Abstract
Coordinated rearrangements of cytoskeletal structures are the principal source of forces that govern cell and tissue morphogenesis1,2. However, unlike for actin-based mechanical forces, our knowledge about the contribution of forces originating from other cytoskeletal components remains scarce. Here, we establish microtubules as central components of cell mechanics during tissue morphogenesis. We find that individual cells are mechanically autonomous during early Drosophila wing epithelium development. Each cell contains a polarized apical non-centrosomal microtubule cytoskeleton that bears compressive forces, whereby acute elimination of microtubule-based forces leads to cell shortening. We further establish that the Fat planar cell polarity (Ft-PCP) signalling pathway3,4 couples microtubules at adherens junctions (AJs) and patterns microtubule-based forces across a tissue via polarized transcellular stability, thus revealing a molecular mechanism bridging single cell and tissue mechanics. Together, these results provide a physical basis to explain how global patterning of microtubules controls cell mechanics to coordinate collective cell behaviour during tissue remodelling. These results also offer alternative paradigms towards the interplay of contractile and protrusive cytoskeletal forces at the single cell and tissue levels.





Similar content being viewed by others
Data availability
Data supporting the conclusions are available from the corresponding author upon reasonable request.
References
Heisenberg, C. P. & Bellaiche, Y. Forces in tissue morphogenesis and patterning. Cell 153, 948–962 (2013).
Lecuit, T., Lenne, P. F. & Munro, E. Force generation, transmission, and integration during cell and tissue morphogenesis. Annu. Rev. Cell. Dev. Biol. 27, 157–184 (2011).
Matis, M., Russler-Germain, D. A., Hu, Q., Tomlin, C. J. & Axelrod, J. D. Microtubules provide directional information for core PCP function. eLife 3, e02893 (2014).
Harumoto, T. et al. Atypical cadherins Dachsous and Fat control dynamics of noncentrosomal microtubules in planar cell polarity. Dev. Cell. 19, 389–401 (2010).
Siedlik, M. J. & Nelson, C. M. Regulation of tissue morphodynamics: an important role for actomyosin contractility. Curr. Opin. Genet. Dev. 32, 80–85 (2015).
Foster, P. J., Fürthauer, S., Shelley, M. J. & Needleman, D. Active contraction of microtubule networks. eLife 4, e10837 (2015).
Laan, L., Husson, J., Munteanu, E. L., Kerssemakers, J. W. & Dogterom, M. Force-generation and dynamic instability of microtubule bundles. Proc. Natl Acad. Sci. USA 105, 8920–8925 (2008).
Zhao, T., Graham, O. S., Raposo, A. & St Johnston, D. Growing microtubules push the oocyte nucleus to polarize the Drosophila dorsal–ventral axis. Science 336, 999–1003 (2012).
Takeda, M., Sami, M. M. & Wang, Y. C. A homeostatic apical microtubule network shortens cells for epithelial folding via a basal polarity shift. Nat. Cell Biol. 20, 36–45 (2018).
Bouchet, B. P. et al. Mesenchymal cell invasion requires cooperative regulation of persistent microtubule growth by SLAIN2 and CLASP1. Dev. Cell 39, 708–723 (2016).
Ranft, J. et al. Fluidization of tissues by cell division and apoptosis. Proc. Natl Acad. Sci. USA 107, 20863–20868 (2010).
Blanchard, G. B. et al. Tissue tectonics: morphogenetic strain rates, cell shape change and intercalation. Nat. Methods 6, 458–464 (2009).
Bonnet, I. et al. Mechanical state, material properties and continuous description of an epithelial tissue. J. R. Soc. Interface 9, 2614–2623 (2012).
Aigouy, B. et al. Cell flow reorients the axis of planar polarity in the wing epithelium of Drosophila. Cell 142, 773–786 (2010).
Shimada, Y., Yonemura, S., Ohkura, H., Strutt, D. & Uemura, T. Polarized transport of Frizzled along the planar microtubule arrays in Drosophila wing epithelium. Dev. Cell 10, 209–222 (2006).
