Abstract
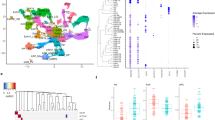
Major depressive disorder (MDD) has an enormous impact on global disease burden, affecting millions of people worldwide and ranking as a leading cause of disability for almost three decades. Past molecular studies of MDD employed bulk homogenates of postmortem brain tissue, which obscures gene expression changes within individual cell types. Here we used single-nucleus transcriptomics to examine ~80,000 nuclei from the dorsolateral prefrontal cortex of male individuals with MDD (n = 17) and of healthy controls (n = 17). We identified 26 cellular clusters, and over 60% of these showed differential gene expression between groups. We found that the greatest dysregulation occurred in deep layer excitatory neurons and immature oligodendrocyte precursor cells (OPCs), and these contributed almost half (47%) of all changes in gene expression. These results highlight the importance of dissecting cell-type-specific contributions to the disease and offer opportunities to identify new avenues of research and novel targets for treatment.






Similar content being viewed by others
Data availability
Raw sequencing data, annotated gene–barcode matrix and lists of cells used for differential gene expression analysis are accessible on GEO using the accession number GSE144136. RNAScope and high-throughput qPCR data are available upon request.
Code availability
A sample custom R script (Supplementary_R_Script_1.R) used for analyzing high-throughput qPCR data is provided and an R script used to test the statistical significance of CCInx interactions is provided (Supplementary_R_Script_2.R) along with this paper.
References
Depression and Other Common Mental Disorders: Global Health Estimates (World Health Organization, 2017).
Wray, N. R. et al. Genome-wide association analyses identify 44 risk variants and refine the genetic architecture of major depression. Nat. Genet. 50, 668–681 (2018).
Jansen, R. et al. Gene expression in major depressive disorder. Mol. Psychiatry 21, 339–347 (2016).
Sequeira, A. et al. Global brain gene expression analysis links glutamatergic and GABAergic alterations to suicide and major depression. PLoS ONE 4, e6585 (2009).
Abdallah, C. G., Sanacora, G., Duman, R. S. & Krystal, J. H. The neurobiology of depression, ketamine and rapid-acting antidepressants: is it glutamate inhibition or activation? Pharmacol. Ther. 190, 148–158 (2018).
Pantazatos, S. P. et al. Whole-transcriptome brain expression and exon-usage profiling in major depression and suicide: evidence for altered glial, endothelial and ATPase activity. Mol. Psychiatry 22, 760–773 (2016).
Edgar, N. & Sibille, E. A putative functional role for oligodendrocytes in mood regulation. Transl Psychiatry 2, e109 (2012).
Nagy, C. et al. Astrocytic abnormalities and global DNA methylation patterns in depression and suicide. Mol. Psychiatry 20, 320–328 (2015).
Duman, R. S., Aghajanian, G. K., Sanacora, G. & Krystal, J. H. Synaptic plasticity and depression: new insights from stress and rapid-acting antidepressants. Nat. Med. 22, 238–249 (2016).
Lake, B. B. et al. Integrative single-cell analysis of transcriptional and epigenetic states in the human adult brain. Nat. Biotechnol. 36, 70–80 (2018).
Habib, N. et al. Massively parallel single-nucleus RNA-seq with DroNc-seq. Nat. Methods 14, 955–958 (2017).
Mancarci, B. O. et al. Cross-laboratory analysis of brain cell type transcriptomes with applications to interpretation of bulk tissue data. eNeuro 4, ENEURO.0212-17.2017 (2017).
Zheng, G. X. Y. et al. Massively parallel digital transcriptional profiling of single cells. Nat. Commun. 8, 14049 (2017).
Lake, B. B. et al. A comparative strategy for single-nucleus and single-cell transcriptomes confirms accuracy in predicted cell-type expression from nuclear RNA. Sci. Rep. 7, 6031 (2017).
Renthal, W. et al. Characterization of human mosaic Rett syndrome brain tissue by single-nucleus RNA sequencing. Nat. Neurosci. 21, 1670–1679 (2018).
Mathys, H. et al. Single-cell transcriptomic analysis of Alzheimer’s disease. Nature 570, 332–337 (2019).
Velmeshev, D. et al. Single-cell genomics identifies cell type-specific molecular changes in autism. Science 364, 685–689 (2019).
Northoff, G. & Sibille, E. Why are cortical GABA neurons relevant to internal focus in depression? A cross-level model linking cellular, biochemical and neural network findings. Mol. Psychiatry 19, 966–977 (2014).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Sofroniew, M. & Vinters, H. Astrocytes: biology and pathology. Acta Neuropathol. 119, 7–35 (2010).
Anderson, M. A., Ao, Y. & Sofroniew, M. V. Heterogeneity of reactive astrocytes. Neurosci. Lett. 565, 23–29 (2014).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Thomas, P. D. et al. Applications for protein sequence-function evolution data: mRNA/protein expression analysis and coding SNP scoring tools. Nucleic Acids Res. 34, W645–W650 (2006).
