiv preprint arXiv:2210.06385 (2022).
Hammernik, K. et al. Learning a variational network for reconstruction of accelerated MRI data. Magnetic resonance in medicine 79(6), 3055–3071 (2018).
Aggarwal, H. K., Mani, M. P. & Jacob, M. MoDL: Model-based deep learning architecture for inverse problems. IEEE transactions on medical imaging 38(2), 394–405 (2018).
Zhao, R. et al. fastMRI+, Clinical pathology annotations for knee and brain fully sampled magnetic resonance imaging data. Scientific Data 9(1), 152 (2022).
Knoll, F. et al. fastMRI: A publicly available raw k-space and DICOM dataset of knee images for accelerated MR image reconstruction using machine learning. Radiology: Artificial Intelligence 2(1), e190007 (2020).
Tibrewala R, et al. FastMRI Prostate: A public, biparametric MRI dataset to advance machine learning for prostate cancer imaging. Scientific Data 11(1), 404 (2024).
Chen, C. et al. OCMR (v1. 0)–open-access multi-coil k-space dataset for cardiovascular magnetic resonance imaging. Preprint at https://doi.org/10.48550/arXiv.2008.03410 (2020).
El‐Rewaidy, H. et al. Multi‐domain convolutional neural network (MD‐CNN) for radial reconstruction of dynamic cardiac MRI. Magnetic Resonance in Medicine 85(3), 1195–1208 (2021).
Wang, C. et al. CMRxRecon: An open cardiac MRI dataset for the competition of accelerated image reconstruction. Preprint at https://doi.org/10.48550/arXiv.2309.10836 (2023).
Wang, C. et al. CMRxRecon: A publicly available k-space dataset. Synapse https://doi.org/10.7303/syn52965326.1 (2023).
Griswold, M. A. et al. Generalized autocalibrating partially parallel acquisitions (GRAPPA). Magnetic Resonance in Medicine: An Official Journal of the International Society for Magnetic Resonance in Medicine 47(6), 1202–1210 (2002).
Uecker, M. et al. ESPIRiT-an eigenvalue approach to autocalibrating parallel MRI: where SENSE meets GRAPPA. Magnetic resonance in medicine 71(3), 990–1001 (2014).
Acknowledgements
This study was supported in part by the National Natural Science Foundation of China (No. 62331021, 62001120, 62122064, 61971361) and the Royal Society (IEC\NSFC\211235).
Author information
Authors and Affiliations
Contributions
Chengyan Wang, Jun Lyu, Chen Qin, Shuo Wang, Xiahai Zhuang, Wenjia Bai and Xiaobo Qu designed the research; Xutong Kuang, Mengting Sun and Meng Liu performed the segmentations; Chengyan Wang and Xinyu Zhang performed data anonymization; Xutong Kuang and Yinghua Chu collected data; Chengyan Wang, Fanwen Wang, Yan Li, Jianhua Jin, Ziqiang Xu, Mengting Sun, Meng Liu, He Wang and Xiaobo Qu analyzed data; Shuo Wang, Kang Wang, Haoran Wang and Guang Yang performed quality control of the data; Chengyan Wang, Jun Lyu, Jin Qin and Xiaobo Qu wrote the paper; All authors revised and corrected the manuscript. Chengyan Wang and Jun Lyu contributed equally to the paper and Jing Qin, Xiaobo Qu are senior authors of this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, C., Lyu, J., Wang, S. et al. CMRxRecon: A publicly available k-space dataset and benchmark to advance deep learning for cardiac MRI. Sci Data 11, 687 (2024). https://doi.org/10.1038/s41597-024-03525-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03525-4
- Springer Nature Limited