Abstract
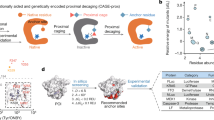
Cellular context is crucial for understanding the complex and dynamic kinase functions in health and disease. Systematic dissection of kinase-mediated cellular processes requires rapid and precise stimulation (‘pulse’) of a kinase of interest, as well as global and in-depth characterization (‘chase’) of the perturbed proteome under living conditions. Here we developed an optogenetic ‘pulse-chase’ strategy, termed decaging kinase coupled proteomics (DeKinomics), for proteome-wide profiling of kinase-driven phosphorylation at second-timescale in living cells. We took advantage of the ‘gain-of-function’ feature of DeKinomics to identify direct kinase substrates and further portrayed the global phosphorylation of understudied receptor tyrosine kinases under native cellular settings. DeKinomics offered a general activation-based strategy to study kinase functions with high specificity and temporal resolution under living conditions.






Similar content being viewed by others
Data availability
The mass spectrometry proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the iProX partner repository72,73 with the dataset identifier PXD039316. Source data are provided with this paper.
Code availability
The custom computer code has been deposited to Zenodo at https://doi.org/10.5281/zenodo.8365329.
References
Altelaar, A. F. M., Munoz, J. & Heck, A. J. R. Next-generation proteomics: towards an integrative view of proteome dynamics. Nat. Rev. Genet. 14, 35–48 (2013).
Hubbard, S. R. & Till, J. H. Protein tyrosine kinase structure and function. Annu. Rev. Biochem. 69, 373–398 (2000).
Karginov, A. V., Ding, F., Kota, P., Dokholyan, N. V. & Hahn, K. M. Engineered allosteric activation of kinases in living cells. Nat. Biotechnol. 28, 743–747 (2010).
Zhou, X. X., Fan, L. Z., Li, P., Shen, K. & Lin, M. Z. Optical control of cell signaling by single-chain photoswitchable kinases. Science 355, 836–842 (2017).
Li, Y., Zhang, Y., Li, X., Yi, S. & Xu, J. Gain-of-function mutations: an emerging advantage for cancer biology. Trends Biochem. Sci. 44, 659–674 (2019).
Zhang, G. et al. Bioorthogonal chemical activation of kinases in living systems. ACS Cent. Sci. 2, 325–331 (2016).
Wang, J. et al. Time-resolved protein activation by proximal decaging in living systems. Nature 569, 509–513 (2019).
Chin, J. W. Expanding and reprogramming the genetic code of cells and animals. Annu. Rev. Biochem. 83, 379–408 (2014).
Gautier, A. et al. Genetically encoded photocontrol of protein localization in mammalian cells. J. Am. Chem. Soc. 132, 4086–4088 (2010).
Arbely, E., Torres-Kolbus, J., Deiters, A. & Chin, J. W. Photocontrol of tyrosine phosphorylation in mammalian cells via genetic encoding of photocaged tyrosine. J. Am. Chem. Soc. 134, 11912–11915 (2012).
Walker, O. S. et al. Photoactivation of mutant isocitrate dehydrogenase 2 reveals rapid cancer-associated metabolic and epigenetic changes. J. Am. Chem. Soc. 138, 718–721 (2016).
Liaunardy-Jopeace, A., Murton, B. L., Mahesh, M., Chin, J. W. & James, J. R. Encoding optical control in LCK kinase to quantitatively investigate its activity in live cells. Nat. Struct. Mol. Biol. 24, 1155–1163 (2017).
Lake, D., Corrêa, S. A. L. & Müller, J. Negative feedback regulation of the ERK1/2 MAPK pathway. Cell. Mol. Life Sci. 73, 4397–4413 (2016).
Hanks, S. K. & Hunter, T. Protein kinases 6. The eukaryotic protein kinase superfamily: kinase (catalytic) domain structure and classification. FASEB J. 9, 576–596 (1995).
Wan, P. T. C. et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 116, 855–867 (2004).
Sharma, K. et al. Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell Rep. 8, 1583–1594 (2014).
