Abstract
Euglena gracilis (E. gracilis), pivotal in the study of photosynthesis, endosymbiosis, and chloroplast development, is also an industrial microalga for paramylon production. Despite its importance, E. gracilis genome exploration faces challenges due to its intricate nature. In this study, we achieved a chromosome-level de novo assembly (2.37 Gb) using Illumina, PacBio, Bionano, and Hi-C data. The assembly exhibited a contig N50 of 619 Kb and scaffold N50 of 1.12 Mb, indicating superior continuity. Approximately 99.83% of the genome was anchored to 46 chromosomes, revealing structural insights. Repetitive elements constituted 58.84% of the sequences. Functional annotations were assigned to 39,362 proteins, enhancing interpretative power. BUSCO analysis confirmed assembly completeness at 80.39%. This first high-quality E. gracilis genome offers insights for genetics and genomics studies, overcoming previous limitations. The impact extends to academic and industrial research, providing a foundational resource.
Similar content being viewed by others
Background & Summary
Euglena, a genus of single-celled flagellate eukaryotes, is ubiquitously distributed in both freshwater and saltwater environments. Possessing photosynthetic chloroplasts, Euglena exhibits autotrophic characteristics akin to plants, while also displaying heterotrophic attributes similar to animals1,2,3. E. gracilis, a prominent species within the genus, serves as a widely utilized model organism in both academic and industrial research due to its rich array of valuable compounds, including pigments, unsaturated fatty acids, vitamins, amino acids, and the distinctive β-1,3-glucan, paramylon—an advantageous functional food ingredient4,5,6. Notably, recent studies, such as Wu et al.’s pilot-scale fermentation achieving maximal biomass and paramylon content7, underscore the industrial potential of E. gracilis.
Despite substantial advancements in genetic modification8,9,10,11,12,13, hindered by the absence of a high-quality genome, E. gracilis remains a subject of limited genetic engineering tools and applications. In 2019, Ebenezer et al. presented an initial genome assembly of E. gracilis (1.43 Gb), which, though informative, proved significantly fragmented14. Consequently, researchers have resorted to omics approaches, including de novo transcriptome assembly14,15 and proteomic analysis1,14, to explore physiological and genomic aspects. Nevertheless, a definitive high-quality genome assembly remains a critical prerequisite for advancing genetic engineering and synthetic biology applications in E. gracilis6.
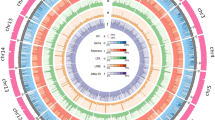
This study addresses the existing gap by introducing a chromosome-level genome assembly of E. gracilis through the integration of Illumina, PacBio, Bionano, and Hi-C technologies (Table 1). The resulting assembly, spanning 2.37 Gb, with contig N50 of 619 Kb and scaffold N50 of 1.12 Mb, exhibits superior continuity (Table 2). Anchoring to 46 chromosomes (Fig. 1a) achieved a remarkable 99.83% rate, unveiling structural insights. Repetitive elements, constituting 58.84% of the genome, contribute to its complexity. The annotation of 39,362 protein-coding gene models and the assessment of 80.39% gene completeness attest to the high quality of this genome. This achievement marks a pivotal step in enhancing our comprehension of E. gracilis, offering a genetic foundation for both experimental and computational inquiries in this species.
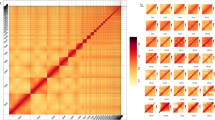
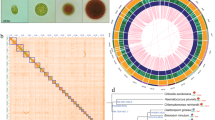
Chromosome-level assembly of the E. gracilis genome. (a) Genome landscape of the E. gracilis. From the outer ring to the inner ring are the distributions of chromosome length, gene density, transposable element (TE) density, tandem repeat (TR) density, and GC content, with densities calculated within a 1 Mb window. (b) Distribution estimation of 19-kmer. (c) Estimation based on flow cytometry. (d) Hi-C interaction heatmap illustrating the genomic interactions within the E. gracilis genome. The colour bar indicates contact density, ranging from red (high) to white (low).
Methods
Sample collection and sequencing
Sample preparation
The E. gracilis Z strain (CCAP 1224/5Z) was purchased from CCAP (Culture Collection of Algae and Protozoa, United Kingdom) and cultivated in our laboratory under autotrophic conditions using CM medium at 26 °C, with a continuous white light intensity of 80 μmol photons·m−2·s−1. Cellular samples were harvested during the mid-log phase, rapidly frozen with liquid nitrogen, and subsequently preserved at −80 °C for subsequent sequencing library preparation.
