Abstract
Mitotic defects activate the spindle-assembly checkpoint, which inhibits the anaphase-promoting complex co-activator CDC20 to induce a prolonged cell cycle arrest1,2. Once errors are corrected, the spindle-assembly checkpoint is silenced, allowing anaphase onset to occur. However, in the presence of persistent unresolvable errors, cells can undergo ‘mitotic slippage’, exiting mitosis into a tetraploid G1 state and escaping the cell death that results from a prolonged arrest. The molecular logic that enables cells to balance these duelling mitotic arrest and slippage behaviours remains unclear. Here we demonstrate that human cells modulate the duration of their mitotic arrest through the presence of conserved, alternative CDC20 translational isoforms. Downstream translation initiation results in a truncated CDC20 isoform that is resistant to spindle-assembly-checkpoint-mediated inhibition and promotes mitotic exit even in the presence of mitotic perturbations. Our study supports a model in which the relative levels of CDC20 translational isoforms control the duration of mitotic arrest. During a prolonged mitotic arrest, new protein synthesis and differential CDC20 isoform turnover create a timer, with mitotic exit occurring once the truncated Met43 isoform achieves sufficient levels. Targeted molecular changes or naturally occurring cancer mutations that alter CDC20 isoform ratios or its translational control modulate mitotic arrest duration and anti-mitotic drug sensitivity, with potential implications for the diagnosis and treatment of human cancers.





Similar content being viewed by others
Data availability
The data supporting the findings of this study are available from the corresponding author on reasonable request. Source data are provided with this paper.
References
Musacchio, A. The molecular biology of spindle assembly checkpoint signaling dynamics. Curr. Biol. 25, R1002–R1018 (2015).
Lara-Gonzalez, P., Pines, J. & Desai, A. Spindle assembly checkpoint activation and silencing at kinetochores. Semin. Cell Dev. Biol. 117, 86–98 (2021).
Sivakumar, S. & Gorbsky, G. J. Spatiotemporal regulation of the anaphase-promoting complex in mitosis. Nat. Rev. Mol. Cell Biol. 16, 82–94 (2015).
Eichhorn, J. M., Sakurikar, N., Alford, S. E., Chu, R. & Chambers, T. C. Critical role of anti-apoptotic Bcl-2 protein phosphorylation in mitotic death. Cell Death Dis. 4, e834 (2013).
Li, M., York, J. P. & Zhang, P. Loss of Cdc20 causes a securin-dependent metaphase arrest in two-cell mouse embryos. Mol. Cell. Biol. 27, 3481–3488 (2007).
Lim, H. H., Goh, P. Y. & Surana, U. Cdc20 is essential for the cyclosome-mediated proteolysis of both Pds1 and Clb2 during M phase in budding yeast. Curr. Biol. 8, 231–234 (1998).
McKinley, K. L. & Cheeseman, I. M. Large-scale analysis of CRISPR/Cas9 cell-cycle knockouts reveals the diversity of p53-dependent responses to cell-cycle defects. Dev. Cell 40, 405–420 (2017).
GTEx Consortium. The Genotype-Tissue Expression (GTEx) pilot analysis: multitissue gene regulation in humans. Science 348, 648–660 (2015).
Hartwell, L. H., Mortimer, R. K., Culotti, J. & Culotti, M. Genetic control of the cell division cycle in yeast: V. Genetic analysis of cdc mutants. Genetics 74, 267–286 (1973).
Tsherniak, A. et al. Defining a cancer dependency map. Cell 170, 564–576 (2017).
Smith, L. M., Kelleher, N. L. & Consortium for Top Down Proteomics. Proteoform: a single term describing protein complexity. Nat. Methods 10, 186–187 (2013).
Hinnebusch, A. G. The scanning mechanism of eukaryotic translation initiation. Annu. Rev. Biochem. 83, 779–812 (2014).
Kozak, M. Initiation of translation in prokaryotes and eukaryotes. Gene 234, 187–208 (1999).