Matis, M. & Axelrod, J. D. Regulation of PCP by the Fat signaling pathway. Genes Dev. 27, 2207–2220 (2013).
Bardet, P. L. et al. PTEN controls junction lengthening and stability during cell rearrangement in epithelial tissue. Dev. Cell 25, 534–546 (2013).
Etournay, R. et al. Interplay of cell dynamics and epithelial tension during morphogenesis of the Drosophila pupal wing. eLife 4, e07090 (2015).
Halder, G. & Johnson, R. L. Hippo signaling: growth control and beyond. Development 138, 9–22 (2011).
Feng, Y. & Irvine, K. D. Fat and expanded act in parallel to regulate growth through warts. Proc. Natl Acad. Sci. USA 104, 20362–20367 (2007).
Baena-Lopez, L. A., Baonza, A. & Garcia-Bellido, A. The orientation of cell divisions determines the shape of Drosophila organs. Curr. Biol. 15, 1640–1644 (2005).
Diaz de la Loza, M. C. & Thompson, B. J. Forces shaping the Drosophila wing. Mech. Dev. 144, 23–32 (2017).
Ray, R. P. et al. Patterned anchorage to the apical extracellular matrix defines tissue shape in the developing appendages of Drosophila. Dev. Cell 34, 310–322 (2015).
Bertet, C., Sulak, L. & Lecuit, T. Myosin-dependent junction remodelling controls planar cell intercalation and axis elongation. Nature 429, 667–671 (2004).
Zallen, J. A. & Wieschaus, E. Patterned gene expression directs bipolar planar polarity in Drosophila. Dev. Cell 6, 343–355 (2004).
Gittes, F., Mickey, B., Nettleton, J. & Howard, J. Flexural rigidity of microtubules and actin filaments measured from thermal fluctuations in shape. J. Cell. Biol. 120, 923–934 (1993).
Brangwynne, C. P. et al. Microtubules can bear enhanced compressive loads in living cells because of lateral reinforcement. J. Cell. Biol. 173, 733–741 (2006).
Stamenovic, D., Mijailovich, S. M., Tolic-Norrelykke, I. M., Chen, J. & Wang, N. Cell prestress. II. Contribution of microtubules. Am. J. Physiol. Cell. Physiol. 282, C617–C624 (2002).
Gupton, S. L., Salmon, W. C. & Waterman-Storer, C. M. Converging populations of F-actin promote breakage of associated microtubules to spatially regulate microtubule turnover in migrating cells. Curr. Biol. 12, 1891–1899 (2002).
Schaefer, A. W., Kabir, N. & Forscher, P. Filopodia and actin arcs guide the assembly and transport of two populations of microtubules with unique dynamic parameters in neuronal growth cones. J. Cell. Biol. 158, 139–152 (2002).
Mao, Y. et al. Dachs: an unconventional myosin that functions downstream of Fat to regulate growth, affinity and gene expression in Drosophila. Development 133, 2539–2551 (2006).
Cao, Y., White, H. D. & Li, X. D. Drosophila myosin-XX functions as an actin-binding protein to facilitate the interaction between Zyx102 and actin. Biochemistry 53, 350–360 (2014).
Olofsson, J., Sharp, K. A., Matis, M., Cho, B. & Axelrod, J. D. Prickle/spiny-legs isoforms control the polarity of the apical microtubule network in planar cell polarity. Development 141, 2866–2874 (2014).
Gomez, J. M., Chumakova, L., Bulgakova, N. A. & Brown, N. H. Microtubule organization is determined by the shape of epithelial cells. Nat. Commun. 7, 13172 (2016).
Borowiak, M. et al. Photoswitchable inhibitors of microtubule dynamics optically control mitosis and cell death. Cell 162, 403–411 (2015).
Pettit, G. R. et al. Isolation and structure of the strong cell growth and tubulin inhibitor combretastatin A-4. Experientia 45, 209–211 (1989).
Picone, R. et al. A polarised population of dynamic microtubules mediates homeostatic length control in animal cells. PLoS Biol. 8, e1000542 (2010).