Library preparation and sequencing
Genomic DNA of high quality was extracted using the CTAB method. Paired-end libraries were constructed using NEBNext Ultra II DNA Library Prep Kit for Illumina (NEB, USA) and sequenced on an Illumina HiSeq2500 platform (Illumina, USA), which generated a total of 264.2 Gb Illumina data, providing approximately 111-fold coverage of the genome (Table 1). In total of 50 mg DNA were used to construct the PacBio Sequel sequencing libraries, then sequencing was performed to produce raw reads. For Bionano sequencing, high molecular weight DNA with a fragment distribution greater than 150 kb were isolated and used for DNA nicking using Nb.BssSI (NEB). The nicks were labelled and then loaded onto the Saphyr Chip nanochannel array (Bionano Genomics) and imaged using the Saphyr system and associated software (Bionano Genomics) according to the Saphyr System User Guide. The PacBio Sequel and Bionano platforms contributed 377.5 Gb and 306.6 Gb data, achieving coverages of approximately 159X and 129X, respectively (Table 1). Hi-C libraries was prepared with the standard procedure described. After digesting the genomic DNA with a restriction enzyme MboI, the sticky ends of the digested fragments were biotinylated, diluted, and then ligated to each other randomly. The prepared sequencing library was sequenced on a NovaSeq platform (Illumina, USA), which yielded a total of 402.3 Gb data with the Illumina sequencing platform (Table 1). Library preparation and sequencing of Illumina survey libraries, PacBio Sequel libraries, Bionano libraries, and all transcriptome libraries were executed by Nowbio Biotechnology Company (Yunnan, China). Frasergen Bioinformatics Co., Ltd (Wuhan, China) undertook the preparation and sequencing of Hi-C libraries on their sequencing platform.
Genome survey and assembly
K-mer frequency analysis
K-mer frequencies (K = 19) were computed from filtered Illumina reads using Jellyfish16 (v2.2.10), serving as the basis for a genome survey conducted with GenomeScope17 (v2.0). The estimated genome size for E. gracilis was determined to be 2.25 Gb (Fig. 1b), aligning closely with the genome size estimations derived from flow cytometry analysis (2.14–2.34 Gb) (Fig. 1c).
Genome assembly
To assemble the genome, NextDenovo18 (v2.2-beta.0) was employed to generate contigs utilizing PacBio reads, followed by three rounds of Illumina read correction using NextPolish19 (v1.0.1). The corrected contigs underwent assembly with Bionano data using Sovle (v3.3). Subsequently, the assembled scaffolds were organized into chromosomes utilizing the 3D-DNA pipeline20 (v201008), followed by manual curation with JuiceBox21 (v2.20.00). The final assembly comprised 46 chromosomes (Fig. 1d), collectively spanning 2.37 Gb, accounting for approximately 99.83% of the entire genome assembly (Table 2), while the individual chromosome lengths ranged from 121.4 Mb (Chr4) to 22.7 Mb (Chr35) (Table 3). Comparing with the previous genome assembly14 of E. gracilis presented by Ebenezer et al., our assembly has much longer N50 and higher BUSCO completeness score (Table 2), which fully suggested that our result is a high-quality assembly, with superior continuity.
Genome repeat and ncRNA analysis
Repeat sequence prediction
A hybrid approach, incorporating both ab initio and homology-based methodologies, was employed to predict repeat sequences within the genome. For ab initio prediction, LTR_FINDER22 (v1.07) and ltrharvest23 (v1.5.10) were used to predict LTR retrotransposons, and the results were integrated using LTR_retriever24 (v2.8). Meanwhile, RepeatModeler25 (v2.0) was also used to identify repeats. Then the results of LTR_retriever and RepeatModeler were merged as a custom library and fed to Repeatmasker26 (v.4.0.9) to predict TEs. Simultaneously, homology-based annotation employed RepeatMasker26 (v.4.0.9) and RepeatProteinMask26 (v.4.0.9) against Repbase27 (Release 20181026). TRF28 (v4.0.9) was used for searching tandem repeats. Following redundancy elimination, a total of 1.4 Gb of repeat sequences were identified, constituting 58.84% of the E. gracilis genome. The repeat sequences predicted by TRF, Repeatmasker, Proteinmask and ab initio pipeline covered 9.85%, 1.89%, 2.07% and 52.75% of the genome sequence, respectively. Within the repeat elements, 32.73% remained unclassified, while long terminal repeats (LTRs) represented 32.81% of the genome. DNA elements, long interspersed nuclear elements (LINEs), and short interspersed nuclear elements (SINEs) accounted for 4.60%, 1.49%, and 0.11% of the genome, respectively (Table 4).
