Abstract
The human placenta has a vital role in ensuring a successful pregnancy. Despite the growing body of knowledge about its cellular compositions and functions, there has been limited research on the heterogeneity of the billions of nuclei within the syncytiotrophoblast (STB), a multinucleated entity primarily responsible for placental function. Here we conducted integrated single-nucleus RNA sequencing and single-nucleus ATAC sequencing analyses of human placentas from early and late pregnancy. Our findings demonstrate the dynamic heterogeneity and developmental trajectories of STB nuclei and their correspondence with human trophoblast stem cell (hTSC)-derived STB. Furthermore, we identified transcription factors associated with diverse STB nuclear lineages through their gene regulatory networks and experimentally confirmed their function in hTSC and trophoblast organoid-derived STBs. Together, our data provide insights into the heterogeneity of human STB and represent a valuable resource for interpreting associated pregnancy complications.
Similar content being viewed by others
Main
The placenta, which is an integral part of the maternal–fetal interface, serves the critical function of supplying nutrition and oxygen to the developing fetus and removes waste materials and carbon dioxide. It is a transient organ that responds to developmental cues and environmental stimuli during pregnancy1,2,3. The human placenta develops during implantation when trophoblasts differentiate from the trophectoderm (TE) of the blastocyst4,5. Following implantation, the TE gives rise to the following three critical trophoblast cell types: cytotrophoblast cells (CTBs), extravillous trophoblast cells (EVTs) and multinucleated syncytiotrophoblast (STB). CTBs can act as progenitor cells that differentiate into EVTs or fuse to the multinucleated STB6. The STB is in direct contact with the maternal tissue and blood, carrying out various functions such as hormone synthesis, immunological defense and active transport during pregnancy5, and thus has an essential role in maintaining a successful pregnancy7,8,9. Insufficient placental development can lead to conditions such as preeclampsia and intrauterine growth restriction10,11,12,13.
The development of the multinucleated STB involves complex structural arrangements and functionalization14. The process of multinucleation in the STB is crucial for protecting the placenta against pathogens. The absence of multinucleated STB increases susceptibility to infections at the maternal–fetal barrier. As pregnancy progresses, aged nuclei known as syncytial knots are gradually replaced by the CTB nuclei underneath15,16,17,18. At term, the number of STB nuclei reaches ~58 billion19,20. Previous histological studies suggested the existence of subpopulations of nuclei within the STB21,22,23,24. However, whether the STB nuclei exhibit development-dependent transcriptional and functional dynamics remains under-appreciated.
Our knowledge of the cell-type-specific transcriptional landscape in the human placenta has been reinterpreted in light of single-cell RNA-sequencing (scRNA-seq) data25,26,27,28,29,30,31. Past scRNA-seq studies have described the transcriptional landscape of the placenta, such as nonproliferative EVTs27. However, scRNA-seq has not been applied to STB due to its multinucleated nature, and furthermore, isolating single STB nuclear is difficult, leaving fundamental questions about the heterogeneity of the STB nuclei, gene expression and regulators largely unanswered32. The utilization of single-nucleus RNA sequence (snRNA-seq) has transformed our ability to identify various nuclear types, states and genetic diversity within the shared cytoplasm of tissues like muscle, as well as other hard-to-dissociated samples like brain. More recently, snRNA-seq has been used to investigate placental samples during early pregnancy33.
Here we constructed a comprehensive map of nuclear identities in the human STB during early and late pregnancy by integrating snRNA-seq and single-nucleus ATAC sequencing (snATAC-seq) on 101,543 placental nuclei. Our study reveals the dynamic heterogeneity of the STB nuclei, their differentiation routes and their potential roles during early and late pregnancy. We identified master transcription factors (TFs) that are biased toward diverse STB nuclear lineages, including MITF, STAT5A, CEBPB and FOSL2. We described their key gene regulatory networks at different stages of pregnancy. We confirmed the presence of STB nuclear subtypes in human trophoblast stem cell (hTSC)-derived STB. Our results indicated that the critical functions of master TFs were partially retained in in vitro models of STB differentiated from hTSCs and trophoblast organoids. Taken together, our work provides a comprehensive analysis of STB nuclear heterogeneity, highlights new regulatory mechanisms and represents a valuable resource for interpreting associated pregnancy complications.
Results
snRNA-seq and snATAC-seq profiling of human placentas revealed dynamic features and heterogeneity of STB nuclei in early and late pregnancy
To comprehensively understand the human placental nuclear repertoire and its regulatory mechanisms, we applied 10x Genomics snRNA-seq and snATAC-seq techniques to nuclei isolated from 12 healthy placentas, six each in early (6–9 weeks of gestation) and late pregnancy (38–39 weeks of gestation). We took measures to eliminate potential sex bias by collecting an equal number of placentas from each sex before sequencing (Supplementary Fig. 1a,c). Detailed information on the sample collection is presented in Supplementary Table 1 and Supplementary Note 1. A graphical overview of the methodology is presented in Fig. 1a.
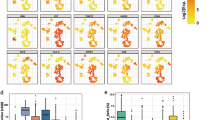
a, Schematic representation of the samples used in this study, nuclear isolation, sequencing experiments, downstream bioinformatic analyses and experimental validation. b, UMAP embeddings of snRNA-seq nuclei where dots correspond to individual nuclei. c, Annotation of the snRNA-seq clusters based on stage information and marker gene expression project on UMAP embeddings. The relative expression levels are presented with color intensities. Gene expression raw counts were normalized by depth, logarithmized and z score scaled, and finally smoothed with imputation. d, Scatter plot shows DEGs in the STB nuclei in early and late pregnancy. DEGs are identified with the Wilcoxon test (Padj ≤ 0.01, log2(FC) ≥ 0.25). Top 20 genes are highlighted and labeled. e. GO enrichment associated with DEGs for the STB nuclei in early and late pregnancy. GO enrichment analysis is tested by one-sided Fisher’s exact test with a default P < 0.1. f. UMAP embeddings of snATAC-seq nuclei where dots correspond to individual nuclei. Abbreviation rules as in b. g, snATAC-seq genomic tracks along denoting chromatin accessibility peaks of marker genes for each cluster. Gray boxes highlight specific accessible regions associated with marker genes. h, TFs overlay on snATAC-seq UMAP (TF activity scores (top) and gene-activity scores (middle)) and snRNA-seq (gene expression level (bottom)) for TF genes, FOSL2, SREBF1, STAT5A and TWIST. The TF activity, gene activity and relative expression levels are presented with color intensities. i. The integrated UAMP plot of STB nuclei from both the snRNA-seq data and the snATAC-seq data. Abbreviation rules as in b. eSTB unintegrated, unintegrated snRNA-seq data and the snATAC dataset from eSTB nuclei in early pregnancy; lSTB unintegrated, unintegrated snRNA-seq data and the snATAC dataset from eSTB nuclei in late pregnancy.
After stringent filtering, we retained 50,850 nuclei and 50,693 nuclei for snRNA-seq and snATAC-seq, respectively, for subsequent analyses (Supplementary Table 1; Methods). Sample preparation batch correction methods were applied to both datasets, resulting in strong correlation (Supplementary Fig. 1d) and high-quality data (Supplementary Fig. 1e,f). Batch-corrected transcriptomic and epigenomic datasets were clustered using the Scanpy34 and SnapATAC v2 (ref. 35) package accordingly and visualized using uniform manifold approximation and projection (UMAP) dimension reduction (Fig. 1b,f). Seven major nuclear clusters were annotated using combined pregnancy stage information and canonical markers, identifying early and late populations of CTB (eCTB, lCTB), STB (eSTB, lSTB) and stromal cells (eSTR, lSTR), as well as an early EVT (eEVT)—e for early pregnancy and l for late pregnancy (Fig. 1c, Supplementary Fig. 1g–i and Supplementary Note 2). The ratio of spliced and unspliced RNA was relatively similar among those annotated nuclear types, indicating that nuclei in the STB have comparable transcriptional capabilities to other cell types (Supplementary Fig. 1b and Supplementary Note 3). Interestingly, CTB, EVT and STR nuclei at different gestational stages clustered closely together, while STB nuclear clusters exhibited distinct differences between early and late pregnancy, suggesting greater functional disparities (Fig. 1b).
Differentially expressed genes (DEGs) and gene ontology (GO) analysis were conducted on the STB nuclei during early and late pregnancy to identify functional differences (Fig. 1d,e, Supplementary Table 2 and Supplementary Note 4). We found the enrichment of GO terms in the early pregnancy-related STB DEGs included cell migration, response to hypoxia and positive regulation of angiogenesis. Those functions are crucial for the proper function of the human placenta in a physiological low-oxygen environment in early pregnancy before the maternal blood flows into the intervillous space2,14. In late pregnancy, enrichment of GO terms included positive regulation of GTPase activity, viral protein processing and intracellular signal transduction. These GO functions together with others might reflect the need for transportation, resistance against viral infection and maximum nutrient requirements for successful fetal delivery.
We next sought to determine whether dynamic STB gene expression in early and late pregnancy correlated with accessible chromatin sites. We analyzed the snATAC-seq data and detected consistent nuclear types based on gene-activity scores (a quantitative measure specific to snATAC-seq data that signifies the extent of gene accessibility) of marker genes from our snRNA-seq analysis, including eCTB, lCTB, eSTB, lSTB, eEVT, eSTR and lSTR (Fig. 1f). Cluster annotation based on gene-activity score was validated by aggregated snATAC-seq tracks around marker genes for each cluster (Fig. 1g and Supplementary Fig. 1j). Notably, the UMAP analysis revealed distinct distributions of STB clusters in early and late pregnancy, a pattern reminiscent of the findings obtained from snRNA-seq analysis. These results indicate a consistent gene expression and chromatin accessibility in placental nuclei in early and late pregnancy, reflecting varied regulatory mechanisms and functions in the STB at different gestational stages.
