Abstract
Type-III effector proteins are major virulence determinants that most gram-negative bacteria inject into host cells to manipulate cellular processes for infection. Because effector-targeted cells are embedded and underrepresented in infected plant tissues, it is technically challenging to isolate them for focused studies of effector-induced cellular changes. This protocol describes a novel technique, effector-inducible isolation of nuclei tagged in specific cell types (eINTACT), for isolating biotin-labeled nuclei from Arabidopsis plant cells that have received Xanthomonas bacterial effectors by using streptavidin-coated magnetic beads. This protocol is an extension of the existing Nature Protocols Protocol of the INTACT method for the affinity-based purification of nuclei of specific cell types in the context of developmental biology. In a phytopathology scenario, our protocol addresses how to obtain eINTACT transgenic lines and compatible bacterial mutants, verify the eINTACT system and purify nuclei of bacterial effector-recipient cells from infected tissues. Differential analyses of purified nuclei from plants infected by bacteria expressing the effector of interest and those from plants infected by effector-deletion bacterial mutants will reveal the effector-dependent nuclear changes in targeted host cells. Provided that the eINTACT system is available, the infection experiment takes 5 d, and the procedures, from collecting bacteria-infected leaves to obtaining nuclei of effector-targeted cells, can be completed in 4 h. eINTACT is a unique method for isolating high-quality nuclei from bacterial effector-targeted host cells in native infection contexts. This method is adaptable to study the functions of type-III effectors from numerous gram-negative bacteria in host plants that are amenable to transformation.
Key points
-
This protocol describes a method for purifying nuclei from live plant cells that have been specifically targeted by bacterial effectors during bacterial infection.
-
Compared with similar affinity-based nuclei-isolation techniques that isolate cell type-specific nuclei such as INTACT, this method specifically isolates nuclei from host cells that have received bacterial effectors, allowing the interrogation of the effect of bacterial effectors on these cells in live infection models.



Similar content being viewed by others
Data availability
All data supporting this protocol are available in the main text or supplementary materials or are accessible via the original research article12. Source data are provided with this paper.
References
Melotto, M., Zhang, L., Oblessuc, P. R. & He, S. Y. Stomatal defense a decade later. Plant Physiol. 174, 561–571 (2017).
Cerutti, A. et al. Immunity at cauliflower hydathodes controls systemic infection by Xanthomonas campestris pv campestris. Plant Physiol. 174, 700–716 (2017).
Buttner, D. Behind the lines—actions of bacterial type III effector proteins in plant cells. FEMS Microbiol. Rev. 40, 894–937 (2016).
El Kasmi, F., Horvath, D. & Lahaye, T. Microbial effectors and the role of water and sugar in the infection battle ground. Curr. Opin. Plant Biol. 44, 98–107 (2018).
Garcia-Ruiz, H., Szurek, B. & Van den Ackerveken, G. Stop helping pathogens: engineering plant susceptibility genes for durable resistance. Curr. Opin. Biotechnol. 70, 187–195 (2021).
Wu, D. et al. A plant pathogen type III effector protein subverts translational regulation to boost host polyamine levels. Cell Host Microbe 26, 638–649.e5 (2019).
Li, Z., Variz, H., Chen, Y., Liu, S. L. & Aung, K. Plasmodesmata-dependent intercellular movement of bacterial effectors. Front. Plant Sci. 12, 640277 (2021).
Tan, C. M. et al. Arabidopsis HFR1 is a potential nuclear substrate regulated by the Xanthomonas type III effector XopD (Xcc8004). PLoS ONE 10, e0117067 (2015).
Tan, L., Rong, W., Luo, H., Chen, Y. & He, C. The Xanthomonas campestris effector protein XopDXcc8004 triggers plant disease tolerance by targeting DELLA proteins. N. Phytol. 204, 595–608 (2014).
Ngou, B. P. M., Ahn, H. K., Ding, P. & Jones, J. D. G. Mutual potentiation of plant immunity by cell-surface and intracellular receptors. Nature 592, 110–115 (2021).
Zhang, J., Coaker, G., Zhou, J. M. & Dong, X. Plant immune mechanisms: from reductionistic to holistic points of view. Mol. Plant 13, 1358–1378 (2020).
You, Y. et al. The eINTACT system dissects bacterial exploitation of plant osmosignalling to enhance virulence. Nat. Plants 9, 128–141 (2023).
Kim, J. G. et al. XopD SUMO protease affects host transcription, promotes pathogen growth, and delays symptom development in Xanthomonas-infected tomato leaves. Plant Cell 20, 1915–1929 (2008).