Butts, B. D., Houde, C. & Mehmet, H. Maturation-dependent sensitivity of oligodendrocyte lineage cells to apoptosis: implications for normal development and disease. Cell Death Differ. 15, 1178–1186 (2008).
Jakel, S. et al. Altered human oligodendrocyte heterogeneity in multiple sclerosis. Nature 566, 543–547 (2019).
Li, Q. S., Tian, C., Seabrook, G. R., Drevets, W. C. & Narayan, V. A. Analysis of 23andMe antidepressant efficacy survey data: implication of circadian rhythm and neuroplasticity in bupropion response. Transl Psychiatry 6, e889 (2016).
Gutierrez-Sacristan, A. et al. PsyGeNET: a knowledge platform on psychiatric disorders and their genes. Bioinformatics 31, 3075–3077 (2015).
Pinero, J. et al. DisGeNET: a comprehensive platform integrating information on human disease-associated genes and variants. Nucleic Acids Res. 45, D833–D839 (2017).
Szklarczyk, D. et al. STRING v11: protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 47, D607–D613 (2019).
Wochnik, G. M. et al. FK506-binding proteins 51 and 52 differentially regulate dynein interaction and nuclear translocation of the glucocorticoid receptor in mammalian cells. J. Biol. Chem. 280, 4609–4616 (2005).
Birey, F. et al. Genetic and stress-induced loss of NG2 glia triggers emergence of depressive-like behaviors through reduced secretion of FGF2. Neuron 88, 941–956 (2015).
Mason, J. L. & Goldman, J. E. A2B5+ and O4+ cycling progenitors in the adult forebrain white matter respond differentially to PDGF-AA, FGF-2, and IGF-1. Mol. Cell. Neurosci. 20, 30–42 (2002).
Shulha, H. P. et al. Human-specific histone methylation signatures at transcription start sites in prefrontal neurons. PLoS Biol. 10, e1001427 (2012).
Spitzer, S. O. et al. Oligodendrocyte progenitor cells become regionally diverse and heterogeneous with age. Neuron 101, 459–471.e5 (2019).
Psachoulia, K., Jamen, F., Young, K. M. & Richardson, W. D. Cell cycle dynamics of NG2 cells in the postnatal and ageing brain. Neuron Glia Biol. 5, 57–67 (2009).
Bergles, D. E., Jabs, R. & Steinhauser, C. Neuron–glia synapses in the brain. Brain Res. Rev. 63, 130–137 (2010).
Birey, F., Kokkosis, A. G. & Aguirre, A. Oligodendroglia-lineage cells in brain plasticity, homeostasis and psychiatric disorders. Curr. Opin. Neurobiol. 47, 93–103 (2017).
Ge, W. P. et al. Long-term potentiation of neuron–glia synapses mediated by Ca2+-permeable AMPA receptors. Science 312, 1533–1537 (2006).
Ueno, H., Huang, X., Tanaka, Y. & Hirokawa, N. KIF16B/Rab14 molecular motor complex is critical for early embryonic development by transporting FGF receptor. Dev. Cell 20, 60–71 (2011).
Turner, C. A., Watson, S. J. & Akil, H. The fibroblast growth factor family: neuromodulation of affective behavior. Neuron 76, 160–174 (2012).
Turecki, G. & Meaney, M. J. Effects of the social environment and stress on glucocorticoid receptor gene methylation: a systematic review. Biol. Psychiatry 79, 87–96 (2016).
Zuehlke, A. D., Beebe, K., Neckers, L. & Prince, T. Regulation and function of the human HSP90AA1 gene. Gene 570, 8–16 (2015).
Huang, J. Y., Lynn Miskus, M. & Lu, H. C. FGF–FGFR mediates the activity-dependent dendritogenesis of layer IV neurons during barrel formation. J. Neurosci. 37, 12094–12105 (2017).
Pittenger, C. & Duman, R. S. Stress, depression, and neuroplasticity: a convergence of mechanisms. Neuropsychopharmacology 33, 88–109 (2008).
Caiati, M. D. et al. PrPC controls via protein kinase A the direction of synaptic plasticity in the immature hippocampus. J. Neurosci. 33, 2973–2983 (2013).
Sevilla, L. M., Nachat, R., Groot, K. R. & Watt, F. M. Kazrin regulates keratinocyte cytoskeletal networks, intercellular junctions and differentiation. J. Cell Sci. 121, 3561–3569 (2008).
Bribian, A. et al. Role of the cellular prion protein in oligodendrocyte precursor cell proliferation and differentiation in the developing and adult mouse CNS. PLoS ONE 7, e33872 (2012).
Liu, J. et al. Impaired adult myelination in the prefrontal cortex of socially isolated mice. Nat. Neurosci. 15, 1621–1623 (2012).
Morel, E. et al. The cellular prion protein PrP(c) is involved in the proliferation of epithelial cells and in the distribution of junction-associated proteins. PLoS ONE 3, e3000 (2008).
Labonte, B. et al. Sex-specific transcriptional signatures in human depression. Nat. Med. 23, 1102–1111 (2017).