To identify potential TFs that may regulate dynamic STB nuclear gene expression and functions during different gestation stages, we used chromVAR to identify enriched TF binding motifs from the JASPAR 2020 database that were present within the accessible chromatin36. Motif enrichment analysis revealed gestation stage-specific enrichment of binding motif on several TFs, including FOSL2, SREBF1, HES2, CEBPA, HIF1A and MITF in early pregnancy and STAT5A, TWIST, STAT6, GCM1, RORA and NR3C2 in late pregnancy (Fig. 1h and Supplementary Fig. 1k). We further demonstrated concordance by direct transcript measurement from the snRNA-seq dataset, inferred gene-activity scores representing the degree of gene accessibility from the snATAC-seq dataset and TF activity using chromVAR, which identified enriched binding motifs of key TFs on the UMAP (Fig. 1h and Supplementary Fig. 1k). In addition, we observed specific enrichment of the HIF1A binding motif in STB during early pregnancy, suggesting potential responsiveness to HIF1A. This could be in response to the normal low-oxygen concentration in the intervillous space in the early placenta, but HIF1A can also be activated by growth-promoting kinases37,38. It is worth mentioning that oxygen levels in the early placenta, although relatively low compared to those in the later stages of pregnancy, still fall within the normal range. We also identified STAT6 and STAT5A as key TFs in STB in late pregnancy. STAT signaling pathway was reported to be involved in regulating PAPPA, a serum marker used in screening for developmental abnormalities, in vitro39.
The correlation between chromatin accessibility and downstream gene expression signatures emphasizes the potential of integrating snATAC-seq with snRNA-seq data. To accomplish this, we combined several recently developed methods, which are as follows: (1) we removed the batch effects using Harmony40; (2) we analyzed our snATAC-seq datasets in early and late pregnancy using SnapATAC v2 (ref. 35), and snRNA-seq datasets were analyzed through Scanpy34; and (3) the final merged snRNA-seq and snATAC-seq data were further integrated using liger (Supplementary Fig. 2a)41. Our analysis revealed that nuclear clusters classified based on chromatin data and transcriptome data were grouped together in the integrated UMAP space (Fig. 1i, Supplementary Fig. 2b–d and Supplementary Note 5).
Nuclear heterogeneity and a differentiation trajectory bifurcation in the human placental STB in early pregnancy
We initially observed the segregation of STB nuclei into subclusters using snRNA-seq, snATAC-seq and integrated UMAP (Fig. 1b,f,i). To investigate the details of nuclear heterogeneity within the STB during early pregnancy, we conducted a focused analysis of 23,702 nuclei from the eSTB and its progenitors, the eCTB, as shown in Fig. 1b. We identified ten subclusters based on snRNA-seq datasets with four eCTB subclusters (clusters 7, 9, 6 and 11, DNMT1, in early pregnancy) and six eSTB subclusters (clusters 8, 2, 10, 1, 3, 4 and 5, PSG8 and CGA, in early pregnancy) (Fig. 2a and Supplementary Fig. 3a). For the eCTB subclusters, we annotated eCTB proliferation (cluster 7, MKI67), eCTB fusion (cluster 11, ERVFRD-1) and two intermediate clusters, eCTB1 (cluster 9) and eCTB2 (cluster 6). Our eCTB subclusters annotations aligned with those of a previous study27. The consistency between scRNA-seq and snRNA-seq results confirmed the reliability of our analytical method.
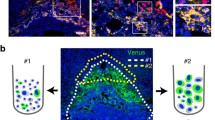
a, UMAP shows CTB and STB nuclei profiled with snRNA-seq in early pregnancy. The upper right UMAP indicates the expression patterns of representative marker genes for STB nuclear subclusters. The expression levels are presented with color intensities. b, UMAP shows CTB and STB nuclei profiled with snATAC-seq in early pregnancy. The upper right UMAP indicates the gene-activity scores of representative marker genes for STB nuclear subclusters. The activity scores are presented with color intensities. c, smFISH staining of indicated marker genes (PAPPA, FLT1 and hCG) characterizes eSTB mature 1 and eSTB mature 2 in early pregnancy. Statistical analysis of the proportion of eSTB mature 1 and eSTB mature 2 (middle). Data are shown as mean ± s.d. Unpaired two-tailed t test. n = 6 donors. The schematic represents the distribution of eSTB mix, eSTB mature 1 and eSTB mature 2 in the early placental STB (right). d, Pseudotime ordering of CTB and STB nuclei profiled with snRNA-seq reveals two differential trajectories (upper). The differentiation time is presented with color intensities. Quantitative classifications for each cluster by differentiation states are shown (bottom). e, Differential trajectories visualization on snATAC-seq UMAP (upper) and pseudotime ordering of the STB differentiation trajectories on snATAC-seq (bottom). The differentiation time is presented with color intensities. This trajectory was inferred by a supervised algorithm (Method). f, GO enrichment reveals STB nuclear functions. Three DEG sets of the two trajectories are used for the GO enrichment test. The −log10 P value of the enrichment is presented with color intensities. GO enrichment analysis is tested by one-sided Fisher’s exact test and P values are adjusted for multiple comparisons with the Benjamini–Hochberg method. **Padj < 0.01; ***Padj < 0.001. g, Functional networks visualize the GO enrichment results. Nodes represent GO terms and DEGs, whereas edges represent a gene’s membership in the GO term. Node color, log(FC) of DEGs; edge width, the signification of enrichment test P adjust value. GO enrichment analysis is tested with the same statistic method as f. NS, not significant.
Interestingly, we observed the distinct gene expression patterns in the eSTB nuclear subclusters (Fig. 2a and Supplementary Fig. 3a). Cluster 8, which expressed SH3TC2, was located in proximity to cluster 11 (eCTB fusion) on the UMAP plot and was annotated as eSTB nascent. Other nuclei, with low SH3TC2 expression and high LEP expression, were considered nuclear subclusters of the eSTB that gradually matured. Clusters 10 and 3, located at the edge of the UMAP, were considered the most matured STB clusters and were annotated as eSTB mature 1 (cluster 10 with highly expressed PAPPA) and eSTB mature 2 (cluster 3 with highly expressed FLT1). Clusters 2, 1, 4 and 5 were annotated as premature or mixed STB nuclei, with lower expression of PAPPA or FLT1 than nulcei in cluster 10 or cluster 3, and were annotated as eSTB premature 1 (cluster 2), eSTB premature 2 (cluster 1) and eSTB mixed (clusters 4 and 5). Overall, our results demonstrate considerable transcriptional heterogeneity within the STB nuclei in early pregnancy. To make cell/nuclear type nomenclature clear, we have included a table outlining the abbreviations, full names and detailed explanations of the eSTB nuclear subclusters (Supplementary Table 3).
To investigate the correlation between chromatin accessibility and eSTB nuclear gene expression, we analyzed snATAC-seq data for 22,785 eSTB and eCTB nuclei. Nine nuclear clusters were identified based on their gene-activity scores of known markers identified in snRNA-seq (Fig. 2b and Supplementary Fig. 3b). Notably, our analysis showed high concordance between the assigned subclusters in the snATAC-seq and snRNA-seq datasets, except for the proliferative eCTB cluster, potentially due to the loss of accessible chromatin during mitosis (Fig. 2a,b)42,43. We further confirmed the presence of distinct subtypes of STB nuclei during early pregnancy through single-molecule fluorescence in situ hybridization (smFISH; Fig. 2c and Supplementary Note 6). Overall, our multi-omic analysis revealed the heterogeneous nature of STB nuclei in early pregnancy at both the gene expression and epigenetic levels, which was missed by previous scRNA-seq analyses of placentas.
The maturation and development of STB nuclei remain poorly understood although there are suggestions that the STB nuclei may be of varying ages22. Pseudotime trajectory analysis of snRNA-seq data and differentiation pseudotime from snATAC-seq data, using a supervised method described in ref. 44, suggested the existence of a bifurcated developmental trajectory rooted from a common progenitor population for STB nuclei (eSTB nascent) toward two mature populations, namely eSTB mature 1 and eSTB mature 2. Notably, there is also a premature state that connects these STB subclusters at the transcriptional and epigenetic levels, which has not been described previously (Fig. 2d,e, Supplementary Fig. 3c and Supplementary Note 7).
Specifically, DEGs and GO enrichment analysis unveiled the involvement of nascent STB nuclei in diverse biosynthetic processes, possibly coordinated by the ERBB and EGFR signaling cascades. Interestingly, eSTB mature 1 nuclei were associated with various growth hormone signaling pathways (Fig. 2f,g, Supplementary Fig. 3d–g, Supplementary Table 4 and Supplementary Note 8), while eSTB mature 2 nuclei are linked to the oxygen response, lipid localization and carboxylic acid transportation.
Taken together, our results demonstrated nuclear and functional heterogeneity, as well as the route by which these STB nuclei differentiate during early pregnancy.