Deal, R. B. & Henikoff, S. The INTACT method for cell type-specific gene expression and chromatin profiling in Arabidopsis thaliana. Nat. Protoc. 6, 56–68 (2011).
Moreno-Romero, J., Santos-Gonzalez, J., Hennig, L. & Kohler, C. Applying the INTACT method to purify endosperm nuclei and to generate parental-specific epigenome profiles. Nat. Protoc. 12, 238–254 (2017).
Romer, P. et al. Plant pathogen recognition mediated by promoter activation of the pepper Bs3 resistance gene. Science 318, 645–648 (2007).
Slane, D., Kong, J., Schmid, M., Jurgens, G. & Bayer, M. Profiling of embryonic nuclear vs. cellular RNA in Arabidopsis thaliana. Genom. Data 4, 96–98 (2015).
Zaghlool, A. et al. Characterization of the nuclear and cytosolic transcriptomes in human brain tissue reveals new insights into the subcellular distribution of RNA transcripts. Sci. Rep. 11, 4076 (2021).
Palovaara, J. et al. Transcriptome dynamics revealed by a gene expression atlas of the early Arabidopsis embryo. Nat. Plants 3, 894–904 (2017).
You, Y. et al. Phloem companion cell-specific transcriptomic and epigenomic analyses identify MRF1, a regulator of flowering. Plant Cell 31, 325–345 (2019).
Reynoso, M. A. et al. Nuclear transcriptomes at high resolution using retooled INTACT. Plant Physiol. 176, 270–281 (2018).
Kajala, K. et al. Innovation, conservation, and repurposing of gene function in root cell type development. Cell 184, 3333–3348.e19 (2021).
You, Y. et al. Temporal dynamics of gene expression and histone marks at the Arabidopsis shoot meristem during flowering. Nat. Commun. 8, 15120 (2017).
Yadav, V. K., Santos-Gonzalez, J. & Kohler, C. INT-Hi-C reveals distinct chromatin architecture in endosperm and leaf tissues of Arabidopsis. Nucleic Acids Res. 49, 4371–4385 (2021).
Sijacic, P., Bajic, M., McKinney, E. C., Meagher, R. B. & Deal, R. B. Changes in chromatin accessibility between Arabidopsis stem cells and mesophyll cells illuminate cell type-specific transcription factor networks. Plant J. 94, 215–231 (2018).
Maher, K. A. et al. Profiling of accessible chromatin regions across multiple plant species and cell types reveals common gene regulatory principles and new control modules. Plant Cell 30, 15–36 (2018).
Boch, J. & Bonas, U. Xanthomonas AvrBs3 family-type III effectors: discovery and function. Annu. Rev. Phytopathol. 48, 419–436 (2010).
de Lange, O. et al. Breaking the DNA-binding code of Ralstonia solanacearum TAL effectors provides new possibilities to generate plant resistance genes against bacterial wilt disease. N. Phytol. 199, 773–786 (2013).
Pandey, S. P., Benstein, R. M., Wang, Y. & Schmid, M. Epigenetic regulation of temperature responses—past successes and future challenges. J. Exp. Bot. 72, 7482–7497 (2021).
Denyer, T. & Timmermans, M. C. P. Crafting a blueprint for single-cell RNA sequencing. Trends Plant Sci. 27, 92–103 (2022).
Marques-Bueno, M. D. M. et al. A versatile multisite gateway-compatible promoter and transgenic line collection for cell type-specific functional genomics in Arabidopsis. Plant J. 85, 320–333 (2016).
Winter, D. et al. An "Electronic Fluorescent Pictograph" browser for exploring and analyzing large-scale biological data sets. PLoS ONE 2, e718 (2007).
Klepikova, A. V., Kasianov, A. S., Gerasimov, E. S., Logacheva, M. D. & Penin, A. A. A high resolution map of the Arabidopsis thaliana developmental transcriptome based on RNA-seq profiling. Plant J. 88, 1058–1070 (2016).
Mockler, T. C. et al. The DIURNAL project: DIURNAL and circadian expression profiling, model-based pattern matching, and promoter analysis. Cold Spring Harb. Symp. Quant. Biol. 72, 353–363 (2007).
Ron, M. et al. Hairy root transformation using Agrobacterium rhizogenes as a tool for exploring cell type-specific gene expression and function using tomato as a model. Plant Physiol. 166, 455–469 (2014).
Marois, E., Van den Ackerveken, G. & Bonas, U. The Xanthomonas type III effector protein AvrBs3 modulates plant gene expression and induces cell hypertrophy in the susceptible host. Mol. Plant Microbe Interact. 15, 637–646 (2002).
