Abstract
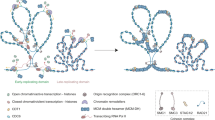
Recent studies have implicated the nucleosome acidic patch in the activity of ATP-dependent chromatin remodeling machines. We used a photocrosslinking-based nucleosome profiling technology (photoscanning) to identify a conserved basic motif within the catalytic subunit of ISWI remodelers, SNF2h, which engages this nucleosomal epitope. This region of SNF2h is essential for chromatin remodeling activity in a reconstituted biochemical system and in cells. Our studies suggest that the basic motif in SNF2h plays a critical role in anchoring the remodeler to the nucleosomal surface. We also examine the functional consequences of several cancer-associated histone mutations that map to the nucleosome acidic patch. Kinetic studies using physiologically relevant heterotypic nucleosomal substrates (‘Janus’ nucleosomes) indicate that these cancer-associated mutations can disrupt regularly spaced chromatin structure by inducing ISWI-mediated unidirectional nucleosome sliding. These results indicate a potential mechanistic link between oncogenic histones and alterations to the chromatin landscape.





Similar content being viewed by others
Data availability
The authors declare that data supporting the finding of this study are available within the article and its Supplementary Information. Additional data are available from the corresponding author upon reasonable request.
References
Jiang, C. & Pugh, B. F. Nucleosome positioning and gene regulation: advances through genomics. Nat. Rev. Genet. 10, 161–172 (2009).
Clapier, C. R. & Cairns, B. R. The biology of chromatin remodeling complexes. Annu. Rev. Biochem. 78, 273–304 (2009).
Becker, P. B. & Workman, J. L. Nucleosome remodeling and epigenetics. Cold Spring Harb. Perspect. Biol. 5, a017905–a017905 (2013).
Bartholomew, B. Regulating the chromatin landscape: structural and mechanistic perspectives. Annu. Rev. Biochem. 83, 671–696 (2014).
Clapier, C. R., Iwasa, J., Cairns, B. R. & Peterson, C. L. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat. Rev. Mol. Cell Biol. 18, 407–422 (2017).
Tsukiyama, T., Becker, P. B. & Wu, C. ATP-dependent nucleosome disruption at a heat-shock promoter mediated by binding of GAGA transcription factor. Nature 367, 525–532 (1994).
Pazin, M. J., Kamakaka, R. T. & Kadonaga, J. T. ATP-dependent nucleosome reconfiguration and transcriptional activation from preassembled chromatin templates. Science 266, 2007–2011 (1994).
Deuring, R. et al. The ISWI chromatin-remodeling protein is required for gene expression and the maintenance of higher order chromatin structure in vivo. Mol. Cell 5, 355–365 (2000).
Poot, R. A. et al. HuCHRAC, a human ISWI chromatin remodelling complex contains hACF1 and two novel histone-fold proteins. EMBO J. 19, 3377–3387 (2000).
Stopka, T. & Skoultchi, A. I. The ISWI ATPase Snf2h is required for early mouse development. Proc. Natl Acad. Sci. USA 100, 14097–14102 (2003).
He, X., Fan, H.-Y., Narlikar, G. J. & Kingston, R. E. Human ACF1 alters the remodeling strategy of SNF2h. J. Biol. Chem. 281, 28636–28647 (2006).
Clapier, C. R., Längst, G., Corona, D. F., Becker, P. B. & Nightingale, K. P. Critical role for the histone H4 N terminus in nucleosome remodeling by ISWI. Mol. Cell. Biol. 21, 875–883 (2001).
Clapier, C. R. & Cairns, B. R. Regulation of ISWI involves inhibitory modules antagonized by nucleosomal epitopes. Nature 492, 280–284 (2012).
Hamiche, A., Kang, J. G., Dennis, C., Xiao, H. & Wu, C. Histone tails modulate nucleosome mobility and regulate ATP-dependent nucleosome sliding by NURF. Proc. Natl Acad. Sci. USA 98, 14316–14321 (2001).
Dang, W., Kagalwala, M. N. & Bartholomew, B. Regulation of ISW2 by concerted action of histone H4 tail and extranucleosomal DNA. Mol. Cell. Biol. 26, 7388–7396 (2006).
Yang, J. G., Madrid, T. S., Sevastopoulos, E. & Narlikar, G. J. The chromatin-remodeling enzyme ACF is an ATP-dependent DNA length sensor that regulates nucleosome spacing. Nat. Struct. Mol. Biol. 13, 1078–1083 (2006).
Deindl, S. et al. ISWI remodelers slide nucleosomes with coordinated multi-base-pair entry steps and single-base-pair exit steps. Cell 152, 442–452 (2013).