Subtype-specific enhancer-gene regulatory network in STB in early pregnancy
To further investigate the regulatory mechanisms driving the nuclear heterogeneity and lineage specification of eSTB, we performed cross-modality integrative analyses using liger45 and divided all eSTB nuclei into nine clusters based on the integrated analysis (Fig. 3a). Major STB nuclear subtypes identified by jointed datasets are comparable to those identified by snATAC-seq or by snRNA-seq separately (Supplementary Fig. 5a,b and Supplementary Note 9).
a, UMAP shows CTB and STB nuclei profiled with integration of snRNA-seq and snATAC-seq in early pregnancy. b, Heatmap shows pseudotime ordering of the 23,746 DARs in early pregnancy (left). Zoomed-in genomic tracks show the cis-element accessibility of representative genes. The normalized accessibility of DARs is presented with color intensities. c, TF-mining heatmap shows candidate master TFs of each nucleus type. The NES and expression z score are presented with dot color and dot size, respectively. Dots with bold edges show selected TFs used for network construction. d, Regulatory networks with three layers (TF, cis-element and target gene) represent the cis-regulatory architecture covering the complete STB differentiation process. The gene percentage, peak accessibility and gene expression score are presented with circle sizes, edge width and color intensities, respectively. e, smFISH staining (left) and fluorescence intensities (right) show the colocalization of TFs (STAT5A and FOSL2) and representative genes (PAPPA and FLT1) of eSTB mature 1 and eSTB mature 2. hCG, STB marker. The white dotted lines show the outline of CTB. The white lines represent the plotted tracks of fluorescence intensity. f, UMAP shows CTB and STB nuclei profiled with integration of snRNA-seq of STB-CT30, STB-BL and placenta villi in early pregnancy. g, Heatmaps show marker gene expression in STB-BL, STB-CT30 and placental villi in early pregnancy. The expression levels are presented with color intensities. Binarized gene expression levels (positive or negative) are calculated and visualized beside heatmaps for the two marker genes, PAPPA and FLT1. h, Pseudotime ordering shows three differential trajectories of CTB and STB nuclei (upper left). The differentiation time is presented with color intensities. Quantitative classifications for each cluster (upper right) and specific pseudotime of cluster 9 and cluster 7 on the integrated UMAP (bottom) are shown. i, Heatmap shows the different expression patterns of marker genes, master TFs and representative hormones among in vitro trophoblast models and in vivo placental villi in early pregnancy. The z scores are presented with color intensities. Examples of genes with similar expression patterns are highlighted in bold black, and those with different patterns are colored in red.
We sought to determine the master regulators of distinct STB nuclear subtypes and delineate their regulatory landscape along the inferred differentiation trajectories in early pregnancy. We used MACS2 to identify candidate cis-regulatory elements (cCREs)46. To accomplish this, we conducted peak calling for each cell type individually. Subsequently, we merged all identified peaks, resulting in the identification of a total of 274,189 cis-elements across all CTB subclusters and STB subclusters. Among these elements, ~8.66% (23,746) exhibited nuclear-subtype-specific accessibility (Fig. 3b, Supplementary Fig. 4, Supplementary Table 5 and detailed descriptions are shown in Supplementary Note 10). Accessibility dynamics of cCREs associated to genes SH3TC2, FLT1, PAPPA, ESRRG, OVOL2, STAT5A, HIF1A and CEBPB along the STB nuclear differentiation path of STB nuclei were revealed (Fig. 3b, Supplementary Fig. 4 and Supplementary Note 10). Potential TFs that may serve as master regulators in guiding the differentiation of STB nuclei into two trajectories were also nominated (Fig. 3c and Supplementary Note 11).
To link the nuclear-specific master TFs regulated gene networks and enhancers for the key TFs (enhancer-gene regulatory network (eGRNs)), we calculated the correlation between gene expression and region accessibility for a symmetrical window of 500 kb around each gene transcription start site, leading to an average of six regions per gene and a total of 100,682 raw peak-to-gene links44 (Supplementary Fig. 5c). We further examined focused eGRNs for the eCTB fusion, eSTB mature 1 and eSTB mature 2 populations (Fig. 3d, Supplementary Fig. 3e and Supplementary Table 6), including 12 activator TFs with 2,081 enhancers linked to 1,566 genes. Those cistrome TFs we highlighted here, including ZNF217, RELA, MAFK and GRHL3 for eCTB fusion, STAT5A, STAT5B, STAT4 and MITF for eSTB mature 1 and FOSL2, CEBPB, CEBPG and SP3 for eSTB mature 2, may act as master lineage-determining regulators (Fig. 3d and Supplementary Fig. 5d,e). MITF enhancers were linked to several known hormone genes such as CSH2 and LEP. Subcellular colocalization of STAT5A with PAPPA and FOSL2 with FLT1 supported their cooperative function in defining eSTB mature 1 and eSTB mature 2 gene expression and function (Fig. 3e).
hTSCs and trophoblast organoids based on hTSCs could partially imitate placental STB heterogeneity, regulatory mechanisms and function
The derivation of hTSCs by us and others represent experimentally tractable and gene-editable in vitro model systems for examining aspects of human placental development47. However, the extent to which those models recapitulate the heterogeneity of the CTB and the STB nuclear subtypes, as well as the associated regulatory mechanisms we have identified in vivo, remains largely unexplored. Therefore, we used our large snRNA-seq and snATAC-seq datasets from human early placentas as references to better characterize these in vitro models. We used two hTSC cell lines derived from a human early blastocyst (hereinafter referred to as hTSCs-BL)47 and first-trimester placental CTBs (hereinafter referred to as hTSCs-CT30)48 to show the generalizability of our findings (Supplementary Note 12).
To determine multinucleated STB features in vitro, we conducted snRNA-seq analysis of STB derived from both hTSCs-BL and hTSCs-CT30 (hereinafter referred to as STB-BL and STB-CT30, respectively). We integrated in vitro and in vivo snRNA-seq data from placentas to assess whether our in vitro trophoblast models (hTSCs-BL, hTSCs-CT30, STB-BL and STB-CT30) could mimic the CTB subtypes and STB nuclear subclusters we identified in this study. Using canonical trophoblast markers identified in vivo, we annotated newly generated nuclear/cellular subclusters in the integrated datasets including intCTB proliferation, intCTB1, intCTB2 and intCTB fusion subclusters, as well as intSTB nascent, intSTB premature 1, intSTB mature 1/mature 2 and intSTB mixed subclusters (Fig. 3f,g,i, Supplementary Fig. 6a–c and Supplementary Note 13). However, eSTB mature 1 and eSTB mature 2 were not observed in STB-BL and STB-CT30 as revealed along the integrated differentiation trajectory (Fig. 3h, Supplementary Fig. 6a–f and Supplementary Note 13).
We investigated the regulatory mechanisms between in vitro and in vivo STB in hTSCs-BL and hTSCs-CT30 with doxycycline (DOX)-inducible overexpression of STAT5A and MITF. The full-length CDS with a Flag-tag was used due to low expression levels of these genes in in vitro STB compared to their in vivo counterparts in eSTB mature 1 (Fig. 4a–e and Supplementary Table 7). We next aimed to determine whether regulatory mechanisms are conserved between in vitro and in vivo STB. The significant increase in the gene expression such as PAPPA and others (Supplementary Note 14) under STAT5A and MITF overexpression in hTSC-BL confirmed our eGRN as shown in Fig. 3d. Similar results were obtained for hTSC-CT30 (Supplementary Fig. 6h–k). CUT&Tag and chromatin immunoprecipitation followed by sequencing (CHIP–seq) experiments were performed and confirmed the consistent regulatory events between the eGRN in placental STB in early pregnancy and STB differentiated from hTSCs (Fig. 4f and Supplementary Note 14). To test these in vitro models, we investigated the role of MITF in the developing trophoblast organoids constructed from hTSCs-BL. We generated trophoblast organoids using hTSCs-BL with DOX-inducible overexpression of MITF. ELISA assays showed a positive correlation between CSH2 secretion and the overexpression of MITF in the trophoblast organoids (Fig. 4g–i).
a, The schematic illustrates the lentiviral vector used for STAT5A and MITF overexpression. b, Bright-field images show hTSCs-BL-STAT5AOE and STB-BL-STAT5AOE. Scale bar: 100 μm. c, RT–qPCR analysis of STAT5A and target gene expression in STB-BL with DOX-inducible overexpression of STAT5A. Data are shown as mean ± s.d. P values by multiple unpaired two-tailed t test. n = 3 independent experiments. The dotted lines in c and e represent the baseline 1. d, Bright-field images show hTSCs-BL-MITFOE and STB-BL-MITFOE. Scale bar: 100 μm. e, RT–qPCR analysis of MITF and target gene expression in STB-BL with DOX-inducible overexpression of MITF. Data are shown as mean ± s.d. P values by multiple unpaired two-tailed t test. n = 3 independent experiments. f, Verification and visualization of eGRN enhancer-gene regulatory events near two target genes of two STB mature 2 specific TF regulators, CEBPB and FOSL2. Genomic tracks are calculated and compared with CEBPB and FOSL2 CUT&Tag, CEBPB ChIP–seq datasets, which are prepared with STB-BL and hTSC-BL cell lines. IgG mock is used for negative control. Regulatory events are highlighted with the following criteria: eGRN target gene regions (enhancers) should overlap STB-BL CUT&Tag peak(s) and/or ChIP–seq peak(s) but are depleted in hTSC-BL. g, Schematic representation of trophoblast organoids derivation. hTSC-BL are transferred to matrigel droplets on day 0 (D0) and maintained in trophoblast organoid medium to analyze on day 6 (D6). h, Bright fields and immunofluorescence analysis of trophoblast markers (CDH1 and hCG) in trophoblast organoids derived from hTSCs-BL with DOX-inducible overexpression of MITF. Scale bars: 250 μm for bright-field images, 20 μm for immunofluorescence images. i, ELISA analysis of CSH2 expression in trophoblast organoids derived from hTSCs-BL with DOX-inducible overexpression of MITF. Data are shown as mean ± s.d. P values by unpaired two-tailed t test. n = 3 independent experiments. CDS, coding sequence; hPGK, human phosphoglycerate kinase 1 promoter; P2A: porcine teschovirus-1 2A peptide; Puro, puromycin; TRE3GS, TRE3GS inducible promoter; Tet-on 3G, Tet-on 3G transactivator protein; WPRE, Woodchuck hepatitis virus’s post-transcriptional regulatory element; OE, overexpression.
