Abstract
Intestinal health relies on the immunosuppressive activity of CD4+ regulatory T (Treg) cells1. Expression of the transcription factor Foxp3 defines this lineage, and can be induced extrathymically by dietary or commensal-derived antigens in a process assisted by a Foxp3 enhancer known as conserved non-coding sequence 1 (CNS1)2,3,4. Products of microbial fermentation including butyrate facilitate the generation of peripherally induced Treg (pTreg) cells5,6,7, indicating that metabolites shape the composition of the colonic immune cell population. In addition to dietary components, bacteria modify host-derived molecules, generating a number of biologically active substances. This is epitomized by the bacterial transformation of bile acids, which creates a complex pool of steroids8 with a range of physiological functions9. Here we screened the major species of deconjugated bile acids for their ability to potentiate the differentiation of pTreg cells. We found that the secondary bile acid 3β-hydroxydeoxycholic acid (isoDCA) increased Foxp3 induction by acting on dendritic cells (DCs) to diminish their immunostimulatory properties. Ablating one receptor, the farnesoid X receptor, in DCs enhanced the generation of Treg cells and imposed a transcriptional profile similar to that induced by isoDCA, suggesting an interaction between this bile acid and nuclear receptor. To investigate isoDCA in vivo, we took a synthetic biology approach and designed minimal microbial consortia containing engineered Bacteroides strains. IsoDCA-producing consortia increased the number of colonic RORγt-expressing Treg cells in a CNS1-dependent manner, suggesting enhanced extrathymic differentiation.




Similar content being viewed by others
Data availability
RNA-sequencing and 16S amplicon sequencing data are available under BioProject (https://www.ncbi.nlm.nih.gov/bioproject/) identification codes PRJNA600898 and PRJNA600979. Source data for Figs. 1–4 and Extended Data Figs. 1–8 are available as .xsl tables with the paper. Other relevant data are available from the corresponding authors upon reasonable request.
References
Josefowicz, S. Z. et al. Extrathymically generated regulatory T cells control mucosal TH2 inflammation. Nature 482, 395–399 (2012).
Zheng, Y. et al. Role of conserved non-coding DNA elements in the Foxp3 gene in regulatory T-cell fate. Nature 463, 808–812 (2010).
Nutsch, K. et al. Rapid and efficient generation of regulatory T cells to commensal antigens in the periphery. Cell Rep. 17, 206–220 (2016).
Kim, K. S. et al. Dietary antigens limit mucosal immunity by inducing regulatory T cells in the small intestine. Science 351, 858–863 (2016).
Arpaia, N. et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 504, 451–455 (2013).
Furusawa, Y. et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 504, 446–450 (2013).
Smith, P. M. et al. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 341, 569–573 (2013).
Ridlon, J. M., Kang, D. J. & Hylemon, P. B. Bile salt biotransformations by human intestinal bacteria. J. Lipid Res. 47, 241–259 (2006).
Ridlon, J. M., Harris, S. C., Bhowmik, S., Kang, D. J. & Hylemon, P. B. Consequences of bile salt biotransformations by intestinal bacteria. Gut Microbes 7, 22–39 (2016).
Wahlström, A., Sayin, S. I., Marschall, H. U. & Bäckhed, F. Intestinal crosstalk between bile acids and microbiota and its impact on host metabolism. Cell Metab. 24, 41–50 (2016).
Devlin, A. S. & Fischbach, M. A. A biosynthetic pathway for a prominent class of microbiota-derived bile acids. Nat. Chem. Biol. 11, 685–690 (2015).
Eyssen, H., De Pauw, G., Stragier, J. & Verhulst, A. Cooperative formation of omega-muricholic acid by intestinal microorganisms. Appl. Environ. Microbiol. 45, 141–147 (1983).
Eyssen, H. J., De Pauw, G. & Van Eldere, J. Formation of hyodeoxycholic acid from muricholic acid and hyocholic acid by an unidentified gram-positive rod termed HDCA-1 isolated from rat intestinal microflora. Appl. Environ. Microbiol. 65, 3158–3163 (1999).
Hamilton, J. P. et al. Human cecal bile acids: concentration and spectrum. Am. J. Physiol. Gastrointest. Liver Physiol. 293, G256–G263 (2007).
Hofmann, A. F. & Small, D. M. Detergent properties of bile salts: correlation with physiological function. Annu. Rev. Med. 18, 333–376 (1967).
Kretschmer, K. et al. Inducing and expanding regulatory T cell populations by foreign antigen. Nat. Immunol. 6, 1219–1227 (2005).
Haxhinasto, S., Mathis, D. & Benoist, C. The AKT-mTOR axis regulates de novo differentiation of CD4+Foxp3+ cells. J. Exp. Med. 205, 565–574 (2008).