Kozak, M. Pushing the limits of the scanning mechanism for initiation of translation. Gene 299, 1–34 (2002).
Lischetti, T., Zhang, G., Sedgwick, G. G., Bolanos-Garcia, V. M. & Nilsson, J. The internal Cdc20 binding site in BubR1 facilitates both spindle assembly checkpoint signalling and silencing. Nat. Commun. 5, 5563 (2014).
Di Fiore, B. et al. The ABBA motif binds APC/C activators and is shared by APC/C substrates and regulators. Dev. Cell 32, 358–372 (2015).
Izawa, D. & Pines, J. The mitotic checkpoint complex binds a second CDC20 to inhibit active APC/C. Nature 517, 631–634 (2015).
Nilsson, J., Yekezare, M., Minshull, J. & Pines, J. The APC/C maintains the spindle assembly checkpoint by targeting Cdc20 for destruction. Nat. Cell Biol. 10, 1411–1420 (2008).
Zhang, S. et al. Molecular mechanism of APC/C activation by mitotic phosphorylation. Nature 533, 260–264 (2016).
Kimata, Y., Baxter, J. E., Fry, A. M. & Yamano, H. A role for the Fizzy/Cdc20 family of proteins in activation of the APC/C distinct from substrate recruitment. Mol. Cell 32, 576–583 (2008).
Ji, Z., Gao, H., Jia, L., Li, B. & Yu, H. A sequential multi-target Mps1 phosphorylation cascade promotes spindle checkpoint signaling. eLife 6, e22513 (2017).
Piano, V. et al. CDC20 assists its catalytic incorporation in the mitotic checkpoint complex. Science 371, 67–71 (2021).
Lara-Gonzalez, P., Kim, T., Oegema, K., Corbett, K. & Desai, A. A tripartite mechanism catalyzes Mad2-Cdc20 assembly at unattached kinetochores. Science 371, 64–67 (2021).
Zeng, X. et al. Pharmacologic inhibition of the anaphase-promoting complex induces a spindle checkpoint-dependent mitotic arrest in the absence of spindle damage. Cancer Cell 18, 382–395 (2010).
Visintin, R., Prinz, S. & Amon, A. CDC20 and CDH1: a family of substrate-specific activators of APC-dependent proteolysis. Science 278, 460–463 (1997).
Yudkovsky, Y., Shteinberg, M., Listovsky, T., Brandeis, M. & Hershko, A. Phosphorylation of Cdc20/fizzy negatively regulates the mammalian cyclosome/APC in the mitotic checkpoint. Biochem. Biophys. Res. Commun. 271, 299–304 (2000).
Labit, H. et al. Dephosphorylation of Cdc20 is required for its C-box-dependent activation of the APC/C. EMBO J. 31, 3351–3362 (2012).
Hein, J. B., Hertz, E. P. T., Garvanska, D. H., Kruse, T. & Nilsson, J. Distinct kinetics of serine and threonine dephosphorylation are essential for mitosis. Nat. Cell Biol. 19, 1433–1440 (2017).
Orth, J. D. et al. Quantitative live imaging of cancer and normal cells treated with Kinesin-5 inhibitors indicates significant differences in phenotypic responses and cell fate. Mol. Cancer Ther. 7, 3480–3489 (2008).
Varetti, G., Guida, C., Santaguida, S., Chiroli, E. & Musacchio, A. Homeostatic control of mitotic arrest. Mol. Cell 44, 710–720 (2011).
Lok, T. M. et al. Mitotic slippage is determined by p31comet and the weakening of the spindle-assembly checkpoint. Oncogene 39, 2819–2834 (2020).
Gascoigne, K. E. & Taylor, S. S. Cancer cells display profound intra- and interline variation following prolonged exposure to antimitotic drugs. Cancer Cell 14, 111–122 (2008).
Pestova, T. V. & Kolupaeva, V. G. The roles of individual eukaryotic translation initiation factors in ribosomal scanning and initiation codon selection. Genes Dev. 16, 2906–2922 (2002).