Overall, our findings indicate that our datasets offer a reliable reference for evaluating in vitro models recapitulating certain aspects of STB heterogeneity, regulatory mechanisms and functions during early pregnancy.
Matured STB nuclear heterogeneity and function were revealed in late pregnancy
To systematically investigate the development of human placental STB in late pregnancy, we examined our large-scale single-nucleus transcriptomic (23,981 nuclei) and epigenomic (24,692 nuclei) datasets from lSTB and lCTB nuclei. Quality control (Supplementary Fig. 7a–d) and annotation of lCTB subtypes are described in Supplementary Note 15. Gene sets previously used to classify nuclear types in early pregnancy demonstrated notable consistency in late pregnancy samples (Fig. 5a,b, Supplementary Fig. 7e,f and Supplementary Table 3). Cluster 11 and cluster 6 were classified as two nascent nuclear types of STB in late pregnancy with high expression of SH3TC2 (lSTB nascent 1 and lSTB nascent 2). PAPPA expression was substantially increased in late pregnancy and was present in five STB subclusters, including cluster 4 (lSTB premature 1-a), cluster 1 (lSTB premature 1-b), cluster 3 (lSTB mature 1-a), cluster 2 (lSTB mature 1-b) and cluster 7 (lSTB mature 1-c). In contrast, FLT1 expression was only detected in cluster 5 (lSTB mature 2-a) and partially in cluster 8 (lSTB mature 2-a), which represents a small fraction of nuclei in STB in late pregnancy (Fig. 5a and Supplementary Fig. 7e). Detailed information on the annotation of STB nuclear subtypes is shown in Supplementary Notes 16 and 17. The decreasing expression of FLT1-positive STB nuclei may indicate an intriguing regulatory effect of the altered environmental cues from hypoxia in early pregnancy to normoxia in late pregnancy and dynamic developmental demands. Interestingly, the small population of FLT1-expressing lSTB nuclei showed dramatic closure of the chromatin locus around the FLT1 gene when compared with early pregnancy (Supplementary Fig. 8b,d). The low chromatin accessibility around the FLT1 gene expression locus during late pregnancy led us to consider detectable FLT1 RNA in clusters 5 and 8 as residual RNA of the FLT1 gene from an earlier pregnancy stage.
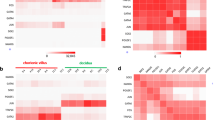
a, UMAP shows CTB and STB nuclei profiled with snRNA-seq in late pregnancy. The bottom UMAP indicates the expression patterns of representative marker genes for STB nuclear subclusters in late pregnancy. The expression levels are presented with color intensities. Gene expression raw counts were normalized by depth, logarithmized and z score scaled, and finally smoothed with imputation. b, UMAP shows CTB and STB nuclei profiled with snATAC-seq in late pregnancy. The bottom UMAP indicates the gene-activity scores of representative marker genes for STB nuclear subclusters in late pregnancy. The activity scores are presented with color intensities. c, Pseudotime ordering of lSTB nuclei profiled with snRNA-seq shows the differentiation trajectory in late pregnancy by monocle 2 DDRTree algorithm. The differentiation time is presented with color intensities. Nuclei of each cluster are aligned above and ordered by pseudotime values. d, Trajectory heatmap shows 16 typical new marker genes (row) expressed dynamically along pseudotime ordering (column). The pseudotime time is presented with color intensities. e, Functional networks visualize the GO enrichment results. Nodes represent GO terms and DEGs, whereas edges represent a gene’s membership in the GO term. Node color, log(FC) of DEGs; edge width, the signification of enrichment test P adjust value. GO enrichment analysis is tested by one-sided Fisher’s exact test and P values are adjusted for multiple comparisons with the BH method. f, smFISH staining of indicated marker genes (PAPPA, FLT1 and hCG) characterizes lSTB mature 1 and lSTB mature 2 in late pregnancy (left). Statistical analysis of the proportion of lSTB mature 1 and lSTB mature 2 (middle). Data are shown as mean ± s.d. P values by unpaired two-tailed t test. n = 6 donors. The schematic represents the distribution of lSTB mature 1-a, lSTB mature 1-b and lSTB mature 2 in the late placental STB (right). g, UMAP shows lCTB and lSTB nuclei profiled with snRNA-seq and snATAC-seq after the python package 'GLUE' integration (Methods). h, Heatmap shows the activity of TF-target regulatory modules elicited from the integration of snRNA-seq and snATAC-seq data using the FigR package. Genes represent target genes and candidate TF regulators in rows and columns, respectively. i, TF-regulatory network construction with a combination of modules lSTB mature 1-a and lSTB mature 1-b in late pregnancy. The Domains of Regulatory Chromatin (DORC) accessibility and expression level are presented with circle sizes and color intensities, respectively. The network is built with a different method for late pregnancy.
To further uncover the molecular mechanisms driving lSTB nuclear heterogeneity and place the newly annotated lSTB nuclear subclusters in a defined trajectory, we performed pseudotime analysis (Fig. 5c,d). A linear transcriptional trajectory was revealed for all PAPPA-positive nuclei in late pregnancy. Following the order of pseudotime, we observed that the first cluster of lSTB nuclei was largely composed of nascent nuclei we annotated with specific gene expression of PDE4D, BACE2 and SH3TC2 (Fig. 5c and Supplementary Fig. 7h–j). lSTB nuclei with high expression of LAMA3 and located at the end of the trajectory were the most matured nuclear type with BMP1 highly expressed nuclei located in the middle (Fig. 5c and Supplementary Fig. 7h–j). We further revealed dynamic gene expression throughout a continuum of nuclear-state transition in Fig. 5d and identified DEGs along the trajectory, including PDE4D, SH3TC2, PAPPA, BMP1, STAT5A, PTCHD4, CCDC30, INPP5D and CROT. GO enrichment analysis revealed that extracellular matrix organization (ECM) was highly associated with nuclei in lSTB mature 1-a and lSTB mature 1-b subcluster (Fig. 5e and Supplementary Table 8).
Placenta in late pregnancy was stained with probes targeting genes PSG8, SH3TC2, LEP, PAPPA and FLT1. The smFISH results for FLT1 and PAPPA suggest that a substantial portion of STB nuclei exhibit PAPPA expression, aligning with our bioinformatic analysis (Fig. 5f and Supplementary Note 17).
The integration of snRNA-seq and snATAC-seq through analysis of peaks and gene regulatory proximity to the linear genome reveals strong connections of gene expression patterns and accessibility dynamics of regulatory elements (Fig. 5g, Supplementary Fig. 9a–g and Supplementary Note 18). Subsequently, we performed a TF-mining and TF-regulatory network analysis (FigR, v0.1.0) and identified seven unique modules of genes that are regulated by distinct TFs (Fig. 5h,i and Supplementary Figs. 10 and 11). Notably, key TFs that were enriched in the most matured and functional lSTB mature 1-a and lSTB mature 1-b subclusters, such as STAT6, STAT5A, STAT4 and MITF (module 1 and module 2 as shown in Figs. 5h and 5i), were also important in the eSTB mature 1 subcluster in early pregnancy. GO enrichment revealed a strong association between ECM and the nuclear subtypes of STB in the late pregnancy stage (Supplementary Note 19).
Discussion
Recently, nuclear-specific functions in the syncytia of various organisms, including the fungus Ashbya gossypii49, muscle fibers50 and other tissue systems, have been documented. However, the nuclear heterogeneity and differentiation in the human placental STB remains largely unexplored16,18,19,22,51. Here we leveraged the power of snRNA-seq and snATAC-seq to profile the nuclear diversity and regulatory mechanism from both transcriptomic and epigenetic perspectives. Our analysis demonstrated the dynamic STB nuclear subtypes during early and late pregnancy. It provided evidence of significant heterogeneity in the STB nuclei from a gene expression and regulation perspective, allowing us to identify a dynamic bifurcating trajectory of STB nuclei with distinct biological functions. These findings together with the eGRNs we uncovered lay the groundwork for further exploration of mechanisms driving human placental STB development in supporting successful pregnancy (Supplementary Note 20).
Specifically, integrative snRNA-seq and snATAC-seq analysis of STB nuclear populations in early pregnancy, revealed some TFs, including STAT5A, that might serve as a driving force in guiding the differentiation of STB nuclei towards a more mature state, which is characterized by high levels of PAPPA expression. PAPPA, a serum marker used for screening developmental abnormalities in early pregnancy, was detected at increasing concentrations in the maternal circulation during pregnancy52. We anticipate that future work will clarify the relationship of TFs we identified here and the conditions associated with PAPPA. Moreover, a direct regulatory network FLT1 controlled by CEBPB and FOSL2 was revealed. This is especially important in the clinical context, as CEBPB was recently reported to be involved in EVT dysfunction in the severe preeclampsia placenta53,54. The abnormally overexpressed soluble FLT1 protein in the placenta, which is secreted into maternal blood, is also known to contribute to the pathogenesis of preeclampsia55. Therefore further investigation is necessary to fully understand the regulatory roles of CEBPB and FOSL2 in FLT1 expression in the STB during pregnancy and in related diseases.
Newly established hTSCs have been considered a substantial breakthrough in the field48,56,57,58. Our previous work has already proven that hTSCs could be used as a model for human trophoblast differentiation during implantation59. Manipulation of the key STB nuclear lineage-determining TFs that we identified here, in hTSCs, could provide a better experimental model in the interpretation of human placenta STB in later stages beyond implantation.
