Abstract
The Drosophila trachea, as the functional equivalent of mammalian blood vessels, senses hypoxia and oxygenates the body. Here, we show that the adult intestinal tracheae are dynamic and respond to enteric infection, oxidative agents and tumours with increased terminal branching. Increased tracheation is necessary for efficient damage-induced intestinal stem cell (ISC)-mediated regeneration and is sufficient to drive ISC proliferation in undamaged intestines. Gut damage or tumours induce HIF-1α (Sima in Drosophila), which stimulates tracheole branching via the FGF (Branchless (Bnl))–FGFR (Breathless (Btl)) signalling cascade. Bnl–Btl signalling is required in the intestinal epithelium and the trachea for efficient damage-induced tracheal remodelling and ISC proliferation. Chemical or Pseudomonas-generated reactive oxygen species directly affect the trachea and are necessary for branching and intestinal regeneration. Similarly, tracheole branching and the resulting increase in oxygenation are essential for intestinal tumour growth. We have identified a mechanism of tracheal–intestinal tissue communication, whereby damage and tumours induce neo-tracheogenesis in Drosophila, a process reminiscent of cancer-induced neoangiogenesis in mammals.








Similar content being viewed by others
Data availability
Source data are provided with this paper. All other data supporting the findings of this study are available from the corresponding authors upon reasonable request.
Code availability
No custom algorithms or software were generated for this study.
References
Ghabrial, A., Luschnig, S., Metzstein, M. M. & Krasnow, M. A. Branching morphogenesis of the Drosophila tracheal system. Annu. Rev. Cell Dev. Biol. 19, 623–647 (2003).
Hayashi, S. & Kondo, T. Development and function of the Drosophila tracheal system. Genetics 209, 367–380 (2018).
Jarecki, J., Johnson, E. & Krasnow, M. A. Oxygen regulation of airway branching in Drosophila is mediated by branchless FGF. Cell 99, 211–220 (1999).
Centanin, L. et al. Cell autonomy of HIF effects in Drosophila: tracheal cells sense hypoxia and induce terminal branch sprouting. Dev. Cell 14, 547–558 (2008).
Best, B. T. Single-cell branching morphogenesis in the Drosophila trachea. Dev. Biol. 451, 5–15 (2019).
Eilken, H. M. & Adams, R. H. Dynamics of endothelial cell behavior in sprouting angiogenesis. Curr. Opin. Cell Biol. 22, 617–625 (2010).
Fraisl, P., Mazzone, M., Schmidt, T. & Carmeliet, P. Regulation of angiogenesis by oxygen and metabolism. Dev. Cell 16, 167–179 (2009).
Apidianakis, Y., Tamamouna, V., Teloni, S. & Pitsouli, C. Chapter 5 - Intestinal stem cells: a decade of intensive research in Drosophila and the road ahead. Adv. Insect Physiol. (ed. Ligoxygakis, P.) vol. 52, 139–178 (Academic Press, Elsevier 2017).
Jiang, H., Tian, A. & Jiang, J. Intestinal stem cell response to injury: lessons from Drosophila. Cell Mol. Life Sci. 73, 3337–3349 (2016).
Lemaitre, B. & Miguel-Aliaga, I. The digestive tract of Drosophila melanogaster. Annu. Rev. Genet. 47, 377–404 (2013).
Jasper, H. Intestinal stem cell aging: origins and interventions. Annu. Rev. Physiol. 82, 203–226 (2020).
Micchelli, C. A. & Perrimon, N. Evidence that stem cells reside in the adult Drosophila midgut epithelium. Nature 439, 475–479 (2006).
Ohlstein, B. & Spradling, A. The adult Drosophila posterior midgut is maintained by pluripotent stem cells. Nature 439, 470–474 (2006).
Zeng, X. & Hou, S. X. Enteroendocrine cells are generated from stem cells through a distinct progenitor in the adult Drosophila posterior midgut. Development 142, 644–653 (2015).
Lin, G., Xu, N. & Xi, R. Paracrine Wingless signalling controls self-renewal of Drosophila intestinal stem cells. Nature 455, 1119–1123 (2008).
Xu, N. et al. EGFR, Wingless and JAK/STAT signaling cooperatively maintain Drosophila intestinal stem cells. Dev. Biol. 354, 31–43 (2011).