Xu, Z. et al. Microtubules acquire resistance from mechanical breakage through intralumenal acetylation. Science 356, 328–332 (2017).
Aumeier, C. et al. Self-repair promotes microtubule rescue. Nat. Cell Biol. 18, 1054–1064 (2016).
Schaedel, L. et al. Microtubules self-repair in response to mechanical stress. Nat. Mater. 14, 1156–1163 (2015).
Acknowledgements
We thank S. Blair, D. Brunner, V. Gelfand, N. Sherwood, M. Simon, K. Irvine, Y. Bellaieche and the Developmental Studies Hybridoma Bank for reagents. We also thank members of the Galic Lab and the Matis Lab for critical reading of the manuscript. This work was supported by funds from the DFG to M.G. (EXC-1003 and GA 2268-2/1), M.M. (EXC-1003, SPP-1782 MA 6726/1-1 and FF-2015-07) and to O.T.-S. (SFB-1032, Emmy Noether TH 2231/1-1) and from the IZKF to M.M. (Mat2/019/16). A.S. and T.H. are members of the joint graduate school Cells-in-Motion Cluster of Excellence (EXC-1003) and the International Max Planck Research School—Molecular Biomedicine, Münster, Germany.
Author information
Authors and Affiliations
Contributions
A.S. and M.M. designed the project, carried out most experiments and performed image analyses. A.R., H.N. and J.K. prepared and imaged the TEM samples. T.S., I.B. and M.G. developed the curvature quantification software, carried out the microtubule alignment simulations and designed the three-dimensional model. O.T-S. designed and synthesized the microtubule-destabilizing drug, and M.M. wrote the manuscript with feedback from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Integrated supplementary information
Supplementary Figure 1 Non-centrosomal MTs form an apical array of parallel MT bundles aligned along P/D axis in 18 hrs APF wild-type wings.
(a) TEM micrograph showing a vertical section of 18 hrs APF wild-type wing cell and (b) a cross-section at the level of AJs (yellow line in a). Note that MTs are enriched at the apical sites and are bent (red arrowheads). Images shown in (a, b) are representative of n = 3 independent experiments. (c) 3D reconstruction of 18 hrs APF wild-type wing expressing MyoII-GFP and stained for MTs (α-Tub). Apical is at the top. (d) Tyr-TUB, α−TUB and EOS-TUB show no difference in MT labeling. Scale bars, (a) 500 nm, (b) 200 nm, (d) 5 μm. (e) MT alignment precede cell elongation. MyoII-GFP and α−TUB show aligned MTs along the P/D axis in 16 hrs APF wing. Note that some cells are still round, but MTs are already aligned. (f) Histogram of angular distribution of MTs in round cells show alignment of MT along P/D axis. Orientation of MTs (panel on the right) is color-coded according to angles. Scale bars, (e, f) 5 μm. (g-j) Tissue shape changes during early pupal wing development. Whole wings expressing Arm-GFP at (g) 15 hrs APF, (h) 16 hrs APF, (i) 18 hrs APF and (j) 24 hrs APF, respectively. Scale bars, (top) 50 μm, (below) 5 μm. Images shown in (c-f) are representative of n = 3 independent experiments. Images shown in (g-j) are representative of n = 5 independent experiments.
Supplementary Figure 2 Ft-PCP signaling pathway affects wing morphogenesis.
