Abstract
Although dengue virus (DENV) infection typically causes asymptomatic disease, DENV-infected patients can experience severe complications. A risk factor for symptomatic disease is pre-existing anti-DENV IgG antibodies. Cellular assays suggested that these antibodies can enhance viral infection of Fcγ receptor (FcγR)-expressing myeloid cells. Recent studies, however, revealed more complex interactions between anti-DENV antibodies and specific FcγRs by demonstrating that modulation of the IgG Fc glycan correlates with disease severity. To investigate the in vivo mechanisms of antibody-mediated dengue pathogenesis, we developed a mouse model for dengue disease that recapitulates the unique complexity of human FcγRs. In in vivo mouse models of dengue disease, we discovered that the pathogenic activity of anti-DENV antibodies is exclusively mediated through engagement of FcγRIIIa on splenic macrophages, resulting in inflammatory sequelae and mortality. These findings highlight the importance of IgG–FcγRIIIa interactions in dengue, with important implications for the design of safer vaccination approaches and effective therapeutic strategies.





Similar content being viewed by others
Data availability
Raw data for all main and extended data figures are included in the paper as source files. Source data are provided with this paper.
References
Liu, S. Y. et al. A bibliometric analysis on dengue outbreaks in tropical and sub-tropical climates worldwide since 1950. Int. J. Environ. Res. Public Health https://doi.org/10.3390/ijerph18063197 (2021).
Dengue: Guidelines for Diagnosis, Treatment, Prevention and Control (World Health Organization, 2009).
Bhatt, S. et al. The global distribution and burden of dengue. Nature 496, 504–507 (2013).
Guzman, M. G. & Harris, E. Dengue. Lancet 385, 453–465 (2015).
Morens, D. M., Larsen, L. K. & Halstead, S. B. Study of the distribution of antibody-dependent enhancement determinants on dengue 2 isolates using dengue 2-derived monoclonal antibodies. J. Med. Virol. 22, 163–167 (1987).
Sangkawibha, N. et al. Risk factors in dengue shock syndrome: a prospective epidemiologic study in Rayong, Thailand. I. The 1980 outbreak. Am. J. Epidemiol. 120, 653–669 (1984).
Kliks, S. C., Nimmanitya, S., Nisalak, A. & Burke, D. S. Evidence that maternal dengue antibodies are important in the development of dengue hemorrhagic fever in infants. Am. J. Trop. Med. Hyg. 38, 411–419 (1988).
Thulin, N. K. et al. Maternal anti-dengue IgG fucosylation predicts susceptibility to dengue disease in infants. Cell Rep. 31, 107642 (2020).
Brown, M. G., King, C. A., Sherren, C., Marshall, J. S. & Anderson, R. A dominant role for FcgammaRII in antibody-enhanced dengue virus infection of human mast cells and associated CCL5 release. J. Leukoc. Biol. 80, 1242–1250 (2006).
Goncalvez, A. P., Engle, R. E., St Claire, M., Purcell, R. H. & Lai, C. J. Monoclonal antibody-mediated enhancement of dengue virus infection in vitro and in vivo and strategies for prevention. Proc. Natl Acad. Sci. USA 104, 9422–9427 (2007).
Kontny, U., Kurane, I. & Ennis, F. A. Gamma interferon augments Fc gamma receptor-mediated dengue virus infection of human monocytic cells. J. Virol. 62, 3928–3933 (1988).
Littaua, R., Kurane, I. & Ennis, F. A. Human IgG Fc receptor II mediates antibody-dependent enhancement of dengue virus infection. J. Immunol. 144, 3183–3186 (1990).
Yamanaka, A. et al. Antibody-dependent enhancement representing in vitro infective progeny virus titer correlates with the viremia level in dengue patients. Sci. Rep. 11, 12354 (2021).
Katzelnick, L. C. et al. Antibody-dependent enhancement of severe dengue disease in humans. Science 358, 929–932 (2017).
Bournazos, S. & Ravetch, J. V. Fcγ receptor pathways during active and passive immunization. Immunol. Rev. 268, 88–103 (2015).