Ivanov, I. P., Loughran, G., Sachs, M. S. & Atkins, J. F. Initiation context modulates autoregulation of eukaryotic translation initiation factor 1 (eIF1). Proc. Natl Acad. Sci. USA 107, 18056–18060 (2010).
Partscht, P., Simon, A., Chen, N. P., Erhardt, S. & Schiebel, E. The HIPK2/CDC14B-MeCP2 axis enhances the spindle assembly checkpoint block by promoting cyclin B translation. Sci. Adv. 9, eadd6982 (2023).
Sloss, O., Topham, C., Diez, M. & Taylor, S. Mcl-1 dynamics influence mitotic slippage and death in mitosis. Oncotarget 7, 5176–5192 (2016).
Brito, D. A. & Rieder, C. L. Mitotic checkpoint slippage in humans occurs via cyclin B destruction in the presence of an active checkpoint. Curr. Biol. 16, 1194–1200 (2006).
Jordan, M. A. & Wilson, L. Microtubules as a target for anticancer drugs. Nat. Rev. Cancer 4, 253–265 (2004).
Weaver, B. A. How taxol/paclitaxel kills cancer cells. Mol. Biol. Cell 25, 2677–2681 (2014).
Huang, H. C., Shi, J., Orth, J. D. & Mitchison, T. J. Evidence that mitotic exit is a better cancer therapeutic target than spindle assembly. Cancer Cell 16, 347–358 (2009).
Cheng, B. & Crasta, K. Consequences of mitotic slippage for antimicrotubule drug therapy. Endocr. Relat. Cancer 24, T97–T106 (2017).
Kochetov, A. V. Alternative translation start sites and hidden coding potential of eukaryotic mRNAs. Bioessays 30, 683–691 (2008).
Lee, S. et al. Global mapping of translation initiation sites in mammalian cells at single-nucleotide resolution. Proc. Natl Acad. Sci. USA 109, E2424–E2432 (2012).
Van Damme, P., Gawron, D., Van Criekinge, W. & Menschaert, G. N-terminal proteomics and ribosome profiling provide a comprehensive view of the alternative translation initiation landscape in mice and men. Mol. Cell Proteom. 13, 1245–1261 (2014).
Chen, J. et al. Pervasive functional translation of noncanonical human open reading frames. Science 367, 1140–1146 (2020).
Cornelis, S. et al. Identification and characterization of a novel cell cycle-regulated internal ribosome entry site. Mol. Cell 5, 597–605 (2000).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Morgenstern, J. P. & Land, H. Advanced mammalian gene transfer: high titre retroviral vectors with multiple drug selection markers and a complementary helper-free packaging cell line. Nucleic Acids Res. 18, 3587–3596 (1990).
Qian, K. et al. A simple and efficient system for regulating gene expression in human pluripotent stem cells and derivatives. Stem Cells 32, 1230–1238 (2014).
Sliedrecht, T., Zhang, C., Shokat, K. M. & Kops, G. J. Chemical genetic inhibition of Mps1 in stable human cell lines reveals novel aspects of Mps1 function in mitosis. PLoS ONE 5, e10251 (2010).
Berg, S. et al. ilastik: interactive machine learning for (bio)image analysis. Nat. Methods 16, 1226–1232 (2019).
McQuin, C. et al. CellProfiler 3.0: next-generation image processing for biology. PLoS Biol. 16, e2005970 (2018).
Papadopoulos, J. S. & Agarwala, R. COBALT: constraint-based alignment tool for multiple protein sequences. Bioinformatics 23, 1073–1079 (2007).
Yates, A. D. et al. Ensembl 2020. Nucleic Acids Res. 48, D682–D688 (2020).
Cheeseman, I. M. & Desai, A. A combined approach for the localization and tandem affinity purification of protein complexes from metazoans. Sci. STKE 2005, pl1 (2005).