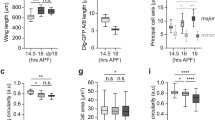
(a-c) Adult wing shape analysis of individual PCP mutants compared to wild-type (a). The Ft-PCP mutant flies ftl(2) fd/ftGRV; ActP-Gal4/UAS-FtΔECDΔN-1 (N1) (b) and ds38k fjN7/dsUA071 fjd1; UAS-Ds/TubP-Gal4 (c) show a shorter wing and abnormal round shape. (d) Overlay of wild-type (black) and mutants (b in orange and c in red) adult wings show shorter and rounder mutant wings. Images shown in (a-d) are representative of n = 4 independent experiments. (e) Ft-PCP mutant wings have perturbed cell elongation. Arm in 18 hrs APF wild-type, dachs mutant (d), ftl(2)fdd1/ ftl(2)fddGC13 double mutant (ftd) and ftl(2) fd/ftGRV; ActP-Gal4/UAS-FtΔECDΔN-1 (N1) mutant wing. Images are representative of n = 3 independent experiments. (f) Quantification of cell EI in wild-type and mutant wings. Cell EI of ft d and N1 mutant is significantly smaller than that of wild-type and d mutant. (Kruskal-Wallis test, from left to right: n.s. p > 0.9999, **** p < 0.0001, **** p < 0.0001, **** p < 0.0001, **** p < 0.0001, n.s. p > 0.9999, n (wt) = 496 cells, 5 pupae, n (d1/ dGC13) = 329 cells, 4 pupae, n (ftd) = 232 cells, 3 pupae, n (N1) = 259 cells, 4 pupae). Scale bars, 5 μm. (g) Quantification of MT alignment along P/D axis in d1/ dGC13 mutant wings (Kruskal-Wallis test, from left to right: **** p < 0.0001, **** p < 0.0001, **** p < 0.0001, n = 445 cells, 4 pupae). (h) Wing epithelial cells are mechanically autonomous. Cell isolation experiment, showing cell expressing Arm-GFP just before, during and after circular ablation of junctions (24 s and 60 s). Red dotted line marks where AJs were ablated. (i) Time series of single cell ablation expressing Arm-GFP just before, during and after circular ablation of junctions (24 s, 60 s and 135 s) (see Supplementary Movie 2) and (j) Cross-sections along apical basal axis (from the basal to the apical site) of the same ablated area. (Right) 3D reconstruction of the ablated area. (k) Quantification of cell EI in patch before (gray box) and after ablation (red box). Note that there is no significant change in cell shape after isolation of cell from the rest of the tissue (two-tailed Wilcoxon matched-pairs test, p = 0.5988; n.s., non-significant; n = 35 cells, 5 pupae). (l) Graph showing the cell maximum Ferret’s diameter before (t0) and 60 s after ablation (t60) for single cell isolation (S) and small patches of cells (P). (n(S) = 26 cells, 9 pupae and n(P) = 25 cells, 3 pupae). Scale bars, (h, i) 5 μm. Images shown in (h-j) are representative of n = 7 independent experiments. Boxes in all box plots extend from the 25th to 75th percentiles, with a line at the median. Whiskers show min and max values.
Supplementary Figure 3 dumpy mutants show no difference in wing shape compared to wild-type wings at 18 hrs APF.
(a) Wing shape analysis of wild-type (top), and dumpy mutant (below) wings at 18 hrs APF. Scale bars, 50 μm. (b) Quantification of wild-type and dumpy mutant wing area shows no significant difference at 18 hrs APF. (two-tailed Mann-Whitney U test, p = 0.5476; n.s., n(wt) = 5 pupae and n(dpov1) = 5 pupae). (c) α−TUB and Arm in 18 hrs APF dpov1 wing show normal wing development at 18 hrs APF. (d) Left: Histogram of angular distribution of MTs in dpov1 wing show alignment of MT along P/D axis. Right: Quantification of MT alignment along P/D axis (Ordinary one-way ANOVA, from left to right: **** p < 0.0001, **** p < 0.0001, * p = 0.0107, n = 361 cells, 3 pupae). (e) MyoII is polarized in dumpy mutant wing at 18 hrs APF. MyoII-GFP in 18 hrs APF dpov1 wing show normal polarization pattern at 18 hrs APF. (f) Mean intensities of junctional MyoII as a function of the angle (θ) of the junctions with respect to the P/D axis. Right: Quantification of MyoII polarity (two-tailed Mann-Whitney U test, **** p < 0.0001; n(P/D) = 185 cells and n(A/P) = 107 cells, 4 pupae). (g) MyoII-GFP in 18 hrs APF wild-type wing. (h) Initial recoil velocities for the A/P and P/D oriented junctions show increased tension along P/D oriented junctions. (two-tailed t-test, ** p = 0.001; n(P/D) = 10 cells and n(A/P) = 10 cells, 5 pupae). Scale bars, (d, f, h left) 50 μm, (d, f, h right) 10 μm. Images shown in (a, c, e, g) are representative of n = 3 independent experiments. Boxes in all box plots extend from the 25th to 75th percentiles, with a line at the median. Whiskers show min and max values.
