Abstract
The extent of tissue regeneration varies widely between species. Mammals have a limited regenerative capacity whilst lower vertebrates such as the zebrafish (Danio rerio), a freshwater teleost, can robustly regenerate a range of tissues, including the spinal cord, heart, and fin. The molecular and cellular basis of this altered response is one of intense investigation. In this review, we summarise the current understanding of the association between zebrafish regeneration and Hippo pathway function, a phosphorylation cascade that regulates cell proliferation, mechanotransduction, stem cell fate, and tumorigenesis, amongst others. We also compare this function to Hippo pathway activity in the regenerative response of other species. We find that the Hippo pathway effectors Yap/Taz facilitate zebrafish regeneration and that this appears to be latent in mammals, suggesting that therapeutically promoting precise and temporal YAP/TAZ signalling in humans may enhance regeneration and hence reduce morbidity.
Similar content being viewed by others
Introduction
Many different organisms have the ability to regenerate, although the robustness, efficiency, and scope of this regeneration is varied. Invertebrates such as planarians and Hydra regenerate their entire body such that, when cut in half, each section forms an entire new organism1,2,3,4. At the other end of the scale, mammalian regeneration is limited, with adult animals often responding to injury with fibrotic scarring rather than regeneration5,6. Some mammalian tissues do regenerate, including the skin, intestine, liver, peripheral nervous system, and blood7,8,9,10,11, as well as foetal tissues12 but this capability is impaired in ageing systems13,14, which, along with a general lack of regenerative ability in most tissues, causes high morbidity in humans.
Midway on the scale from complete (invertebrate) to limited (mammalian) regeneration are lower vertebrates, including amphibians and fish. The zebrafish Danio rerio has the potential to completely regenerate multiple adult and embryonic organs, including the heart, fin, and many nervous system components15,16,17,18,19,20. First explored in the 1980s by Streisinger21,22, the zebrafish is regularly utilised in the study of adult and embryonic regeneration due to their rapid external development, relative low cost, ease of genetic manipulation, scalability, transparent juveniles, and high rate of regeneration, none of which are present in the mouse.
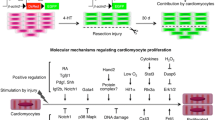
The cellular and molecular drivers of zebrafish regeneration have been the subject of intense research5,19. Effective replacement of lost or damaged cells requires a large pool of available healthy cells. Cell pools can be formed by multiple sources, including the activation of resident stem or progenitor cells (differentiation), the reversion of differentiated cells to a more immature pluripotent state (dedifferentiation), or the conversion of one differentiated cell type into another mature cell type (transdifferentiation)5,23. On a molecular level, these can be driven by epigenetic and gene expression changes, such as alterations in DNA methylation5,23,24,25, histone modifications5,26,27,28,29, regeneration-responsive enhancers28,30,31,32,33, and the activation of a range of key developmental signalling pathways, including Bmp, Fgf, Notch, RA, Shh, and Wnt/β-catenin (summarised in Table 1)34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67,68,69,70,71,72,73,74,75,76,77. In recent years, it has become evident that Hippo signalling (Fig. 1) plays a critical role in developmental and regenerative processes in both zebrafish and mammals. This is associated with the Hippo pathway’s role in regulating cell proliferation and migration, detecting and responding to changes in tissue tension, extracellular matrix, chemical cues, which consequently alter cell fates78,79,80,81,82.
The Hippo pathway is regulated by the integration of a range of upstream stimuli. This includes mechanotransductive elements (such as caveolae and Piezo signalling), metabolism, extracellular matrix and integrin signalling, transduction of extracellular stimuli via mitogenic growth factor signalling and GPCRs, cell polarity and cell–cell contacts. Activation of the Hippo pathway triggers a phosphorylation cascade that leads to the phosphorylation of the Hippo pathway effectors YAP/TAZ. Phosphorylation of YAP/TAZ redistributes YAP/TAZ to the cytoplasm, blocking TEAD-mediated gene expression. Hippo pathway inactivation prevents YAP/TAZ phosphorylation, allowing their nuclear translocation and hence TEAD-mediated gene expression. Note that MST1/2 (mammalian STE20-like kinase1/2) are encoded by STK4/3, and TAZ by WWTR1. Figure 1 is created in BioRender.com.
The core Hippo signalling pathway is comprised of a serine/threonine kinase phosphorylation cascade (Fig. 1), most of which were identified in genetic screens of Drosophila melanogaster for tumour suppressor genes83,84. Activity of this pathway is regulated by a range of stimuli, including mechanical signalling, cell shape, ECM stiffness, cell polarity, metabolism, and cell:cell contacts78,79,82,85,86,87,88,89,90, which are integrated to stimulate key kinases MST1/2 (the fly Hippo orthologs), STK25, and MAP4Ks when the Hippo pathway is active87,89,91,92,93. These kinases then phosphorylate, and so activate, LATS1/2, which phosphorylate the core Hippo effectors transcriptional co-activator YAP1 and its paralog TAZ on multiple conserved serine residues86,87,91,92,94,95. YAP1/TAZ phosphorylation triggers their retention in the cytoplasm via binding to protein 14-3-3, or ubiquitin-mediated degradation86,87,94,95,96,97. When the Hippo pathway is inactive these phosphorylations do not occur, resulting in YAP1/TAZ nuclear localisation, where they outcompete VGLL4 and bind to transcription factors TEAD1-487,98,99,100,101. Binding to TEADs stimulate the expression of a range of pro-proliferative, -oncogenic, -stemness, and -EMT genes, such as CTGF and CYR6178,87,90,98,99,100,102,103,104. Additional YAP1/TAZ transcription factors have also been identified87, but the most extensively studied are the TEADs. Zebrafish Hippo pathway genes have high genetic orthology to human genes, suggesting that this is an appropriate model in which to study Hippo pathway function (Fig. 2). Here we review the role of the Hippo pathway in the regeneration of a range of organs, including heart, spinal cord, tail fin, lateral line, and liver regeneration, with a focus on the zebrafish.
Direct gene sequence comparison between a sample of human and zebrafish Hippo pathway members and transcriptional targets shows a range of similarity scores, emphasizing a high degree of similarities between fish and human genes, while also highlighting that some Hippo pathway components appear to have no direct orthologs present in both species. WWTR1 encodes TAZ. STK4 encodes MST1 and STK3 encodes MST2 (in accordance with the consensus of the Hippo pathway field). CYR61 is also known as CCN1 and CTGF as CCN2. % gene sequence similarity identified using ensembl.org under orthology tab. ctgfb, nf2b, map4k2, and rhoaa-c could not be identified as orthologues in this manner, so manual BLAST comparison of genomic sequence (from GRCz11) was performed to give the values indicated.
Heart regeneration
Cardiovascular diseases are the primary cause of morbidity and mortality globally, with around half of these deaths caused by ischaemic heart disease leading to heart failure105. This is due to the limited regeneration capacity of the adult human heart, which responds to heart muscle damage with fibrosis and scarring rather than the reformation of contractile muscle106. A similar response is seen in other mammals (such as the mouse), which also show limited cardiac regeneration after experimental injury paradigms107. An exception is an enhanced heart regeneration potential in neonatal mice, but this is transient and is lost within the first week of life108, coinciding with a decrease in YAP1 transcriptional activity78, and the withdrawal of cardiomyocytes (CMs) from the cell cycle109. However, this regenerative ability in neonatal mice108 highlights that there may be therapeutic potential in reactivating the regenerative capacity in humans.
In contrast to restricted mammalian regeneration, both adult and embryonic zebrafish regenerate their heart fully following injury and even after multiple insults5,6,15,19,110,111,112,113 (Fig. 3). This extensive heart regeneration is the result of two key characteristics: a high level of existing CM proliferation (around 3% per week, compared to <1% per year in adult mice114 and humans115), and a permissive extracellular environment that stimulates it19,116. One major hurdle and pathological driver in mammalian heart regeneration is the formation of a fibrotic scar and non-permissive ECM at the injury area, replacing dead CMs with non-contractile elements such as collagen or fibroblasts rather than new CMs106,117 (Fig. 3B). However, in the zebrafish, although collagen and fibronectin does accumulate and a scar is formed, it is eliminated to allow effective regeneration113,118,119,120. This scarring is regulated by Hippo signalling, with cav-1, yap1, and ctgfa mutants having disrupted scar formation and hence regeneration121,122,123 (see Table 2 for a summary of these phenotypes). Heart injury promotes Ctgfa secretion into the ECM from endocardial cells, where it promotes the expression of pro-regenerative ECM genes (such as fibronectins and collagens)121. This expression allows for a transient scar, as shown by ctgfa mutants having a larger and more persistent scar, whilst ctgfa overexpression speeds scar resolution121. Similarly, yap1 mutants have an altered ECM composition at the injury site, resulting in increased scarring and impaired regeneration at early time points122. This alteration of the scar microenvironment by secretion of Hippo pathway transcriptional targets suggests that Hippo signalling may also indirectly regulate the infiltration and proliferation of CMs through a cell non-autonomous mechanism.
a Structure of the uninjured zebrafish adult heart. b Injury at the ventricle apex induces collagen and fibronectin deposition and scar formation. yap1, ctgfa, and cav-1 promote appropriate and transient scar formation. c Heart epicardium undergoes EMT and inflammatory cells (blue) infiltrate into the scar. yap1 and ctgfa inhibit inflammatory cell infiltration. d New coronary vessels form to revascularize the injury site. e Mature cardiomyocytes (CMs) (pink) dedifferentiate into progenitor cells (yellow) and migrate along the new coronary vessels into the injury site. ctgfa promotes CM migration. f CM progenitors proliferate to create a progenitor cell pool, which matures back to CMs to reform the heart muscle. ctgfa and cav-1 promote cell proliferation.
The immune system creates a permissive microenvironment for regeneration (Fig. 3C). This is clearly demonstrated in the medaka, a teleost species closely related to the zebrafish. The medaka displays limited heart regeneration, a finding that is surprising considering their evolutionary similarity to the zebrafish124,125. This limited regeneration is, at least in part, due to the medaka’s delay in macrophage recruitment to the injury site124. When recapitulated in the zebrafish by clodronate liposome-mediated macrophage depletion, these macrophage defects cause compromised neovascularisation and CM proliferation and consequently severe defects in heart regeneration124. This is due to the role of macrophages and other immune cell components such as Treg cells in many areas of cardiac regeneration, including enhancing neovascularisation, CM proliferation, and scar resolution via the production of pro-regenerative factors, with inhibition of inflammation and timely immune cell recruitment inhibiting regeneration124,126,127,128,129.
