Abstract
Insulators play a critical role in spatiotemporal gene regulation in animals. The evolutionarily conserved CCCTC-binding factor (CTCF) is required for insulator function in mammals, but not all of its binding sites act as insulators. Here we explore the sequence requirements of CTCF-mediated transcriptional insulation using a sensitive insulator reporter in mouse embryonic stem cells. We find that insulation potency depends on the number of CTCF-binding sites in tandem. Furthermore, CTCF-mediated insulation is dependent on upstream flanking sequences at its binding sites. CTCF-binding sites at topologically associating domain boundaries are more likely to function as insulators than those outside topologically associating domain boundaries, independently of binding strength. We demonstrate that insulators form local chromatin domain boundaries and weaken enhancer–promoter contacts. Taken together, our results provide genetic, molecular and structural evidence connecting chromatin topology to the action of insulators in the mammalian genome.





Similar content being viewed by others
Data availability
All next-generation sequencing data are available under GEO accession GSE153403. Raw images of multiplexed FISH experiments and raw FACS data for specific mESC colonies are available on request. Source data are provided with this paper.
Code availability
Code for analyses of multiplexed FISH data can be found on Github at https://github.com/epigen-UCSD/huang-natgen2021.
References
Hnisz, D., Day, D. S. & Young, R. A. Insulated neighborhoods: structural and functional units of mammalian gene control. Cell 167, 1188–1200 (2016).
Kellis, M. et al. Defining functional DNA elements in the human genome. Proc. Natl Acad. Sci. USA 111, 6131–6138 (2014).
Levine, M., Cattoglio, C. & Tjian, R. Looping back to leap forward: transcription enters a new era. Cell 157, 13–25 (2014).
West, A. G., Gaszner, M. & Felsenfeld, G. Insulators: many functions, many mechanisms. Genes Dev. 16, 271–288 (2002).
Geyer, P. K. & Corces, V. G. DNA position-specific repression of transcription by a Drosophila zinc finger protein. Genes Dev. 6, 1865–1873 (1992).
Recillas-Targa, F., Bell, A. C. & Felsenfeld, G. Positional enhancer-blocking activity of the chicken β-globin insulator in transiently transfected cells. Proc. Natl Acad. Sci. USA 96, 14354–14359 (1999).
Stief, A., Winter, D. M., Stratling, W. H. & Sippel, A. E. A nuclear DNA attachment element mediates elevated and position-independent gene activity. Nature 341, 343–345 (1989).
Gurudatta, B. V. & Corces, V. G. Chromatin insulators: lessons from the fly. Brief. Funct. Genomic Proteomic 8, 276–282 (2009).
Chung, J. H., Bell, A. C. & Felsenfeld, G. Characterization of the chicken β-globin insulator. Proc. Natl Acad. Sci. USA 94, 575–580 (1997).
Lobanenkov, V. V. et al. A novel sequence-specific DNA binding protein which interacts with three regularly spaced direct repeats of the CCCTC-motif in the 5'-flanking sequence of the chicken c-myc gene. Oncogene 5, 1743–1753 (1990).
Bell, A. C. & Felsenfeld, G. Methylation of a CTCF-dependent boundary controls imprinted expression of the Igf2 gene. Nature 405, 482–485 (2000).
Flavahan, W. A. et al. Insulator dysfunction and oncogene activation in IDH mutant gliomas. Nature 529, 110–114 (2016).
Katainen, R. et al. CTCF/cohesin-binding sites are frequently mutated in cancer. Nat. Genet. 47, 818–821 (2015).
Ohlsson, R., Renkawitz, R. & Lobanenkov, V. CTCF is a uniquely versatile transcription regulator linked to epigenetics and disease. Trends Genet. 17, 520–527 (2001).
Filippova, G. N. et al. An exceptionally conserved transcriptional repressor, CTCF, employs different combinations of zinc fingers to bind diverged promoter sequences of avian and mammalian c-myc oncogenes. Mol. Cell. Biol. 16, 2802–2813 (1996).