Noncoding RNA annotation
To annotate noncoding RNA (ncRNA), tRNAScan-SE29 (v1.3.1) and blast30 (v2.2.26) were applied for tRNA and rRNA prediction, respectively. Additionally, Rfam31 (v9.1) and INFERNAL32 (v0.81) were utilized for miRNA and snRNA prediction on the genome. This comprehensive approach identified four types of ncRNAs within the E. gracilis genome, encompassing 188 miRNAs, 4882 tRNAs, 223 rRNAs, and 165 snRNAs.
Gene prediction and annotation
Pre-processing and de novo assembly
The Illumina RNA-seq data underwent initial filtration utilizing Trimmomatic33 (v0.32) to obtain clean reads, subsequently employed in Trinity34 (v2.1.1) for de novo assembly. The Pacbio full-length RNA-seq dataset was refined to derive consensus sequences using smrtlink (v6.0.0).
Transcript integration and ab initio prediction
The two distinct sets of transcripts were amalgamated via PASA35 (v2.4.1) for ab initio gene prediction, utilizing Augustus36 (v2.5.5) and SNAP37 (2006-07-28). Homology annotation was conducted with ten representative species, including Bodo saltans, Naegleria gruberi, Phytomonas sp., Chlamydomonas reinhardtii, Leishmania major Friedlin, Nannochloropsis gaditana, Trypanosoma brucei, Cyanidioschyzon merolae, Leptomonas pyrrhocoris, and Perkinsela sp., downloaded from NCBI. The comprehensive integration of all data and generation of the predicted gene set were accomplished using MAKER38 (v3.01.02). The ensuing analysis revealed a total of 32,806 genes and 39,362 coding DNA sequences (CDSs) within the E. gracilis genome, with an average CDS length of 1,149 bp and an average of 8 exons per gene.
Functional annotation
For functional annotation, blastp30 (v2.2.26) was applied to align protein-coding genes with KEGG39 database. The GO Ontology40 (GO) and InterPro41 function were obtained using InterProScan. The subsequent functional annotation of CDSs demonstrated coverage of 28.2%, 40.6%, and 50.2% across the GO, InterPro, and KEGG databases, respectively, with a cumulative 57.3% of CDSs annotated in at least one database.
Data Records
Sequencing data deposit
The comprehensive E. gracilis genome project has been archived in the Genome Sequence Archive42,43 (GSA) under the accession44 CRA013190, except that the Illumina RNA-seq data have been archived in the SRA at NCBI SRP35377445.
Assembly deposit
The assembly of the E. gracilis genome, along with its corresponding annotation file, is available at figshare46 and NCBI GenBank with accession number GCA_039621445.147.
Technical Validation
Genome assembly quality assessment
The quality assessment of the E. gracilis genome assembly was executed through two distinct methodologies. Firstly, the completeness of the assembly was rigorously validated utilizing compleasm48 (v0.2.2), an improved BUSCO49 workflow based on miniprot, with specific parameters (-m lite–min_identity 0.8–min_length_percent 0.9–min_rise 0.9), and employing the eukaryota_odb10 (v5, 2020-09-10) reference gene set (n = 255). The final BUSCO analysis yielded a completeness score of 80.39%, comprised of 162 (63.53%) single-copy BUSCOs, 43 (16.86%) duplicated BUSCOs, 11 (4.31%) fragmented BUSCOs, and 39 (15.29%) missing BUSCOs. Secondly, to affirm the accuracy and integrity of the genome survey, the filtered Illumina short reads utilized were aligned back to the E. gracilis genome utilizing the Burrows-Wheeler aligner50 (BWA, v0.7.17-r1188). This meticulous alignment process revealed an impressive mapping rate of 99.42% for the short reads against the genome. The combination of these validated results attests to the high-quality nature of the E. gracilis genome assembly.
Code availability
All commands and pipelines employed for data processing adhered strictly to the guidelines specified in the manuals of the pertinent bioinformatics software, with the parameters explicitly detailed in the Methods section. In instances where no specific parameters were explicitly stated for a particular software, default parameters were applied. It is noteworthy that no bespoke scripts or custom code were formulated or utilized throughout the course of this study.
References
Chen, Z. et al. Proteomic Responses of Dark-Adapted Euglena gracilis and Bleached Mutant Against Light Stimuli. Frontiers in bioengineering and biotechnology 10, 843414 (2022).