In summary, we have presented a comprehensive characterization of STB nuclear heterogeneity during pregnancy (see the summary model in Supplementary Fig. 13). The data we generated not only enhanced our understanding of the regulatory mechanisms that govern the dynamic development of STB nuclear identities but also established a valuable framework for future studies aimed at interpreting the roles of key molecules associated with placental development and disorders in humans.
Methods
Description of human placental donors and ethical approval
The placental specimens used in this study of snRNA-seq and snATAC-seq were collected from six early pregnant women (6–9 weeks of pregnancy) and six late pregnant women (38–39 weeks of pregnancy). Additional three healthy placental tissues from the three early pregnant women (7–8 weeks of pregnancy) and three placental tissues from the three late pregnant women (38 weeks of pregnancy) were used for experimental validation (RNAscope). The placental specimens were all collected from Peking University Third Hospital. Sex identification for samples was analyzed with PCR of the following two sex-linked genes: SRY and NLGN4, before sequencing. This research was approved by the Ethics Committee of the Institute of Zoology, the Chinese Academy of Sciences, and the Ethics Committee of the Peking University Third Hospital under research license (2019) JLS (242-01). All patients participating in the study were healthy pregnant women without any exclusion. Every patient involved in the project has signed the informed consent. All the patients in our research project shared one ethical approval license number.
Library construction and sequencing
Single-nucleus isolation
The villi were separated from the early or term placenta and cut up after removing the blood with PBS. Ice-cold Nuclei EZ lysis buffer (Sigma-Aldrich, NUC-101) containing RNAase inhibitor (Takara, 2313B) and protease inhibitor (Sigma-Aldrich, P8340) was added to resuspend the tissue and grinded with a Dounce tissue grinder (Sigma-Aldrich, D8938). The lysates were bathed in ice and lysed twice. After washing with PBS, the nuclei were split in half and filtered through a 40 μm sieve to proceed with ATAC-seq and RNA-seq.
snRNA-seq
Single Cell B Chip Kit (10x Genomics, 1000074) and the nucleus suspension (600 nuclei per microliter determined by CountStar) were loaded onto the Chromium single cell controller (10x Genomics) to generate single-nucleus gel beads in the emulsion (GEMs) according to the manufacturer’s protocol. In brief, single nuclei were suspended in PBS containing 0.04% BSA. About 16,000 nuclei were added to each channel, with 8,000 target nuclei estimated to be recovered. Captured nuclei were lysed, and the released RNA was barcoded through reverse transcription in individual GEMs. Reverse transcription was performed on an S1000TM Touch Thermal Cycler (Bio-Rad) at 53 °C for 45 min, followed by 85 °C for 5 min and held at 4 °C. The cDNA was generated and then amplified, and quality was assessed using an Agilent 4200 (performed by CapitalBio Technology). Next, snRNA-seq library was constructed using Single Cell 3′ Library and Gel Bead Kit V3.1 according to the manufacturer’s instructions. The libraries were finally sequenced using an Illumina NovaSeq 6000 sequencer with a sequencing depth of at least 100,000 reads per nuclei with a pair-end 150 bp (PE150) reading strategy (performed by CapitalBio Technology).
snATAC-seq
Following the 10x Genomics single-cell ATAC solution, by using Chromium Chip E Single Cell Kit (1000156) and Chromium Single Cell ATAC Library and Gel Bead Kit (1000110), the nuclei in a bulk sample were partitioned into nanoliter-scale GEMs and a pool of ~750,000 10× Barcodes was sampled to separately and uniquely index the transposed DNA of each nucleus. Libraries were then generated (performed by CapitalBio Technology). The libraries were sequenced using an Illumina Novaseq sequencer with a sequencing depth of at least 25k read pairs per nuclei with a pair-end 50 bp (PE50) reading strategy.
Massive integration analysis of snRNA-seq and snATAC-seq from two pregnancy periods
Identification of nucleus types at a global level
To understand the nuclei type consistent of snRNA-seq and snATAC-seq of the early and late pregnancy, we started the analysis with the Cellranger ‘aggre’ command (without dept normalization) to aggregate libraries. For a quick and general analysis, we used the Python packages Scanpy v1.6.1 (ref. 34) and SnapATAC v2.2.0 (ref. 35) to perform the upstream analysis. Then each data modality was applied with Harmony40 to remove the batch effect, and we then achieved dimension reduction, nuclei cluster partition and annotation with known marker genes. We used customized R codes to visualize the results. To assist cluster annotation in snATAC-seq, the gene-activity score of each gene was quantified by make_gene_matrix function from the package SnapATAC. For integrating the 24 libraries from two modalities at two pregnancy stages, the liger package (v0.5.0)41 was finally applied. The nuclear major types from different modalities and pregnancy stages were visualized by marker genes in the integration space.
DEG identification of the STB population
To understand the diversity of the STB nuclei in early and late pregnancy, we identified the DEGs according to the snRNA-seq analysis results. We first subset all STB nuclear clusters from the early and late pregnancy data and performed DEG test at the single-nucleus level with the following three methods: Seurat findMarkers (Wilcoxon test, default parameter), Scanpy rank_gene_groups (Wilcoxon test, default parameter) and method from ref. 60 (analysis of variance (ANOVA), fold change (FC) > 0.5, P < 0.05). We then found a shared set within the top 200 of each sorted DEG gene sets. To directly visualize and compare the gene expression difference between pregnancy stages, we aggregated and calculated a pseudobulk RNA-seq data by stage and donor. We then averaged logarithm of counts per million reads, log2(CPM) expression values of the six donors per stage and plotted with scatter plot and labeled the top 20 genes. GO of Biological Process enrichment was analyzed with DAVID61 of the knowledge base v2023q2 with share DEG genes.
Differential TF identification of the STB population
We applied chromVAR62 to perform a quick scan for snATAC-seq data of two pregnancy stages against the JASPAR 2020 motif database36. In brief, the genomic positions of each motif were identified in each peak. A deviation score of each motif was calculated for each nucleus, producing a deviation score by nuclei matrix. Then this matrix was z-score normalized. We used FindMarkers in the Seurat package to identify differentially enriched motifs with default parameters between early and late pregnancy. TF motifs of interest were selected from the top 50 list and further aligned and visualized with gene-activity score and gene expression data.
snRNA-seq analysis for STB subtypes in early pregnancy
Dimension reduction and graphic clustering for nucleus-type annotation
We used the R package Seurat (v3)63 to conduct the general upstream analysis. Generally, we used principal component analysis for dimension reduction with a maximum dimension of 30. We then performed graphic clustering by the ‘Louvain’ algorithm with a start resolution of 0.9. The final resolution was chosen after an iterative tuning for best interpreting of the biological complexity of the placenta sample. We annotated each cluster by marker gene expression.
Differential gene expression analysis and GO enrichment analysis
To identify DEGs, we applied Seurat FindAllMarkers with parameters of min.pct = 0.1, logfc.threshold = 0.25. Top 50 DEGs were visualized by heatmap. Clusterprofiler (v3.18.1) was applied for GO enrichment analysis and visualized by customized R script. All DEGs were saved and used for further analysis.
Differentiation trajectory reconstruction
We used the R package Monocle2 (ref. 64) to conduct pseudotime analysis. DEGs identified before were used as the informative gene set, which would encode the cell differential trajectory. To focus on the STB nuclei differentiation, we excluded the CTB nuclei and started at the fusion-competent CTB nuclei (ERVFRD-1 positive). We set DEGs mentioned above as ordering genes for unbiased trajectory inference. The trajectory tree was then visualized as ‘DDRTree,’ and essential genes during differentiation were examined by heatmap. Each nuclear cluster type was classified along the trajectory path and visualized by customized R scripts.
snATAC-seq analysis for STB subtypes in early pregnancy
Dimension reduction and graphic clustering for nucleus-type annotation
We applied the SnapATAC v2 (ref. 35,65) package to perform the general upstream analysis because we found this package used a spectral clustering method as the dimension reduction algorithm, which can better interpret the intrinsic population structure of the STB clusters. Harmony40 was applied for batch effect removal. We also compared the dimension reduction results with cisTopic (v0.3.0) with the Latent Dirichlet Allocation (LDA) algorithm, ArchR (v1.0.1) with the iterative Latent Semantic Indexing (LSI) algorithm and Signac (v1.1.0) with the LSI algorithm (data not shown). All these tools produced similar dimension reduction results. For nuclei clustering, we applied the ‘Leiden’ algorithm with a resolution of 0.9 for the initiative graphic clustering. To gain reasonable cluster numbers (which is essential to interpret the nuclei types), we iteratively tested the resolution to optimize the final clustering results. Several known marker genes were used to verify and annotate the clusters.
Differentially accessible regions (DARs) identification
We used the function ‘rank_genes_groups’ in the scanpy package with ‘t test’ methods to identify DARs. This function used a raw peak count matrix as the input data. We applied a strict filtering threshold of P value < 0.001 and a minimal log2(FC) of 1. The top 5,000 DARs were kept and used for further analysis.
Differentiation trajectory reconstruction
We applied a supervised trajectory inference method44 to illustrate the developmental paths of STB nuclei in early pregnancy. Briefly, we tried to fit a candidate trajectory path on a UMAP map and then pseudotime values were assigned to each nucleus. It is based on the hypothesis that UMAP coordinates per se harbor the intrinsic differential information.
Matching snRNA-seq and snATAC-seq clusters in early pregnancy
Calculate gene-activity score for snATAC-seq nuclei
We aggregated and counted the snATAC-seq Tn5 insertion sites within the gene body and a 2 kb extended upstream region to calculate the gene-activity score for each gene of the human gene annotation database (Gencode v41) by SnapATAC v2 function make_gene_matrix.