However, this pro-regenerative effect of the immune system is not simple. Yap1-Ctgfa signalling, shown to enhance cardiac regeneration, also negatively regulates the migration and infiltration of macrophages into the injury site121,122, suggesting that inhibiting macrophage infiltration promotes cardiac regeneration. Similarly, yap1 KO fish have increased macrophage infiltration in the scar and increased monocyte chemotactic gene expression122, and ctgfa KO promotes the chemokine receptor gene cxcr3.1 in the heart to increase M1 macrophage polarisation and so enhance inflammatory signalling121, and both KO lines have defective regeneration. This apparent contradiction may be due to differences between experimental paradigms in investigating immune cell function in regeneration—it has been shown that the type of immune cells recruited, and the different regenerative stages alter the functional role of the immune system in regeneration128. An alternative explanation for this apparent discrepancy could be due to the requirement for tight spatio-temporal control of the immune system function during regeneration. This is shown by disruption of reparative regeneration after both immune system hyperactivation121,122 and excessive inhibition124,126,127,129. Another potential reason for the inconsistency is that various immune cell types likely react differently to the injury, and so the Hippo pathway may respond in a range of ways to the same trigger. Therefore, the extent of activation or inhibition in these studies will greatly impact the results. Further in-depth studies are needed in order to fully elucidate the detailed spatiotemporal inflammatory response including revealing the exact immune cell types involved in regeneration and thereby the role of the Hippo pathway in the immune system’s contribution to cardiac regeneration.
Hippo pathway signalling has also been linked to the epicardium, which is activated after heart injury in the zebrafish (Fig. 3C)15,130. The epicardium promotes regeneration, potentially by functioning as a cellular scaffold that generates epicardial-derived cells which differentiate into myofibroblasts and perivascular fibroblasts in the injured myocardium131. This may then act in a paracrine manner to induce CM proliferation and neoangiogenesis131. Epicardial activation has not yet been linked to the Hippo pathway in zebrafish heart regeneration. However, in the developing mouse, Hippo components are expressed in both the proepicardium and epicardium, and deletion of either Yap or Taz in the mouse gives coronary defects and impacts on epicardial cell proliferation, EMT, and specification of cell fate132. Similar developmental cardiac defects can be seen in a range of Hippo pathway component mutants in the zebrafish112,133,134,135,136,137,138,139,140,141,142,143,144,145,146,147,148,149,150,151,152, suggesting that this role of the Hippo pathway may be conserved between mammals and teleosts.
After injury, existing differentiated CMs undergo limited dedifferentiation, upregulate the embryonic cardiogenesis gene gata4, and proliferate113,153,154,155,156. These CMs migrate to the injury site along newly-formed coronary vasculature157,158,159,160,161,162 (Fig. 3E), where they proliferate further and differentiate to replace dead CMs and form new functional heart muscle163 (Fig. 3F). CM proliferation is promoted by a range of signalling pathways, including Nrg, Tgfβ, Igf, and the Hippo pathway (Table 1). Disruption of the Hippo pathway-regulated genes cav-1α and ctgfa inhibits CM proliferation and repopulation of the injury area121,123. Similarly, TGFβ-mediated activation of regulatory elements upstream of ctgfa promotes CM proliferation at the injury site32. cav-1α and ctgfa are induced after injury in epicardial and endocardial cells respectively, suggesting a role for these cells in cell non-autonomous regulation of CM function, such as in ECM secretion in response to extracellular stimuli121,123. Disruption of cav-1α and ctgfa results in defective heart regeneration121,123, whilst overexpression of ctgfa and yap1 has the opposite effect121,122. Disrupting Hippo signalling in mammals gives comparable results. In pigs, CM-specific knockdown of Sav (which results in increased YAP activity164,165) increases CM proliferation and improves heart function after myocardial infarction165. Similar outcomes are observed when Yap1 is disrupted in mice, causing heart regeneration defects through decreased CM proliferation166,167,168,169,170,171, whilst heart regeneration (and CM proliferation) is stimulated after Yap1 activation167,168,169, potentially due to the Hippo pathway’s link to cytoskeletal and ECM regulation170. However, the opposite effect is observed when Hippo signalling is disrupted in murine cardiac fibroblasts172,173. Deletion of Yap1/Taz in these fibroblasts results in improved cardiac function after myocardial infarction through modulation of the fibrotic and fibroinflammatory response172. Enhanced Yap1/Taz signalling (through either Yap1 overexpression or Lats1/2 deletion) has the opposing effect, with mice displaying elevated fibrotic responses173,172. This apparent contradiction between the role of the Hippo pathway in CMs and cardiac fibroblasts supports a model where the Hippo pathway functions differently in different cell types.
CMs in zebrafish ctgfa mutants also fail to migrate along the coronary vasculature to infiltrate the wound, despite no changes in revascularisation, potentially as a result of alterations in cytoskeletal gene expression in a cell autonomous regulation of CM infiltration121,123. Supporting this, data using in vitro primary rat cultures of cardiac fibroblasts show that Yap1 siRNA-mediated knockdown reduces expression of factors associated with cytoskeletal motility and ECM adhesion, although these results have not been recapitulated in zebrafish, CMs, or in vivo122.
In summary, the Hippo signalling pathway enhances cardiac regeneration by temporal activation of Yap1/Taz and promotes normal cardiovascular development. Yap1/Taz promote appropriate scar formation and potentially prevent overactivation of the immune response, which, when combined, increase scar resolution, spatiotemporal CM proliferation, and thereby cardiac regeneration. Taking advantage of this regenerative capacity may hold therapeutic potential in the treatment of human MI. For example, pharmacological regulation of the Hippo pathway could modulate CM proliferation and fate plasticity156,174, promoting scarless healing in the adult heart and reducing disease burden. Recent work disrupting Hippo signalling in pigs after myocardial infarction165 suggests, in a clinically relevant model system, that this could be possible. However, precise cell type-specific modulation of the Hippo pathway will be vital to realise its full potential, as the Hippo pathway has been shown to have different functions in the cell types involved. For example, heart function is improved after injury in mammals when YAP activity is increased in CMs165 but also when Yap1/Taz is deleted in cardiac fibroblasts172.
Spinal cord regeneration
The Hippo pathway is also associated with regeneration after spinal cord injury (SCI) in the zebrafish. After SCI in humans and other mammals, the affected axons and neurons are destroyed and a non-permissive scar is formed in the place of new cells, commonly resulting in lifelong disability175,176,177. However, both adult and larval zebrafish robustly and effectively regenerate their spinal cords after injury, with viable axon regrowth over the lesion site and return of full swimming function within weeks after injury18,19,178,179,180,181,182 (Fig. 4).
a Structure of the uninjured spinal cord, with ependymal radial glia (ERG) (green) lining the central canal and motor neurons (yellow). b Spinal cord transection disrupts neuronal processes. c ERGs undergo EMT to form ERG progenitors (blue) and migrate to the site of injury. yap1 promotes EMT of ERGs, and yap1 and ctgfa promote progenitor proliferation. d ERG progenitors extend processes across the injury site to form a glial bridge (grey). yap1 and ctgfa promote the formation of the glial bridge. e Neuronal processes extend across the injury site, guided by the glial bridge to promote remodelling and reformation of the spinal cord.
For functional recovery in the spinal cord, new and existing cells must proliferate, migrate to the injury site, bridge the lesion, and differentiate to reintegrate with existing distal neuronal circuitry183. Neurogenesis from tissue-resident progenitors is a vital step for this to occur in zebrafish, which is promoted by multiple signalling pathways, including Wnt/β-catenin, Fgf, Shh, and is inhibited by Notch signalling (Table 1). The tissue-resident progenitors responsible for cell proliferation and bridging are thought to be the ventral ependymal radial glia (ERG)181,184,185,186. These cells have general functions during development and adulthood in maintaining spinal cord homoeostasis such as sealing the blood-brain barrier and maintaining ion balance, but also proliferate and differentiate into a range of neuronal cell types after injury181,183,187.
To allow new cell processes to traverse the lesion site, a glial bridge is formed. After injury, ERGs migrate to the lesion and elongate to form an astroglial bridge over the lesion, along which axons can grow to innervate distal targets (Fig. 4C–E). This is driven by pro-regenerative gene expression (e.g. col12a1a/b and tenascin-c), interactions with other cell types such as Schwann cells, and additional environmental cues180,182,184,188,189,190. Zebrafish glial bridging shares clear morphological and functional similarities with the bridging observed during mammalian peripheral nerve regeneration (which occurs to a much greater extent than mammalian CNS regeneration)183,191,192,193, indicating that this common process may be manipulated in the human for therapeutic benefit.
In order to induce glial cells to undergo bridging, ventral ERGs undergo an epithelial-to-mesenchymal transition (EMT)193 (Fig. 4C). EMT is a common feature of many cells activated by injury, and is linked to stem cell activation, increased cellular plasticity, and tissue remodelling194,195,196. Glial EMT is both necessary and sufficient to induce glial bridging, and is linked to Yap1-Ctgfa signalling193,197. yap1, wwtr1 (gene encoding Taz), and ctgfa are upregulated following SCI, with yap1 and ctgfa expression localised to bridging glia and ventral ERGs193,197. As well as inducing ctgfa expression in ventral ERGs, Yap1 promotes twist1a expression193. twist1a is an established EMT marker, activation of which directs a mesenchymal transition in Ctgfa+ ERGs, promoting glial bridging and functional spinal cord repair193.
Similar to heart regeneration, one major difference between the zebrafish and mammalian response to SCI is the formation of a glial scar. SCI causes vascular damage, oedema, and inflammation, resulting in widespread gliosis, necrosis, and apoptosis that eventually forms a glial/fibrotic scar in mammals, stretching beyond the site of the initial trauma and acting to prevent secondary damage but also preventing axon regrowth176,198,199. There is no significant scarring in the zebrafish, so there is no experimental work linking the Hippo pathway in zebrafish to scar resolution, however siRNA-mediated knockdown of the YAP1/TAZ-TEAD target gene Ctgf in rats reduces the glial scar and hence improves regeneration after SCI200, suggesting that YAP1/TAZ signalling may promote scar formation or impair scar resolution and outlining a potential therapeutic target for SCI treatment in mammals.