Kay, S., Hahn, S., Marois, E., Hause, G. & Bonas, U. A bacterial effector acts as a plant transcription factor and induces a cell size regulator. Science 318, 648–651 (2007).
Li, T., Huang, S., Zhou, J. & Yang, B. Designer TAL effectors induce disease susceptibility and resistance to Xanthomonas oryzae pv. oryzae in rice. Mol. Plant 6, 781–789 (2013).
Guy, E. et al. Natural genetic variation of Xanthomonas campestris pv. campestris pathogenicity on Arabidopsis revealed by association and reverse genetics. mBio 4, e00538–e00512 (2013).
Van den Ackerveken, G., Marois, E. & Bonas, U. Recognition of the bacterial avirulence protein AvrBs3 occurs inside the host plant cell. Cell 87, 1307–1316 (1996).
Rivero, L. et al. Handling Arabidopsis plants: growth, preservation of seeds, transformation, and genetic crosses. Methods Mol. Biol. 1062, 3–25 (2014).
Acknowledgements
We thank T. Lahaye (Eberhard-Karls-University Tübingen, Germany) for supplying the pDS300F and pDSK plasmids and a preliminary idea of a TALE-inducible INTACT system, L. D. Noël (Université de Toulouse, France) for contributing the Xcc bacterial strains and A.-G. Andrade-Galan for technical assistance. Our research benefits from the infrastructure at the ZMBP at the Eberhard-Karls-University Tübingen. This work was supported by the Institutional Strategy Program of the University of Tübingen (DFG, ZUK 63) and the German Research Foundation (DFG, no. 427105396) (both to Y.Y).
Author information
Authors and Affiliations
Contributions
Y.Y. acquired funding, supervised the research and performed most experiments; Z.J. performed western blotting and microscopic analysis. Y.Y. analyzed the overall results and wrote the manuscript with input from Z.J.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Adam Bogdanove, Roger Deal and Jordi Moreno-Romero for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key reference using this protocol
You, Y. et al. Nat. Plants 9, 128–141 (2023): https://doi.org/10.1038/s41477-022-01302-y
This protocol is an extension to: Nat. Protoc. 6, 56–68 (2011): https://doi.org/10.1038/nprot.2010.175 and Nat. Protoc. 12, 238–254 (2017): https://doi.org/10.1038/nprot.2016.167
Extended data
Extended Data Fig. 1 XopD suppresses wilting and necrotic symptoms caused by Xcc∆xopD* infection.
Representative Arabidopsis leaves infected with Xcc∆xopD*AvrBs3 (−XopD) and Xcc*AvrBs3 (+XopD) at 7 DPI.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2
Source data
Source Data Fig. 2
Statistical data for Fig. 2b. Bacterial population per cm2 of Xcc*-inoculated leaves of the wild-type Col-0 plants and Xcc*AvrBs3-inoculated leaves of the eINTACT transgenic line at 7 DPI.
Source Data Fig. 2
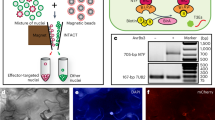
Uncropped original gel pictures for Fig. 2c,d. c, Semi-quantitative RT-PCR analysis of RedNTF expression in leaves of the Arabidopsis eINTACT transgenic line infected with Xcc*AvrBs3 and Xcc*vec at 5 DPI. The expression of TUB2 and the genomic DNA of RedNTF and TUB loci (gRedNTF and gTUB) in the eINTACT transgenic line are used as controls. d, Uncropped original western blots. Enrichment of the RedNTF protein in eINTACT-purified nuclei. RedNTF and H3 protein detection via western blotting of the total protein extracts from Xcc*AvrBs3-infected and Xcc*vec-infected infected leaves and eINTACT-purified nuclei (nuceINTACT) from Xcc*AvrBs3-infected leaves. The blotted membrane was divided into two at 25 kDa, according to size marks in the protein ladder. The upper part (25–180 kDa) was used for detecting biotinylated-RedNTF protein (42 kDa) by using streptavidin alkaline phosphatase. The lower part (10–25 kDa) was used for detecting H3 protein by using an anti-H3 antibody. The abundance of H3 protein serves as an internal control of the number of nuclei in each sample.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
You, Y., Jiang, Z. The eINTACT method for studying nuclear changes in host plant cells targeted by bacterial effectors in native infection contexts. Nat Protoc 18, 3173–3193 (2023). https://doi.org/10.1038/s41596-023-00879-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-023-00879-8
- Springer Nature Limited