Tan, C. S. H. et al. Thermal proximity coaggregation for system-wide profiling of protein complex dynamics in cells. Science 359, 1170–1177 (2018).
Olsen, J. V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648 (2006).
Kim, L. C., Song, L. & Haura, E. B. Src kinases as therapeutic targets for cancer. Nat. Rev. Clin. Oncol. 6, 587–595 (2009).
Bian, Y. et al. Ultra-deep tyrosine phosphoproteomics enabled by a phosphotyrosine superbinder. Nat. Chem. Biol. 12, 959–966 (2016).
Mariner, D. J. et al. Identification of Src phosphorylation sites in the catenin p120. J. Biol. Chem. 276, 28006–28013 (2001).
Songyang, Z. et al. Catalytic specificity of protein-tyrosine kinases is critical for selective signalling. Nature 373, 536–539 (1995).
Francavilla, C. et al. Multilayered proteomics reveals molecular switches dictating ligand-dependent EGFR trafficking. Nat. Struct. Mol. Biol. 23, 608–618 (2016).
Allen, J. J. et al. A semisynthetic epitope for kinase substrates. Nat. Methods 4, 511–516 (2007).
Embogama, D. M. & Pflum, M. K. H. K-BILDS: a kinase substrate discovery tool. Chem. Bio. Chem. 18, 136–141 (2017).
Ferrando, I. M. et al. Identification of targets of c-Src tyrosine kinase by chemical complementation and phosphoproteomics. Mol. Cell. Proteom. 11, 355–369 (2012).
Holt, L. J. et al. Global analysis of Cdk1 substrate phosphorylation sites provides insights into evolution. Science 325, 1682–1686 (2009).
Elsässer, S. J., Ernst, R. J., Walker, O. S. & Chin, J. W. Genetic code expansion in stable cell lines enables encoded chromatin modification. Nat. Methods 13, 158–164 (2016).
Barghout, S. H. & Schimmer, A. D. E1 enzymes as therapeutic targets in cancer. Pharmacol. Rev. 73, 1–56 (2021).
Hann, Z. S. et al. Structural basis for adenylation and thioester bond formation in the ubiquitin E1. Proc. Natl Acad. Sci. USA 116, 15475–15484 (2019).
He, X., Ma, B., Chen, Y., Guo, J. & Niu, W. Genetic encoding of a nonhydrolyzable phosphotyrosine analog in mammalian cells. Chem. Commun. 58, 5897–5900 (2022).
Lv, Z. et al. Domain alternation and active site remodeling are conserved structural features of ubiquitin E1. J. Biol. Chem. 292, 12089–12099 (2017).
Frangini, A. et al. The aurora B kinase and the polycomb protein Ring1B combine to regulate active promoters in quiescent lymphocytes. Mol. Cell 51, 647–661 (2013).
Zheng, N. & Shabek, N. Ubiquitin ligases: structure, function, and regulation. Annu. Rev. Biochem. 86, 129–157 (2017).
Mevissen, T. E. T. & Komander, D. Mechanisms of deubiquitinase specificity and regulation. Annu. Rev. Biochem. 86, 159–192 (2017).
Zhimin, L. & Tony, H. Degradation of activated protein kinases by ubiquitination. Annu. Rev. Biochem. 78, 435–475 (2009).
Pottier, C. et al. Tyrosine kinase inhibitors in cancer: breakthrough and challenges of targeted therapy. Cancers 12, 731 (2020).
Manning, G., Whyte, D. B., Martinez, R., Hunter, T. & Sudarsanam, S. The protein kinase complement of the human genome. Science 298, 1912–1934 (2002).
Oh, D.-Y. & Bang, Y.-J. HER2-targeted therapies—a role beyond breast cancer. Nat. Rev. Clin. Oncol. 17, 33–48 (2020).
Cocco, E., Scaltriti, M. & Drilon, A. NTRK fusion-positive cancers and TRK inhibitor therapy. Nat. Rev. Clin. Oncol. 15, 731–747 (2018).