Acknowledgements
We thank K.-C. Su for his support, insights and guidance; K. McKinley for initial observations; E. Spooner for assistance with the mass spectrometry analysis; G. Bell for analysing the altORF conservation; and A. Amon, D. Bartel and the members of the Cheeseman laboratory for their input and suggestions. This work was supported by grants to IMC from the NIH/National Institute of General Medical Sciences (R35GM126930), National Science Foundation (2029868), the Gordon and Betty Moore Foundation, the American Cancer Society (Theory Lab Collaborative Grant (TLC-20-117-01-TLC)) and a Hope Funds for Cancer Research fellowship (HFCR-18-03-02) to M.-J.T.
Author information
Authors and Affiliations
Contributions
Conceptualization: M.-J.T. and I.M.C. Methodology: M.-J.T. Validation: M.-J.T. Investigation: M.-J.T. Writing—original draft preparation: M.-J.T. and I.M.C. Writing—review and editing: M.-J.T. and I.M.C. Visualization: M.-J.T. Supervision: I.M.C. Funding acquisition: M.-J.T. and I.M.C.
Corresponding author
Ethics declarations
Competing interests
A US patent (application number 17/878,774) has been filed on all aspects of this work by Whitehead Institute, on which I.M.C. and M.-J.T. are listed as inventors. The patent and initial disclosure have been filed, but the consideration is still pending.
Peer review
Peer review information
Nature thanks Jakob Nilsson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Human cells express alternative isoforms of Cdc20. Related to Fig. 1.
(A) Western blot showing mitotic HeLa cells collected by shake-off after overnight treatment with 330 nM nocodazole. Lysates alone, with buffer only, or with buffer and lambda phosphatase treatment were probed using antibodies recognizing the C-terminus of human Cdc20 (aa 450–499). GAPDH was used as loading control. (B) Western blot showing multiple human cancer cell lines (A549, DLD-1, and U2OS) and the non-transformed human hTERT RPE-1 cells treated with control or Cdc20 siRNAs, as well as the mouse NIH/3T3 cell line. Cdc20 protein species were detected with antibodies recognizing the C-terminus of human Cdc20 (aa 450–499). β-actin or GAPDH was used as loading control. (C) Sequence information for the homozygous ∆M1 mutant cell line lacking the canonical M1 ATG start codon. The DNA sequence of the genomic locus was determined by next-generation sequencing. (D) Sequence information for the M1-stop mutant cell line showing insertions of 53 nt and 105 nt respectively after the L14 residue. Underlined are premature stop codons that are in-frame with the M1 ATG start codon for both CDC20 alleles. The DNA sequence of the genomic locus was determined by next-generation sequencing. (E) Growth curves of control HeLa compared to the ∆M1 and M1-stop mutant cell lines. (F) Graph showing mitotic duration for control HeLa compared to the ∆M1 and M1-stop mutant cell lines. Each point represents a single cell. The bars correspond to the mean. Indicated are the mean mitotic duration ± standard deviation across two experimental replicates, with replicates shaded-coded. The total number of cells analysed is indicated. Statistics from Student’s two-sample t-Test with two-tailed distribution (****p < 0.0001, NS not significant).
Extended Data Fig. 2 Cdc20 isoforms are produced by alternative translation initiation at downstream in-frame start codons. Related to Fig. 1.