Clusters matching with liger
Similar to the previous massive integration analysis, we applied the liger method for modality integration of the early pregnancy data. We selected the variable genes set from snRNA-seq data with var.thresh = 0.2 and set the k = 35. Finally, the Integrative Non-negative Matrix Factorization (iNMF) dimension reduction matrix was calculated with all nucleus ids as the col-names. Louvain clustering was then conducted with resolution = 0.8.
To visualize the integration results, we merged every liger modality dataset and built a Seurat object with both gene-activity score and gene expression data. Then we normalized the data slot with ‘normalizeData’ function in Seurat. We visualized the marker genes in the integration space to annotate liger clusters. snRNA-seq clusters and snATAC-seq clusters were joined and paired according to the integration clusters. A confusion matrix of cluster pairs was also made from paired nuclei with annotation labels in previous results.
Quantify and visualize the between-modality alignment result
We applied a customized K-Nearest Neighbors (KNN) algorithm to quantify the nucleus-to-nucleus matching result. In brief, we first randomly picked 50 snATAC-seq nuclei in the liger integration UMAP space and then iteratively found the nearest snRNA-seq nuclei. The results were compared and visualized with a combination of snRNA-seq and snATAC-seq clusters by a scatter plot.
Peak-to-gene linkage analysis
We used the liger iNMF dimension reduction matrix to pair nuclei of the snRNA-seq data to nuclei from the snATAC-seq data by a simple nearest Euclidean distance strategy44,66. This method found the closest RNA nuclei for each ATAC nuclei in the liger iNMF space and built the nuclei pairs. Accordingly, the pairing result was used to impute the expression matrix (gene × nuclei of snRNA) to the imputedRNA matrix (gene × nuclei of snATAC), resulting in two matrics with paired nucleus ids. Then all-to-all correlation was calculated among all gene rows of the imputedRNA matrix against all peak rows of the peak count matrix (peak × nuclei of snATAC). We then filtered the correlation table with a cutoff of correlation co-efficiency of 0.45 and an False Discovery Rate (FDR) cutoff of 0.0001. The resulting peak-versus-gene pairs were defined as peak-to-gene links. Then all peak-to-gene links were aligned with a combination of peak accessibility and gene expression data and visualized as a heatmap. To illustrate the local enhancer–promoter regulatory landscape of essential genes (PAPPA and FLT1), we plotted genomic regions around genes with the pseudobulk ATAC tracks matching single nuclear gene expression data.
TF mining and construction of a regulatory network
To decipher the TF regulation network for STB with a multi-omics-based strategy67, we reinterpreted the core steps with homemade R codes with slight modifications to perform the TF mining and construct a precise and robust ‘TF—cis-element—target gene’ regulatory network (Supplementary Fig. 6g). In brief, the TF motif enrichment tool i-cisTarget68 and pycistarget69 were used to scan TF motif enrichment in previously identified cluster-specific DARs (coordinates were lifted from hg38 to hg19), with default parameters except for an area under the curve (AUC) threshold of 0.001. We used the motif Position Weight Matrix (PWM) database only. For pycistarget, we used the nonredundant motif database ‘hg38_screen_v10_clust’ for clearer TF gene assignment. The TF motif enrichment results of the above mentioned two packages were collected, merged and parsed, and a maximal normalized enrichment score (NES) was assigned to each TF motif that passed the AUC threshold 3.0. According to the hypothesis that a TF regulator is a true-positive TF regulator only if the TF gene expression is positively correlated with the accessibility of motifs66, we calculated Pearson’s correlation for all TF motif NES against TF gene expression. We ranked the correlations for TF genes and visualized all selected cluster-specific TF regulators in a heatmap.
To build the ‘TF—cis-element—target gene’ regulatory network, we extracted all candidate target genes for each TF motif in the enrichment table and searched for the linked peaks (cis-element) for each target gene within all predefined peak-to-gene links. The final networks were constructed with Cytoscape (v3.9.1)70, integrating node and edge attributions according to snRNA-seq or snATAC-seq data.
Culture of hTSCs
The culture of human trophoblast was performed as described previously47. Briefly, the plate was coated with 5 μg ml−1 Collagen I (Corning) at 37 °C at least for 1 h. hTSCs were cultured in hTSCs medium (DMEM/F12 (Gibco) supplemented with 0.1 mM 2-mercaptoethanol (Gibco), 0.2% FBS (Gibco), 0.5% Penicillin–Streptomycin (Gibco), 0.3% BSA (Sigma-Aldrich), 1% ITS-X supplement (Gibco), 1.5 μg ml−1 l-ascorbic acid (Sigma-Aldrich), 50 ng ml−1 Epidermal Growth Factor (EGF, MedChemExpress), 2 μM CHIR99021 (MedChemExpress), 0.5 μM A83-01 (MedChemExpress), 1 μM SB431542(MedChemExpress), 0.8 mM valproic acid (VPA, Wako) and 5 μM Y27632 (MedChemExpress)). hTSCs were dissociated with TrypLE (Gibco) for 8 min at 37 °C, and the cells were passaged to a new Collagen I-coated plate. hTSCs were routinely passaged every 4–5 d at a 1:4–1:6 ratio.
Lentivirus production
The full-length coding sequence of STAT5A and MITF were cloned into the pLVX-TetOne-TRE3GS-MCS-2A-PURO lentiviral vector, respectively. The overexpressing lentivirus vector together with packaging plasmids pPAX2 and pMD2.G were cotransfected into HEK293T cells using a polyethyleneimine transfection protocol (Proteintech). The viral supernatants were collected at 48 and 72 h post-transfection and then filtered through a 0.45 μm polyethersulfone (PES) filter.
Generation of overexpressing hTSCs
hTSCs were preseeded into 12-well plate at a density of 5 × 104 cells per well and were infected with overexpressing lentivirus at a confluency of 30% for 12 h. Then the infected hTSCs were treated with 2 μg ml−1 puromycin for 48 h to establish stable overexpressing hTSC lines.
Differentiation of hTSCs into STB
hTSCs were grown to 80% confluence in the hTSCs medium and dissociated with TrypLE for 8 min at 37 °C. For the induction of STB, hTSCs were seeded in a six-well plate precoated with 2.5 μg ml−1 Collagen I at a density of 1 × 105 cells per well and cultured in STB medium (DMEM/F12 supplemented with 0.1 mM 2-mercaptoethanol, 0.5% Penicillin–Streptomycin, 0.3% BSA, 1% ITS-X supplement, 2.5 μM Y27632, 2 μM forskolin (MedChemExpress) and 4% Knockout Serum Replacement (KSR, Gibco). The medium of overexpressing STB was supplemented with 5 μM DOX (MedChemExpress). The medium was replaced every 2 d and analyzed on day 6.
Generation of trophoblast organoid
The establishment of trophoblast organoids was performed as described previously71. Briefly, dissociated hTSCs were suspended in 100 μl Matrigel at a density of 1 × 104. One droplet per well was added to an eight-well ibidi chamber or eight droplets per well were added to a six-well plate. After setting the plate at 37 °C for 15 min, 200 µl or 2 ml hTSC culture medium was loaded per well. In this case, it is called trophoblast organoid medium. The medium of overexpressing organoids was supplemented with 5 μM DOX. The medium was replaced every 2 d and analyzed on day 6.
Comparison of in vivo and in vitro snRNA-seq
We first analyzed and annotated two datasets of snRNA-seq obtained from hTSCs-BL and hTSCs-CT30 using the Seurat algorithm, similar to the analysis we performed on placenta samples in early pregnancy. We used known marker genes, including GATA3, TEAD4, TOP2A, PCNA and ITGA6, to annotate proliferative hTSC and CTB. In addition, we annotated STB-BL and STB-CT30 subclusters using markers that were previously used to annotate eCTB fusion (ERVFRD-1), eSTB nascent (SH3TC2) and eSTB mature (CGA, PAPPA and FLT1). We used the Seurat CCA algorithm63 to identify anchor genes and integrated two snRNA-seq datasets from hTSCs-BL and hTSCs-CT30 with snRNA-seq datasets from placental villus in early pregnancy. Subsequently, we jointly annotated the clusters using marker genes and aligned them along the differentiation trajectory using Monocle2 (v2.14.0)64,72.
CUT&Tag
The experiment was conducted according to the instructions provided by Vazyme (TD904). In brief, approximately 5 × 104 cells were collected and counted, then mixed with activated ConA beads and incubated overnight with primary antibodies (CEBPB—Proteintech, 23431-1-AP; FOSL2—USBiological, 351814; rabbit IgG—Abcam, ab172730). The secondary antibody (VazymeAb207) was incubated at room temperature with rotation for 1 h. After removing the supernatant, pA/G-Tnp Pro and TTBL were separately incubated with rotation at room temperature and 37 °C for 1 h. The supernatant was then mixed with DNA Extract Beads Pro to dissolve DNA with ddH2O and used directly for PCR amplification. Subsequently, VAHTS DNA Clean Beads were added to purify the DNA, which was then subjected to next-generation sequencing using the Illumina NovaSeq platform.