Li, Z., Zhang, Y., Han, L., Shi, L. & Lin, X. Trachea-derived Dpp controls adult midgut homeostasis in Drosophila. Dev. Cell 24, 133–143 (2013).
Miguel-Aliaga, I., Jasper, H. & Lemaitre, B. Anatomy and physiology of the digestive tract of Drosophila melanogaster. Genetics 210, 357–396 (2018).
Kux, K. & Pitsouli, C. Tissue communication in regenerative inflammatory signaling: lessons from the fly gut. Front. Cell Infect. Microbiol. 4, 49 (2014).
Jiang, H. et al. Cytokine/Jak/Stat signaling mediates regeneration and homeostasis in the Drosophila midgut. Cell 137, 1343–1355 (2009).
Buchon, N., Broderick, N. A., Poidevin, M., Pradervand, S. & Lemaitre, B. Drosophila intestinal response to bacterial infection: activation of host defense and stem cell proliferation. Cell Host Microbe 5, 200–211 (2009).
Apidianakis, Y., Pitsouli, C., Perrimon, N. & Rahme, L. Synergy between bacterial infection and genetic predisposition in intestinal dysplasia. Proc. Natl Acad. Sci. USA 106, 20883–20888 (2009).
Amcheslavsky, A., Jiang, J. & Ip, Y. T. Tissue damage-induced intestinal stem cell division in Drosophila. Cell Stem Cell 4, 49–61 (2009).
Biteau, B., Hochmuth, C. E. & Jasper, H. JNK activity in somatic stem cells causes loss of tissue homeostasis in the aging Drosophila gut. Cell Stem Cell 3, 442–455 (2008).
Markstein, M. et al. Systematic screen of chemotherapeutics in Drosophila stem cell tumors. Proc. Natl Acad. Sci. USA 111, 4530–4535 (2014).
Linneweber, G. A. et al. Neuronal control of metabolism through nutrient-dependent modulation of tracheal branching. Cell 156, 69–83 (2014).
Campbell, K. et al. Collective cell migration and metastases induced by an epithelial-to-mesenchymal transition in Drosophila intestinal tumors. Nat. Commun. 10, 2311 (2019).
Brand, A. H. & Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118, 401–415 (1993).
Shiga, Y., Tanaka-Matakatsu, M. & Hayashi, S. A nuclear GFP/β-galactosidase fusion protein as a marker for morphogenesis in living Drosophila. Dev. Growth Differ. 38, 99–106 (1996).
Potter, C. J., Tasic, B., Russler, E. V., Liang, L. & Luo, L. The Q system: a repressible binary system for transgene expression, lineage tracing, and mosaic analysis. Cell 141, 536–548 (2010).
Sutherland, D., Samakovlis, C. & Krasnow, M. A. branchless encodes a Drosophila FGF homolog that controls tracheal cell migration and the pattern of branching. Cell 87, 1091–1101 (1996).
Grifoni, D., Sollazzo, M., Fontana, E., Froldi, F. & Pession, A. Multiple strategies of oxygen supply in Drosophila malignancies identify tracheogenesis as a novel cancer hallmark. Sci. Rep. 5, 9061 (2015).
Wang, C. W., Purkayastha, A., Jones, K. T., Thaker, S. K. & Banerjee, U. In vivo genetic dissection of tumor growth and the Warburg effect. eLife 5, e18126 (2016).
Kaelin, W. G. Jr. The von Hippel–Lindau protein, HIF hydroxylation, and oxygen sensing. Biochem. Biophys. Res. Commun. 338, 627–638 (2005).
Klimova, T. & Chandel, N. S. Mitochondrial complex III regulates hypoxic activation of HIF. Cell Death Differ. 15, 660–666 (2008).
Semenza, G. L. Hypoxia-inducible factors: coupling glucose metabolism and redox regulation with induction of the breast cancer stem cell phenotype. EMBO J. 36, 252–259 (2017).
Movafagh, S., Crook, S. & Vo, K. Regulation of hypoxia-inducible factor-1a by reactive oxygen species: new developments in an old debate. J. Cell Biochem. 116, 696–703 (2015).