Qin, H. et al. Occurrence and light response of residual plastid genes in a Euglena gracilis bleached mutant strain OflB2. Journal of Oceanology and Limnology 38, 1858–1866 (2020).
Shao, Q. et al. Metabolomic response of Euglena gracilis and its bleached mutant strain to light. PLoS One 14, e0224926 (2019).
Gissibl, A., Sun, A., Care, A., Nevalainen, H. & Sunna, A. Bioproducts from Euglena gracilis: synthesis and applications. Frontiers in bioengineering and biotechnology 7, 108 (2019).
Kottuparambil, S., Thankamony, R. L. & Agusti, S. Euglena as a potential natural source of value-added metabolites. A review. Algal research 37, 154–159 (2019).
Chen, Z. et al. A Synthetic Biology Perspective on the Bioengineering Tools for an Industrial Microalga: Euglena gracilis. Frontiers in Bioengineering and Biotechnology 10 (2022).
Wu, M. et al. A new pilot-scale fermentation mode enhances Euglena gracilis biomass and paramylon (β-1,3-glucan) production. Journal of Cleaner Production 321, 128996 (2021).
Becker, I. et al. Agrobacterium tumefaciens-mediated nuclear transformation of a biotechnologically important microalga—Euglena gracilis. International Journal of Molecular Sciences 22, 6299 (2021).
Chen, Z. et al. High‐throughput sequencing revealed low-efficacy genome editing using Cas9 RNPs electroporation and single‐celled microinjection provided an alternative to deliver CRISPR reagents into Euglena gracilis. Plant Biotechnology Journal 20, 2048 (2022).
Gao, P. & Sun, C. Fast and efficient molecule delivery into Euglena gracilis mediated by cell‐penetrating peptide or dimethyl sulfoxide. FEBS Open bio 13, 597–605 (2023).
Khatiwada, B., Kautto, L., Sunna, A., Sun, A. & Nevalainen, H. Nuclear transformation of the versatile microalga Euglena gracilis. Algal Research 37, 178–185 (2019).
Nakazawa, M. et al. Stable nuclear transformation methods for Euglena gracilis and its application to a related Euglenida. Algal Research 75, 103292 (2023).
Nomura, T. et al. Highly efficient transgene‐free targeted mutagenesis and single‐stranded oligodeoxynucleotide‐mediated precise knock‐in in the industrial microalga Euglena gracilis using Cas9 ribonucleoproteins. Plant biotechnology journal 17, 2032 (2019).
Ebenezer, T. E. et al. Transcriptome, proteome and draft genome of Euglena gracilis. BMC Biol 17, 11 (2019).
Cordoba, J. et al. De Novo Transcriptome Meta-Assembly of the Mixotrophic Freshwater Microalga Euglena gracilis. Genes (Basel) 12 (2021).
Marçais, G. & Kingsford, C. A fast, lock-free approach for efficient parallel counting of occurrences of k-mers. Bioinformatics 27, 764–770 (2011).
Ranallo-Benavidez, T. R., Jaron, K. S. & Schatz, M. C. GenomeScope 2.0 and Smudgeplot for reference-free profiling of polyploid genomes. Nature communications 11, 1432 (2020).
Hu, J. et al. An efficient error correction and accurate assembly tool for noisy long reads. bioRxiv, (2023). 2023.03.09.531669.
Hu, J., Fan, J., Sun, Z. & Liu, S. NextPolish: a fast and efficient genome polishing tool for long-read assembly. Bioinformatics 36, 2253–2255 (2020).
Dudchenko, O. et al. De novo assembly of the Aedes aegypti genome using Hi-C yields chromosome-length scaffolds. Science 356, 92–95 (2017).
Durand, N. C. et al. Juicebox Provides a Visualization System for Hi-C Contact Maps with Unlimited Zoom. Cell Syst 3, 99–101 (2016).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res 35, W265–8 (2007).
Ellinghaus, D., Kurtz, S. & Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinformatics 9, 18 (2008).
Ou, S. & Jiang, N. LTR_retriever: A Highly Accurate and Sensitive Program for Identification of Long Terminal Repeat Retrotransposons. Plant Physiol 176, 1410–1422 (2018).
Flynn, J. M. et al. RepeatModeler2 for automated genomic discovery of transposable element families. Proc Natl Acad Sci USA 117, 9451–9457 (2020).