Lupianez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Shukla, S. et al. CTCF-promoted RNA polymerase II pausing links DNA methylation to splicing. Nature 479, 74–79 (2011).
Vostrov, A. A. & Quitschke, W. W. The zinc finger protein CTCF binds to the APBβ domain of the amyloid β-protein precursor promoter. Evidence for a role in transcriptional activation. J. Biol. Chem. 272, 33353–33359 (1997).
Zhang, X. et al. Fundamental roles of chromatin loop extrusion in antibody class switching. Nature 575, 385–389 (2019).
Guo, Y. et al. CTCF/cohesin-mediated DNA looping is required for protocadherin alpha promoter choice. Proc. Natl Acad. Sci. USA 109, 21081–21086 (2012).
Guo, Y. et al. CRISPR inversion of CTCF sites alters genome topology and enhancer/promoter function. Cell 162, 900–910 (2015).
Ghirlando, R. & Felsenfeld, G. CTCF: making the right connections. Genes Dev. 30, 881–891 (2016).
Phillips-Cremins, J. E. & Corces, V. G. Chromatin insulators: linking genome organization to cellular function. Mol. Cell 50, 461–474 (2013).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the X-inactivation centre. Nature 485, 381–385 (2012).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Nora, E. P. et al. Targeted degradation of CTCF decouples local insulation of chromosome domains from genomic compartmentalization. Cell 169, 930–944 (2017).
Luppino, J. M. et al. Cohesin promotes stochastic domain intermingling to ensure proper regulation of boundary-proximal genes. Nat. Genet. 52, 840–848 (2020).
Wutz, G. et al. Topologically associating domains and chromatin loops depend on cohesin and are regulated by CTCF, WAPL, and PDS5 proteins. EMBO J. 36, 3573–3599 (2017).
Alipour, E. & Marko, J. F. Self-organization of domain structures by DNA-loop-extruding enzymes. Nucleic Acids Res. 40, 11202–11212 (2012).
Davidson, I. F. et al. DNA loop extrusion by human cohesin. Science 366, 1338–1345 (2019).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Haarhuis, J. H. I. et al. The cohesin release factor WAPL restricts chromatin loop extension. Cell 169, 693–707 (2017).
Kim, Y., Shi, Z., Zhang, H., Finkelstein, I. J. & Yu, H. Human cohesin compacts DNA by loop extrusion. Science 366, 1345–1349 (2019).
Rao, S. S. P. et al. Cohesin loss eliminates all loop domains. Cell 171, 305–320 (2017).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Vian, L. et al. The energetics and physiological impact of cohesin extrusion. Cell 173, 1165–1178 (2018).
Wutz, G. et al. ESCO1 and CTCF enable formation of long chromatin loops by protecting cohesinSTAG1 from WAPL. eLife 9, e52091 (2020).
Brackley, C. A. et al. Nonequilibrium chromosome looping via molecular slip links. Phys. Rev. Lett. 119, 138101 (2017).
Barbieri, M. et al. Complexity of chromatin folding is captured by the strings and binders switch model. Proc. Natl Acad. Sci. USA 109, 16173–16178 (2012).
Bianco, S. et al. Polymer physics predicts the effects of structural variants on chromatin architecture. Nat. Genet. 50, 662–667 (2018).
Brackley, C. A., Taylor, S., Papantonis, A., Cook, P. R. & Marenduzzo, D. Nonspecific bridging-induced attraction drives clustering of DNA-binding proteins and genome organization. Proc. Natl Acad. Sci. USA 110, E3605–E3611 (2013).
Buckle, A., Brackley, C. A., Boyle, S., Marenduzzo, D. & Gilbert, N. Polymer simulations of heteromorphic chromatin predict the 3D folding of complex genomic loci. Mol. Cell 72, 786–797 (2018).
Conte, M. et al. Polymer physics indicates chromatin folding variability across single-cells results from state degeneracy in phase separation. Nat. Commun. 11, 3289 (2020).
Di Pierro, M., Zhang, B., Aiden, E. L., Wolynes, P. G. & Onuchic, J. N. Transferable model for chromosome architecture. Proc. Natl Acad. Sci. USA 113, 12168–12173 (2016).