Loss of function mutations of yap1, wwtr1, and ctgfa all result in impaired functional recovery after SCI, with ctgfa and yap1 disruption causing a glia-specific cell proliferation reduction, resulting in impaired bridging and axon regeneration across the lesion site193,197. Exogenous administration of human CTGF to these ctgfa mutants reversed this defect197. This finding, and the similar finding that heart scar formation is larger and more persistent in ctgfa, yap1, and cav-1 mutants121,122,123 appears in contrast to that seen in the rat glial scar200, which found that knockdown of CTGF increased recovery through the clearance of scarring, and the current clinical trials which are targeting CTGF to reduce fibrosis and scarring201. This may be due to species differences in the function of the Hippo pathway, but this is not supported by the relatively high translatability of other studies between rodent and zebrafish. An alternative explanation might be that Yap/Taz-Ctgfa signalling has opposing effects at different stages of spinal cord regeneration, or that strictly regulated temporal activation/repression of signalling is key, although studies of this in mammals must be performed after the scar has been resolved, which currently presents an experimental challenge.
Yap1 signalling is also associated with the regenerative role of glial cells in other parts of the CNS, such as the retina. In the zebrafish, retinal damage induces reprogramming events where Müller glia are converted to a highly proliferative progenitor-like state, dividing asymmetrically to replace lost photoreceptors202,203,204,205,206,207. Yap1 knockdown blocks Müller glial cell proliferation and neurogenesis after light damage of the zebrafish retina208, suggesting a common role for yap1 in the regenerative functions of glial cells. Mammalian retinas usually do not have a proliferative, pro-regenerative, Müller glia response to injury. However, in the mouse, YAP promotes glial reprogramming, with YAP activation inducing Müller glia reprogramming to a highly proliferative, progenitor-like cell202,204. This suggests that promoting Yap1 signalling therapeutically may also promote CNS regeneration in humans.
These findings propose a model in which Yap1 senses the mechanical stress caused by SCI, enhancing ctgfa and twist1a expression to activate a pro-EMT and pro-proliferative transcriptional programme in ventral ERGs, promoting glial bridging, axon regeneration, and, consequently, functional recovery193. This model suggests that enhancing scar resolution, promoting EMT, enhancing CTGF signalling at later stages of regeneration, and identifying CTGF-responsive spinal cord cells may allow for the identification of a therapeutic target to promote mammalian spinal cord regeneration197. Targeting CTGF has been investigated in a variety of preclinical and clinical trials for multiple conditions, including muscular dystrophy and pancreatic cancer. For example, the monoclonal antibody Pamrevlumab has shown promise in trials for idiopathic pulmonary fibrosis209,210. However, these trials involve the inhibition of CTGF activity, rather than the enhancement that may be required to promote recovery201,211. Consequently, further insights must be obtained before translating these findings into an effective treatment option in humans.
Tail fin regeneration
Zebrafish and other teleosts regenerate their fins completely after multiple consecutive amputations212, a phenomenon that was studied as early as the 18th century213, and by the regeneration pioneer T. H. Morgan at the turn of the 20th century214,215,216. Fin regeneration occurs through epimorphic regeneration, a process characterised by the presence of a blastema early in regeneration (Fig. 5). This mass of undifferentiated proliferating progenitor cells at the site of injury is formed by mature cell dedifferentiation, which can then differentiate back into mature cells to generate an actively growing tissue that replaces the lost appendage217.
a The uninjured tail fin of the adult zebrafish is formed of many bony rays, which each consist of epidermis surrounding mature osteoblasts (purple) in the mesenchyme. b Amputation of the tail fin disrupts the bony ray segment. c In the initial stages of tail fin regeneration the epidermis covers the wound. d Osteoblasts and other mature cells dedifferentiate and proliferate at the wound tip to form a blastema with osteoprogenitors (green). yap1 inhibits osteoblast dedifferentiation and bmp4 enhances blastema cell proliferation. e The bony ray segment extends through maturation of the progenitor cells back to their original cell type. yap1 promotes osteoprogenitor maturation.
There is not yet direct evidence for a role for the Hippo pathway in dedifferentiation in the zebrafish caudal fin blastema, but in other in vivo models, both mammalian and invertebrate, the Hippo pathway maintain stemness, promote proliferation, and revert differentiated cells to a progenitor cell state81,218,219,220,221,222,223,224. In the zebrafish, dedifferentiated cells proliferate to form a large pool of progenitor cells in the blastema (Fig. 5D).
Blastema formation is enhanced and maintained by a range of developmental signalling pathways, including Hippo, Wnt/β-catenin, Igf, Notch, Fgf, Shh, Tgfβ, (Table 1) as well as inflammatory signals such as Il1β and Hsp90α225,226,227. The concentration gradient of these signalling pathways gives positional information along the proximodistal axis of the injured tissue, ensuring that structures are reformed at the correct location and that the tissue grows at an appropriate rate, halting when the previous size and shape is reached16,228,229,230,231. Hippo signalling is one such signalling pathway with activity changes in proximodistal expression. In the high cell density distal blastema, Yap1 is mainly cytoplasmic (and so inactive), whilst in the low density proximal blastema, it becomes mainly nuclear (active)232. Yap1 is also localised to α-catenin and F-actin when in the cytoplasm232. This suggests that the heterogeneous cell densities within the blastema could be transduced through cell junctions and the cytoskeleton232. These mechanical properties then impact Yap1 localisation, which alters the regenerative capacity of the fin232. For example, yap1 disruption impairs cell proliferation and alters key signalling pathways, including promoting Wnt and reducing Bmp signalling after fin injury232,233. This results in an accumulation of osteoprogenitors and prevention of osteoblast differentiation, and so defective regeneration233. Ctgfa levels are also increased following fin injury, and disruption of its regulatory sequences induces increased tissue stiffness and ECM deposition234.
Tail fin progenitor cells are not multipotent. Instead, cells remain lineage restricted235,236. The osteoblast is one such cell type. After injury, these cells dedifferentiate, proliferate, and mature to only give rise to osteoblasts in the regenerate (Fig. 5)236,237,238. More specifically, injury induces differentiated mature osteoblasts close to the injury site, which usually form the bony rays of the fin, to lose expression of late and intermediate osteoblast differentiation markers (such as osteocalcin and osterix) and undergo a Wnt/β-catenin-mediated EMT to gain progenitor markers and generate osteoprogenitor cells, which migrate to the blastema and proliferate in a Fgf-dependent manner237,239. These progenitors then undergo Bmp-mediated maturation into osteoblasts239 (Fig. 5D), a process that is associated with the Hippo pathway233,239. This link to osteoblast formation and function is most dramatically illustrated by wwtr1 disruption in embryonic zebrafish, which results in a complete lack of skeletal ossification149. Similarly, disruption of yap1 results in major bone defects and impaired fin regeneration, caused by an inhibition of osteoprogenitor cell maturation, giving an increased osteoprogenitor pool with a downregulation of intermediate and mature gene markers233. These defects are mediated by a reduction in Bmp signalling (which usually promotes maturation into osteoblasts239). In wild-type fish, Yap1 promotes Bmp signalling in a cell non-autonomous manner, restricting osteoprogenitors to the distal blastema (where Yap1 is inactive), and promotes osteoblast formation in the proximal blastema (where Yap1 is active)233. bmp4 is also associated with tail fin regeneration. Bmp4 is expressed in the distal blastema, and its inhibition reduces fin outgrowth after injury due to reduced proliferation of blastema cells240,241. This data suggests that Yap1 functions in the blastema to mechanotransduce tension changes and control the fate and migration of specific cell types in the amputated fin, regulating the precise control of tissue growth, potentially through the expression of ECM factors such as Ctgfa232,234.
The Hippo pathway is also associated with the differentiation of osteoblasts from mesenchymal stem cells (MSCs) during development, which generate neurons, adipocytes, skeletal muscle, and osteoblasts242. In in vitro studies, TAZ promotes osteoblast differentiation from MSCs via activation of Runx2-dependent gene transcription whilst inhibiting adipocyte differentiation via repression of PPARγ signalling149. CTGF also promotes osteoblast differentiation from MSCs in vitro243. Similar data are observed in mice, where YAP1 and TAZ promote bone formation and repair through their regulation of the osteoblast lineage244,245. Osteoblast lineage-specific Yap1 KO mice have reduced osteoblast differentiation and increased adipocyte formation, an effect that is diminished following increased β-catenin expression, demonstrating the importance of Wnt/β-catenin signalling in this process245. However, the role of the Hippo pathway in osteoblast differentiation is contested, with some in vitro studies suggesting that YAP1/TAZ suppress osteoblast differentiation and bone formation, and increase adipogenesis246,247, so more work is required to elucidate this complexity.
Zebrafish tail fin regeneration is most closely associated with limb regeneration, which does not occur in mammals or other higher vertebrates, although the mouse has been found in some instances to regenerate the digit tip in both newborns and adults248. Appendage regeneration does occur in certain amphibians such as salamanders as well as some invertebrates, and the Drosophila yap1 ortholog yki has been shown to promote wing disc regeneration249. Regeneration of an entire limb in mammals appears unlikely, but work in the zebrafish tail fin and other systems suggests that Hippo signalling may play an important role and promoting it could enhance regenerative capacity of specific aspects of limb regeneration, such as enhanced bone regeneration after breaks.
Hair cell regeneration in the lateral line
The lateral line is a mechanosensitive organ in fish and other aquatic amphibians that detects motion of the external liquid, aiding feeding and social behaviour as well as orientation in currents. In zebrafish, this rapidly developing organ is formed of sixty small clusters of cells (termed neuromasts) in adulthood (expanded from an initial eight in larvae)250, located along with the head (anterior lateral line) and trunk (posterior lateral line, pLL) in stereotyped positions17. Neuromasts consist of a group of hair cells with stereocilia projecting out of the skin and into the surrounding water, mechanical movement of which triggers sensation, and surrounding interdigitating supporting cells and mantle cells (Fig. 6A). Hair cells are innervated by ribbon synapses with afferent sensory neurons17 that project to the hindbrain, where they exhibit a somatotopy similar to the tonotopy seen in mammalian cochlear afferent projections251.
a Uninjured neuromasts consist of hair cells (green) with cilia projecting into the external liquid, support cells (blue), mantle cells (orange), and afferent sensory neurons (red) that project to the brain. b Administration of aminoglycosides or Cu2+ causes specific hair cell death. c Support cell proliferation increases and cells transdifferentiate into hair cells. yap1 promotes support cell transdifferentiation. d Hair cell cilia regrowth restores neuromast function.