Hang, S. et al. Bile acid metabolites control TH17 and Treg cell differentiation. Nature 576, 143–148 (2019).
Song, X. et al. Microbial bile acid metabolites modulate gut RORγ+ regulatory T cell homeostasis. Nature 577, 410–415 (2020).
Sefik, E. et al. Individual intestinal symbionts induce a distinct population of RORγ+ regulatory T cells. Science 349, 993–997 (2015).
Ohnmacht, C. et al. The microbiota regulates type 2 immunity through RORγt+ T cells. Science 349, 989–993 (2015).
Parks, D. J. et al. Bile acids: natural ligands for an orphan nuclear receptor. Science 284, 1365–1368 (1999).
Campbell, C. et al. Extrathymically generated regulatory T cells establish a niche for intestinal border-dwelling bacteria and affect physiologic metabolite balance. Immunity 48, 1245–1257 (2018).
White, J. et al. Two better cell lines for making hybridomas expressing specific T cell receptors. J. Immunol. 143, 1822–1825 (1989).
Hooijberg, E., Bakker, A. Q., Ruizendaal, J. J. & Spits, H. NFAT-controlled expression of GFP permits visualization and isolation of antigen-stimulated primary human T cells. Blood 96, 459–466 (2000).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Anders, S., Pyl, P. T. & Huber, W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31, 166–169 (2015).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Schmidt, J. et al. NSAIDs ibuprofen, indometacin, and diclofenac do not interact with farnesoid X receptor. Sci. Rep. 5, 14782 (2015).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl Acad. Sci. USA 108 (Suppl 1), 4516–4522 (2011).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Edgar, R. C. SINTAX: a simple non-Bayesian taxonomy classifier for 16S and ITS sequences. bioRxiv https://doi.org/10.1101/074161 (2016).
Cole, J. R. et al. Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 42, D633–D642 (2014).
Whitaker, W. R., Shepherd, E. S. & Sonnenburg, J. L. Tunable expression tools enable single-cell strain distinction in the gut microbiome. Cell 169, 538–546 (2017).
Acknowledgements
We thank F. Gonzalez (National Institutes of Health, NIH) for providing Nr1h4fl mice and F. Geissmann (MSKCC) for Csf1rcre mice. We thank J. Sonnenburg (Stanford University) for providing genetic tools to engineer B. theta. We thank S.-E. Brown, M. Rosenthal, T. Nguyen and P. Gonzales (Boehringer Ingelheim) and R. Pinedo (Weill Cornell Medical College) for assistance with germ-free mice. We thank O. Ouerfelli and the staff at the Organic Synthesis core at MSKCC for producing 6-oxoMCA. We thank A. Pickard (MKSCC) for assistance with metabolomic data analyses. We thank E. D. D’Andrea (University of Arizona) for assistance with DSF experiments. We thank J. van der Veeken for discussion of gene-expression data and all other members of the Rudensky laboratory for suggestions and technical assistance. This study was supported by NIH grant R37, the Ludwig Institute for Cancer Research, the Hilton Foundation, and Research Beyond Borders at Boehringer Ingelheim. W.G. and C.J.G were supported by NIH grant 1DP2HD101401-01. A.Y.R is an investigator with the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
C.C., P.T.M and A.Y.R. conceived the study, designed experiments and wrote the manuscript; C.C. and P.T.M performed experiments and analysed data; D.K. designed and cloned the original B. theta constructs; K.K. and C.M. provided technical assistance with experiments; M.S. and O.I.I. analysed gene-expression data sets; J.V. maintained germ-free mouse strains; C.-J.G. and W.-B.J. analysed bacterial bile acid transformation by mass spectrometry; S.V., R.J.R. and J.R.C. quantified SCFAs by mass spectrometry; J.H. and A.Y.R. supervised the study.
Corresponding authors
Ethics declarations
Competing interests
An invention disclosure has been filed based on the data generated in this study. P.T.M. and A.Y.R. received funding from Boehringer Ingelheim. A.Y.R. is a co-founder and member of the scientific advisory board of, and holds stock options in, Vedanta Biosciences. P.T.M. receives licensing royalties from Seres Therapeutics and is a co-inventor on patent applications US20170087196A1, US20180256653A1 and WO2018195467A1.
Additional information
Peer review information Nature thanks Richard Steven Blumberg and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Effects of iso- and oxo-bile acids on T cell differentiation and proliferation.