Pincetic, A. et al. Type I and type II Fc receptors regulate innate and adaptive immunity. Nat. Immunol. 15, 707–716 (2014).
Bournazos, S. et al. Antibody fucosylation predicts disease severity in secondary dengue infection. Science 372, 1102–1105 (2021).
Wang, T. T. et al. IgG antibodies to dengue enhanced for FcγRIIIA binding determine disease severity. Science 355, 395–398 (2017).
Bournazos, S. IgG Fc receptors: evolutionary considerations. Curr. Top. Microbiol. Immunol. 423, 1–11 (2019).
Smith, P., DiLillo, D. J., Bournazos, S., Li, F. & Ravetch, J. V. Mouse model recapitulating human Fcγ receptor structural and functional diversity. Proc. Natl Acad. Sci. USA 109, 6181–6186 (2012).
Yauch, L. E. & Shresta, S. Mouse models of dengue virus infection and disease. Antivir. Res. 80, 87–93 (2008).
Kularatne, S. A. et al. Extensive haemorrhagic necrosis of liver is an unpredictable fatal complication in dengue infection: a postmortem study. BMC Infect. Dis. 14, 141 (2014).
Rathi, K. R. et al. Autopsy findings in fatal dengue haemorrhagic fever - 06 Cases. Med. J. Armed Forces India 69, 254–259 (2013).
Nimmerjahn, F. & Ravetch, J. V. Divergent immunoglobulin g subclass activity through selective Fc receptor binding. Science 310, 1510–1512 (2005).
Yamin, R. et al. Fc-engineered antibody therapeutics with improved anti-SARS-CoV-2 efficacy. Nature 599, 465–470 (2021).
Dejnirattisai, W. et al. A new class of highly potent, broadly neutralizing antibodies isolated from viremic patients infected with dengue virus. Nat. Immunol. 16, 170–177 (2015).
de Alwis, R. et al. Identification of human neutralizing antibodies that bind to complex epitopes on dengue virions. Proc. Natl Acad. Sci. USA 109, 7439–7444 (2012).
Chan, K. R. et al. Ligation of Fc gamma receptor IIB inhibits antibody-dependent enhancement of dengue virus infection. Proc. Natl Acad. Sci. USA 108, 12479–12484 (2011).
Bournazos, S., Corti, D., Virgin, H. W. & Ravetch, J. V. Fc-optimized antibodies elicit CD8 immunity to viral respiratory infection. Nature 588, 485–490 (2020).
Palermo, M. S., Alves Rosa, M. F., Van Rooijen, N. & Isturiz, M. A. Depletion of liver and splenic macrophages reduces the lethality of Shiga toxin-2 in a mouse model. Clin. Exp. Immunol. 116, 462–467 (1999).
Masood, K. I. et al. Role of TNF α, IL-6 and CXCL10 in Dengue disease severity. Iran J. Microbiol. 10, 202–207 (2018).
Nanda, J. D. et al. IL-18: the forgotten cytokine in dengue immunopathogenesis. J. Immunol. Res. 2021, 8214656 (2021).
Ngono, A. E. & Shresta, S. Immune response to dengue and Zika. Annu. Rev. Immunol. 36, 279–308 (2018).
Orozco, S. et al. Characterization of a model of lethal dengue virus 2 infection in C57BL/6 mice deficient in the alpha/beta interferon receptor. J. Gen. Virol. 93, 2152–2157 (2012).
Zellweger, R. M. & Shresta, S. Mouse models to study dengue virus immunology and pathogenesis. Front. Immunol. 5, 151 (2014).
Uciechowski, P. et al. IFN-gamma induces the high-affinity Fc receptor I for IgG (CD64) on human glomerular mesangial cells. Eur. J. Immunol. 28, 2928–2935 (1998).
Bardina, S. V. et al. Enhancement of Zika virus pathogenesis by preexisting antiflavivirus immunity. Science 356, 175–180 (2017).
Boonnak, K., Slike, B. M., Donofrio, G. C. & Marovich, M. A. Human FcγRII cytoplasmic domains differentially influence antibody-mediated dengue virus infection. J. Immunol. 190, 5659–5665 (2013).