(A) Protein sequence of the N-terminal region of human Cdc20 with conserved motifs indicated. Methionine residues are highlighted in red. Arrow indicates the location of indel mutations in the CDC20_M1-fs-M43 cell line. (B) Cdc20 tryptic peptides with N-terminal acetylation indicative of translation initiation were identified following GFP immunoprecipitation-mass spectrometry of mitotically-enriched HeLa cells containing one allele of CDC20 with an endogenous C-terminal mEGFP tag. The identified peptide sequence, number of peptide-spectrum matches (#PSMs), and the cross-correlation (Xcorr) value from the SEQUEST search are indicated. Due to the presence of neighbouring arginine residues, we did not detect the Ac-M43 peptide in this tryptic digest. (C) Cdc20 peptides as in (B), except using the endopeptidase LysC alone and isolated using a C-terminal mEGFP tag in a cell line lacking the full-length Cdc20 protein (CDC20_M1-fs-M43) to maximize identification of alternative isoforms. (D) Analysis of human CDC20 nucleic acid sequence reveals two alternative out-of-frame start codons between Met1 and Met43. The amino acid sequence of the predicted alternative open reading frame (altORF) is indicated in blue, with the methionines bolded in cyan. Methionine residues are highlighted in red. (E) Conservation analysis for the presence of Cdc20 altORF across various mammalian species. The altORF was detected in 77 out of 83 species analysed. For each species, start and stop codons are mapped relative to human Cdc20 protein sequence and indicated in cyan and magenta respectively. (F) Western blot following GFP immunoprecipitation of mEGFP-Cdc20 fusions expressed in mitotically-enriched M1-stop cells. Antibodies against GFP, the C-terminus of Cdc20, the N-terminus of Cdc20, APC7, and Bub3 were used to detect the mEGFP-fusions, endogenous Cdc20, APC/C, and MCC respectively.
Extended Data Fig. 3 Translation initiation at alternative out-of-frame start codons in HeLa cells. Related to Fig. 1.
(A) Schematic illustrating the strategy to assess whether translation initiation occurs at the alternative out-of-frame start codons. Cdc20 protein indicated in black; altORF peptide indicated in blue and cyan. Cell lines with indel mutations after the alternative start codons at the endogenous locus in all CDC20 alleles disrupt translation of the full-length Cdc20 protein. Some indel mutations resulted in a frame shift that would be predicted to connect the altORF peptide with amino acid sequences encoding downstream regions of Cdc20. If the altORF is translated, this would produce a chimeric protein (altATG-Cdc20) that is shorter than full-length Cdc20, but is detectable with antibodies against the Cdc20 C-terminus. In contrast, insertions upstream of the alternative out-of-frame start codons, such as those in the M1-stop mutant, should only abrogated expression of the full-length Cdc20 protein without generating a chimeric altATG-Cdc20 protein. (B) Analysis of wild-type human CDC20 nucleic acid sequence reveals two alternative out-of-frame start codons between Met1 and Met43. The amino acid sequence of the predicted alternative open reading frame (altORF) is indicated below the nucleic acid sequence, with the methionines bolded. Sequence information is shown for representative clones with indel mutations where at least one CDC20 allele results in a frame shift that connects the altORF peptide with amino acid sequences encoding downstream regions of Cdc20. The DNA sequence of the genomic locus was determined by next-generation sequencing. Insertions are highlighted in red. When present, the amino acid sequence of the resulting altATG-Cdc20 peptide produced is shown. (C) Western blot showing mitotically-enriched control HeLa, M1-stop mutant, and representative clones with indel mutations where at least one CDC20 allele resulted in a frame shift that connects the altORF peptide with amino acid sequences encoding downstream regions of Cdc20. Endogenous Cdc20 protein was detected using antibodies recognizing the C-terminus of human Cdc20 (aa 450–499). β-actin was used as loading control.
Extended Data Fig. 4 Truncated Cdc20 isoforms are inefficient targets of the SAC and promote mitotic slippage. Related to Fig. 2.