Lavista-Llanos, S. et al. Control of the hypoxic response in Drosophila melanogaster by the basic helix-loop-helix PAS protein Similar. Mol. Cell Biol. 22, 6842–6853 (2002).
Centanin, L., Ratcliffe, P. J. & Wappner, P. Reversion of lethality and growth defects in Fatiga oxygen-sensor mutant flies by loss of hypoxia-inducible factor-α/Sima. EMBO Rep. 6, 1070–1075 (2005).
Majmundar, A. J., Wong, W. J. & Simon, M. C. Hypoxia-inducible factors and the response to hypoxic stress. Mol. Cell 40, 294–309 (2010).
Tamamouna, V. & Pitsouli, C. The hypoxia-inducible factor-1α in angiogenesis and cancer: insights from the Drosophila model. Gene Expression and Regulation in Mammalian Cells—Transcription Toward the Establishment of Novel Therapeutics (ed. Uchiumi, F.) 209–241 (IntechOpen, 2018).
Deziel, E. et al. The contribution of MvfR to Pseudomonas aeruginosa pathogenesis and quorum sensing circuitry regulation: multiple quorum sensing-regulated genes are modulated without affecting lasRI, rhlRI or the production of N-acyl-l-homoserine lactones. Mol. Microbiol. 55, 998–1014 (2005).
Xiao, G. et al. MvfR, a key Pseudomonas aeruginosa pathogenicity LTTR-class regulatory protein, has dual ligands. Mol. Microbiol. 62, 1689–1699 (2006).
Liberati, N. T. et al. An ordered, nonredundant library of Pseudomonas aeruginosa strain PA14 transposon insertion mutants. Proc. Natl Acad. Sci. USA 103, 2833–2838 (2006).
Lee, K. A. et al. Bacterial-derived uracil as a modulator of mucosal immunity and gut-microbe homeostasis in Drosophila. Cell 153, 797–811 (2013).
Hochmuth, C. E., Biteau, B., Bohmann, D. & Jasper, H. Redox regulation by Keap1 and Nrf2 controls intestinal stem cell proliferation in Drosophila. Cell Stem Cell 8, 188–199 (2011).
Patel, P. H. et al. Damage sensing by a Nox–Ask1–MKK3–p38 signaling pathway mediates regeneration in the adult Drosophila midgut. Nat. Commun. 10, 4365 (2019).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Zhai, Z. et al. Accumulation of differentiating intestinal stem cell progenies drives tumorigenesis. Nat. Commun. 6, 10219 (2015).
McGuire, S. E., Mao, Z. & Davis, R. L. Spatiotemporal gene expression targeting with the TARGET and gene-switch systems in Drosophila. Sci. STKE 2004, pI6 (2004).
Buchon, N. et al. Morphological and molecular characterization of adult midgut compartmentalization in Drosophila. Cell Rep. 3, 1725–1738 (2013).
Marianes, A. & Spradling, A. C. Physiological and stem cell compartmentalization within the Drosophila midgut. eLife 2, e00886 (2013).
Potente, M., Gerhardt, H. & Carmeliet, P. Basic and therapeutic aspects of angiogenesis. Cell 146, 873–887 (2011).
Trinchieri, G. Cancer and inflammation: an old intuition with rapidly evolving new concepts. Annu. Rev. Immunol. 30, 677–706 (2012).
Folkman, J. Tumor angiogenesis: therapeutic implications. N. Engl. J. Med. 285, 1182–1186 (1971).
Folkman, J. Fundamental concepts of the angiogenic process. Curr. Mol. Med. 3, 643–651 (2003).
Weis, S. M. & Cheresh, D. A. Tumor angiogenesis: molecular pathways and therapeutic targets. Nat. Med. 17, 1359–1370 (2011).
Choudhry, H. & Harris, A. L. Advances in hypoxia-inducible factor biology. Cell Metab. 27, 281–298 (2018).
Lee, P., Chandel, N. S. & Simon, M. C. Cellular adaptation to hypoxia through hypoxia inducible factors and beyond. Nat. Rev. Mol. Cell Biol. 6, 268–283 (2020).
Takenaga, K. Angiogenic signaling aberrantly induced by tumor hypoxia. Front Biosci. (Landmark Ed.) 16, 31–48 (2011).