Zellweger, R. M., Prestwood, T. R. & Shresta, S. Enhanced infection of liver sinusoidal endothelial cells in a mouse model of antibody-induced severe dengue disease. Cell Host Microbe 7, 128–139 (2010).
Casey, E. et al. A new mouse expressing human Fcγ receptors to better predict therapeutic efficacy of human anti-cancer antibodies. Leukemia 32, 547–549 (2018).
Bournazos, S., DiLillo, D. J., Goff, A. J., Glass, P. J. & Ravetch, J. V. Differential requirements for FcγR engagement by protective antibodies against Ebola virus. Proc. Natl Acad. Sci. USA 116, 20054–20062 (2019).
Weitzenfeld, P., Bournazos, S. & Ravetch, J. V. Antibodies targeting sialyl Lewis A mediate tumor clearance through distinct effector pathways. J. Clin. Invest. 129, 3952–3962 (2019).
DiLillo, D. J. & Ravetch, J. V. Differential Fc-receptor engagement drives an anti-tumor vaccinal effect. Cell 161, 1035–1045 (2015).
Santiago, G. A. et al. Analytical and clinical performance of the CDC real time RT-PCR assay for detection and typing of dengue virus. PLoS Negl. Trop. Dis. 7, e2311 (2013).
Smith, P., DiLillo, D. J., Bournazos, S., Li, F. & Ravetch, J. V. Mouse model recapitulating human Fcgamma receptor structural and functional diversity. Proc. Natl Acad. Sci. USA 109, 6181–6186 (2012).
Dekkers, G. et al. Multi-level glyco-engineering techniques to generate IgG with defined Fc-glycans. Sci. Rep. 6, 36964 (2016).
Kraus, A. A., Messer, W., Haymore, L. B. & de Silva, A. M. Comparison of plaque- and flow cytometry-based methods for measuring dengue virus neutralization. J. Clin. Microbiol. 45, 3777–3780 (2007).
Chauhan, A. et al. Splenectomy protects aged mice from injury after experimental stroke. Neurobiol. Aging 61, 102–111 (2018).
Acknowledgements
We thank P. Smith, E. Lam, R. Peraza, R. Francis and J. Edgar for technical assistance; all the members of the Laboratory of Molecular Genetics and Immunology (Rockefeller University) for discussions; S. Carrasco, S. St Jean and staff from the Laboratory of Comparative Pathology for histopathology support (with funding from NIH Core Grant P30CA008748); M. Mack (University Hospital Regensburg) for providing the anti-CCR2 antibody (clone MC-21); the CRISPR and Genome Editing Center at Rockefeller University for help with the design of the Ifnar1−/− KO mice; and The Rockefeller University for continued institutional support. The following reagents were obtained through BEI Resources, NIAID, NIH: Dengue Virus Type 2 (DENV2), New Guinea C (NGC) NR-84. Research reported in this publication was supported by National Institute of Allergy and Infectious Diseases Grants R01AI137276 (to S.B.), U19AI111825 (to J.V.R., C.M.R and S.B.) and R01AI124690 (to C.M.R.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH.
Author information
Authors and Affiliations
Contributions
R.Y., J.V.R. and S.B. conceptualized the project. R.Y. and K.S.K. developed the methodology. R.Y., K.S.K. and T.C. conducted investigations and acquired resources. R.Y., J.V.R. and S.B. wrote the paper. R.Y. and S.B. performed visualization. M.R.M. and C.M.R. provided intellectual input. J.V.R. and S.B. supervised the project. J.V.R. and S.B. acquired funding.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Microbiology thanks Leah Katzelnick and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 FcγR expression profile of engineered cell lines and assessment of in vitro ADE activity.