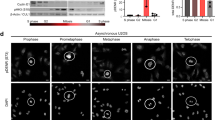
(A) Cumulative frequency distribution showing the fraction of mitotic cells over time post-mitotic entry for HeLa, ∆M1, and M1-stop cells treated with 10 µM STLC alone or in combination with the APC/C-inhibitor proTAME (12 µM). The total number of cells included in the analyses are indicated in brackets. (B) Cumulative frequency distribution as in (A) for HeLa, ∆M1, and M1-stop cells treated with 10 µM STLC and 100nM of either control siRNAs or Cdh1 siRNAs. (C) Cumulative frequency distribution as in (A) for HeLa, ∆M1, and M1-stop cells treated with 10 µM STLC and either expressing endogenous Cdc20 protein and treated with control siRNAs or upon Cdc20 replacement with ectopic wild-type CDC20 cDNA. (D) Mitotic duration of individual HeLa cells expressing doxycycline-inducible Cas9 and sgRNAs recognizing different regions within the CDC20 gene. Unperturbed mitotic progression or mitotic arrest behaviour were monitored upon treatment with DMSO or 10 µM STLC respectively. Cells entering mitosis in the first 350–400 min of time lapse experiments were included in the analyses. Open red circles indicate cells that exit mitosis. Closed black circles indicate cells that remained arrested in mitosis till the end of the time lapse. Bars correspond to the median. (E) Representative immunofluorescence images of Bub1 or Mad2 localization to kinetochores immuno-stained with anti-centromere antibodies (ACA). Images are maximum intensity projections of deconvolved Z-stacks of selected mitotic cells from control HeLa or mutant ∆M1 or M1-stop cell lines treated with nocodazole. Images were scaled individually to highlight kinetochore localization. Scale bar, 5 µm. (F) Cumulative frequency distribution as in (A) except for HeLa, ∆M1, and M1-stop cells treated with 10 µM STLC alone or in combination with the Mps1-inhibitor AZ3146 (4 µM). (G) Cumulative frequency distribution as in (A) except for HeLa, ∆M1, and M1-stop cells treated with 10 µM STLC and either control siRNAs or Mad2 siRNAs.
Extended Data Fig. 5 Loss of amino acid sequences and protein interactions within the Cdc20 N-terminus results in SAC defect. Related to Fig. 2.
(A) Protein sequence of the N-terminal region of human Cdc20 with conserved motifs and CDK phosphorylation sites indicated. Targeted mutations are shown in magenta. (B) Mitotic arrest duration in the presence of 10 µM STLC for individual HeLa cells in which the endogenous Cdc20 protein is replaced with either wild-type siRNA-resistant CDC20 cDNA or ∆M1 or Box1-Ala mutant constructs. Cells entering mitosis in the first 450 min of time lapse experiments were included in analyses. Open red circles indicate cells that exit mitosis. Closed black circles indicate cells that remained arrested in mitosis until the end of the time lapse. Bars correspond to median. Indicated are the median mitotic duration times across two experimental replicates, with replicates shaded-coded. The total number of cells analysed is indicated. Statistics are from a Mann-Whitney Test (**p < 0.01, ***p < 0.001, ****p < 0.0001). (C) Mitotic arrest duration analysis as in (B) comparing the wild-type Cdc20 construct with R132A or ∆M1 alone or ∆M1 R132A double mutant constructs. (D) Mitotic arrest duration analysis as in (B) comparing the wild-type Cdc20 construct with either ∆M1, S41A, or 4A mutant constructs.
Extended Data Fig. 6 Cdc20 translational isoforms modulate mitotic arrest duration. Related to Fig. 3.
(A) Mitotic arrest duration of individual HeLa or U2OS cells treated with 10 µM STLC alone or with a combination of STLC and siRNA treatment (either control siRNAs or Cdc20 siRNAs). Cells entering mitosis in the first 600 min (HeLa) or 700 min (U2OS) of time lapse experiments were included in analyses. Open red circles indicate cells that exit mitosis. Closed black circles indicate cells that remained arrested in mitosis till the end of the time lapse. Blue circles indicate cells that die in mitosis. Bars correspond to median. Indicated are the median mitotic duration times across two experimental replicates, with replicates shaded-coded. The total number of cells analysed is indicated. (B) Western blot of mitotically-enriched HeLa or U2OS cells expressing endogenous Cdc20 protein or upon Cdc20 replacement with either wild-type CDC20 cDNA or a Cdc20 M43L M88L mutant construct. Cells were enriched in mitosis with 10 µM STLC for 18 h. Cdc20 protein was detected using antibodies recognizing the human Cdc20 C-terminus (aa 450–499). β-actin was used as loading control. (C) Similar mitotic arrest duration analysis as in (A) except comparing HeLa cells in which the endogenous Cdc20 protein is replaced with siRNA-resistant CDC20(R132A) cDNA or a CDC20(M43L M88L R132A) mutant construct. NOTE: data for the CDC20(R132A) mutant is duplicated from Extended Data Fig. 5c. Statistics are from Mann-Whitney Test (*p < 0.05). (D) Competitive proliferation assay with long-term conditional Cdc20 replacement. Left, schematic of competitive proliferation assay. BFP-expressing HeLa cells with an ectopic siRNA-resistant CDC20(WT) construct were mixed 1:1 with mCherry-expressing HeLa cells with similar CDC20(WT) or CDC20(M43L M88L) constructs. Endogenous Cdc20 protein was depleted by siRNA transfection every 3-4 days for 24 days. Right, graph monitoring proliferation by flow cytometry analysis. The percentage of mCherry-positive cells at the indicated timepoint was quantified and normalized to the initial levels determined two days after the first siRNA transfection. (E) Western blot of single-thymidine synchronized HeLa cells expressing GFP alone or GFP-eIF1 post-release into medium with STLC for 16 h. Cdc20 isoform levels in serial diluted lysates were quantified for the full-length and M43 isoforms from short exposure and long exposure images respectively with antibodies recognizing the two Cdc20 isoforms. The levels in the undiluted HeLa sample expressing GFP alone (40 µg) were set to 1.