ChIP–seq
The experiment of ChIP–seq was performed as previously described73. Briefly, about 5 × 106 hTSCs-BL and STB-BL were used for cross-linking by 1% formaldehyde and quenched by 125 mM glycine. Cells were lysed and nuclei were sonicated and diluted to a final concentration of 0.1% by SDS. Protein A/G magnetic beads were added and incubated at 4 °C for 1 h. Precleared chromatin was incubated with primary antibody (CEBPB, 23431-1-AP, Proteintech) or monoclonal rabbit isotype control (Abcam, ab172730) overnight at 4 °C. Then protein A/G magnetic beads were added to capture the specific chromatin complexes. The supernatants were collected with 500 mM NaCl and incubated at 65 °C overnight for reverse cross-linking. Samples were incubated at 37 °C for 1 h with RNase A and were incubated at 50 °C for 1 h with proteinase-K. The released DNA was purified by the QIAquick PCR Purification Kit and processed for sequencing library preparation. Multiplexed ChIP–seq libraries were prepared by NEBNext Ultra II DNA Library Prep Kit (NEB, E7645S). The final libraries and multiplexed libraries were subjected to Illumina NovaSeq 6000 using PE150 (paired-end 150 nt) sequencing.
Analysis of CUT&Tag and ChIP–seq data
We applied bowtie2 (v2.4.2) with default parameters for raw Fastq reads mapping to human (hg38) reference genome and used MACS2 to call peaks for CUT&Tag and ChIP–seq datasets. We used the ‘reduce’ function from the R package ‘GenomicRanges’ for the merge of peaks of each replication. We then intersected and counted overlap peaks from CUT&Tag and previous eGRN peaks that were assigned to nearby genes by peak-to-gene links. We quantitatively summarized the validation result by the overlap percentage of these two peak sets. Finally, we visualized the regulatory events with CUT&Tag, ChIP–seq tracks and eGRN TF-target peaks tracks for two randomly picked genes within the eGRN target gene list.
smFISH with RNAscope
Fresh tissues were collected and fixed in 10% neutral-buffered formalin (Sigma-Aldrich) for 18 h at room temperature followed by dehydration in ascending series of ethanol, clearing in xylene, embedding in paraffin and section for smFISH with RNA probes targeting genes SH3TC2, LEP, FLT1, PAPPA, PSG8, STAT5A and FOSL2 following instructions from RNAscope Multiplex Fluorescent Reagent Kit V2 (Advanced Cell Diagnostics, 323100). First, sections were pretreated with RNAscope hydrogen peroxide, the process of target retrieval and protease plus to unmask target RNA and permeabilize the cells. Second, probes were carefully prepared and used for the hybridization with the target RNA molecules. Third, RNAscope detection reagents were used to amplify the hybridization signals through sequential hybridization of amplifiers and label probes and stained with different fluorescent dyes. Finally, sections were counterstained with DAPI, mounted with fluorescent mounting medium and stored at −20 °C for long-term storage. Images were collected under a Zeiss LSM 880 confocal laser scanning microscope, and the image processing was performed with ZEN software. Additional methods are described in detail in Supplementary Methods.
Statistics
Clustering result refinement
We tuned the clustering result with different graph construction parameters of k and community discovery methods (Louvain or Leiden) for snATAC-seq data. After a fix of k = 30 and setting the clustering algorithm as ‘Leiden,’ we iterated the community detection resolution and chose the most frequent stable cluster assignment. We then evaluate the cluster results by the Dunn index and Silhouette coefficient.
Statistical analysis was performed using GraphPad Prism software (v9.0.0) by unpaired t test or one-way ANOVA analysis. Results were shown as means ± s.d.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
Sequencing data generated for this study have been deposited in the Gene Expression Omnibus with the accession GSE247038. There are no restrictions on data availability or use. We used the human genome version hg38 for all the analysis steps. The processed results were deposited on the figshare repository, with links: https://figshare.com/s/61db5dc9030e6363267e (early pregnancy snATAC-seq datasets), https://figshare.com/s/ccec028e5a3ec5b20106 (early pregnancy snRNA-seq datasets), https://figshare.com/s/05fdcc5d0c6b46786047 (late pregnancy snATAC-seq datasets), https://figshare.com/s/5a435d5ea48b3d51b40b (late pregnancy snRNA-seq datasets) and https://figshare.com/s/40bc9295d533d163b738 (scRNA-seq and snRNA-seq datasets for cell lines). We also uploaded raw result tables for the construction of eGRN networks (https://figshare.com/s/a3ec1630f75de248698b). Source data are provided with this paper.
Code availability
Codes for the generation of main result figures can be accessed at GitHub74 (https://github.com/wangmeijiao/Human_placenta_multi-omics_snRNA-seq_snATAC-seq; https://doi.org/10.5281/zenodo.10212321), and details are provided upon contact.
References
Hemberger, M., Hanna, C. W. & Dean, W. Mechanisms of early placental development in mouse and humans. Nat. Rev. Genet. 21, 27–43 (2020).
Aplin, J. D., Myers, J. E., Timms, K. & Westwood, M. Tracking placental development in health and disease. Nat. Rev. Endocrinol. 16, 479–494 (2020).
Burton, G. J., Fowden, A. L. & Thornburg, K. L. Placental origins of chronic disease. Physiol. Rev. 96, 1509–1565 (2016).
Li, Q., Wu, H., Wang, Y. & Wang, H. Current understanding in deciphering trophoblast cell differentiation during human placentation. Biol. Reprod. 107, 317–326 (2022).
Maltepe, E. & Fisher, S. J. Placenta: the forgotten organ. Annu. Rev. Cell Dev. Biol. 31, 523–552 (2015).
Xiao, Z., Yan, L., Liang, X. & Wang, H. Progress in deciphering trophoblast cell differentiation during human placentation. Curr. Opin. Cell Biol. 67, 86–91 (2020).
Villee, D. B. Development of endocrine function in the human placenta and fetus (second of two parts). N. Engl. J. Med. 281, 533–541 (1969).
Burton, G. J. & Jauniaux, E. What is the placenta? Am. J. Obstet. Gynecol. 213, S6 (2015).
Couzin-Frankel, J. Mysteries of development. How does fetal environment influence later health? Science 340, 1160–1161 (2013).
Burton, G. J., Redman, C. W., Roberts, J. M. & Moffett, A. Pre-eclampsia: pathophysiology and clinical implications. BMJ 366, l2381 (2019).
Walker, J. J. Pre-eclampsia. Lancet 356, 1260–1265 (2000).
Colson, A., Sonveaux, P., Debieve, F. & Sferruzzi-Perri, A. N. Adaptations of the human placenta to hypoxia: opportunities for interventions in fetal growth restriction. Hum. Reprod. Update 27, 531–569 (2021).
Gong, S. et al. The RNA landscape of the human placenta in health and disease. Nat. Commun. 12, 2639 (2021).
Aplin, J. D. & Jones, C. J. P. Cell dynamics in human villous trophoblast. Hum. Reprod. Update 27, 904–922 (2021).
Aghababaei, M., Hogg, K., Perdu, S., Robinson, W. P. & Beristain, A. G. ADAM12-directed ectodomain shedding of E-cadherin potentiates trophoblast fusion. Cell Death Differ. 22, 1970–1984 (2015).
Burton, G. J. & Jones, C. J. Syncytial knots, sprouts, apoptosis, and trophoblast deportation from the human placenta. Taiwan J. Obstet. Gynecol. 48, 28–37 (2009).
Fogarty, N. M., Ferguson-Smith, A. C. & Burton, G. J. Syncytial knots (Tenney-Parker changes) in the human placenta: evidence of loss of transcriptional activity and oxidative damage. Am. J. Pathol. 183, 144–152 (2013).
Loukeris, K., Sela, R. & Baergen, R. N. Syncytial knots as a reflection of placental maturity: reference values for 20 to 40 weeks’ gestational age. Pediatr. Dev. Pathol. 13, 305–309 (2010).
Simpson, R. A., Mayhew, T. M. & Barnes, P. R. From 13 weeks to term, the trophoblast of human placenta grows by the continuous recruitment of new proliferative units: a study of nuclear number using the disector. Placenta 13, 501–512 (1992).
Lu, X. et al. Fine-tuned and cell-cycle-restricted expression of fusogenic protein syncytin-2 maintains functional placental syncytia. Cell Rep. 21, 1150–1159 (2017).
Martin, B. J. & Spicer, S. S. Ultrastructural features of cellular maturation and aging in human trophoblast. J. Ultrastruct. Res. 43, 133–149 (1973).
Ellery, P. M., Cindrova-Davies, T., Jauniaux, E., Ferguson-Smith, A. C. & Burton, G. J. Evidence for transcriptional activity in the syncytiotrophoblast of the human placenta. Placenta 30, 329–334 (2009).
Fogarty, N. M., Mayhew, T. M., Ferguson-Smith, A. C. & Burton, G. J. A quantitative analysis of transcriptionally active syncytiotrophoblast nuclei across human gestation. J. Anat. 219, 601–610 (2011).
Fogarty, N. M., Burton, G. J. & Ferguson-Smith, A. C. Different epigenetic states define syncytiotrophoblast and cytotrophoblast nuclei in the trophoblast of the human placenta. Placenta 36, 796–802 (2015).
Tsang, J. C. H. et al. Integrative single-cell and cell-free plasma RNA transcriptomics elucidates placental cellular dynamics. Proc. Natl Acad. Sci. USA 114, E7786–E7795 (2017).
Vento-Tormo, R. et al. Single-cell reconstruction of the early maternal-fetal interface in humans. Nature 563, 347–353 (2018).
Liu, Y. et al. Single-cell RNA-seq reveals the diversity of trophoblast subtypes and patterns of differentiation in the human placenta. Cell Res. 28, 819–832 (2018).
Wang, Q. et al. Single-cell transcriptional profiling reveals cellular and molecular divergence in human maternal-fetal interface. Sci. Rep. 12, 10892 (2022).
Suryawanshi, H. et al. A single-cell survey of the human first-trimester placenta and decidua. Sci. Adv. 4, eaau4788 (2018).
Pique-Regi, R. et al. Single cell transcriptional signatures of the human placenta in term and preterm parturition. eLife 8, e52004 (2019).