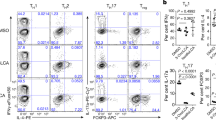
a, Effects of Treg cell-inducing bile acids on the in vitro generation of TH17 cells. Naive CD4+ T cells were activated by DCs in TH17-polarizing conditions (2 ng ml−1 TGF-β, 1 μg ml−1 CD3 antibody and 20 ng ml− IL-6). On day 3, cocultures were restimulated with phorbol myristate (PMA) and ionomycin in the presence of brefeldin A and monensin for 3 h before FACS analysis of IL-17 production. b, The 6β-OH group of ω-MCA is required for its Treg cell-inducing activity. Naive CD4+ T cells were activated by DCs in suboptimal Treg cell-inducing conditions (1 ng ml−1 TGF-β, 1 μg ml−1 CD3 antibody and 100 U ml−1 IL-2) and exposed to ω-MCA or 6-oxoMCA at the indicated concentrations. Foxp3 induction was assessed by FACS on day 3. c, Assessment of cell division in the presence of isoDCA and 3-oxoDCA (100 μM). Naive CD4+ T cells were labelled with Cell Trace Violet and activated with CD3/CD28 antibody-coated beads in the presence of TGF-β and IL-2 for three days before FACS analysis. Data shown are means ± s.d. of replicates (a–c, n = 3). Statistical significance determined by one-way (a, b) or two-way (c) ANOVA followed by a Dunnet’s (a) or Tukey’s (b, c) multiple comparison test. *P < 0.05, **P < 0.01, ***P < 0.001 versus vehicle; hash symbol, P < 0.05 vs ω-MCA (paired concentration); plus symbol, P < 0.05 versus isoDCA (paired concentration); ns, not significant. Data are representative of at least two independent experiments.
Extended Data Fig. 2 Characterization of mice with FXR deficiency in the myeloid compartment.
a–e, WT (Csf1rWT Nr1h4fl/fl) and DCΔFXR (Csf1rcre Nr1h4fl/fl) littermate mice were analysed between six to eight weeks of age. a, Gating strategy and b, quantification of conventional (c)DC1s (live CD45+ Lin− (dump: CD90, CD3; CD64−, Ly6C−, Siglec-F−) CD11c+ MHC class IIhi CD11b− XCR1+) and cDC2 (live CD45+ Lin− (dump: CD90, CD3; CD64−, Ly6C−, Siglec-F−) CD11c+ MHC class IIhi CD11b+ XCR1−) in the spleen (Spl), mesenteric lymph node (MLN) and LILP. c, Gating strategy for d, e. d, Number of total Foxp3+ Treg cells in the indicated organs. e, Quantification of RORγt+ Foxp3+ Treg cells in the LILP. Data shown are means ± s.d. (n = 5), representative of two independent cohorts of mice. Statistical significance determined by a two-tailed t-test. **P < 0.01.
Extended Data Fig. 3 Anti-inflammatory effects of isoDCA treatment on DCs.
a, DCs (1 × 105) were stimulated for 18 h with various TLR agonists (x-axes) in the presence or absence of 50 μM isoDCA. Levels of the indicated cytokines (TNF-α and IL-6) in the culture supernatant were determined by ELISA. b, DCs (1 × 104) were pulsed with ovalbumin (OVA, 1 mg ml−1) in the presence of various concentrations of isoDCA for 1 h in serum-free medium and allowed to process antigen for 4 h in complete medium before addition of an NFAT–GFP reporter cell line expressing the MHC-II-restricted OT-II TCR recognizing the ISQAVHAAHAEINEAGR peptide of OVA. The frequency of GFP+ cells was determined by FACS analysis after 24 h. Cocultures treated with CD3 antibody (1 μg ml−1) served as controls for DC-dependent, antigen-processing-independent effects of isoDCA on the activation of reporter cells. Activation with CD3/CD28 antibody beads in the presence of isoDCA served as a control for DC-independent effects on reporter gene expression. Shown are means ± s.d. of replicates in a and fold-change relative to vehicle (0 μM isoDCA) within each condition (OVA, CD3 or CD3/CD28 antibody-coated beads) in b. Statistical significance in a was determined by multiple t-tests using the Holm–Sidak correction method with α = 0.05. ****P < 0.001 versus vehicle. Statistical significance in b was determined by a two-way ANOVA followed by Dunnet’s multiple comparison’s test. *P < 0.05; ****P < 0.001 versus vehicle in each condition. Data are representative of three independent experiments.
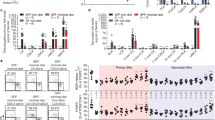
Extended Data Fig. 4 Liquid chromatography–mass spectrometry (LC–MS)-based analysis of isoDCA production by engineered B. theta strains.
Bacteria were grown to exponential phase and transferred to media containing DCA. Following incubation for 24 h, media was extracted with methanol and supernatants were analysed by liquid chromatography-mass spectrometry (LC–MS). Shown are traces for spike-in controls with DCA and isoDCA standards, and for media conditioned by B. thetaeWT, B. thetaeCD or the parental, unmanipulated B. thetaVPI strain VPI-5482. Data are representative of two independent experiments carried out in triplicate.