Perochon, J., Yu, Y., Aughey, G. N., Southall, T. D. & Cordero, J. B. Dynamic adult tracheal plasticity drives stem cell adaptation to changes in intestinal homeostasis. Nat. Cell Bio. (2021).
Ohshiro, T. & Saigo, K. Transcriptional regulation of breathless FGF receptor gene by binding of TRACHEALESS/dARNT heterodimers to three central midline elements in Drosophila developing trachea. Development 124, 3975–3986 (1997).
Santabarbara-Ruiz, P. et al. ROS-induced JNK and p38 signaling is required for unpaired cytokine activation during Drosophila regeneration. PLoS Genet. 11, e1005595 (2015).
André-Lévigne, D., Modarressi, A., Pepper, M. S. & Pittet-Cuénod, B. Reactive oxygen species and NOX enzymes are emerging as key players in cutaneous wound repair. Int. J. Mol. Sci. 18, 2149 (2017).
Jia, Y.-T. et al. Activation of p38 MAPK by reactive oxygen species is essential in a rat model of stress-induced gastric mucosal injury. J. Immunol. 179, 7808–7819 (2007).
Warren, C. M., Ziyad, S., Briot, A., Der, A. & Iruela-Arispe, M. L. A ligand-independent VEGFR2 signaling pathway limits angiogenic responses in diabetes. Sci. Signal. 7, ra1 (2014).
Nezu, M. et al. Nrf2 inactivation enhances placental angiogenesis in a preeclampsia mouse model and improves maternal and fetal outcomes. Sci. Signal 10, eaam5711 (2017).
Reczek, C. & Chandel, N. The two faces of reactive oxygen species in cancer. Annu. Rev. Cancer Biol. 1, 79–98 (2017).
Perez, E., Lindblad, J. L. & Bergmann, A. Tumor-promoting function of apoptotic caspases by an amplification loop involving ROS, macrophages and JNK in Drosophila. eLife 6, e26747 (2017).
Ha, E.-M. et al. An antioxidant system required for host protection against gut infection in Drosophila. Dev. Cell 8, 125–132 (2005).
Lee, W. J. & Brey, P. T. How microbiomes influence metazoan development: insights from history and Drosophila modeling of gut–microbe interactions. Annu. Rev. Cell Dev. Biol. 29, 571–592 (2013).
Jones, R. M. et al. Symbiotic lactobacilli stimulate gut epithelial proliferation via Nox-mediated generation of reactive oxygen species. EMBO J. 32, 3017–3028 (2013).
Kim, S. H. & Lee, W. J. Role of DUOX in gut inflammation: lessons from Drosophila model of gut–microbiota interactions. Front. Cell Infect. Microbiol. 3, 116 (2014).
Jang, S. et al. Dual oxidase enables insect gut symbiosis by mediating respiratory network formation. Proc. Natl Acad. Sci. USA 118, e2020922118 (2021).
Pitsouli, C. & Perrimon, N. Embryonic multipotent progenitors remodel the Drosophila airways during metamorphosis. Development 137, 3615–3624 (2010).
Gervais, L. & Casanova, J. The Drosophila homologue of SRF acts as a boosting mechanism to sustain FGF-induced terminal branching in the tracheal system. Development 138, 1269–1274 (2011).
Bardin, A. J., Perdigoto, C. N., Southall, T. D., Brand, A. H. & Schweisguth, F. Transcriptional control of stem cell maintenance in the Drosophila intestine. Development 137, 705–714 (2010).
Sato, M. & Kornberg, T. B. FGF is an essential mitogen and chemoattractant for the air sacs of the Drosophila tracheal system. Dev. Cell 3, 195–207 (2002).
Kumar, J. P. & Moses, K. EGF receptor and Notch signaling act upstream of Eyeless/Pax6 to control eye specification. Cell 104, 687–697 (2001).
Ha, E. M., Oh, C. T., Bae, Y. S. & Lee, W. J. A direct role for dual oxidase in Drosophila gut immunity. Science 310, 847–850 (2005).
Thibault, S. T. et al. A complementary transposon tool kit for Drosophila melanogaster using P and piggyBac. Nat. Genet. 36, 283–287 (2004).