Extended Data Fig. 7 Cdc20 isoform levels change during a prolonged mitotic arrest to promote mitotic slippage. Related to Fig. 4.
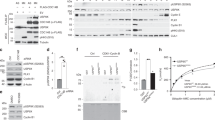
(A) Western blot showing HeLa cells that were synchronized using double-thymidine block and collected at different time points post-release with STLC treatment to induce a prolonged mitotic arrest. Cdc20 isoform levels were determined with antibodies recognizing the full-length and M43 isoforms. GAPDH was used as loading control. The ratio of full-length:M43 isoforms was quantified at the indicated time points using a LI-COR system. (B) Western blot of immunoprecipitated APC/C from mitotically-arrested HeLa cells collected by shake-off after treatment with STLC for 6 h (short mitotic arrest) or 16 h (long mitotic arrest). APC/C-bound Cdc20 proteins were detected using antibodies against the N-terminus of Cdc20 antibodies recognizing the full-length and M43 isoforms. (C) Western blot showing mitotically-arrested U2OS cells collected at various time points after addition of 50 µg/ml cycloheximide to inhibit new protein synthesis. Serial dilutions of the initial untreated cells were quantified for the full-length and M43 isoforms using a LI-COR system to determine the linear range of the N-terminal Cdc20 antibody recognizing the two Cdc20 isoforms. (D) Western blot as in (C) showing U2OS cells treated with 10 µM MG-132 alone to assess new protein synthesis or together with 50 µg/ml cycloheximide to inhibit new protein synthesis. Cdc20 isoform levels were quantified for the full-length and M43 isoform using a LI-COR system and GAPDH was used as loading control. The levels in the initial MG-132 only cells were set to 1. (E) Western blot as in (C) showing U2OS cells treated with 50 µg/ml cycloheximide alone or together with 10 µM MG-132 to inhibit proteosome-mediated degradation. (F) Representative phase contrast images of mitotically-arrested U2OS cells at 3 h post-cycloheximide addition or the corresponding untreated cells. NOTE: Larger field view of images in Fig. 4d. Scale bar, 100 µm.
Extended Data Fig. 8 Cdc20 translational isoform levels alter cancer cell anti-mitotic drug sensitivity. Related to Fig. 5.