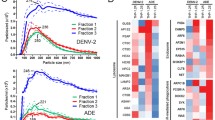
(a) FcγR expression on K562 was assessed by flow cytometry using antibodies against FcγRI, FcγRIIa, FcγRIIb, and FcγRIII (shaded histogram). Corresponding isotype control is shown in open histograms. (b,c) The in vitro ADE activity of the anti-DENV mAb C10 expressed as afucosylated (afuc) human IgG1 was assessed in U937 (black) vs U937-panFcγR (purple) cells. Percent (%) infection was assessed by flow cytometry. Area under curve (AUC) was calculated for each cell type and compared by two-tailed unpaired t-test. Figure shows one representative experiment (mean ± s.e.m) out of three performed in duplicates and normalized AUC from the three independent experiments. (d) The in vitro ADE activity of fucosylated (fuc) human IgG1, afucosylated (afuc) human IgG1, or Fc null variant was determined by multi-step viral yield on 6-, 15-, 24-, and 32-hours post-infection. Percent (%) infection was assessed by flow cytometry and compared between fucosylated and afucosylated hIgG1 by two-way ANOVA (Bonferroni post hoc analysis adjusted for multiple comparisons). Figure shows one representative experiment (mean ± s.e.m) performed in triplicates.
Extended Data Fig. 2 Characterization of Ifnar1−/−/FcγR KO and Ifnar1−/−/ FcγR humanized mice.
(a) Overview of Ifnar1 gene targeting with CRISPR/Cas9 in mice. Single guide RNA (sgRNA) was designed to target a sequence in exon 3 of the mouse Ifnar1 gene (marked in red). CRISPR-Cas9 constructs were injected to FcγR KO mouse or FcγR humanized mouse and resulted in 1 bp insertion or 1 bp deletion, respectively, that led to a premature stop codon. (b) IFNα/βR expression in various leucocyte populations (see gating strategy on the left) was assessed by flow cytometry in Ifnar1−/−/FcγR KO (grey) or Ifnar1−/−/FcγR humanized (blue) mice in comparison to FcγR humanized mice (red). (c) FcγR expression in various leucocyte populations in the blood was assessed by flow cytometry for Ifnar1−/−/FcγR KO mice (grey) and Ifnar1−/−/FcγR humanized mice (blue) and compared to FcγR expression in FcγR humanized mice. Matching isotype controls are indicated by open histograms. (d) Ifnar1−/−/FcγR KO mice were pre-treated with anti-DENV antibodies and infected with DENV. Pathological changes related to DENV infection were assessed on day 4 post-infection by H&E staining and histological evaluation of mesenteric lymph node (upper panel, 40x), lung (middle panel, 20x) and liver (lower panel, 20x). Images are representative of two infected mice.
Extended Data Fig. 3 Characterization of Fc domain variants with differential FcγR binding to assess in vivo ADE of dengue disease.
(a) Affinity of human IgG1 Fc domain variants for the various classes of human FcγRs and type II mouse FcRs. Numbers indicate the fold change in affinity compared with wild-type human IgG1. n.d.b.: no detectable binding. (b-d) Ifnar1−/−/FcγR humanized mice were administered i.v. with 20 μg or 200 μg of the ALIE variant of anti-DENV 2D22 mAb 8h before DENV2 challenge. Weight loss (b), survival (c), and platelet counts (d) were compared between the two doses by two-way ANOVA (Bonferroni post hoc analysis adjusted for multiple comparisons), log-rank (Mantel-Cox) test and two-tailed t-test, respectively. n = 6 mice per group in two independent experiments. Data are presented as mean ± s.e.m. *P<0.03; ***P<0.0008; ****P<0.0001. (e) Serum was obtained from DENV-infected mice (pre-treated with the different anti-DENV Fc variants) on day 3 post-infection and analyzed by ELISA to quantify antibody levels. IgG levels of the various Fc variants were compared to WT human IgG1 by two-way ANOVA (Bonferroni post hoc analysis adjusted for multiple comparisons). n = 7 (V11, ALIE, GAALIE), n = 8 (GA, afucosylated), n = 9 (GRLR), n = 11 (WT) mice per group from at least two independent experiments. Data are presented as mean ± s.e.m. (f) Ifnar1−/−/FcγR humanized mice were administered with 20 μg of anti-DENV mAb Fc-variants (GRLR or GAALIE) and platelet counts (mean ± s.e.m.) were measured 3 days post treatment and compared to mice that were treated with WT mAb by one-way ANOVA (Bonferroni post hoc analysis adjusted for multiple comparisons). n = 3 (WT, GRLR), n = 4 (GAALIE) mice per group from one independent experiment. NS, not significant.