Shannon, M. J. et al. Cell trajectory modeling identifies a primitive trophoblast state defined by BCAM enrichment. Development 149, dev199840 (2022).
Jaremek, A., Jeyarajah, M. J., Jaju Bhattad, G. & Renaud, S. J. Omics approaches to study formation and function of human placental syncytiotrophoblast. Front. Cell Dev. Biol. 9, 674162 (2021).
Arutyunyan, A. et al. Spatial multiomics map of trophoblast development in early pregnancy. Nature 616, 143–151 (2023).
Wolf, F. A., Angerer, P. & Theis, F. J. SCANPY: large-scale single-cell gene expression data analysis. Genome Biol. 19, 15 (2018).
Zhang, K. et al. A single-cell atlas of chromatin accessibility in the human genome. Cell 184, 5985–6001 (2021).
Fornes, O. et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 48, D87–D92 (2020).
Caniggia, I. et al. Hypoxia-inducible factor-1 mediates the biological effects of oxygen on human trophoblast differentiation through TGF-β3. J. Clin. Invest. 105, 577–587 (2000).
Semenza, G. Signal transduction to hypoxia-inducible factor 1. Biochem. Pharm. 64, 993–998 (2002).
Suman, P., Malhotra, S. S. & Gupta, S. K. LIF-STAT signaling and trophoblast biology. JAKSTAT 2, e25155 (2013).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Welch, J. D. et al. Single-cell multi-omic integration compares and contrasts features of brain cell identity. Cell 177, 1873–1887 (2019).
Naumova, N. et al. Organization of the mitotic chromosome. Science 342, 948–953 (2013).
Zheng, H. & Xie, W. The role of 3D genome organization in development and cell differentiation. Nat. Rev. Mol. Cell Biol. 20, 535–550 (2019).
Granja, J. M. et al. Single-cell multiomic analysis identifies regulatory programs in mixed-phenotype acute leukemia. Nat. Biotechnol. 37, 1458–1465 (2019).
Liu, J. et al. Jointly defining cell types from multiple single-cell datasets using LIGER. Nat. Protoc. 15, 3632–3662 (2020).
Morabito, S. et al. Single-nucleus chromatin accessibility and transcriptomic characterization of Alzheimer’s disease. Nat. Genet. 53, 1143–1155 (2021).
Wang, Y. et al. LMNA determines nuclear morphology during syncytialization of human trophoblast stem cells. Front. Cell Dev. Biol. 10, 836390 (2022).
Okae, H. et al. Derivation of human trophoblast stem cells. Cell Stem Cell 22, 50–63 (2018).
Wasserstrom, L., Lengeler, K. B., Walther, A. & Wendland, J. Molecular determinants of sporulation in Ashbya gossypii. Genetics 195, 87–99 (2013).
Dos Santos, M. et al. Single-nucleus RNA-seq and FISH identify coordinated transcriptional activity in mammalian myofibers. Nat. Commun. 11, 5102 (2020).
Huppertz, B. IFPA award in placentology lecture: biology of the placental syncytiotrophoblast—myths and facts. Placenta 31, S75–S81 (2010).
Oxvig, C. The role of PAPP-A in the IGF system: location, location, location. J. Cell Commun. Signal. 9, 177–187 (2015).
Nuzzo, A. M. et al. JunB/cyclin-D1 imbalance in placental mesenchymal stromal cells derived from preeclamptic pregnancies with fetal-placental compromise. Placenta 35, 483–490 (2014).
Zou, Y. et al. Promotion of trophoblast invasion by lncRNA MVIH through inducing Jun-B. J. Cell Mol. Med. 22, 1214–1223 (2018).
Redman, C. W. G., Staff, A. C. & Roberts, J. M. Syncytiotrophoblast stress in preeclampsia: the convergence point for multiple pathways. Am. J. Obstet. Gynecol. 226, S907–S927 (2022).
Karvas, R. M. et al. Stem-cell-derived trophoblast organoids model human placental development and susceptibility to emerging pathogens. Cell Stem Cell 29, 810–825 (2022).
Sheridan, M. A. et al. Establishment and differentiation of long-term trophoblast organoid cultures from the human placenta. Nat. Protoc. 15, 3441–3463 (2020).
Io, S. et al. New era of trophoblast research: integrating morphological and molecular approaches. Hum. Reprod. Update 26, 611–633 (2020).
Wang, Y. et al. A single-cell characterization of human post-implantation embryos cultured in vitro delineates morphogenesis in primary syncytialization. Front. Cell Dev. Biol. 10, 835445 (2022).
Zemmour, D. et al. Single-cell gene expression reveals a landscape of regulatory T cell phenotypes shaped by the TCR. Nat. Immunol. 19, 291–301 (2018).
Dennis, G. Jr et al. DAVID: database for annotation, visualization, and integrated discovery. Genome Biol. 4, P3 (2003).
Schep, A. N., Wu, B., Buenrostro, J. D. & Greenleaf, W. J. chromVAR: inferring transcription-factor-associated accessibility from single-cell epigenomic data. Nat. Methods 14, 975–978 (2017).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Qiu, X. et al. Reversed graph embedding resolves complex single-cell trajectories. Nat. Methods 14, 979–982 (2017).
Fang, R. et al. Comprehensive analysis of single cell ATAC-seq data with SnapATAC. Nat. Commun. 12, 1337 (2021).
Granja, J. M. et al. ArchR is a scalable software package for integrative single-cell chromatin accessibility analysis. Nat. Genet. 53, 403–411 (2021).
Janssens, J. et al. Decoding gene regulation in the fly brain. Nature 601, 630–636 (2022).
Herrmann, C., Van de Sande, B., Potier, D. & Aerts, S. i-cisTarget: an integrative genomics method for the prediction of regulatory features and cis-regulatory modules. Nucleic Acids Res. 40, e114 (2012).
Bravo Gonzalez-Blas, C. et al. SCENIC+: single-cell multiomic inference of enhancers and gene regulatory networks. Nat. Methods 20, 1355–1367 (2023).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Wu, H., Wang, Y. & Wang, H. Generation of human trophoblast stem cell-dependent placental in vitro models. Methods Mol. Biol. https://doi.org/10.1007/7651_2022_463 (2022).
Trapnell, C. et al. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 32, 381–386 (2014).
Yu, X. et al. A selective WDR5 degrader inhibits acute myeloid leukemia in patient-derived mouse models. Sci. Transl. Med 13, eabj1578 (2021).
Wang, M. et al. wangmeijiao/Human_placenta_multi-omics_snRNA-seq_snATAC-seq: 2023.11.28 released (v0.1). Zenodo https://doi.org/10.5281/zenodo.10212321 (2023).
Acknowledgements
We are grateful to the donors and their families, whose generous contributions made this work possible. We extend our gratitude to N. Shyh-Chang, F. Guo and J. Guo from the Institute of Zoology, Chinese Academy of Sciences, for their invaluable bioinformatic expertise and insightful comments throughout the revision process. We thank Dr. Y. Shan, Nanfang Hospital, Southern Medical University, for assistance during the experimental validation. Financial support was provided by the National Key Technologies R&D Program of China (2021YFC2700302 and 2022YFA1103601), the National Natural Science Foundation of China-Major program (82192870), the Chinese Academy of Sciences (CAS) Project for Young Scientists in Basic Research (YSBR-012 to F.L.), the National Natural Science Foundation of China (82001562 to Z.X. and 31900602 to Y.L.) and the Strategic Collaborative Research Program of the Ferring Institute of Reproductive Medicine. We thank Wang lab members for their discussions.
Author information
Authors and Affiliations
Contributions
H.W., Z.X., F.L. and R.L. conceptualized this project and supervised the overall experiments. Y.L., R.S., J.Z., J.L., X.J., X.X. and F.L. performed isolation of the STB nuclei and snRNA-seq, snATAC-seq library construction and sequencing. D.L. performed the ChIP–seq experiments. M.W., Y.L. and F.L. designed and performed bioinformatics analyses with suggestions from H.W., Z.X., R.L. and Y.L. Y.L., R.S. J.L., X.X. and Y.W. performed verification experiments. H.W., Z.X. and Y.L. performed manuscript writing, review and editing with input and feedback from all authors. S.L. and X.Z. performed technical direction in confocal photography.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Genetics thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–13, Supplementary Notes 1–21, Supplementary Methods, Supplementary References and Supporting data for Supplementary Figs. 1, 4 and 9.
Supplementary Tables
Supplementary Tables 1–9.
Source data
Source Data Fig. 1
Gene expression matrix for visualization of stage-specific DEGs.
Source Data Fig. 2
smFISH staining of indicated marker genes (PAPPA, FLT1 and hCG) characterizes eSTB mature 1 and eSTB mature 2 in early pregnancy.
Source Data Fig. 3
Normalized gene expression matrix for comparison of in vivo and in vitro STB population transcriptional signature.
Source Data Fig. 3
smFISH staining of TFs (STAT5A and FOSL2) and representative genes (PAPPA and FLT1) of eSTB mature 1 and eSTB mature 2.
Source Data Fig. 4
Immunofluorescence images of trophoblast markers (CDH1 and hCG) in TO derived from hTSCs-BL and MITF-overexpressing hTSCs-BL.
Source Data Fig. 5
smFISH staining of indicated marker genes (PAPPA, FLT1 and hCG) characterizes lSTB mature 1 and lSTB mature 2 in late pregnancy.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, M., Liu, Y., Sun, R. et al. Single-nucleus multi-omic profiling of human placental syncytiotrophoblasts identifies cellular trajectories during pregnancy. Nat Genet 56, 294–305 (2024). https://doi.org/10.1038/s41588-023-01647-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-023-01647-w
- Springer Nature America, Inc.