Lee, T. & Luo, L. Mosaic analysis with a repressible cell marker (MARCM) for Drosophila neural development. Trends Neurosci. 24, 251–254 (2001).
Pitsouli, C. & Delidakis, C. The interplay between DSL proteins and ubiquitin ligases in Notch signaling. Development 132, 4041–4050 (2005).
Acknowledgements
The authors would like to thank the BDSC, the VDRC, the Kyoto Stock Center and the TRiP for fly stocks; A. Bardin, S. Hou, J. Casanova, T. Kornberg, C. Potter, P. Wappner and A. Ignatiou for fly stocks; E. Snyder for use of the Nikon Ti microscope; and the DSHB for antibodies. This project was supported by FP7-PEOPLE-2011-CIG-303727, the Fondation Santé and the Cyprus Research and Innovation Foundation EXCELLENCE/0918/0082 to C.P., and by ERC AdG 268515, DFG SFB873, the Huntsman Cancer Foundation and NIH GM124434 to B.A.E.
Author information
Authors and Affiliations
Contributions
V.T., M.M.R. and M.P. designed and performed experiments, and analysed data. I.C. and K.K. performed experiments and analysed data. H.M., V.B. and B.I. assisted with experiment execution. B.A.E. and C.P. conceived and supervised the project. C.P. compiled the data and wrote the manuscript with the help of M.M.R., V.T. and B.A.E.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Cell Biology thanks Rongwen Xi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Infection and oxidative damage increase esg > GFP + cells in the midgut and associate with increased TTC branching.
a, Adult midgut intestinal progenitors labelled with esgNP5130-Gal4 > UAS-srcGFP were imaged in unchallenged conditions (4% sucrose) and upon oral P.a. infection (48hrs), and feeding with H2O2 (48hrs) and PQ (24hrs). DAPI (blue) in the upper panels stains all midgut nuclei. The bottom panels show the GFP-labeled progenitors separately. P.a. and PQ expanded the intestinal progenitors with a posterior midgut bias, whereas H2O2 exhibited an anterior midgut bias. b-c, Quantification of midgut mitosis (b, n = 10 each) and TTC branching (c, n = 7,6) in PQ-treated flies. d-e, Posterior midgut (R4) images of btl-Gal4 > UAS-srcGFP flies in baseline conditions (sucrose) and upon PQ feeding. DAPI (blue) staining all the nuclei. Single channel images of the GFP are shown in d’-e’. f-g, Posterior midgut images of QF6 > QUAS-mtdTomato flies in baseline conditions exhibit tracheal expression of the reporter. Midgut epithelial ECs with low expression of the reporter are visible is zoomed image (g). Single channel images of the Tomato are shown in f’-g’. Scale bars: 300 μm in a, 75 μm in d-g. Data are presented as mean values ± SD. Statistical significance (t-tested, two-sided for b, and U-tested for c): ns, not significant, * 0.01 < p ≤ 0.05, ** 0.001 < p ≤ 0.01 and *** p ≤ 0.001.
Extended Data Fig. 2 The FGFR/Btl is necessary and sufficient for midgut TTC branching and ISC mitosis.
a-b, Brightfield images of the tracheae of P.a. infected R5 regions of the midgut in trh-Gal4 control (a) and trh-Gal4 > UAS-btlDN (b). c-d, Brightfield images of the tracheae of uninfected R5 regions of the midgut in trh-Gal4 control (c) and trh-Gal4 > UAS-λbtl (d). e-g, Quantification of TTCs (e, n = 10,11,10,9,10), TTC branching (f, n = 10,11,10,8,10), and midgut mitosis (g, n = 8,6,10,9,6) upon trh-Gal4-driven btl manipulation with or without P.a. infection. h-i, Brightfield images of the tracheae of P.a.-infected R5 regions of the midgut in dSRF-Gal4 control (h) and dSRF-Gal4 > UAS-btlDN (i). j-k, Brightfield images of the tracheae of uninfected R5 regions of the midgut in dSRF-Gal4 control (j) and dSRF-Gal4 > UAS-λbtl (k). l-n, Quantification of TTCs (l, n = 10,10,8,8,11), TTC branching (m, n = 10,10,8,8,11), and midgut mitosis (n, n = 11,9,12,12,9) upon dSRF-Gal4-driven btl manipulation with or without P.a. infection. All scale bars: 75 μm. Data are presented as mean values ± SD. Statistical significance (t-tested, two-sided): ns, not significant, * 0.01 < p ≤ 0.05, ** 0.001 < p ≤ 0.01 and *** p ≤ 0.001.