Schwarzer, W. et al. Two independent modes of chromatin organization revealed by cohesin removal. Nature 551, 51–56 (2017).
Despang, A. et al. Functional dissection of the Sox9–Kcnj2 locus identifies nonessential and instructive roles of TAD architecture. Nat. Genet. 51, 1263–1271 (2019).
Gribnau, J., Hochedlinger, K., Hata, K., Li, E. & Jaenisch, R. Asynchronous replication timing of imprinted loci is independent of DNA methylation, but consistent with differential subnuclear localization. Genes Dev. 17, 759–773 (2003).
Li, Y. et al. CRISPR reveals a distal super-enhancer required for Sox2 expression in mouse embryonic stem cells. PLoS ONE 9, e114485 (2014).
Zhou, H. Y. et al. A Sox2 distal enhancer cluster regulates embryonic stem cell differentiation potential. Genes Dev. 28, 2699–2711 (2014).
Kentepozidou, E. et al. Clustered CTCF binding is an evolutionary mechanism to maintain topologically associating domains. Genome Biol. 21, 5 (2020).
Frith, M. C., Saunders, N. F., Kobe, B. & Bailey, T. L. Discovering sequence motifs with arbitrary insertions and deletions. PLoS Comput. Biol. 4, e1000071 (2008).
Nakahashi, H. et al. A genome-wide map of CTCF multivalency redefines the CTCF code. Cell Rep. 3, 1678–1689 (2013).
Xu, D. et al. Dynamic nature of CTCF tandem 11 zinc fingers in multivalent recognition of DNA as revealed by NMR spectroscopy. J. Phys. Chem. Lett. 9, 4020–4028 (2018).
Yin, M. et al. Molecular mechanism of directional CTCF recognition of a diverse range of genomic sites. Cell Res. 27, 1365–1377 (2017).
Yan, J. et al. Histone H3 lysine 4 monomethylation modulates long-range chromatin interactions at enhancers. Cell Res. 28, 204–220 (2018).
Fang, R. et al. Mapping of long-range chromatin interactions by proximity ligation-assisted ChIP-seq. Cell Res. 26, 1345–1348 (2016).
Mumbach, M. R. et al. HiChIP: efficient and sensitive analysis of protein-directed genome architecture. Nat. Methods 13, 919–922 (2016).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Bintu, B. et al. Super-resolution chromatin tracing reveals domains and cooperative interactions in single cells. Science 362 (2018).
Mateo, L. J. et al. Visualizing DNA folding and RNA in embryos at single-cell resolution. Nature 568, 49–54 (2019).
Wang, S. et al. Spatial organization of chromatin domains and compartments in single chromosomes. Science 353, 598–602 (2016).
Su, J. H., Zheng, P., Kinrot, S. S., Bintu, B. & Zhuang, X. Genome-scale imaging of the 3D organization and transcriptional activity of chromatin. Cell 182, 1641–1659 (2020).
Alexander, J. M. et al. Live-cell imaging reveals enhancer-dependent Sox2 transcription in the absence of enhancer proximity. eLife 8, e41769 (2019).
Jia, Z. et al. Tandem CTCF sites function as insulators to balance spatial chromatin contacts and topological enhancer-promoter selection. Genome Biol. 21, 75 (2020).
Cai, H. N. & Shen, P. Effects of cis arrangement of chromatin insulators on enhancer-blocking activity. Science 291, 493–495 (2001).
Muravyova, E. et al. Loss of insulator activity by paired Su(Hw) chromatin insulators. Science 291, 495–498 (2001).
Rhee, H. S. & Pugh, B. F. Comprehensive genome-wide protein-DNA interactions detected at single-nucleotide resolution. Cell 147, 1408–1419 (2011).
Benabdallah, N. S. et al. Decreased enhancer-promoter proximity accompanying enhancer activation. Mol. Cell 76, 473–484 (2019).
Hnisz, D., Shrinivas, K., Young, R. A., Chakraborty, A. K. & Sharp, P. A. A phase separation model for transcriptional control. Cell 169, 13–23 (2017).