Supplementary Figure 4 MTs and MyoII remain polarized in isolated cells.
(a) Time-lapse images of EOS-Tub in 17hrs APF wing cells during ablation. (b) Cross-sections along apical basal axis (from the basal to the apical site). (c)Top: Histogram of angular distribution of MTs in isolated patch of cells. Bottom: Quantification of MT alignment before and after cell ablation shows that MTs stay aligned after cell isolation. (Kruskal-Wallis test, from left to right: **** p < 0.0001, *** p = 0.0008, **** p < 0.0001, **** p < 0.0001, *** p = 0.0003, *** p < 0.0001, n = 71 cells, 3 pupae). Angles are measured with respect to the P/D axis. (d) Time-lapse images of MyoII-3XmKate2 (color-coded grayscale) in 17 hrs APF wing cells during ablation. (e) Cross-section along apical basal axis (from the basal to the apical site). (f) Top: Mean intensities of junctional MyoII as a function of the angle (θ) of the junctions with respect to the P/D axis (top). Bottom: Quantification of the MyoII polarity. (Kruskal-Wallis test, from left to right: ** p = 0.0011, *** p = 0.0002, n = 166 junctions, 3 pupae). (g) Local ablation of MTs triggers rapid translocation of the adjacent junction. Time-lapse images of ECad-mTomato in 17 hrs APF wing cell (see Supplementary Movie 4). Note that immediately after MT ablation (red line) the adjacent junction moves towards the cell center. (h) Kymograph showing junction movement after MT ablation along the yellow dotted line in (i). (i) Quantitative analysis of junction movements in control cell (con) and cells were MTs were locally ablated (cut). (two-tailed Mann-Whitney U test, **** p < 0.0001, n(con) = 15 cells, 3 pupae and n(cut) = 15 cells, 5 pupae). In all experiments (15/15) junctions moved towards the center of the cell. Scale bars (a, b, d, e), 5 μm, (h) 1 μm. Images shown in (a, b, d, e) are representative of n = 3 independent experiments. Images shown in (g, h) are representative of n = 5 independent experiments. Boxes in all box plots extend from the 25th to 75th percentiles, with a line at the median. Whiskers show min and max values.
Supplementary Figure 5 Quantification of MT dynamic inhibition by PST1 in vivo.
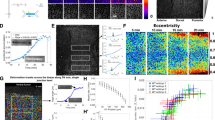
(a) Time-series of EB1-Cherry movement before and after photoconversion of the MT-destabilizing drug. Note the rapid drop in EB1 intensity upon photoactivation within the activation region but not in adjacent regions, arguing that the activated drug locally inhibits MT polymerization. (b) Ratio of EOS-Tub mean intensities at the time of activation and after 60 s in cells were PST-1 was activated, and in adjacent cells that were not illuminated (control cells). Note that no significant drop in signal intensity was observed before a second activation. (Right) images of EOS-Tub at the time of subsequent activation (red arrowhead). (two-tailed t-test, n.s. n(con) = 33 cells and n(PST1) = 33 cells, 3 pupae). Scale bar, 5 μm. (c) Kymograph of EOS-Tub signal after first activation of PST1 by UV illumination. (d) Kymographs show no change in cell shape (marked with E-Cad-Tomato) along the P/D and A/P axes in control wing upon UV illumination. (e) Kymographs show contraction of the wing along the P/D axis (marked with E-Cad-Tomato) upon photoconversion of PST1. (f) Elongation-index of cell-shape before (t = 0 sec) and after (t = 60 sec) photoconversion show a significant change in EI for PST1 treated wings (red) but not in control wings (grey). Top, two-tailed Wilcoxon matched-pairs test, Control: p = 0.1615 and PST-1: **** p < 0.0001; n = 142 cells, 3 pupae for control and n = 202 cells, 3 pupae for PST1. Bottom, scatter plot of EI values for individual cells before (x-axis) and after (y-axis) UV activation. Colored segments indicate decrease ( = yellow) and increase ( = blue) of EI upon UV activation. Scale bars, (a, b) 3 μm. (g-k) Depletion of MTs by Spastin expression leads to cell shape heterogeneity. (g) Cartoon depicting strategy for temporal MT depletion from the anterior compartment of pupal wings using the GAL80ts system. To express Spastin only during the desired time window, larvae are reared at 25 °C (that is where Gal4 activity is blocked by GAL80ts). 