Extended Data Fig. 3 Infection and oxidative damage induce FGF/bnl in the midgut epithelium.
Adult midgut bnl-expressing cells labeled with the reporter bnl-Gal4 > UAS-srcGFP were imaged in unchallenged conditions (4% sucrose) and upon oral P.a. infection (48hrs), feeding with H2O2 (48hrs) and PQ (24hrs). DAPI (blue) in the upper panels stained all midgut nuclei. The bottom panels show the GFP-labeled bnl expressing cells separately. P.a. and PQ induced the reporter throughout the midgut, whereas H2O2 exhibited an anterior midgut bias. Scale bar: 300 μm.
Extended Data Fig. 4 Btl/Bnl signaling in the epithelial cells is necessary for efficient tracheal remodelling and mitosis in response to infection.
a-b, Quantification of TTC branching upon progenitor- (a) and EC-specific (b) silencing of bnl (bnlRNAi3) and btl (btlRNAi) (a, n = 10,8,5,10,7,7 and b, n = 10,7,9,10,9,8). c-d, Quantification of midgut mitosis upon progenitor- (c) and EC-specific (d) silencing of bnl (bnlRNAi3) (c, n = 6,8,12,12 and d, n = 9,9,11,13). e, Quantification of esg + progenitors as a percent of total number of cells in the posterior regions of the midgut upon progenitor-specific knockdown of btl (btlRNAi) and bnl (bnlRNAi3) (n = 12,15,15). f-g, Quantification of midgut mitosis upon progenitor- (f) and EC-specific (g) silencing of btl (btlRNAi) (f, n = 8,13,11,13 and g, n = 12,11,11,16). h, Quantification of esg + progenitor cells/total number of cells in the posterior midgut upon progenitor-specific knockdown of btl (btlDN, n = 9,9). Data are presented as mean values ± SD. Statistical significance (t-tested, two-sided): ns, not significant, * 0.01 < p ≤ 0.05, ** 0.001 < p ≤ 0.01 and *** p ≤ 0.001.
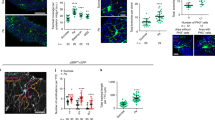
Extended Data Fig. 5 Infection and oxidative damage activate Hif-1α/Sima in the midgut epithelium and the visceral TTCs.
Hif-1α/Sima activation was monitored via the ldh-Gal4 > UAS-nlsGFP reporter expression in the adult midgut epithelium and the intestinal trachea of the R5 region in unchallenged flies (sucrose) and upon P.a. and PQ treatment (a, c, e), and of the R2 region in unchallenged flies (sucrose) and upon H2O2 feeding (b, d). Epithelial sections (a-d) and trachea surface sections (a’-d’) of the same midguts were imaged. DAPI (blue) in a-d and a’-d’ stains all the nuclei. a”-d” and a”’-d”’ correspond to separated channels for reporter expression in the epithelium and the intestinal trachea, respectively. The ldh-Gal4 > UAS-nlsGFP reporter was expressed in cells of the midgut epithelium and in the midgut TTCs in baseline conditions in the anterior (R2 in b, b’) and posterior (R5 in a, a’) midgut. P.a. (c, c’), H2O2 (d, d’) and PQ (e, e’) induced the reporter in the epithelium and the trachea at varying degrees. All images were acquired at the same confocal settings as their respective controls. Scale bar: 75 μm.
Extended Data Fig. 6 Hif-1a/Sima is necessary in the intestinal epithelium and the trachea for TTC branching.
a-b, Brightfield images of the midgut TTCs (R5 region) upon trachea-specific (via btl-Gal4) sima knockdown in the background of heterozygous simaKG in baseline conditions. c-d, Bright-field images of the midgut TTCs (R5 region) upon EC-specific (via Myo1A-Gal4) sima knockdown in the background of heterozygous simaKG in baseline conditions. e-f, Bright-field images of the midgut TTCs (R5 region) upon trachea-specific (via btl-Gal4) sima knockdown in the background of heterozygous simaKG in P.a.-infected conditions. g-h, Bright-field images of the midgut TTCs (R5 region) upon EC-specific (via Myo1A-Gal4) sima knockdown in the background of heterozygous simaKG in P.a.-infected conditions. The images correspond to examples of those quantified for Fig. 4e,i. Scale bar: 75 μm.