During early zebrafish development, a pLL primordium (pLLP) is generated behind the otic vesicle, forming a mass of cells that migrates along the flank beneath the skin, depositing protoneuromasts at periodic intervals252,253,254,255. The deposition of protoneuromasts and their development into mature neuromasts is mediated by Wnt/β-catenin, Notch, and Fgf signalling pathways, and is reviewed elsewhere17,256. These migrating cells must maintain a cohesive structure through high levels of expression of E-cadherin and tight junctions. In mammalian epithelial cells, E-Cadherin is a key upstream regulator of YAP1/TAZ257,258, indicating a potential role for Hippo signalling in this process. In fact, the Hippo pathway is linked to lateral line development in the zebrafish, as indicated by the induced expression of Yap1, Amotl2a, and Cav-1α in the developing lateral line136,259,260,261, and how disrupting these proteins functions impact lateral line formation. Downregulation of Cav-1α reduces the number of neuromasts formed136 and disruption of yap1 triggers a range of phenotypes, including a reduction in primordium size, reduced number of neuromasts, and a decrease in hair cell number259,260,261,262. Amotl2a negatively regulates Yap1 in the developing lateral line, limiting proliferation and so restricting the size of the pLLP, coupling with Notch signalling (which upregulates Yap1 to promote proliferation) to ensure correct pLLP size is reached259,261.
Analysis of the transcriptome of yap1-deficient embryos shows multiple gene expression changes, including those involved in the Wnt/β-catenin signalling pathway260, and lysophosphatidic acid (LPA)262. One of these factors is Prox1a, a target of β-catenin that aids hair cell differentiation in the lateral line260,263. Analysis of yap1- and prox1a-deficient embryos shows that yap1 deficiency recapitulates the prox1a deficiency phenotype of reduced hair cell number and impaired mechanoreceptor differentiation. These yap1 phenotypes are rescued by the administration of prox1a mRNA, suggesting that Yap1 functions by promoting Prox1a activity, so regulating hair cell maturation260. In the lateral line, the LPA receptor Lpar2b is expressed in the pLL and neuromasts, and its loss-of-function phenocopies yap1 KD262. LPA inhibits the Hippo kinase module, consequently activating YAP1/TAZ264,265 and stimulating cell proliferation, migration, and differentiation262,266. In the zebrafish specifically, LPA affects early development, promoting vascular and midline development, left-right patterning, and cell migration during gastrulation, amongst others267,268,269,270. In the pLL Lpar2b regulates Yap1 phosphorylation, suggesting that LPA signalling controls both primordium size and neuromast number by regulating Yap1 activity262. These results suggest that the Hippo pathway promotes appropriate size and cell function in lateral line development through a range of signalling pathways, many of which are also associated with other developmental processes.
The high regenerative capacity of the amphibian lateral line was first observed in the salamander271,272 but has since been observed in multiple organisms, including the zebrafish273 (Fig. 6). This is in contrast to the limited regeneration of mammalian hair cells, e.g. of the inner ear256. The majority of regenerated lateral line hair cells are formed by symmetric asynchronous division of support cells in the first 20 hours post injury254,274,275, where mitotic division of one support cell gives rise to two hair cells276 (Fig. 6C). The molecular and cellular triggers of this regeneration include pathways involved in lateral line development—Wnt/β-catenin, Notch, and Fgf signalling—as well as novel factors such as the Jak/Stat3 pathway (Table 1), which balance self-renewal, hair cell differentiation, and the risk of overgrowth.
The Hippo pathway links to lateral line regeneration277. The expression pattern of supporting cells during regeneration is reminiscent of expression in the migrating primordium during lateral line development, which is silenced when leading progenitors differentiate into mature supporting cells and hair cells259,278,279. This includes the expression of Hippo components cav-1 and ctgfa, which are upregulated in the support cells of both the zebrafish lateral line and the mouse inner ear280. In addition, after severe hair cell injury, Yap1 is activated in hair cell precursors, and regeneration is impaired in yap1 mutants277. Yap1 activation may occur through cell junction damage and resulting loss of junction-associated proteins such as Amotl2a, which usually restricts Yap1 activity in the lateral line261,277. Activated Yap1 upregulates lin28a transcription, an RNA-binding protein that regulates the translation of mRNAs involved in developmental timing, pluripotency and metabolism281. This promotes a Yap1-lin28a-let7-Wnt signalling axis that is both necessary and sufficient to promote progenitor cell activation and hence neuromast regeneration. The Yap1-lin28a-let7-Wnt signalling axis has other roles in dedifferentiation, including zebrafish retinal regeneration, mammalian embryonic inner ear development, and in vitro reprogramming of stem cell cultures282,283,284,285.
In summary, Yap1/Taz signalling in progenitor support cells is triggered after hair cell injury, promoting their differentiation towards hair cells via a Wnt signalling pathway, and enhancing recovery. Promoting Yap1/Taz signalling may also have therapeutic benefits in humans. The hair cells of the inner ear do not regenerate256 but have high similarity to zebrafish lateral line hair cells. This includes similar expression patterns of mechanosensitive ion channel and tip link genes and responses to key signalling pathways and ototoxic insults256,286,287,288,289,290, and so targeting the Hippo pathway to promote the regeneration of inner ear hair cells to combat age-related hearing decline may be a viable approach.
Liver regeneration
Despite limited mammalian regeneration of many organs, both mammals and zebrafish can regenerate their livers efficiently through the proliferation of differentiated hepatocytes, regaining liver function through epimorphic regrowth and compensatory enlargement of liver lobes291,292 (Fig. 7). However, this capacity of hepatocytes to repopulate the liver in humans can be overwhelmed by chronic or severe injury, resulting in liver failure that is only treatable by liver transplantation. There are many functional, cellular, and structural similarities between mammalian and zebrafish liver, both of which can regenerate their liver after more chronic insults,292 making the zebrafish a useful model to study the development and regeneration of the liver. However, limited research has been performed investigating the role of the Hippo pathway in zebrafish liver regeneration, although much work on this topic has been performed in the mouse. After experimental murine liver injury, YAP1 protein levels increase, with increased nuclear localisation in the liver and enhanced expression of downstream YAP1/TAZ target genes220,293,294. In a mouse model with Yap deletion in hepatocytes, bile duct ligation results in hepatic necrosis, reduced hepatocyte proliferation, and increased mortality220,295 compared to wild-type mice, suggesting a key role for Hippo signalling in mammalian liver regeneration.
a Healthy (uninjured) zebrafish liver consists of multiple cell types hepatocytes (orange) and bile ducts comprising of biliary ductal cells (green). b Minor liver injury such as partial hepatectomy removes portions of the liver and the associated cells. c Liver recovery after minor liver damage involves hypertrophy and increased proliferation of remaining cells. Yap1 promotes hepatocyte proliferation. b’ Chronic or severe liver damage causes widespread cell death and necrosis. c’ Remaining cells dedifferentiate into liver progenitor cells, promoted by Yap1. d’ Progenitor cells proliferate then differentiate into mature hepatocytes and biliary ductal cells.
One method posited to promote liver regeneration is the recapitulation of developmental processes to generate progenitor-like cells that repopulate the liver after hepatocyte loss. Supporting this, after severe liver injury biliary cells have been shown to transdifferentiate into hepatocytes via a dedifferentiated progenitor-like state to repopulate the liver291,292. Hippo signalling is implicated in multiple cell fate transitions during liver regeneration in the mouse296,297. This includes YAP signalling activation by the alteration of cholangiocytes’ epigenome and transcriptome to aid their restoration of normal hepatocyte and cholangiocyte number296. YAP also associates with factors such as Arid1a to promote the induction of liver progenitor-like cell-enriched genes297.
The Hippo pathway is linked to liver development in both mammalian and zebrafish livers, and likely regulates cell fate plasticity in this process. In mice, YAP1 overexpression causes hepatomegaly that is reversible upon cessation of YAP1 signalling, suggesting a function for YAP1 in regulation of cell proliferation and hepatocyte function221,298. Hepatomegaly is also observed in the zebrafish after Yap1 overexpression or nf2a disruption299,300, whilst conversely yap1-/- fish have reduced liver size147. Other structural defects observed when disrupting upstream Hippo pathway components in the zebrafish include dilated bile ducts299, biliary dysgenesis301, and extrahepatic choledochal cysts299. Yap1 has also been linked to metabolism in the zebrafish liver, where it stimulates nucleotide biosynthesis to promote tissue growth through increasing glutamine synthetase and glucose transporter glut1 expression147,300,302.
The Hippo pathway’s role in hepatocyte development is thought to be vital in its role in the liver as hepatocytes are the predominant cell type in the liver and are key to liver function292. Appropriate Hippo pathway function is essential in the maintenance of mature hepatocytes, with hepatocyte-specific Nf2 loss in mice leading to hepatocyte dedifferentiation into highly renewable progenitors303, and overexpression causing a dysplastic hepatocyte morphology221. YAP1 is also associated with the formation of bile ducts in the developing mouse304, and with the function of the bile ducts (which promote immune cell recruitment and function) in the regenerating adult mouse liver305. Similarly, stk3 and sav1 zebrafish mutants (which both result in increased Yap1 activity) display altered hepatocyte morphology and polarity alongside biliary cell disruption301. Overall, these data suggest a conserved role for the Hippo pathway in structural liver, hepatocyte and biliary cell function between mammals and zebrafish. This implies that the Hippo pathway may also have a role in zebrafish liver regeneration, although this research is still in its infancy and will need further detailed investigation before conclusions can be drawn.
Conclusion
The zebrafish is a powerful model system for the study of regeneration due to their rapid external development, relative low cost, transparent juvenile stages and robust reparative regeneration as well as the availability of a range of established genetic tools and other experimental procedures to study these. In this review, the role of the Hippo pathway in zebrafish regeneration is summarised, with the finding that Yap1/Taz signalling often enhances regeneration through the promotion of cell proliferation, progenitor cell dedifferentiation and maturation, EMT, and scar resolution, as well as linking to key developmental pathways. The phenotypes resulting from the disruption of Hippo pathway components is summarised in Table 2.
The positive effect of Yap1/Taz signalling on regeneration in the zebrafish, which appears to be latent in mammals, suggests some therapeutic potential in promoting YAP/TAZ signalling to enhance mammalian regeneration. However, this must be carefully investigated, as many of the processes associated with enhanced regeneration are linked to an increased risk of cancer, such as an elevated cell proliferation rate, cellular heterogeneity, and increased stemness306. In fact, dysregulation of the Hippo pathway and thereby pathological hyperactivation of YAP1/TAZ promotes carcinogenesis in most, if not all, types of solid tumours102,307,308. The zebrafish may therefore be vital in the elucidation of this association between cancer and regeneration, which could allow us to manipulate regenerative potential without impacting carcinogenesis or vice versa. One way to do this could be through the utilisation of zebrafish Hippo pathway-induced cancer models, which recapitulate human findings in that manipulation of Hippo signalling can trigger tumour formation309,310,311. However, the field of Hippo signalling in the zebrafish is still relatively new, and so much work must be performed to bridge the gaps that are currently preventing its translation to the clinic, particularly the study of the molecular and cellular drivers of the Hippo pathway’s effects on both regeneration and development (Box 1).