0 hr APF pupae were collected and at 12 hrs APF shifted to 29 °C. This inactivates Gal80ts and activates CiGal4 to induce Spastin expression. Pupae from the control cross (expressing UAS-GFP, not shown) were simultaneously shifted from 25 °C to 29 °C. (h) 18 hrs APF wing expressing Spastin in the anterior part of the wing. The red dashed line marks the A/P border. (i) Representative images of TUB (grey) show reduction of the polarized MT cytoskeleton in the anterior part of the wing. (j) Representative images of Arm (gray) in 18 hrs APF wing show no effect of Spastin expression on membrane trafficking. (k) Relative Arm-GFP intensities show no difference in anterior and posterior compartments (two-tailed Mann-Whitney U test, p = 0.1059; n.s., n(A) = 284 junctions, 3 pupae n(P) = 275 junctions, 3 pupae). Scale bars, (h) 50 μm, (i, j) 3 μm. Images shown in (a, c-e, h-j) are representative of n = 3 independent experiments. Boxes in all box plots extend from the 25th to 75th percentiles, with a line at the median. Whiskers show min and max values.
Supplementary information
Supplementary Information
Supplementary Figures 1–5 and Supplementary Movie legends.
Supplementary Table 1
List of fly stock donors.
Supplementary Movie 1
Cell isolation experiment.
Supplementary Movie 2
Cell isolation experiment.
Supplementary Movie 3
MyoII remains polarized in 17 h APF wing cells following ablation.
Supplementary Movie 4
3D reconstruction of the ablated area shown in Supplementary Movie 3.
Supplementary Movie 5
MT dynamics in wing cells.
Supplementary Movie 6
MT polymerization experiment showing short wavelength buckling of MTs only following contact with the cell cortex (white arrowhead).
Supplementary Movie 7
MT ablation experiment showing that buckled MTs are under compression.
Supplementary Movie 8
MT ablation experiment showing that buckled MTs are under compression.
Supplementary Movie 9
MT ablation experiment showing that local ablation of MTs (red line) triggers rapid translocation of the adjacent junction (white arrowhead) towards the center of the cell.
Supplementary Movie 10
EB1 comets (marked with EB1-GFP) move along P–D axis generating a directional gradient in pupal wing cell at 18 h APF.
Supplementary Movie 11
MTs growing along A–P axis switch frequently between growth and shrinking, but do not buckle (white arrowhead).
Supplementary Movie 12
Numerical simulation of the MT trap.
Rights and permissions
About this article
Cite this article
Singh, A., Saha, T., Begemann, I. et al. Polarized microtubule dynamics directs cell mechanics and coordinates forces during epithelial morphogenesis. Nat Cell Biol 20, 1126–1133 (2018). https://doi.org/10.1038/s41556-018-0193-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41556-018-0193-1
- Springer Nature Limited
This article is cited by
-
Dynamic interplay of microtubule and actomyosin forces drive tissue extension
Nature Communications (2024)
-
Hemipiperazines as peptide-derived molecular photoswitches with low-nanomolar cytotoxicity
Nature Communications (2022)
-
Foregut organ progenitors and their niche display distinct viscoelastic properties in vivo during early morphogenesis stages
Communications Biology (2022)
-
Microtubule disassembly by caspases is an important rate-limiting step of cell extrusion
Nature Communications (2022)
-
Regulation of microtubule dynamics, mechanics and function through the growing tip
Nature Reviews Molecular Cell Biology (2021)