Smart, S., Vasileiadi, E., Wang, X., DeRyckere, D. & Graham, D. The emerging role of TYRO3 as a therapeutic target in cancer. Cancers 10, 474 (2018).
Hsu, P.-L., Jou, J. & Tsai, S.-J. TYRO3: a potential therapeutic target in cancer. Exp. Biol. Med. 244, 83–99 (2019).
Graham, D. K., DeRyckere, D., Davies, K. D. & Earp, H. S. The TAM family: phosphatidylserine-sensing receptor tyrosine kinases gone awry in cancer. Nat. Rev. Cancer 14, 769–785 (2014).
Chien, C.-W. et al. Targeting TYRO3 inhibits epithelial–mesenchymal transition and increases drug sensitivity in colon cancer. Oncogene 35, 5872–5881 (2016).
Casado, P. et al. Kinase-substrate enrichment analysis provides insights into the heterogeneity of signaling pathway activation in leukemia cells. Sci. Signal. 6, rs6 (2013).
Murtuza, A. et al. Novel third-generation EGFR tyrosine kinase inhibitors and strategies to overcome therapeutic resistance in lung cancer. Cancer Res. 79, 689–698 (2019).
Otto, T. & Sicinski, P. Cell cycle proteins as promising targets in cancer therapy. Nat. Rev. Cancer 17, 93–115 (2017).
Ke, M. et al. Spatiotemporal profiling of cytosolic signaling complexes in living cells by selective proximity proteomics. Nat. Commun. 12, 71 (2021).
Lobingier, B. T. et al. An approach to spatiotemporally resolve protein interaction networks in living cells. Cell 169, 350–360.e12 (2017).
Paek, J. et al. Multidimensional tracking of GPCR signaling via peroxidase-catalyzed proximity labeling. Cell 169, 338–349.e11 (2017).
Levskaya, A., Weiner, O. D., Lim, W. A. & Voigt, C. A. Spatiotemporal control of cell signalling using a light-switchable protein interaction. Nature 461, 997–1001 (2009).
Songyang, Z. et al. Use of an oriented peptide library to determine the optimal substrates of protein kinases. Curr. Biol. CB 4, 973–982 (1994).
Tang, S. et al. Mechanism-based traps enable protease and hydrolase substrate discovery. Nature 602, 701–707 (2022).
Liu, Y. et al. Spatiotemporally resolved subcellular phosphoproteomics. Proc. Natl Acad. Sci. USA 118, e2025299118 (2021).
Shi, Y. et al. Targeting LIF-mediated paracrine interaction for pancreatic cancer therapy and monitoring. Nature 569, 131–135 (2019).
Li, W. et al. Tyrosine phosphorylation of protein kinase C-delta in response to its activation. J. Biol. Chem. 269, 2349–2352 (1994).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Chen, W., Chen, L. & Tian, R. An integrated strategy for highly sensitive phosphoproteome analysis from low micrograms of protein samples. Analyst 143, 3693–3701 (2018).
Yao, Y. et al. An immobilized titanium (IV) ion affinity chromatography adsorbent for solid phase extraction of phosphopeptides for phosphoproteome analysis. J. Chromatogr. A 1498, 22–28 (2017).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
R Core Team R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, 2021); https://www.r-project.org/
Kumar, L. & Futschik, M. E. Mfuzz: a software package for soft clustering of microarray data. Bioinformation 2, 5–7 (2007).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS J. Integr. Biol. 16, 284–287 (2012).
Carlson, M. org.Hs.eg.db: Genome wide annotation for Human. Bioconductor https://www.bioconductor.org/packages/org.Hs.eg.db/ (2019).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Colaert, N., Helsens, K., Martens, L., Vandekerckhove, J. & Gevaert, K. Improved visualization of protein consensus sequences by iceLogo. Nat. Methods 6, 786–787 (2009).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2016).
Swinton, J. Vennerable: Venn and Euler area-proportional diagrams. GitHub https://github.com/js229/Vennerable (2021).
Kolde, R. pheatmap: Pretty Heatmaps. CRAN https://cran.r-project.org/package=pheatmap (2019).