Chen, H. et al. Dynamic interplay between enhancer-promoter topology and gene activity. Nat. Genet. 50, 1296–1303 (2018).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
van de Geijn, B., McVicker, G., Gilad, Y. & Pritchard, J. K. WASP: allele-specific software for robust molecular quantitative trait locus discovery. Nat. Methods 12, 1061–1063 (2015).
Juric, I. et al. MAPS: model-based analysis of long-range chromatin interactions from PLAC-seq and HiChIP experiments. PLoS Comput. Biol. 15, e1006982 (2019).
Durand, N. C. et al. Juicebox provides a visualization system for Hi-C contact maps with unlimited zoom. Cell Syst. 3, 99–101 (2016).
Acknowledgements
We are grateful for comments from members of the Ren laboratory. This study was supported by funding from the Ludwig Institute for Cancer Research and NIH (U54 DK107977 and UM1HG011585, to B.R., X.Z., M.H. and M.N., and 3U54DK107977-05S1 to B.R.). Y.Z. was supported by 1K99CA252020-01. X.Z. is a Howard Hughes Medical Institute Investigator.
Author information
Authors and Affiliations
Contributions
B.R. and H.H. conceived the study. B.R. supervised the study. H.H. performed insulator assays and related analysis. R.H. and M.Y. performed PLAC-seq and Hi-C experiments. I.J. and M.H. analyzed PLAC-seq data. M.T. performed western blot experiments. Y.Z. performed Hi-C analysis. Q.Z. and Y.H. performed chromatin tracing experiments with help from B.B. and X.Z. A.J., B.B., C.K., M.C., S.B., A.M.C. and M.N. analyzed chromatin tracing data. The manuscript was written by H.H. and B.R. with input from all coauthors.
Corresponding author
Ethics declarations
Competing interests
B.R. is a cofounder and consultant for Arima Genomics, Inc. and cofounder of Epigenome Technologies. X.Z. is a cofounder and consultant for Vizgen, Inc. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Genetics thanks Qiang Wu and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Genotyping mESC reporter cell lines.
a, Genotyping egfp and mcherry labeled Sox2 gene. Left, Sanger sequencing results for allele-specific PCR products. Allele-specific SNP is highlighted. Right, the construct of the clone and the SNP information used to distinguish the two alleles. The reverse primer was common, while the forward primer was allele-specific, matching with egfp and mcherry sequence, respectively. b-c, Genotyping the Insulator reporter and control cell lines. Left, Sanger sequencing and SNP information. Right, Construct of the clone and positions of PCR primers. The forward primer is specific to the inserted HyTK gene. b, insulator reporter cell line. c, Insulator control cell line.
Extended Data Fig. 2 Insulation features of CBSs from the Sox9-Kcnj2 TAD boundary.
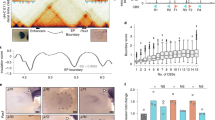
a, Hi-C contact map of the Sox9-Kcnj2 locus in mouse ES cells. Zoom in view shows the four CTCF binding sites cloned for insulator activity test. b, ChIP-seq of CTCF in the no insertion clone and the clone with an extra copy of the four Sox9-Kcnj2 TAD boundary CBS inserted inside the Sox2 domain. c, Reduction in Sox2-eGFP expression by one additional CBS. The comparison was between the clones presented in Fig. 2b. (0 CBS, n = 8; 1 CBS inside, n = 23; 2 CBS inside, n = 18; 3 CBS inside, n = 13; 4 CBS inside, n = 5; Data are mean ± sd). d, Allele-specific Sox2 expression in the no insertion clone (n = 2), the 4CBS clone (n = 3), and the 4CBS downstream clone (n = 2) as measured by RNA-seq. Sox2 expression from the CAST and 129 allele was represented by normalized read counts (rpkm) of the tagged egfp and mcherry gene, respectively. e, Relative Sox2 expression in the 4CBS and the 4CBS downstream clone in d measured by RNA-seq and FACS. The Sox2 expression from the egfp allele was first normalized to the mcherry allele, then compared to the no insertion clone. f, FACS profiling of the no insertion clone and the 4CBS clone. g, FACS profiling of GFPlow, GFPhigh sub-populations, and the unsort total population of the 4CBS insertion clone in f after extended culturing for 8 days. Left, GFP signal, right, mCherry signal from the same cells. h, ChIP-seq of H3K4me3 and H3K27ac in the no insertion clone and the 4CBS clone (n = 2). i, Allelic quantification of H3K27ac signal on the Sox2 super-enhancer of clones in h. H3K27ac ChIP-seq reads on the Sox2 super-enhancer were normalized by the total reads mapped to chromosome 3 for each allele.