Blosser, T. R., Yang, J. G., Stone, M. D., Narlikar, G. J. & Zhuang, X. Dynamics of nucleosome remodelling by individual ACF complexes. Nature 462, 1022–1027 (2009).
Racki, L. R. et al. The chromatin remodeller ACF acts as a dimeric motor to space nucleosomes. Nature 462, 1016–1021 (2009).
Goldman, J. A., Garlick, J. D. & Kingston, R. E. Chromatin remodeling by imitation switch (ISWI) Class ATP-dependent remodelers is stimulated by histone variant H2A.Z. J. Biol. Chem. 285, 4645–4651 (2010).
Dann, G. P. et al. ISWI chromatin remodellers sense nucleosome modifications to determine substrate preference. Nature 548, 607–611 (2017).
Gamarra, N., Johnson, S. L., Trnka, M. J., Burlingame, A. L. & Narlikar, G. J. The nucleosomal acidic patch relieves auto-inhibition by the ISWI remodeler SNF2h. eLife 7, 34270 (2018).
Kalashnikova, A. A., Porter-Goff, M. E., Muthurajan, U. M., Luger, K. & Hansen, J. C. The role of the nucleosome acidic patch in modulating higher order chromatin structure. J. R. Soc. Interface 10, 20121022 (2013).
McGinty, R. K. & Tan, S. Nucleosome structure and function. Chem. Rev. 115, 2255–2273 (2014).
Barbera, A. J. et al. The nucleosomal surface as a docking station for Kaposi’s sarcoma herpesvirus LANA. Science 311, 856–861 (2006).
Makde, R. D., England, J. R., Yennawar, H. P. & Tan, S. Structure of RCC1 chromatin factor bound to the nucleosome core particle. Nature 467, 562–566 (2010).
Armache, K.-J., Garlick, J. D., Canzio, D., Narlikar, G. J. & Kingston, R. E. Structural basis of silencing: Sir3 BAH domain in complex with a nucleosome at 3.0 Å resolution. Science 334, 977–982 (2011).
Kato, H. et al. A conserved mechanism for centromeric nucleosome recognition by centromere protein CENP-C. Science 340, 1110–1113 (2013).
McGinty, R. K., Henrici, R. C. & Tan, S. Crystal structure of the PRC1 ubiquitylation module bound to the nucleosome. Nature 514, 591–596 (2014).
Morgan, M. T. et al. Structural basis for histone H2B deubiquitination by the SAGA DUB module. Science 351, 725–728 (2016).
Götze, M. et al. StavroX—A software for analyzing crosslinked products in protein interaction studies. J. Am. Soc. Mass Spectrom. 23, 76–87 (2011).
Yang, B. et al. Identification of cross-linked peptides from complex samples. Nat. Methods 9, 904–906 (2012).
Ludwigsen, J., Klinker, H. & Planitz, F. M. No need for a power stroke in ISWI‐mediated nucleosome sliding. EMBO Rep. 14, 1092–1097 (2013).
Grüne, T. et al. Crystal structure and functional analysis of a nucleosome recognition module of the remodeling factor ISWI. Mol. Cell 12, 449–460 (2003).
Hwang, W. L., Deindl, S., Harada, B. T. & Zhuang, X. Histone H4 tail mediates allosteric regulation of nucleosome remodelling by linker DNA. Nature 512, 213–217 (2014).
Tsukiyama, T., Palmer, J., Landel, C. C., Shiloach, J. & Wu, C. Characterization of the imitation switch subfamily of ATP-dependent chromatin-remodeling factors in Saccharomyces cerevisiae. Genes Dev. 13, 686–697 (1999).
Zhao, S. et al. Mutational landscape of uterine and ovarian carcinosarcomas implicates histone genes in epithelial-mesenchymal transition. Proc. Natl Acad. Sci. USA 113, 12238–12243 (2016).
Arimura, Y. et al. Cancer-associated mutations of histones H2B, H3.1 and H2A.Z.1 affect the structure and stability of the nucleosome. Nucleic Acids Res. 46, 10007–10018 (2018).
Nacev, B. A. et al. The expanding landscape of ‘oncohistone’ mutations in human cancers. Nature 567, 473–478 (2019).
Alabert, C., Jasencakova, Z. & Groth, A. in DNA Replication. Advances in Experimental Medicine and Biology Vol. 1042 (eds Masai, H. & Foiani, M.) 311-333 (Springer, 2017).
Levendosky, R. F., Sabantsev, A., Deindl, S. & Bowman, G. D. The Chd1 chromatin remodeler shifts hexasomes unidirectionally. eLife 5, 3302 (2016).
Hota, S. K. et al. Nucleosome mobilization by ISW2 requires the concerted action of the ATPase and SLIDE domains. Nat. Struct. Mol. Biol. 20, 222–229 (2013).