Deeb, S. J., D’Souza, R. C. J., Cox, J., Schmidt-Supprian, M. & Mann, M. Super-SILAC allows classification of diffuse large B-cell lymphoma subtypes by their protein expression profiles. Mol. Cell. Proteom. 11, 77–89 (2012).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740 (2016).
Chen, T. et al. iProX in 2021: connecting proteomics data sharing with big data. Nucleic Acids Res. 50, D1522–D1527 (2022).
Ma, J. et al. Iprox: an integrated proteome resource. Nucleic Acids Res. 47, D1211–D1217 (2019).
Gillen, C. M. & Forbush, B. Functional interaction of the K-Cl cotransporter (KCC1) with the Na-K-Cl cotransporter in HEK-293 cells. Am. J. Physiol. Cell Physiol. 276, C328–C336 (1999).
Acknowledgements
This work was supported by grant nos. 2021YFA1302603 (P.R.C.), 2021YFA1301601 (R.T.), 2021YFA1301602 (R.T.), 2021YFA1302603 (R.T.), 2022YFC3401104 (R.T.) and 2020YFE0202200 (R.T.) from the National Key Research and Development Program of China; grant nos. 21937001 (P.R.C.), 22137001 (P.R.C.), 91753000 (P.R.C.), 92253304 (R.T.) and 22125403 (R.T.) from the National Natural Science Foundation of China; grant no. 2019B151502050 (R.T.) from the Guangdong Provincial Fund for Distinguished Young Scholars; grant nos. JCYJ20200109141212325 (R.T.), JSGGZD20220822095200001 (R.T.) and JCYJ20210324120210029 (R.T.) from the Shenzhen Innovation of Science and Technology Commission; grant no. Z200010 from the Beijing Natural Science Foundation (P.R.C.); New Cornerstone Science Foundation through the New Cornerstone Investigator Program (P.R.C.) and the XPLORER PRIZE (P.R.C.). We thank J. Guo (University of Nebraska-Lincoln) for sharing the CMF-RS plasmid. We thank M. Pan and Q. Zheng (Protein Design and Synthesis Center, Institute of Translational Medicine, Shanghai Jiao Tong University) for providing recombinant E2 and Ub-FITC proteins. We thank Y. Sun and P. Bai (Southern University of Science and Technology) for providing primary PDAC cells.
Author information
Authors and Affiliations
Contributions
P.R.C. and R.T. conceived and supervised the study. Y.W. conceived all experiments and performed cell culture and treatment, kinase-ONPK expression and decaging, proteomics data analysis, immunofluorescence experiments, stable cell line constructions, ubiquitin thioester formation assays and other unspecified experiments. Y.W., W.C. and Q.K. performed pTyr proteomics and Flag IP-MS. Y.W., W.C., Y.M. and Y.Q. performed phosphoproteomics experiments. R.Z. and Y.L. synthesized ONPK. A.H. and M.K. performed LC–MS analysis of ONPK decaging rate. Y.W., R.W., W.S.C.N., H.Z. and J.W. performed CMF site-specific incorporation, protein purification and kinase phosphorylation assays. Y.W., R.T. and P.R.C. wrote the paper with inputs from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Chemical Biology thanks Kazuya Machida, Yonghao Yu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of ONPK decaging in vitro and in living cells.
a, Liquid chromatography-mass spectrometry (LC-MS) analysis of remaining ONPK with UV irradiation. Error bars represent mean ± SD (n = 3 independent experiments). b, The MS/MS spectrum of the ONP caged peptide (residue 295–318 of SRC) with neutral loss of ONP group. ONP, ONP modification; OX, oxidation modification.