(A) Sensitivity of HeLa or M1-stop cells to increasing concentrations of Taxol, Nocodazole, or the CENP-E inhibitor GSK923295. Cell viability was determined by MTT assay in triplicate following 72 h drug treatment. Data are mean ± s.e.m from three (Nocodazole) or four (Taxol, GSK923295) experimental replicates. Statistics from Student’s two-sample t-Test with two-tailed distribution comparing HeLa and M1-stop cell viabilities per drug concentration (*p < 0.05, **p < 0.01). (B) Mitotic arrest duration in the presence of 10 µM STLC for individual HeLa, ∆M1, or M1-stop cells expressing the wild-type CDC20 cDNA. Open red circles indicate cells that exit mitosis. Closed black circles indicate cells that remained arrested in mitosis till the end of the time lapse. Bars correspond to the median across two experimental replicates, with replicates shaded-coded. The total number of cells analysed is indicated. (C) Schematic illustrating the approach to isolate clonal cell lines from the polyclonal M1-stop mutant expressing the doxycycline-inducible wild-type CDC20 construct. Multiple clones were analysed to assess the correlation between the mitotic arrest behaviour of a given clone and the expression level of the integrated doxycycline-inducible CDC20 construct (see text for details). (D) Similar sensitivity assay as in (A) except for STLC for representative clones of M1-stop mutant with low, medium, or high expression of the doxycycline-inducible wild-type CDC20 construct without induction or induced with 20 ng/ml doxycycline. Data are mean ± s.e.m from three experimental replicates. Statistics from Student’s two-sample t-Test with two-tailed distribution comparing the uninduced and induced cell viabilities per drug concentration (*p < 0.05, **p < 0.01). (E) Similar mitotic arrest duration as in (B) except for representative clones of M1-stop mutant with low, medium, or high expression of the doxycycline-inducible wild-type CDC20 construct. Cells were treated with Cdc20 siRNAs to deplete endogenous truncated Cdc20 isoforms. Low Cdc20 expression from the inducible CDC20 construct fails to support mitotic progression even before addition of STLC. (F) Cumulative frequency distribution for the fraction of cells in mitosis at the indicated time after entry into mitosis (mitotic arrest duration) for HEC-6 cells treated with 10 µM STLC alone or together with 6 µM proTAME. The total number of cells analysed across two experimental replicates is indicated. (G) Mitotic duration in the presence of increasing concentrations of proTAME for individual HeLa or HEC-6 cells. Symbols are as described in (B). Indicated are the median mitotic duration times across two experimental replicates, with the total number of cells analysed and replicates shaded-coded. (H) Cumulative frequency distribution as in (F) for HEC-6 cells that had the endogenous Cdc20 protein replaced with either CDC20(∆M1) or CDC20(∆M1 M43L) mutant constructs.
Extended Data Fig. 9 Differential turnover of Cdc20 isoforms creates a mitotic arrest timer.
Diagram showing changing Cdc20 isoform ratios across the indicated conditions inferred from the data in the referenced experiments. We propose the changing levels of the Cdc20 isoforms in arrested cells occurs due to differential protein turnover creating a molecular timer to directly control the mitotic arrest duration. When the short M43 isoform reaches a ratio such that it comprises approximately one quarter of the total Cdc20 protein, it will be able to activate a sufficient portion of the APC/C to trigger mitotic exit even in the presence of continued SAC activation. The graph highlights multiple experiments conducted in this paper to alter Cdc20 isoform ratios or prevent new protein synthesis.
Supplementary information
Supplementary Information
Uncropped western blots for each figure (Supplementary Fig. 1); a list of the antibodies used with the corresponding experimental conditions (Supplementary Table 1); and a list of the cell lines used in the study (Supplementary Table 2).
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Tsang, MJ., Cheeseman, I.M. Alternative CDC20 translational isoforms tune mitotic arrest duration. Nature 617, 154–161 (2023). https://doi.org/10.1038/s41586-023-05943-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-05943-7
- Springer Nature Limited
This article is cited by
-
Escaping from CRISPR–Cas-mediated knockout: the facts, mechanisms, and applications
Cellular & Molecular Biology Letters (2024)
-
Functional analysis of Cdc20 reveals a critical role of CRY box in mitotic checkpoint signaling
Communications Biology (2024)
-
The two sides of chromosomal instability: drivers and brakes in cancer
Signal Transduction and Targeted Therapy (2024)
-
Weakened APC/C activity at mitotic exit drives cancer vulnerability to KIF18A inhibition
The EMBO Journal (2024)
-
An unexpected timer for cell division
Nature (2023)