Extended Data Fig. 5 Analyses of microbial community composition in gnotobiotic and conventionalized mice.
GF mice were gavaged with WT or CD engineered consortia (C. scindens plus B. thetaeWT or C. scindens plus B. thetaeCD). Recipients of an FMT or noncolonized mice (PBS) served as references. The OTU composition of the caecal microbiota on day 10 post-colonization was determined by 16S sequencing. Shown are total read counts (left) and relative abundances (right) of bacteria in individual experimental mice, with data pooled from two independent experiments (n = 10).
Extended Data Fig. 6 Effects of isoDCA-producing consortia on colonic lymphocytes.
GF mice were gavaged with engineered consortia (C. scindens plus B. thetaeWT or C. scindens plus B. thetaeCD), PBS or a complex microbial community (FMT) as in Fig. 4b. a–g, Immune cell composition in the LILP was analysed by FACS on day 10 (D10; a–e) or day 30 (f, g) post-colonization. a, b, Frequencies of total Foxp3+ (a) and RORγt+ Foxp3+ (b) Treg cells among CD45+ cells. c, d, Frequency of RORγt+ Foxp3+ cells in the MLN (c) and small intestine lamina propria (SILP, d). e, Frequency of RORγt+ cells among Foxp3− CD4+ T cells (e, f) and Foxp3+ CD4+ T cells (g). Data shown are means ± s.d. (n = 10), pooled from two independent experiments. Statistical significance determined by one-way ANOVA followed by Tukey’s multiple comparison’s test. **P < 0.01; ***P < 0.001; ****P < 0.0001; ns, not significant.
Extended Data Fig. 7 IsoDCA production by engineered Bacteroides sp. strains.
a, Quantification of isoDCA production by engineered and reference strains in vitro. Bacteria were grown to exponential phase and transferred to medium containing DCA. Following incubation for 24 h, medium was extracted with methanol and supernatants were analysed by LC–MS. AUC, area under curve. b, c, GF mice were colonized with consortia containing either the engineered strain of B. frag capable of producing isoDCA or the catalytically dead mutant in combination with C. scindens (C. scindens plus B. frageWT and C. scindens plus B. frageCD, respectively). Recipients of an FMT and noncolonized mice (PBS) served as references. Immune-cell composition and isoDCA quantification were performed 10 days post-colonization. b, FACS analysis of the frequency of RORγt+ Foxp3+ CD4+ T cells in the LILP. c, Quantification of isoDCA in caecal contents. Faecal material was weighed, homogenized and extracted with methanol for LC–MS analysis. In a, c, the AUC is normalized by the weight of the input material. Shown are means ± s.d. (a, n = 3; b, n = 10; c, n = 5). Data in a, c are representative of two independent experiments. Data in b are pooled from two independent experiments. Statistical significance determined by a one-way ANOVA followed by Tukey’s multiple comparison’s test. *P < 0.05; ND, not detected; ns, not significant.
Extended Data Fig. 8 SCFA production by minimal, defined microbial consortia.
GF mice were colonized with consortia containing either the engineered strain of B. frag capable of producing isoDCA or the catalytically dead mutant in combination with C. scindens (C. scindens plus B. frageWT and C. scindens plus B. frageCD, respectively). Recipients of an FMT and noncolonized mice (PBS) served as references. Caecal content material was weighed, homogenized and subjected to organic solvent extraction for GC–MS-based quantification of SCFA levels. Shown are means ± s.d. (n = 6), with data pooled from two independent experiments. Statistical significance determined by a one-way ANOVA followed by Tukey’s multiple comparison’s test. ****P < 0.0001; ns, not significant.
Supplementary information
Rights and permissions
About this article
Cite this article
Campbell, C., McKenney, P.T., Konstantinovsky, D. et al. Bacterial metabolism of bile acids promotes generation of peripheral regulatory T cells. Nature 581, 475–479 (2020). https://doi.org/10.1038/s41586-020-2193-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-020-2193-0
- Springer Nature Limited
This article is cited by
-
Metabolic reprogramming in the tumor microenvironment of liver cancer
Journal of Hematology & Oncology (2024)
-
Cancer immunometabolism: advent, challenges, and perspective
Molecular Cancer (2024)
-
The microbial landscape of colorectal cancer
Nature Reviews Microbiology (2024)
-
Donor regulatory T cells rapidly adapt to recipient tissues to control murine acute graft-versus-host disease
Nature Communications (2024)
-
The maternal gut microbiome in pregnancy: implications for the developing immune system
Nature Reviews Gastroenterology & Hepatology (2024)