Extended Data Fig. 3 Insulation effects of synthetic CTCF binding sites.
a, Additive insulation by synthetic CBS from boundary regions. Left top, compositions of one 139bp-CBS that was synthesized; Left bottom, tandemly arrayed 139bp-CBSs tested for insulator activity. Right, normalized Sox2-eGFP expression of clones with the tandemly arrayed 139bp-CBSs inserted between the Sox2 gene and its super-enhancer. Blue, CBS core motifs were in forward orientation; Red, CBS core motifs were in reverse orientation. Insertions were on the CAST allele only. n = 3, unpaired t-test, two-tailed. Data are mean ± sd. b, Insulation effects of PCR cloned large size CBSs (1–4 kb) and the synthesized 139bp-CBSs that contain the same CTCF motifs. (n = 12, paired t-test, two-tailed, ***P = 0.0007.). c, CTCF binding strength at selected boundary sites and non-boundary sites in mouse ES cells. ChIP-seq signals of CTCF are shown in 2-kb window. d, ChIP-seq of CTCF and Rad21 in clones with the insertion of six (nBd-syn6) or fifteen (nBd-syn15) 139-bp CBSs obtained from non-boundary regions. ChIP-seq reads were mapped to a customized mm10 genome that included the inserted sequence at the target site. Insertion position is highlighted in the red box.
Extended Data Fig. 4 ChIP-seq analysis of CTCF and cohesin binding at the synthetic insulators in various insulator reporter clones.
a, ChIP-seq signal tracks of CTCF in clones with the insertion of different synthetic CBS variants (n = 2). Each insertion consists of six CBSs that were tandemly arrayed in forward orientation (Supplementary Table 4). The insertion location is highlighted in the yellow box. b, CTCF binding strength (counts per million uniquely mapped reads) at the insertion location in the clones in (a). For each clone, ChIP-seq reads were mapped to a specific customized genome that contains the corresponding insertion in the Sox2 locus (n = 2; for bd core with bd adj, n = 4; for nbd core with bd adj n = 3.). c, ChIP-seq signal tracks of Rad21 in the same clones in (a) (n = 2). The insertion location is highlighted in the yellow box. d, Rad21 binding strength (counts per million uniquely mapped reads) at the insertion location in the clones in (c). For each clone, ChIP-seq reads were mapped to a specific customized genome that contains the corresponding insertion in the Sox2 locus (n = 2). e, Sequence alignment of the six boundary CBSs. Each CBS consists of 19-bp core motif plus 20-bp adjacent sequences on both sides. The color indicates the base frequency at each position. The CTCF motifs are highlighted in the red box.
Extended Data Fig. 5 Impact of CTCF ZF-9-11 deletion on transcriptional insulation by a synthetic insulator.