Dumais, A. et al. Risk factors for suicide completion in major depression: a case–control study of impulsive and aggressive behaviors in men. Am. J. Psychiatry 162, 2116–2124 (2005).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2017).
Benaglia, T., Chauveau, D., Hunter, D. R. & Young, D. S. mixtools: an R Package for analyzing mixture models. J. Stat. Soft. 32, 29 (2009).
Paradis, E., Claude, J. & Strimmer, K. APE: analyses of phylogenetics and evolution in R language. Bioinformatics 20, 289–290 (2004).
Hawrylycz, M. J. et al. An anatomically comprehensive atlas of the adult human brain transcriptome. Nature 489, 391–399 (2012).
Lake, B. B. et al. Neuronal subtypes and diversity revealed by single-nucleus RNA sequencing of the human brain. Science 352, 1586–1590 (2016).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Soft. 67, 48 (2015).
Kuznetsova, A., Brockhoff, P. B. & Christensen, R. H. B. lmerTest Package: tests in linear mixed effects models. J. Stat. Soft. 82, 26 (2017).
Lutz, P. E. et al. Association of a history of child abuse with impaired myelination in the anterior cingulate cortex: convergent epigenetic, transcriptional, and morphological evidence. Am. J. Psychiatry 174, 1185–1194 (2017).
Bayega, A. et al. in Gene Expression Analysis: Methods and Protocols (eds Raghavachari, N. & Garcia-Reyero, N.) 121–147 (Springer New York, 2018).
Spurgeon, S. L., Jones, R. C. & Ramakrishnan, R. High throughput gene expression measurement with real time PCR in a microfluidic dynamic array. PLoS ONE 3, e1662 (2008).
Ximerakis, M. et al. Single-cell transcriptomic profiling of the aging mouse brain. Nat. Neurosci. 22, 1696–1708 (2019).
Gong, T. & Szustakowski, J. D. DeconRNASeq: a statistical framework for deconvolution of heterogeneous tissue samples based on mRNA-Seq data. Bioinformatics 29, 1083–1085 (2013).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
G.T. holds a Canada Research Chair (Tier 1) and a NARSAD Distinguished Investigator Award. He is supported by grants from the Canadian Institute of Health Research (CIHR) (FDN148374 and EGM141899). We acknowledge the expert help of the Douglas–Bell Canada Brain Bank staff (J. Prud’homme, M. Bouchouka and A. Baccichet), and H. Djambazian at the MUGQIC. The work was also supported by CFI projects 32557 and 33408 (to J.R.). The Douglas–Bell Canada Brain Bank is supported in part by funding from the Canada First Research Excellence Fund, awarded to McGill University for the Healthy Brains for Healthy Lives project, and from the Fonds de recherche du Québec–Santé (FRQS) through the Quebec Network on Suicide, Mood Disorders and Related Disorders. The present study used the services of the Molecular and Cellular Microscopy Platform (MCMP) at the Douglas Institute.
Author information
Authors and Affiliations
Contributions
C.N. conceptualized, performed experiments and wrote the manuscript. M.M. performed experiments, bioinformatics and wrote the manuscript. A.T., V.Y., M.A.D. and Y.C.W. performed experiments and wrote the manuscript. M.S., K.P., J.-F.T., S.J.T. and P.P. contributed to data analyses and reviewed the manuscript. N.M. contributed to tissue processing, data interpretation and manuscript preparation. J.R. provided technical single-cell expertise and experimental support, and aided in manuscript preparation. G.T. provided general oversight of the study, including in experimental design, data interpretation and manuscript preparation.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Neuroscience thanks Ronald Duman (deceased), Matthew Girgenti, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Methods and Supplementary Figs. 1–14.
Supplementary Tables
Supplementary Tables 1–52.
Supplementary Software
Supplementart_R_Script_1.R for analyzing high-throughput qPCR data and Supplementary_R_Script_2.R for generating P values for CCInx interactions.
Rights and permissions
About this article
Cite this article
Nagy, C., Maitra, M., Tanti, A. et al. Single-nucleus transcriptomics of the prefrontal cortex in major depressive disorder implicates oligodendrocyte precursor cells and excitatory neurons. Nat Neurosci 23, 771–781 (2020). https://doi.org/10.1038/s41593-020-0621-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41593-020-0621-y
- Springer Nature America, Inc.
This article is cited by
-
Single-nucleus transcriptomic analysis reveals the relationship between gene expression in oligodendrocyte lineage and major depressive disorder
Journal of Translational Medicine (2024)
-
Single-cell RNA-seq reveals the role of YAP1 in prefrontal cortex microglia in depression
BMC Neurology (2024)
-
Temporal changes of gene expression in health, schizophrenia, bipolar disorder, and major depressive disorder
Schizophrenia (2024)
-
Shared and unique transcriptomic signatures of antidepressant and probiotics action in the mammalian brain
Molecular Psychiatry (2024)
-
Major depressive disorder: hypothesis, mechanism, prevention and treatment
Signal Transduction and Targeted Therapy (2024)