Extended Data Fig. 2 Validation of DeKinomics’ temporal resolution in phosphoproteome.
a, The numbers of identified phosphoproteins, phosphopeptides and phosphosites in DeKinomics study of BRAF with phosphorylated residue numbers from different classes according to localization probability and distribution of phosphorylated amino acids. b, Quantitative phosphoproteomics revealed the changes of phosphosites in MAPK signaling pathway upon BRAF activation. c, The KEGG pathway enrichment analysis of phosphosites with the Euclidean distance among the smallest 10%. According to the gene ratio, the top 10 pathways were depicted. Two-sided hypergeometric test followed by Benjamini-Hochberg correlation. KEGG, Kyoto Encyclopedia of Genes and Genomes. d, Dynamics of MAPK signaling pathway phosphosites that increased upon BRAF activation were showed in detail. e, Validation of phosphorylation dynamics in MAPK signaling pathway in repetitious experiments. Error bars represent mean ± SD (n = 3 independent experiments).
Extended Data Fig. 3 SRC-driven pTyr proteome dynamics were deeply profiled by DeKinomics.
a, Western blotting validation of rapid activation of tyrosine kinase SRC (n = 2). b, The numbers of identified pTyr proteins, peptides and phosphosites in the SRC study, with the pTyr residue number from different classes according to localization probability. c, Schematic illustration of the time-resolved pTyr proteomics upon UV irradiation as a control experiment. d, Bar graph comparing the unchanged and the significantly changed pTyr sites in UV irradiation experiment. e, Violin plot indicating the increased global pTyr intensity upon SRC activation. The counts of quantified pTyr sites (n) in each time point are indicated in brackets. Box plots show median (center line), the upper and lower quantiles (box), and the minima and maxima of the data (whiskers). f, The Pearson correlation of pTyr intensity between the biological replicates. g, GO enrichment analysis of pTyr sites from different clusters.
Extended Data Fig. 4 Validating SRC substrates with monoclonal stable cell line expressing SRC-ONPK.
a, Volcano plots indicating the increased pTyr sites within 6 s upon SRC activation. n = 3 independent experiments; two-sided Student’s t-test. b, Venn diagram indicating that the SRC substrate candidates were determined with two criteria. c, pPiggyBac plasmid used to generate the HEK293T stable cell line that expressing ONPK-incorporated SRC protein. IRES, internal ribosome entry sites; NeoR, Neomycine Resistance; PuroR, Puromycine Resistance; copGFP, green fluorescent protein from copepod Pontellina plumate. d, Western blotting analysis indicating that the expression level of SRC-ONPK could be tuned by ONPK concentration (n = 2). e, Confocal immunofluorescence imaging confirming same SRC localization between exogenous and endogenous SRC. Scale bars: 5 μm (n = 3). f, Schematic illustration of PRM validation for physiological relevance between SRC and downstream pTyr sites (n = 3). g, Histogram indicating pTyr sites with i) the max intensity of their corresponding precursor, ii) Evidence count > 15 would be conduct to PRM validation. The filters including precursor charge < 4, random selection to shorten the list, manual checking for duplication transition were also applied for determining the final list for PRM assay. h, Veen diagram indicating the number of pTyr sites that were studied, quantified and validated in PRM assay respectively. i, Schematic illustration of biological functions of the validated SRC substrates by PRM.
Extended Data Fig. 5 Control experiments for in vitro kinase phosphorylation assay.
a, The extracted ion chromatograph (XIC) of the fragment ions of ERK1 pY204 (precursor: VpYENVGLMQQQK++) in compare with PTPN11 pY584 (precursor: IADPEHDHTGFLTEpYVATR++). b, Western blotting estimating the SRC concentration in HEK293T cells (n = 2). Loaded protein amount of #1, #2 and #3 HEK293T lysate were 5.2 μg, 6.0 μg and 6.8 μg respectively. Western blotting bands were quantified by image studio software (LI-COR, v5.2). We took 6.5 μL mg−1 as an estimation for the cell volume of the HEK293T cells74.
Extended Data Fig. 6 SRC phosphorylated UBA1 and suppressed its enzymatic activity.
a, Dot plot indicating the number of reference reporting post-translational modification (PTM) sites identification on UBA1 in PhosphoSitePlus database (v6.6.0.4). b, Schematic illustration of modified thioester formation assay for measuring the UBA1’s activity. c, Sequence alignment of region where human UBA1 S140 and Y141 (highlighted) located. Asterisks indicate the conserved residues. d, e, Quantitative fluorescence measurements of E2-Ub-FITC thioester formation assay. Error bars represent mean ± SD (n = 3 independent experiments).