a, A schematic shows the experimental design to delete ZF9-11 of CTCF in mESCs. The exon 10, 11, partial of exon12, and an SV40 polyA signal were inserted into exon 9, resulting in the skip of exon9 in mRNA of the CTCF gene. b, PCR genotyping CTCF ΔZF9-11 mutant colonies. Genotyping primers spanned the insertion in exon 9. Highlighted in red boxes were homozygous mutant colonies evidenced by a single large-sized PCR fragment. PCR products from the homozygous mutant clones were further confirmed by Sanger sequencing. Genotyping of the homozygous mutants was repeated once with similar results. c, Western blot of CTCF in wild type and an exemplary CTCF ΔZF9-11 mutant clone. The primary antibody was the same one used for ChIP-seq (catalog: ab70303, lot GR3281212-7). Bottom, TBP loading control (primary antibody: sc-421, lot #B0304). Western blot was repeated once with similar results. d, Impact of CTCF zinc fingers 9–11 deletion on insulation effects of boundary CBSs with sixty-base-pair adjacent sequences. Left, eGFP profile of exemplary clones expressing wild-type and mutant CTCF protein; middle, mCherry profile of the same cells; right, normalized eGFP signal (eGFP/mCherry) of the wild-type and mutant clones. e, Impact of CTCF zinc fingers 9–11 deletion on insulation effects of boundary CBSs with ten-base-pair adjacent sequences. Left, eGFP profile of exemplary clones expressing wild-type and mutant CTCF protein; middle, mCherry profile of the same cells; right, normalized eGFP signal (eGFP/mCherry) of the wild-type and mutant clones.
Extended Data Fig. 6 Chromatin contacts at inserted CBSs.
a, K-R normalized Hi-C matrix (Observed/Expected) in the clone with the two forward Sox9-Kcnj2 TAD boundary CBSs inserted between the Sox2 promoter and super-enhancer (n = 2, replicates were merged). Hi-C reads were mapped to a customized chromosome 3 containing the insertion of the two forward CBSs. ChIP-seq signal of CTCF and orientations of the inserted CBSs and CBSs around the Sox2 promoter and super-enhancer were shown. The black arrow indicates the interactions between the inserted CBSs and the CBS on the Sox2 super-enhancer. b, Virtual 4C derived from Hi-C contacts in (a) at the viewpoint of the two inserted CBSs. Contacts were counted in each 5kb-bin. Contacts between the inserted CBSs and the Sox2 promoter or super-enhancer were compared to expected values, two-sided Poisson test. c, K-R normalized Hi-C matrix (Observed/Expected) in the clone with the four Sox9-Kcnj2 TAD boundary CBSs inserted between the Sox2 promoter and super-enhancer (n = 2, replicates were merged). Hi-C reads were mapped to a customized chromosome 3 containing the insertion of the four CBSs. ChIP-seq signal of CTCF and orientations of the inserted CBSs and CBSs around the Sox2 promoter and super-enhancer were shown. The black arrows indicate the interactions between the inserted CBSs and the CBS on the Sox2 promoter and super-enhancer. d, Virtual 4C derived from Hi-C contacts in (c) at the viewpoint of the two reverse-orientated CBSs inserted between the Sox2 promoter and super-enhancer. Contacts were counted in each 5kb-bin. Contacts between the two reverse CBSs and the Sox2 promoter or super-enhancer were compared to expected values, two-sided Poisson test.
Extended Data Fig. 7 Allele classification by multiplexed DNA FISH.
a-c, Bar plots showing detect efficiency of the 42 segments of chromatin tracing experiments in the ‘4CBS’ clone (a), the ‘4CBS mutant’ clone (b), and the ‘4CBS downstream’ clone (c). Detect efficiency of each segment was calculated as the fraction of chromosomes that showed a positive fluorescence signal at the specific imaging round. d, Exemplary images of allele classification. Left, nuclei segmentation and the positions of CAST and 129 allele in the nucleus. Right, images of the forty-two 5-kb segments (chr3:34,601,078-34,811,078) of the CAST and 129 allele. The hybridization probes of the 26th segment (highlighted in the red box) specifically targeted the 4CBS sequence. The chromosome positive for the 26th segment (inserted 4CBS) was classified as CAST allele, the negative chromosome in the same cell was classified as 129 allele. Cells with both chromosomes positive or both chromosomes negative for the 26th segment were discarded.