Extended Data Fig. 7 Time-course analysis of NotchDN progenitor-derived midgut tumors.
a-d, The R4a region of control (reared for 4 days at 18 °C) (a) and tumorous midguts (reared for 4, 7 and 10 days at 29 °C) (b-d) of the esg-Gal4 UAS-eGFP tub-Gal80ts > UAS-NotchDN genotype with concomitant expression of QF6 > QUAS-mtdTomato (red) to label the trachea. DAPI (blue in a-d) is used to label all midgut nuclei and Prospero (a”-d”) labels the EEs. a’-d’, a”-d” and a”’-d”’ correspond to the individual channels for eGFP, Prospero and Tomato-labeled trachea, respectively. Scale bars: 75 μm. e-g, Quantification of TTC branching in the R4a of control (NotchDN uninduced) and NotchDN-expressing midguts (e, n = 4,8,4,6,4,7), in the NotchDN tumor-region vs. neighboring non-tumor area on the same image (f, n = 6,6,4,4,6,6), and midgut mitosis of control (NotchDN uninduced) and NotchDN-expressing midguts (g, n = 20 each) during a time-course analysis at 4, 7, and 10 days post-tumor induction. Scale bar: 75 μm. Data are presented as mean values ± SD. Statistical significance (t-tested, two-sided): ** 0.001 < p ≤ 0.01 and *** p ≤ 0.001.
Extended Data Fig. 8 Time-course analysis of RasV12 progenitor-derived midgut tumors.
a-d, The R5 region of control esg-Gal4 UAS-eGFP tub-Gal80ts (reared for 1 day at 29 °C) and esg-Gal4 UAS-eGFP tub-Gal80ts > UAS-RasV12-tumor bearing midguts (reared for 1, 3 and 5 days at 29 °C) with concomitant expression of QF6 > QUAS-mtdTomato (red) to label the trachea. DAPI (blue in a-d) was used to label all midgut nuclei. a’-d’ and a”-d” correspond to the individual channels for the eGFP and the Tomato-labeled trachea, respectively. Scale bar: 75 μm.
Supplementary information
Supplementary Tables
Supplementary Table 1: Drosophila and Pseudomonas strains used in this study. Supplementary Table 2: reagents and antibodies used in this study. Supplementary Table 3: primer sequences used in this study.
Source data
Source Data Fig. 1
Numerical source data for Fig. 1.
Source Data Fig. 1
Additional images for key experiments of Fig. 1.
Source Data Fig. 2
Numerical source data for Fig. 2.
Source Data Fig. 3
Numerical source data for Fig. 3.
Source Data Fig. 4
Numerical source data for Fig. 4.
Source Data Fig. 5
Numerical source data for Fig. 5.
Source Data Fig. 5
Additional images for key experiments of Fig. 5.
Source Data Fig. 6
Numerical source data for Fig. 6.
Source Data Fig. 7
Numerical source data for Fig. 7.
Source Data Fig. 7
Additional images for key experiments of Fig. 7.
Source Data Fig. 8
Numerical source data for Fig. 8.
Source Data Extended Data Fig. 1
Numerical source data for Extended Data Fig. 1.
Source Data Extended Data Fig. 2
Numerical source data for Extended Data Fig. 2.
Source Data Extended Data Fig. 4
Numerical source data for Extended Data Fig. 4.
Source Data Extended Data Fig. 7
Numerical source data for Extended Data Fig. 7.
Rights and permissions
About this article
Cite this article
Tamamouna, V., Rahman, M.M., Petersson, M. et al. Remodelling of oxygen-transporting tracheoles drives intestinal regeneration and tumorigenesis in Drosophila. Nat Cell Biol 23, 497–510 (2021). https://doi.org/10.1038/s41556-021-00674-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-021-00674-1
- Springer Nature Limited
This article is cited by
-
Tracheal remodelling supports stem cells
Nature Cell Biology (2021)