References
Wittlieb, J., Khalturin, K., Lohmann, J. U., Anton-Erxleben, F. & Bosch, T. C. G. Transgenic Hydra allow in vivo tracking of individual stem cells during morphogenesis. Proc. Natl. Acad. Sci. USA 103, 6208–6211 (2006).
van Wolfswinkel, J. C., Wagner, D. E. & Reddien, P. W. Single-cell analysis reveals functionally distinct classes within the planarian stem cell compartment. Cell Stem Cell 15, 326–339 (2014).
Ivankovic, M. et al. Model systems for regeneration: planarians. Development 146, dev167684 (2019).
Vogg, M. C., Galliot, B. & Tsiairis, C. D. Model systems for regeneration: Hydra. Development 146, dev177212 (2019).
Zhao, A., Qin, H. & Fu, X. What determines the regenerative capacity in animals? Bioscience 66, 735–746 (2016).
Poss, K. D., Keating, M. T. & Nechiporuk, A. Tales of regeneration in zebrafish. Dev. Dyn. 226, 202–210 (2003).
Mao, S. A., Glorioso, J. M. & Nyberg, S. L. Liver regeneration. Transl. Res. 163, 352–362 (2014).
Plikus, M. V. et al. Epithelial stem cells and implications for wound repair. Semin. Cell Dev. Biol. 23, 946–953 (2012).
Hong, A. W., Meng, Z. & Guan, K.-L. The Hippo pathway in intestinal regeneration and disease. Nat. Rev. Gastroenterol. Hepatol. 13, 324–337 (2016).
Scheib, J. & Höke, A. Advances in peripheral nerve regeneration. Nat. Rev. Neurol. 9, 668–676 (2013).
Flach, J. & Milyavsky, M. Replication stress in hematopoietic stem cells in mouse and man. Mutat. Res. - Fundam. Mol. Mech. Mutagen. 808, 74–82 (2018).
Larson, B. J., Longaker, M. T. & Lorenz, H. P. Scarless fetal wound healing: a basic science review. Plast. Reconstr. Surg. 126, 1172–1180 (2010).
Sousounis, K., Baddour, J. A. & Tsonis, P. A. Aging and regeneration in vertebrates. Curr. Top. Dev. Biol. 108, 217–246 (2014).
Yun, M. H. Changes in regenerative capacity through lifespan. Int. J. Mol. Sci. 16, 25392–25432 (2015).
Smith, K. A. & Mommersteeg, M. T. M. Talkin’ ‘bout regeneration: new advances in cardiac regeneration using the zebrafish. Curr. Opin. Physiol. 14, 48–55 (2020).
Sehring, I. M. & Weidinger, G. Recent advancements in understanding fin regeneration in zebrafish. Wiley Interdiscip. Rev. Dev. Biol. 9, e367 (2020).
Thomas, E. D., Cruz, I. A., Hailey, D. W. & Raible, D. W. There and back again: development and regeneration of the zebrafish lateral line system. Wiley Interdiscip. Rev. Dev. Biol. 4, 1–16 (2015).
Cigliola, V., Becker, C. J. & Poss, K. D. Building bridges, not walls: spinal cord regeneration in zebrafish. Dis. Model. Mech. 13, dmm044131 (2020).
Gemberling, M., Bailey, T. J., Hyde, D. R. & Poss, K. D. The zebrafish as a model for complex tissue regeneration. Trends Genet. 29, 611 (2013).
Marques, I. J., Lupi, E. & Mercader, N. Model systems for regeneration: zebrafish. Development 146, dev167692 (2019).
Streisinger, G., Singer, F., Walker, C., Knauber, D. & Dower, N. Segregation analyses and gene-centromere distances in zebrafish. Genetics 112, 311–319 (1986).
Streisinger, G., Walker, C., Dower, N., Knauber, D. & Singer, F. Production of clones of homozygous diploid zebra fish (Brachydanio rerio). Nature 291, 293–296 (1981).
Jopling, C., Boue, S. & Belmonte, J. C. I. Dedifferentiation, transdifferentiation and reprogramming: three routes to regeneration. Nat. Rev. Mol. Cell Biol. 12, 79–89 (2011).
Powell, C., Grant, A. R., Cornblath, E. & Goldman, D. Analysis of DNA methylation reveals a partial reprogramming of the Müller glia genome during retina regeneration. Proc. Natl. Acad. Sci. USA 110, 19814–19819 (2013).
Hirose, K., Shimoda, N. & Kikuchi, Y. Transient reduction of 5-methylcytosine and 5-hydroxymethylcytosine is associated with active DNA demethylation during regeneration of zebrafish fin. Epigenetics 8, 899–906 (2013).
Stewart, S., Tsun, Z.-Y. & Belmonte, J. C. I. A histone demethylase is necessary for regeneration in zebrafish. Proc. Natl. Acad. Sci. USA 106, 19889–19894 (2009).
Pfefferli, C., Müller, F., Jazwinska, A. & Wicky, C. Specific NuRD components are required for fin regeneration in zebrafish. BMC Biol. 12, 30 (2014).
Goldman, J. A. et al. Resolving heart regeneration by replacement histone profiling. Dev. Cell 40, 392–404 (2017).
Golenberg, N. et al. Citrullination regulates wound responses and tissue regeneration in zebrafish. J. Cell Biol. 219, e201908164 (2020).
Wang, W. et al. Changes in regeneration-responsive enhancers shape regenerative capacities in vertebrates. Science 309, eaaz3090 (2020).
Kang, J. et al. Modulation of tissue repair by regeneration enhancer elements. Nature 532, 201–206 (2016).
Pfefferli, C. & Jazwinska, A. The careg element reveals a common regulation of regeneration in the zebrafish myocardium and fin. Nat. Commun. 8, 15151 (2017).
Thompson, J. D. et al. Identification and requirements of enhancers that direct gene expression during zebrafish fin regeneration. Development 147, dev191262 (2020).
Wu, C.-C. et al. Spatially Resolved Genome-wide Transcriptional Profiling Identifies BMP Signaling as Essential Regulator of Zebrafish Cardiomyocyte Regeneration. Dev. Cell 36, 36–49 (2016).
Quint, E. et al. Bone patterning is altered in the regenerating zebrafish caudal fin after ectopic expression of sonic hedgehog and bmp2b or exposure to cyclopamine. Proc. Natl Acad. Sci. 99, 8713–8718 (2002).
Schebesta, M., Lien, C.-L., Engel, F. B. & Keating, M. T. Transcriptional Profiling of Caudal Fin Regeneration in Zebrafish. ScientificWorldJournal 6, 38–54 (2006).
Kujawski, S. et al. Calcineurin Regulates Coordinated Outgrowth of Zebrafish Regenerating Fins. Dev. Cell 28, 573–587 (2014).
Goldshmit, Y. et al. Different Fgfs have distinct roles in regulating neurogenesis after spinal cord injury in zebrafish. Neural Dev. 13, (2018).
Poss, K. D. et al. Roles for Fgf Signaling during Zebrafish Fin Regeneration. Dev. Biol. 222, 347–358 (2000).
Whitehead, G. G., Makino, S., Lien, C.-L. & Keating, M. T. fgf20 is essential for initiating zebrafish fin regeneration. Sci. (80-.). 310, 1957–1960 (2005).
König, D., Page, L., Chassot, B. & Jazwinska, A. Dynamics of actinotrichia regeneration in the adult zebrafish fin. Dev. Biol. 433, 416–432 (2018).
Lee, S. G. et al. Myc and Fgf Are Required for Zebrafish Neuromast Hair Cell Regeneration. PLoS One 11, e0157768 (2016).
Choi, W.-Y. et al. In vivo monitoring of cardiomyocyte proliferation to identify chemical modifiers of heart regeneration. Development 140, 660–666 (2013).
Chablais, F. & Jazwinska, A. IGF signaling between blastema and wound epidermis is required for fin regeneration. Development 137, 871–879 (2010).
Fang, Y. et al. Translational profiling of cardiomyocytes identifies an early Jak1/Stat3 injury response required for zebrafish heart regeneration. Proc. Natl Acad. Sci. 110, 13416–13421 (2013).
Liang, J. et al. The stat3/socs3a Pathway Is a Key Regulator of Hair Cell Regeneration in Zebrafish stat3/socs3a Pathway: Regulator of Hair Cell Regeneration. J. Neurosci. 32, 10662–10673 (2012).
Khaliq, M. et al. Stat3 Regulates Liver Progenitor Cell-Driven Liver Regeneration in Zebrafish. Gene Expr. 18, 157–170 (2018).
Karra, R., Knecht, A. K., Kikuchi, K. & Poss, K. D. Myocardial NF-κB activation is essential for zebrafish heart regeneration. Proc. Natl Acad. Sci. 112, 13255–13260 (2015).
Zhao, L. et al. Notch signaling regulates cardiomyocyte proliferation during zebrafish heart regeneration. Proc. Natl Acad. Sci. 111, 1403–1408 (2014).
Dias, T. B., Yang, Y.-J., Ogai, K., Becker, T. & Becker, C. G. Notch Signaling Controls Generation of Motor Neurons in the Lesioned Spinal Cord of Adult Zebrafish. J. Neurosci. 32, 3245–3252 (2012).
Grotek, B., Wehner, D. & Weidinger, G. Notch signaling coordinates cellular proliferation with differentiation during zebrafish fin regeneration. Development 140, 1412–1423 (2013).
Münch, J., González-Rajal, A. & de la Pompa, J. L. Notch regulates blastema proliferation and prevents differentiation during adult zebrafish fin regeneration. Development 140, 1402–1411 (2013).
Romero-Carvajal, A. et al. Regeneration of Sensory Hair Cells Requires Localized Interactions between the Notch and Wnt Pathways. Dev. Cell 34, 267–282 (2015).
Pinto-Teixeira, F. et al. Inexhaustible hair-cell regeneration in young and aged zebrafish. Biol. Open 4, 903–909 (2015).
Huang, M. et al. Antagonistic Interaction Between Wnt and Notch Activity Modulates the Regenerative Capacity of a Zebrafish Fibrotic Liver Model. Hepatology 60, 1753–1766 (2014).