Extended Data Fig. 8 Spatial organization of the Sox2 locus in engineered mESCs.
a, Bulk Hi-C contact matrix (K-R normalized) of the Sox2 locus in the 4CBS clone. b, Median pairwise distance of the same Sox2 region measured by chromatin tracing experiment in the same clone in a, CAST and 129 chromosomes were combined. c, Correlation between the Hi-C contact frequency matrix (a) and median distance matrix(b). d, Normalized Sox2-eGFP expression in the no insertion clone(n = 8), the ‘4CBS’ clone (same cells in a-b, n = 2), and two insertion controls, ‘4CBS mutant’ (n = 3) and ‘4CBS downstream’ (n = 3). One-way analysis of variance with Bonferroni’s multiple comparisons test. Data are mean ± sd. e-f, Median spatial-distance matrix for the 210 kb Sox2 region (chr3: 34601078-34811078) of 129 (left) and CAST (right) chromosomes of the ‘4CBS mutant’ clone(e) and the ‘4CBS downstream clone’(f). The 26th segment was imaged by 4CBS specific probes in e. Similarly, the 38th segment was imaged by 4CBS specific probes in f. g-h, The probability of forming single-chromosome domain boundaries at each segment for the two alleles of the ‘4CBS mutant’ clone (g), and the ‘4CBS downstream’ clone (h). i, The distribution of single-chromosome insulation scores for each of the alleles between Sox2 promoter – 4CBSΔ insertion (segments 10–25) and 4CBSΔ insertion – Sox2 super-enhancer (segments 26–33), respectively. Two-sided Wilcoxon rank-sum test was performed. j, The distribution of single-chromosome insulation scores for each of the alleles between the same two domains (segment 10–25 and segment 26–33) in (i) for the ‘4CBS downstream’ clone. Insulation score was calculated in the same way as in (i). Two-sided Wilcoxon rank-sum test was performed.
Extended Data Fig. 9 Allele differences in median spatial distance.
a-c, Difference of the median distance matrices between the CAST and 129 allele of the ‘4CBS’ clone (a), the ‘4CBS mutant’ clone (b) and the ‘4CBS downstream’ clone(c).
Extended Data Fig. 10 Imaging of both nascent transcripts and chromatin structure at the Sox2 locus.
a, Example images of RNA FISH(left) and chromatin tracing(right) in the same cells. Circles indicate individual nuclei. RNA probes targeting Sox2, efgp, and mcherry are color-coded. Transcripts from the CAST allele are indicated by dots in purple pseudo color. Transcripts from the 129 allele are indicated by dots in cyan pseudo color. Arrows highlight examples of bursting alleles. b, Bursting frequencies of the CAST and 129 allele in the 4CBS clone and the control clone with 4CBS inserted downstream of the Sox2 SE. The numbers of bursting chromosomes and total chromosomes are indicated on the top of each bar. c, Median spatial distance matrices of the CAST and 129 allele in the 4CBS and the control clone (4CBS inserted downstream of the Sox2 super-enhancer). Multiplexed DNA FISH experiments were performed in the same cells following the RNA FISH experiments. d, Enhancer-promoter distances of the bursting and resting chromosomes in the 4CBS and the 4CBS downstream clone. A two-sided KS test between the distributions and a two-sided Wilcoxon test were performed.
Supplementary information
Supplementary Information
Supplementary Figs. 1 and 2 and unprocessed gel of Supplementary Fig. 1b
Supplementary Tables
Supplementary Tables 1–9
Source data
Source Data Extended Data Fig. 5
Unprocessed gels and western blots.
Rights and permissions
About this article
Cite this article
Huang, H., Zhu, Q., Jussila, A. et al. CTCF mediates dosage- and sequence-context-dependent transcriptional insulation by forming local chromatin domains. Nat Genet 53, 1064–1074 (2021). https://doi.org/10.1038/s41588-021-00863-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41588-021-00863-6
- Springer Nature America, Inc.
This article is cited by
-
Bento: a toolkit for subcellular analysis of spatial transcriptomics data
Genome Biology (2024)
-
3C methods in cancer research: recent advances and future prospects
Experimental & Molecular Medicine (2024)
-
Boundary stacking interactions enable cross-TAD enhancer–promoter communication during limb development
Nature Genetics (2024)
-
Spatially organized cellular communities form the developing human heart
Nature (2024)
-
Enhancer–promoter specificity in gene transcription: molecular mechanisms and disease associations
Experimental & Molecular Medicine (2024)