Extended Data Fig. 7 DeKinomics applications on oncogenic RTK variants.
a, Sequence alignment for locating conserved catalytic lysine residue in HER2 and NTRK1. Asterisks indicate the conserved residues. b-c, Schematic illustration of DeKinomics study workflow of HER2-L755S and TPM3-NTRK1 (n = 3). d, Volcano plots indicating the successful profiling of downstream phosphorylation of HER2-L755S. n = 3 independent experiments; two-sided Student’s t-test. e, Fusion protein TPM3-NTRK1 could be rapidly activated and induce downstream Tyr phosphorylation (n = 3 independent experiments). f, Heatmap of pTyr relative abundance indicating autophosphorylation and Tyr phosphorylation on signal transduction protein induced by TPM3-NTRK1 activation.
Extended Data Fig. 8 DeKinomics study of TYRO3.
a, Sequence alignment for locating conserved catalytic lysine residue in TYRO3. Asterisks indicate the conserved residues. b, Confocal immunofluorescence imaging of exogenous TYRO3-K550ONPK (n = 3). Scale bars: 5 μm. c, Heatmap indicating TYRO3 activation induced TYRO3 autophosphorylation. d, The fuzzy c-means clustering of significantly changed pTyr sites upon TYRO3 activation. e, Venn diagram indicating that TYRO3 candidate substrates were determined with 2 criteria: pTyr sites significantly changed in 0–60 sec (ANOVA) and were in cluster 1 with membership > 0.7 in fuzzy c-means clustering.
Extended Data Fig. 9 DeKinomics-depicted TYRO3 function.
a, Volcano plot comparisons between TYRO3-activated cells and control cells. n = 3 independent experiments; two-sided Student’s t-test. b, The fuzzy c-means clustering of significantly changed phosphosites in phosphoproteome. c, An overview of early phosphorylation cascades of TYRO3. The annotations of protein function were adopted from Uniprot database.
Extended Data Fig. 10 Targeting TYRO3-PRKCD axis.
a, Cell proliferation assay to evaluate the effect of inhibitor combination. Primary PDAC cells were isolated from the KPf/fC (KrasLSL-G12D/+;Trp53flox/flox;Pdx1-Cre) PDAC mouse model55. Error bars represent mean ± SD (n = 4 independent experiments); two-sided Student’s t-test. b, Schematic illustration of dual-targeting of the TYRO3-PRKCD signaling axis by inhibitors.
Supplementary information
Supplementary Information
Supplementary Figs. 1–6.
Supplementary Table 1
DeKinomics study of BRAF.
Supplementary Table 2
DeKinomics study of SRC.
Supplementary Table 3
SRC substrate candidates.
Supplementary Table 4
Results of the SRC substrates validation experiment.
Supplementary Table 5
DeKinomics study of HER2 and TPM3-NTRK1.
Supplementary Table 6
TYRO3 study in living cells with DeKinomics.
Supplementary Table 7
KSEA of TYRO3-regulated pSer/pThr sites.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Fig. 1
Unprocessed western blots/gels.
Source Data Fig. 3
Unprocessed western blots/gels.
Source Data Fig. 4
Unprocessed western blots/gels.
Source Data Fig. 5
Unprocessed western blots/gels.
Source Data Extended Data Fig. 3
Unprocessed western blots/gels.
Source Data Extended Data Fig. 4
Unprocessed western blots/gels.
Source Data Extended Data Fig. 5
Unprocessed western blots/gels.
Source Data Extended Data Fig. 6
Unprocessed western blots/gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Weng, Y., Chen, W., Kong, Q. et al. DeKinomics pulse-chases kinase functions in living cells. Nat Chem Biol 20, 615–623 (2024). https://doi.org/10.1038/s41589-023-01497-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-023-01497-x
- Springer Nature America, Inc.
This article is cited by
-
Lighting up kinase contacts in situ
Nature Chemical Biology (2024)