He, J., Lu, H., Zou, Q. & Luo, L. Regeneration of Liver After Extreme Hepatocyte Loss Occurs Mainly via Biliary Transdifferentiation in Zebrafish. Gastroenterology 146, 789–800 (2014).
Ko, S. et al. Hdac1 Regulates Differentiation of Bipotent Liver Progenitor Cells During Regeneration via Sox9b and Cdk8. Gastroenterology 156, 187–202 (2019).
Gemberling, M., Karra, R., Dickson, A. L. & Poss, K. D. Nrg1 is an injury-induced cardiomyocyte mitogen for the endogenous heart regeneration program in zebrafish. Elife 4, e05871 (2015).
Mathew, L. K. et al. Comparative expression profiling reveals an essential role for Raldh2 in epimorphic regeneration. J. Biol. Chem. 284, 33642–33653 (2009).
Kikuchi, K. et al. Retinoic Acid Production by Endocardium and Epicardium Is an Injury Response Essential for Zebrafish Heart Regeneration. Dev. Cell 20, 397–404 (2011).
Blum, N. & Begemann, G. Retinoic acid signaling controls the formation, proliferation and survival of the blastema during adult zebrafish fin regeneration. Development 139, 107–116 (2012).
Blum, N. & Begemann, G. Osteoblast de- and redifferentiation are controlled by a dynamic response to retinoic acid during zebrafish fin regeneration. Development 142, 2894–2918 (2015).
Blum, N. & Begemann, G. Retinoic acid signaling spatially restricts osteoblasts and controls ray-interray organization during zebrafish fin regeneration. Development 142, 2888–2893 (2015).
Han, P. et al. Hydrogen peroxide primes heart regeneration with a derepression mechanism. Cell Res. 24, 1091–1107 (2014).
Gauron, C. et al. Sustained production of ROS triggers compensatory proliferation and is required for regeneration to proceed. Sci. Rep. 3, (2013).
Reimer, M. M. et al. Sonic hedgehog is a polarized signal for motor neuron regeneration in adult zebrafish. J. Neurosci. 29, 15073–15082 (2009).
Reimer, M. M. et al. Dopamine from the Brain Promotes Spinal Motor Neuron Generation during Development and Adult Regeneration. Dev. Cell 25, 478–491 (2013).
Jazwinska, A., Badakov, R. & Keating, M. T. Activin-βA Signaling Is Required for Zebrafish Fin Regeneration. Curr. Biol. 17, 1390–1395 (2007).
Chablais, F. & Jazwinska, A. The regenerative capacity of the zebrafish heart is dependent on TGFβ signaling. Development 139, 1921–1930 (2012).
Dogra, D. et al. Opposite effects of Activin type 2 receptor ligands on cardiomyocyte proliferation during development and repair. Nat. Commun. 8, (2017).
Kawakami, Y. et al. Wnt/β-catenin signaling regulates vertebrate limb regeneration. Genes Dev. 20, 3232–3237 (2006).
Stoick-Cooper, C. L. et al. Distinct Wnt signaling pathways have opposing roles in appendage regeneration. Development 134, 479–489 (2007).
Wehner, D. et al. Wnt/β-catenin Signaling Defines Organizing Centers that Orchestrate Growth and Differentiation of the Regenerating Zebrafish Caudal Fin. Cell Rep. 6, 467–481 (2014).
Head, J. R., Gacioch, L., Pennisi, M. & Meyers, J. R. Activation of Canonical Wnt/Beta-Catenin Signaling Stimulates Proliferation in Neuromasts in the Zebrafish Posterior Lateral Line. Dev. Dyn. 242, 832–846 (2013).
Poulain, M. & Ober, E. A. Interplay between Wnt2 and Wnt2bb controls multiple steps of early foregutderived organ development. Development 138, 3557–3568 (2011).
Goessling, W. et al. APC mutant zebrafish uncover a changing temporal requirement for wnt signaling in liver development. Dev. Comp. Immunol. 320, 161–174 (2008).
Choi, T.-Y., Ninov, N., Stainier, D. Y. R. & Shin, D. Extensive Conversion of Hepatic Biliary Epithelial Cells to Hepatocytes After Near Total Loss of Hepatocytes in Zebrafish. Gastroenterology 146, 776–788 (2014).
Moya, I. M. & Halder, G. Hippo–YAP/TAZ signalling in organ regeneration and regenerative medicine. Nat. Rev. Mol. Cell Biol. 20, 211–226 (2019).
Panciera, T., Azzolin, L., Cordenonsi, M. & Piccolo, S. Mechanobiology of YAP and TAZ in physiology and disease. Nat. Publ. Gr. 18, 758–770 (2017).
Fu, V., Plouffe, S. W. & Guan, K.-L. The Hippo pathway in organ development, homeostasis, and regeneration. Curr. Opin. Cell Biol. 49, 99–107 (2017).
Zhao, B., Tumaneng, K. & Guan, K.-L. The Hippo pathway in organ size control, tissue regeneration and stem cell self-renewal. Nat. Cell Biol. 13, 877–883 (2011).
Ardestani, A., Lupse, B. & Maedler, K. Hippo signaling: key emerging pathway in cellular and whole-body metabolism. Trends Endocrinol. Metab. 29, 492–509 (2018).
Kim, W. & Jho, E. The history and regulatory mechanism of the Hippo pathway. BMB Rep. 51, 106–118 (2018).
Davis, J. R. & Tapon, N. Hippo signalling during development. Development 146, dev167106 (2019).
Dupont, S. et al. Role of YAP/TAZ in mechanotransduction. Nature 474, 179–185 (2011).
Zhao, B. et al. Inactivation of YAP oncoprotein by the Hippo pathway is involved in cell contact inhibition and tissue growth control. Genes Dev. 21, 2747–2761 (2007).
Gram Hansen, C., Moroishi, T. & Guan, K.-L. YAP and TAZ: a nexus for Hippo signaling and beyond. Trends Cell Biol. 25, 499–513 (2015).
Meng, Z., Moroishi, T. & Guan, K.-L. Mechanisms of Hippo pathway regulation. Genes Dev. 30, 1–17 (2016).
Fulford, A., Tapon, N. & Ribeiro, P. S. Upstairs, downstairs: spatial regulation of Hippo signalling. Curr. Opin. Cell Biol. 51, 22–32 (2017).
Rausch, V. & Gram Hansen, C. The Hippo pathway, YAP/TAZ, and the plasma membrane. Trends Cell Biol. 30, 32–48 (2020).
Meng, Z. et al. MAP4K family kinases act in parallel to MST1/2 to activate LATS1/2 in the Hippo pathway. Nat. Commun. 6, 8357 (2015).
Zheng, Y. et al. Identification of happyhour/MAP4K as alternative Hpo/Mst-like kinases in the Hippo kinase cascade. Dev. Cell 34, 642–655 (2015).
Lim, S. et al. Identification of the kinase STK25 as an upstream activator of LATS signaling. Nat. Commun. 10, 1547 (2019).
Zhao, B., Li, L., Tumaneng, K., Wang, C.-Y. & Guan, K.-L. A coordinated phosphorylation by Lats and CK1 regulates YAP stability through SCF^β-TRCP. Genes Dev. 22, 72–85 (2010).
Lei, Q.-Y. et al. TAZ Promotes cell proliferation and epithelial-mesenchymal transition and is inhibited by the Hippo pathway. Mol. Cell. Biol. 28, 2426–2436 (2008).
Liu, C.-Y. et al. The Hippo tumor pathway promotes TAZ degradation by phosphorylating a phosphodegron and recruiting the SCF^β-TRCP E3 ligase. J. Biol. Chem. 285, 37159–37169 (2010).
Wang, S. et al. YAP antagonizes innate antiviral immunity and is targeted for lysosomal degradation through IKKε-mediated phosphorylation. Nat. Immunol. 18, 733–743 (2017).
Zhang, H. et al. TEAD transcription factors mediate the function of TAZ in cell growth and epithelial-mesenchymal transition. J. Biol. Chem. 284, 13355–13362 (2009).
Ota, M. & Sasaki, H. Mammalian Tead proteins regulate cell proliferation and contact inhibition as transcriptional mediators of Hippo signaling. Development 135, 4059–4069 (2008).
Zhao, B. et al. TEAD mediates YAP-dependent gene induction and growth control. Genes Dev. 22, 1962–1971 (2008).
Hillmer, R. E. & Link, B. A. The roles of Hippo signaling transducers Yap and Taz in chromatin remodeling. Cells 8, 502 (2019).
Salem, O. & Gram Hansen, C. The Hippo pathway in prostate cancer. Cells 8, 370 (2019).
Yu, F.-X., Zhao, B. & Guan, K.-L. Hippo pathway in organ size control, tissue homeostasis, and cancer. Cell 163, 811–828 (2015).
Rausch, V. et al. The Hippo pathway regulates caveolae expression and mediates flow response via caveolae. Curr. Biol. 29, 1–14 (2019).
Riching, A. S. & Song, K. Cardiac regeneration: new insights into the frontier of ischemic heart failure therapy. Front. Bioeng. Biotechnol. 8, 637538 (2021).
St. John Sutton, M. G. & Sharpe, N. Left ventricular remodeling after myocardial infarction: pathophysiology and therapy. Circulation 101, 2981–2988 (2000).
Engel, F. B., Hsieh, P. C. H., Lee, R. T. & Keating, M. T. FGF1/p38 MAP kinase inhibitor therapy induces cardiomyocyte mitosis, reduces scarring, and rescues function after myocardial infarction. Proc. Natl. Acad. Sci. USA 103, 15546–15551 (2006).
Porrello, E. R. et al. Transient regenerative potential of the neonatal mouse heart. Science 331, 1078–1080 (2011).
Li, F., Wang, X., Capasso, J. M. & Gerdes, A. M. Rapid transition of cardiac myocytes from hyperplasia to hypertrophy during postnatal development. J. Mol. Cell. Cardiol. 28, 1737–1746 (1996).
Bise, T., Sallin, P., Pfefferli, C. & Jazwinska, A. Multiple cryoinjuries modulate the efficiency of zebrafish heart regeneration. Sci. Rep. 10, 11551 (2020).
Sehring, I. M., Jahn, C. & Weidinger, G. Zebrafish fin and heart: what’s special about regeneration? Curr. Opin. Genet. Dev. 40, 48–56 (2016).
Wang, J. et al. The regenerative capacity of zebrafish reverses cardiac failure caused by genetic cardiomyocyte depletion. Development 138, 3421–3430 (2011).