Tarailo-Graovac, M. & Chen, N. Using RepeatMasker to identify repetitive elements in genomic sequences. Curr Protoc Bioinformatics Chapter 4, 4.10.1–4.10.14 (2009).
Bao, W., Kojima, K. K. & Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mobile DNA 6, 11 (2015).
Benson, G. Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27, 573–80 (1999).
Chan, P. P. & Lowe, T. M. tRNAscan-SE: Searching for tRNA Genes in Genomic Sequences. Methods Mol Biol 1962, 1–14 (2019).
Ye, J., McGinnis, S. & Madden, T. L. BLAST: improvements for better sequence analysis. Nucleic acids research 34, W6–W9 (2006).
Kalvari, I. et al. Non-Coding RNA Analysis Using the Rfam Database. Curr Protoc Bioinformatics 62, e51 (2018).
Nawrocki, E. P. & Eddy, S. R. Infernal 1.1: 100-fold faster RNA homology searches. Bioinformatics 29, 2933–5 (2013).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–20 (2014).
Grabherr, M. G. et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol 29, 644–52 (2011).
Haas, B. J. et al. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res 31, 5654–66 (2003).
Stanke, M. et al. AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic acids research 34, W435–W439 (2006).
Korf, I. Gene finding in novel genomes. BMC Bioinformatics 5, 59 (2004).
Holt, C. & Yandell, M. MAKER2: an annotation pipeline and genome-database management tool for second-generation genome projects. BMC Bioinformatics 12, 491 (2011).
Ogata, H. et al. KEGG: Kyoto Encyclopedia of Genes and Genomes. Nucleic Acids Res 27, 29–34 (1999).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nature Genetics 25, 25–29 (2000).
Blum, M. et al. The InterPro protein families and domains database: 20 years on. Nucleic acids research 49, D344–D354 (2021).
Database Resources of the National Genomics Data Center, China National Center for Bioinformation in 2023. Nucleic Acids Res 51, D18-d28 (2023).
Chen, T. et al. The Genome Sequence Archive Family: Toward Explosive Data Growth and Diverse Data Types. Genomics Proteomics Bioinformatics 19, 578–583 (2021).
Genome Sequence Archive https://ngdc.cncb.ac.cn/gsa/browse/CRA013190 (2024).
NCBI Sequence Read Archive https://identifiers.org/ncbi/insdc.sra:SRP353774 (2022).
Chen, Z. et al. A chromosome-level genome assembly for the paramylon-producing microalga Euglena gracilis, Figshare, https://doi.org/10.6084/m9.figshare.c.7024970.v1 (2024).
NCBI GenBank https://identifiers.org/ncbi/insdc.gca:GCA_039621445.1 (2024).
Huang, N. & Li, H. compleasm: a faster and more accurate reimplementation of BUSCO. Bioinformatics 39 (2023).
Manni, M., Berkeley, M. R., Seppey, M., Simao, F. A. & Zdobnov, E. M. BUSCO Update: Novel and Streamlined Workflows along with Broader and Deeper Phylogenetic Coverage for Scoring of Eukaryotic, Prokaryotic, and Viral Genomes. Mol Biol Evol 38, 4647–4654 (2021).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–60 (2009).
Acknowledgements
This work was partially supported by China’s National Key R&D Programs (2021YFA0910800, 2018YFA0902500, 2020YFA0908703), the National Natural Science Foundation of China (41876188), the Science Technology and Innovation Committee of Shenzhen Municipality (KCXFZ202002011006448), Shenzhen Science and Technology Program (KCXST20221021111206015 and KCXFZ20201221173404012), and Natural Science Foundation of Guangdong Province (2024B1515020034).
Author information
Authors and Affiliations
Contributions
Zixi Chen, Yang Dong and Shengchang Duan analysed the data and wrote the manuscript. Jiayi He, Huan Qin, Zhenfan Chen, Chenchen Liu, Chao Zheng, Ming Du, Rao Yao and Chao Li performed the experiments. Chao Bian, Panpan Jiang and Qiong Shi analysed the data. Yun Wang, Shuangfei Li, Ning Xie, Ying Xu and Zhangli Hu revised the manuscript. Anping Lei, Liqing Zhao and Jiangxin Wang conceived and designed the whole project, and revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, Z., Dong, Y., Duan, S. et al. A chromosome-level genome assembly for the paramylon-producing microalga Euglena gracilis. Sci Data 11, 780 (2024). https://doi.org/10.1038/s41597-024-03404-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41597-024-03404-y
- Springer Nature Limited