Extended Data Fig. 4 Validation of antibody-mediated depletion of various cell populations.
(a-e) The efficiency of antibody-mediated depletion of NK cells, neutrophils, and the targeting of CCR2+ monocytes was determined by flow cytometry 2 days after antibody administration. The abundance of the depleted population in peripheral blood (for NK (n = 3), neutrophils (n = 3) and CCR2+ monocytes (n = 5)) or spleen (for CCR2+ monocytes (n = 5)) was calculated out of total CD45+ leucocytes and compared to matched isotype control by two-tailed unpaired t-test (one independent experiment). NK cells were defined as CD45+/CD11b−/CD3−/B220−/NK1.1+. Neutrophils were defined as CD45+/CD3−/CD11b+/SSChigh/Gr1+ (see Extended Data Fig. 2b for gating strategy). CCR2+ monocytes were defined as CD45+/CD19−/CD3−/NK1.1−/CD11b+/Ly6G−/Ly6C+. (f-i) Ifnar1−/−/FcγR humanized mice were treated with clodronate liposomes or control liposomes to deplete macrophages. Depletion of macrophages in the spleen and liver as well as efficiency of depletion on multiple myeloid subsets in the spleen (macrophages, neutrophils, and monocytes) was determined 2 days post treatment by flow cytometry. Macrophages (Mac) were defined as CD45+/CD19−/CD3−/SiglecF−/Gr1−/CD11b+/F4/80+. Red pulp macrophages (RPM) were defined as CD45+/CD19−/CD3−/SiglecF−/Gr1−/CD11blow/F4/80+. Neutrophils were defined as CD19−/CD3−/SiglecF−/SSChigh/Gr1high. Monocytes were defined as CD19−/CD3−/SiglecF−/SSClow/Gr1intermediate. Groups were compared by two-tailed unpaired t-test. n = 3 mice per group in one experiment. (j) To determine whether splenectomy affect platelet numbers in circulation, platelet counts were determined pre- and post- splenectomy and compared by two-tailed unpaired t-test (n = 5 mice per group in one experiment).
Extended Data Fig. 5 DENV infection in splenic macrophage populations.
(a) DENV infection (DENV+) of macrophage populations in the spleen of Ifnar1−/−/FcγR humanized mice that were pre-treated with the Fc-variants (GRLR, WT or ALIE) of the anti-DENV 2D22 was determined 4 days post infection. Representative flow cytometry histograms and quantification of each population are presented. Marginal zone (MZ) macrophages were defined as CD11b+/F4/80+/Tim4high. MZ metallophilic macrophages were defined as CD11b+/F4/80+/Tim4low. Red pulp macrophages were defined as CD11blow/F4/80+. Groups were compared by one-way ANOVA (Bonferroni post hoc analysis adjusted for multiple comparisons). n = 4 mice per group. In box plots, the center line shows the median, boxes represent the middle quartiles and whiskers show the range of values (minimum to maximum).
Supplementary information
Source data
Source Data Fig. 1
Source data.
Source Data Fig. 2
Source data.
Source Data Fig. 3
Source data.
Source Data Fig. 4
Source data.
Source Data Fig. 5
Source data.
Source Data Extended Data Fig. 1
Source data.
Source Data Extended Data Fig. 2
Source data.
Source Data Extended Data Fig. 4
Source data.
Source Data Extended Data Fig. 5
Source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yamin, R., Kao, K.S., MacDonald, M.R. et al. Human FcγRIIIa activation on splenic macrophages drives dengue pathogenesis in mice. Nat Microbiol 8, 1468–1479 (2023). https://doi.org/10.1038/s41564-023-01421-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41564-023-01421-y
- Springer Nature Limited
This article is cited by
-
Splenic macrophages escalate dengue disease
Nature Microbiology (2023)