Poss, K. D., Wilson, L. G. & Keating, M. T. Heart regeneration in zebrafish. Science 298, 2188–2190 (2002).
Senyo, S. E. et al. Mammalian heart renewal by pre-existing cardiomyocytes. Nature 493, 433–436 (2013).
Bergmann, O. et al. Dynamics of cell generation and turnover in the human heart. Cell 161, 1566–1575 (2015).
Foglia, M. J. & Poss, K. D. Building and re-building the heart by cardiomyocyte proliferation. Development 143, 729–740 (2016).
Bassat, E. et al. The extracellular matrix protein agrin promotes heart regeneration in mice. Nature 547, 179–184 (2017).
González-Rosa, J. M., Martín, V., Peralta, M., Torres, M. & Mercader, N. Extensive scar formation and regression during heart regeneration after cryoinjury in zebrafish. Development 138, 1663–1674 (2011).
Wang, J., Karra, R., Dickson, A. L. & Poss, K. D. Fibronectin is deposited by injury-activated epicardial cells and is necessary for zebrafish heart regeneration. Dev. Biol. 382, 427–435 (2013).
Lowe, V. et al. Neuropilin 1 mediates epicardial activation and revascularization in the regenerating zebrafish heart. Development 146, dev174482 (2019).
Mukherjee, D. et al. Ccn2a is an injury-induced matricellular factor that promotes cardiac regeneration in zebrafish. Development 148, dev193219 (2021).
Flinn, M. A., Jeffery, B. E., O’Meara, C. C. & Link, B. A. Yap is required for scar formation but not myocyte proliferation during heart regeneration in zebrafish. Cardiovasc. Res. 115, 570–577 (2019).
Cao, J. et al. Single epicardial cell transcriptome sequencing identifies caveolin 1 as an essential factor in zebrafish heart regeneration. Development 143, 232–243 (2016).
Lai, S.-L. et al. Reciprocal analyses in zebrafish and medaka reveal that harnessing the immune response promotes cardiac regeneration. Elife 6, e25605 (2017).
Furutani-Seiki, M. & Wittbrodt, J. Medaka and zebrafish, an evolutionary twin study. Mech. Dev. 121, 629–637 (2004).
Huang, W.-C. et al. Treatment of Glucocorticoids inhibited early immune responses and impaired cardiac repair in adult zebrafish. PLoS One 8, e66613 (2013).
de Preux Charles, A.-S., Bise, T., Baier, F., Marro, J. & Jazwinska, A. Distinct effects of inflammation on preconditioning and regeneration of the adult zebrafish heart. Open Biol. 6, 160102 (2016).
Godwin, J. W., Pinto, A. R. & Rosenthal, N. A. Chasing the recipe for a pro-regenerative immune system. Semin. Cell Dev. Biol. 61, 71–79 (2017).
Hui, S. P. et al. Zebrafish regulatory T cells mediate organ-specific regenerative programs. Dev. Cell 43, 659–672 (2017). e5.
Stewart, K. M. R., Walker, S. L., Baker, A. H., Riley, P. R. & Brittan, M. Hooked on heart regeneration: the zebrafish guide to recovery. Cardiovasc. Res. https://doi.org/10.1093/cvr/cvab214 (2021).
González-Rosa, J. M., Peralta, M. & Mercader, N. Pan-epicardial lineage tracing reveals that epicardium derived cells give rise to myofibroblasts and perivascular cells during zebrafish heart regeneration. Dev. Biol. 370, 173–186 (2012).
Singh, A. et al. Hippo signaling mediators Yap and Taz are required in the epicardium for coronary vasculature development. Cell Rep. 15, 1384–1393 (2016).
Miesfeld, J. B. et al. Yap and Taz regulate retinal pigment epithelial cell fate. Development 142, 3021–3032 (2015).
Hildebrand, S. et al. The E-cadherin/AmotL2 complex organizes actin filaments required for epithelial hexagonal packing and blastocyst hatching. Sci. Rep. 7, 9540 (2017).
Hultin, S. et al. Amotl2 links VE-cadherin to contractile actin fibres necessary for aortic lumen expansion. Nat. Commun. 5, 3743 (2014).
Nixon, S. J. et al. Caveolin-1 is required for lateral line neuromast and notochord development. J. Cell Sci. 120, 2151–2161 (2007).
Flinn, M. A. et al. Llgl1 regulates zebrafish cardiac development by mediating Yap stability in cardiomyocytes. Development 147, dev193581 (2020).
Fang, P.-K. et al. Caveolin-1α and -1β perform nonredundant roles in early vertebrate development. Am. J. Pathol. 169, 2209–2222 (2006).
Dai, X. et al. Phosphorylation of angiomotin by Lats1/2 kinases inhibits F-actin binding, cell migration, and angiogenesis. J. Biol. Chem. 288, 34041–34051 (2013).
Nagasawa-Masuda, A. & Terai, K. Yap/Taz transcriptional activity is essential for vascular regression via Ctgf expression and actin polymerization. PLoS One 12, e0174633 (2017).
Nakajima, H. et al. Flow-dependent endothelial YAP regulation contributes to vessel maintenance. Dev. Cell 40, 523–536 (2017). e6.
Astone, M. et al. Zebrafish mutants and TEAD reporters reveal essential functions for Yap and Taz in posterior cardinal vein development. Sci. Rep. 8, 1–15 (2018).
Grimm, L. et al. Yap1 promotes sprouting and proliferation of lymphatic progenitors downstream of Vegfc in the zebrafish trunk. Elife 8, e42881 (2019).
Xu, D. et al. Scribble influences cyst formation in autosomal-dominant polycystic kidney disease by regulating Hippo signaling pathway. FASEB J. 32, 4394–4407 (2018).
Park, M.-H. et al. CCN1 interlinks integrin and hippo pathway to autoregulate tip cell activity. Elife 8, e46012 (2019).
Dooley, C. M. et al. The gene regulatory basis of genetic compensation during neural crest induction. PLOS Genet. 15, e1008213 (2019).
Cox, A. G. et al. Yap regulates glucose utilization and sustains nucleotide synthesis to enable organ growth. EMBO J. 37, e100294 (2018).
Pappalardo, A. et al. Thyroid development in zebrafish lacking Taz. Mech. Dev. 138, 268–278 (2015).
Hong, J.-H. et al. TAZ, a transcriptional modulator of mesenchymal stem cell differentiation. Sci. (80-.) 309, 1074–1078 (2005).
Yi, X. et al. The effector of Hippo signaling, Taz, is required for formation of the micropyle and fertilization in zebrafish. PLoS Genet. 15, e1007408 (2019).
Lo, H. P. et al. The caveolin–cavin system plays a conserved and critical role in mechanoprotection of skeletal muscle. J. Cell Biol. 210, 833–849 (2015).
Kim, M., Kim, M., Lee, M.-S., Kim, C.-H. & Lim, D.-S. The MST1/2-SAV1 complex of the Hippo pathway promotes ciliogenesis. Nat. Commun. 5, 1–14 (2014).
Jopling, C. et al. Zebrafish heart regeneration occurs by cardiomyocyte dedifferentiation and proliferation. Nature 464, 606–609 (2010).
Kikuchi, K. et al. Primary contribution to zebrafish heart regeneration by gata4+ cardiomyocytes. Nature 464, 601–605 (2010).
Gupta, V. et al. An injury-responsive gata4 program shapes the zebrafish cardiac ventricle. Curr. Biol. 23, 1221–1227 (2013).
Zhang, R. et al. In vivo cardiac reprogramming contributes to zebrafish heart regeneration. Nature 498, 497–502 (2013).
Harrison, M. R. M. et al. Chemokine-guided angiogenesis directs coronary vasculature formation in zebrafish. Dev. Cell 33, 442–454 (2015).
Kikuchi, K. et al. tcf21+ epicardial cells adopt non-myocardial fates during zebrafish heart development and regeneration. Development 138, 2895–2902 (2011).
Kim, J. et al. PDGF signaling is required for epicardial function and blood vessel formation in regenerating zebrafish hearts. Proc. Natl Acad. Sci. USA 107, 17206–17210 (2010).
Marín-Juez, R. et al. Coronary revascularization during heart regeneration is regulated by epicardial and endocardial cues and forms a scaffold for cardiomyocyte repopulation. Dev. Cell 51, 503–515 (2019).
Marín-Juez, R. et al. Fast revascularization of the injured area is essential to support zebrafish heart regeneration. Proc. Natl Acad. Sci. USA 113, 11237–11242 (2016).
Lepilina, A. et al. A dynamic epicardial injury response supports progenitor cell activity during zebrafish heart regeneration. Cell 127, 607–619 (2006).
Itou, J. et al. Migration of cardiomyocytes is essential for heart regeneration in zebrafish. Development 139, 4133–4142 (2012).
Tapon, N. et al. Salvador promotes both cell cycle exit and apoptosis in drosophila and is mutated in human cancer cell lines. Cell 110, 467–478 (2002).
Liu, S. et al. Gene therapy knockdown of Hippo signaling induces cardiomyocyte renewal in pigs after myocardial infarction. Sci. Transl. Med. 13, eabd6892 (2021).
Del Re, D. P. et al. Yes-associated protein isoform 1 (Yap1) promotes cardiomyocyte survival and growth to protect against myocardial ischemic injury. J. Biol. Chem. 288, 3977–3988 (2013).
von Gise, A. et al. YAP1, the nuclear target of Hippo signaling, stimulates heart growth through cardiomyocyte proliferation but not hypertrophy. Proc. Natl Acad. Sci. USA 109, 2394–2399 (2012).
Xin, M. et al. Hippo pathway effector Yap promotes cardiac regeneration. Proc. Natl. Acad. Sci. USA 110, 13839–13844 (2013).
Lin, Z. et al. Cardiac-specific YAP activation improves cardiac function and survival in an experimental murine MI model. Circ. Res. 115, 354–363 (2014).
Morikawa, Y., Heallen, T., Leach, J., Xiao, Y. & Martin, J. F. Dystrophin-glycoprotein complex sequesters Yap to inhibit cardiomyocyte proliferation. Nature 547, 227–231 (2017).
Mia, M. M. & Singh, M. K. The Hippo signaling pathway in cardiac development and diseases. Front. Cell Dev. Biol. 7, 211 (2019).
Mia, M. M. et al. Loss of Yap/Taz in cardiac fibroblasts attenuates adverse remodelling and improves cardiac function. Cardiovasc. Res. https://doi.org/10.1093/cvr/cvab205 (2021).
Xiao, Y. et al. Hippo pathway deletion in adult resting cardiac fibroblasts initiates a cell state transition with spontaneous and self-sustaining fibrosis. Genes Dev. 33, 1491–1505 (2019).