Sabantsev, A., Levendosky, R. F., Zhuang, X., Bowman, G. D. & Deindl, S. Direct observation of coordinated DNA movements on the nucleosome during chromatin remodelling. Nat. Commun. 10, 1720 (2019).
Yan, L., Wu, H., Li, X., Gao, N. & Chen, Z. Structures of the ISWI-nucleosome complex reveal a conserved mechanism of chromatin remodeling. Nat. Struct. Mol. Biol. 26, 258–266 (2019).
Lewis, P. W. et al. Inhibition of PRC2 activity by a gain-of-function H3 mutation found in pediatric glioblastoma. Science 340, 857–861 (2013).
Lu, C. et al. Histone H3K36 mutations promote sarcomagenesis through altered histone methylation landscape. Science 352, 844–849 (2016).
Levendosky, R. F. & Bowman, G. D. Asymmetry between the two acidic patches dictates the direction of nucleosome sliding by the ISWI chromatin remodeler. eLife 8, e45472 (2019).
Voigt, P. et al. Asymmetrically modified nucleosomes. Cell 151, 181–193 (2012).
Rhee, H. S., Bataille, A. R., Zhang, L. & Pugh, B. F. Subnucleosomal structures and nucleosome asymmetry across a genome. Cell 159, 1377–1388 (2014).
Nekrasov, M. et al. Histone H2A.Z inheritance during the cell cycle and its impact on promoter organization and dynamics. Nat. Struct. Mol. Biol. 19, 1076–1083 (2012).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539 (2011).
Ikeda, Y. & Behrman, E. J. Improved synthesis of photo-leucine. Syn. Commun. 38, 2276–2284 (2008).
Debelouchina, G. T., Gerecht, K. & Muir, T. W. Ubiquitin utilizes an acidic surface patch to alter chromatin structure. Nat. Chem. Biol. 13, 105–110 (2017).
Luger, K., Rechsteiner, T. J., Flaus, A. J., Waye, M. M. & Richmond, T. J. Characterization of nucleosome core particles containing histone proteins made in bacteria. J. Mol. Biol. 272, 301–311 (1997).
Leonard, J. D. & Narlikar, G. J. A nucleotide-driven switch regulates flanking DNA length sensing by a dimeric chromatin remodeler. Mol. Cell 57, 850–859 (2015).
Zhou, C. Y. & Narlikar, G. J. Analysis of nucleosome sliding by ATP-dependent chromatin remodeling enzymes. Meth. Enzymol. 573, 119–135 (2016).
Brachmann, C. B. et al. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14, 115–132 (1998).
Acknowledgements
We thank R. E. Thompson, K. Diehl, J. D. Bagert, K. Jani and other members of the Muir laboratory for valuable discussions. We thank S. Tharan from the Proteomics and Mass Spectrometry core. Yeast strains (YTT 227, YTT 420-424) were a gift from T. Tsukiyama. H.T.D. was funded by a postdoctoral fellowship from the Jane Coffin Childs Memorial fund. This work was supported by the US National Institutes of Health (NIH grant nos. R37-GM086868 and P01-CA196539).
Author information
Authors and Affiliations
Contributions
H.T.D. synthesized the probe and all photoactivable nucleosomes, heterotypic nucleosomes, dinucleosomes, expressed and purified all remodelers, and performed all biochemical experiments. H.T.D. and B.E.D. performed yeast experiments. G.P.D. expressed and purified the Sir3-BAH domain, and assisted with the expression and purification of the ACF complex. G.P.L. expressed and purified RCC1 and mRCC1. H.T.D. and T.W.M. analyzed all data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Tables 3 and 4, Figs. 1–39 and Notes 1 and 2
Rights and permissions
About this article
Cite this article
Dao, H.T., Dul, B.E., Dann, G.P. et al. A basic motif anchoring ISWI to nucleosome acidic patch regulates nucleosome spacing. Nat Chem Biol 16, 134–142 (2020). https://doi.org/10.1038/s41589-019-0413-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-019-0413-4
- Springer Nature America, Inc.
This article is cited by
-
Synovial sarcoma X breakpoint 1 protein uses a cryptic groove to selectively recognize H2AK119Ub nucleosomes
Nature Structural & Molecular Biology (2024)
-
Structure of the ISW1a complex bound to the dinucleosome
Nature Structural & Molecular Biology (2024)
-
Tracking chromatin state changes using nanoscale photo-proximity labelling
Nature (2023)
-
Nucleosome recognition and DNA distortion by the Chd1 remodeler in a nucleotide-free state
Nature Structural & Molecular Biology (2022)
-
The role of auxiliary domains in modulating CHD4 activity suggests mechanistic commonality between enzyme families
Nature Communications (2022)