Tian, Y. et al. A microRNA-Hippo pathway that promotes cardiomyocyte proliferation and cardiac regeneration in mice. Sci. Transl. Med. 7, 279ra38 (2015).
Courtine, G. & Sofroniew, M. V. Spinal cord repair: advances in biology and technology. Nat. Med. 25, 898–908 (2019).
Tran, A. P., Warren, P. M. & Silver, J. The biology of regeneration failure and success after spinal cord injury. Physiol. Rev. 98, 881–917 (2018).
Hilton, B. J., Moulson, A. J. & Tetzlaff, W. Neuroprotection and secondary damage following spinal cord injury: concepts and methods. Neurosci. Lett. 652, 3–10 (2017).
Becker, T., Wullimann, M. F., Becker, C. G., Bernhardt, R. R. & Schachner, M. Axonal regrowth after spinal cord transection in adult zebrafish. J. Comp. Neurol. 377, 577–595 (1997).
Becker, C. G. et al. L1.1 is involved in spinal cord regeneration in adult zebrafish. J. Neurosci. 24, 7837–7842 (2004).
Becker, T. & Becker, C. G. Dynamic cell interactions allow spinal cord regeneration in zebrafish. Curr. Opin. Physiol. 14, 64–69 (2020).
Becker, C. G. & Becker, T. Neuronal regeneration from ependymo-radial glial cells: cook, little pot, cook! Dev. Cell 32, 516–527 (2015).
Rasmussen, J. P. & Sagasti, A. Learning to swim, again: axon regeneration in fish. Exp. Neurol. 287, 318–330 (2017).
Ghosh, S. & Hui, S. P. Regeneration of zebrafish CNS: adult neurogenesis. Neural Plast. 2016, 5815439 (2016).
Reimer, M. M. et al. Motor neuron regeneration in adult zebrafish. J. Neurosci. 28, 8510–8516 (2008).
Briona, L. K., Poulain, F. E., Mosimann, C. & Dorsky, R. I. Wnt/ß-catenin signaling is required for radial glial neurogenesis following spinal cord injury. Dev. Biol. 403, 15–21 (2015).
Ohnmacht, J. et al. Spinal motor neurons are regenerated after mechanical lesion and genetic ablation in larval zebrafish. Development 143, 1464–1474 (2016).
Kuscha, V. et al. Lesion-induced generation of interneuron cell types in specific dorsoventral domains in the spinal cord of adult zebrafish. J. Comp. Neurol. 520, 3604–3616 (2012).
Goldshmit, Y. et al. Fgf-dependent glial cell bridges facilitate spinal cord regeneration in Zebrafish. J. Neurosci. 32, 7477–7492 (2012).
Yu, Y.-M. et al. The extracellular matrix glycoprotein tenascin-C promotes locomotor recovery after spinal cord injury in adult zebrafish. Neuroscience 183, 238–250 (2011).
Wehner, D. et al. Wnt signaling controls pro-regenerative Collagen XII in functional spinal cord regeneration in zebrafish. Nat. Commun. 8, 126 (2017).
Gomez-Sanchez, J. A. et al. After nerve injury, lineage tracing shows that myelin and Remak Schwann cells elongate extensively and branch to form repair Schwann cells, which shorten radically on remyelination. J. Neurosci. 37, 9086–9099 (2017).
Parrinello, S. et al. EphB signaling directs peripheral nerve regeneration through Sox2-dependent Schwann cell sorting. Cell 143, 145–155 (2010).
Klatt Shaw, D. et al. Localized EMT reprograms glial progenitors to promote spinal cord repair. Dev. Cell 56, 613.e7–626.e7(2021).
Wilson, M. M., Weinberg, R. A., Lees, J. A. & Guen, V. J. Emerging mechanisms by which EMT programs control stemness. Trends Cancer 6, 775–780 (2020).
Ye, X. & Weinberg, R. A. Epithelial-mesenchymal plasticity: a central regulator of cancer progression. Trends Cell Biol. 25, 675–686 (2015).
Jessen, K. R. & Arthur-Farraj, P. Repair Schwann cell update: adaptive reprogramming, EMT, and stemness in regenerating nerves. Glia 67, 421–437 (2019).
Mokalled, M. H. et al. Injury-induced ctgfa directs glial bridging and spinal cord regeneration in zebrafish. Science 354, 630–634 (2016).
Bradbury, E. J. & Burnside, E. R. Moving beyond the glial scar for spinal cord repair. Nat. Commun. 10, 3879 (2019).
Hu, R. et al. Glial scar and neuroregeneration: histological, functional, and magnetic resonance imaging analysis in chronic spinal cord injury. J. Neurosurg. 13, 169–180 (2010).
Wang, Y. et al. Lentivirus-mediated silencing of the CTGF gene suppresses the formation of glial scar tissue in a rat model of spinal cord injury. Spine J. 18, 164–172 (2018).
NIH, U. S. National Library of Medicine. ClinicalTrials.gov https://clinicaltrials.gov/ct2/results?cond=&term=ctgf&cntry=&state=&city=&dist (2021).
Hamon, A. et al. Linking YAP to Müller glia quiescence exit in the degenerative retina. Cell Rep. 27, 1712–1725 (2019).
Hamon, A., Roger, J. E., Yang, X.-J. & Perron, M. Müller glial cell-dependent regeneration of the neural retina: an overview across vertebrate model systems. Dev. Dyn. 245, 727–738 (2016).
Rueda, E. M. et al. The Hippo pathway blocks mammalian retinal Müller glial cell reprogramming. Cell Rep. 27, 1637–1649 (2019).
Wan, J. & Goldman, D. Retina regeneration in zebrafish. Curr. Opin. Genet. Dev. 40, 41–47 (2016).
Bernardos, R. L., Barthel, L. K., Meyers, J. R. & Raymond, P. A. Late-stage neuronal progenitors in the retina are radial Müller glia that function as retinal stem cells. J. Neurosci. 27, 7028–7040 (2007).
Thummel, R., Kassen, S. C., Montgomery, J. E., Enright, J. M. & Hyde, D. R. Inhibition of Müller glial cell division blocks regeneration of the light-damaged zebrafish retina. Dev. Neurobiol. 68, 492–408 (2008).
Hoang, T. et al. Gene regulatory networks controlling vertebrate retinal regeneration. Science 370, eabb8598 (2020).
Richeldi, L. et al. Pamrevlumab, an anti-connective tissue growth factor therapy, for idiopathic pulmonary fibrosis (PRAISE): a phase 2, randomised, double-blind, placebo-controlled trial. Lancet Respir. Med. 8, 25–33 (2020).
Raghu, G. et al. FG-3019 anti-connective tissue growth factor monoclonal antibody: results of an open-label clinical trial in idiopathic pulmonary fibrosis. Eur. Respir. J. 47, 1481–1491 (2016).
Ramazani, Y. et al. Connective tissue growth factor (CTGF) from basics to clinics. Matrx Biol. 68–69, 44–66 (2018).
Azevedo, A. S., Grotek, B., Jacinto, A., Weidinger, G. & Saúde, L. The regenerative capacity of the zebrafish caudal fin is not affected by repeated amputations. PLoS One 6, e22820 (2011).
Broussonet, M. Observations sur la regeneration de quelques parties du corps des poissons. Hist. d. l’Acad. R. des Sci. 105, 625–641 (1786).
Morgan, T. H. Regeneration (Macmillan, 1901).
Morgan, T. H. Regeneration in teleosts. Arch. f.ür. Entwickslungsmech Org. 10, 120–131 (1900).
Sunderland, M. E. Regeneration: Thomas hunt Morgan’s window into development. J. Hist. Biol. 43, 325–361 (2010).
Pfefferli, C. & Jazwinska, A. The art of fin regeneration in zebrafish. Regeneration 2, 72–83 (2015).
Cao, X., Pfaff, S. L. & Gage, F. H. YAP regulates neural progenitor cell number via the TEA domain transcription factor. Genes Dev. 22, 3320–3334 (2008).
Panciera, T. et al. Induction of expandable tissue-specific stem/progenitor cells through transient expression of YAP/TAZ. Cell Stem Cell 19, 725–737 (2016).
Patel, S. H., Camargo, F. D. & Yimlamai, D. Hippo signaling in the liver regulates organ size, cell fate, and carcinogenesis. Gastroenterology 152, 533–545 (2017).
Camargo, F. D. et al. YAP1 increases organ size and expands undifferentiated progenitor cells. Curr. Biol. 17, 2054–2060 (2007).
Karpowicz, P., Perez, J. & Perrimon, N. The Hippo tumor suppressor pathway regulates intestinal stem cell regeneration. Development 138, 4135–4145 (2010).
Mo, J.-S., Park, H. W. & Guan, K.-L. The Hippo signaling pathway in stem cell biology and cancer. EMBO Rep. 15, 642–656 (2014).
Lian, I. et al. The role of YAP transcription coactivator in regulating stem cell self-renewal and differentiation. Genes Dev. 24, 1106–1118 (2010).
Hasegawa, T. et al. Transient inflammatory response mediated by interleukin-1β is required for proper regeneration in zebrafish fin fold. Elife 6, e22716 (2017).
Li, J. et al. Expression analysis of Hsp90α and cytokines in zebrafish caudal fin regeneration. Dev. Comp. Immunol. 116, 103922 (2021).
Nguyen-Chi, M. et al. TNF signaling and macrophages govern fin regeneration in zebrafish larvae. Cell Death Dis. 8, e2979 (2017).
Tornini, V. A. et al. Live monitoring of Blastemal cell contributions during appendage regeneration. Curr. Biol. 26, 2981–2991 (2016).
Lee, Y., Grill, S., Sanchez, A., Murphy-Ryan, M. & Poss, K. D. Fgf signaling instructs position-dependent growth rate during zebrafish fin regeneration. Development 132, 5173–5183 (2005).
Shibata, E., Liu, Z., Kawasaki, T., Sakai, N. & Kawakami, A. Robust and local positional information within a fin ray directs fin length during zebrafish regeneration. Dev. Growth, Differ. 60, 354–364 (2018).
Nechiporuk, A. & Keating, M. T. A proliferation gradient between proximal and msxb-expressing distal blastema directs zebrafish fin regeneration. Development 129, 2607–2617 (2002).
Mateus, R. et al. Control of tissue growth by Yap relies on cell density and F-actin in zebrafish fin regeneration. J. Cell Sci. 128, e1.2–e1.2 (2015).
Brandão, A. S. et al. Yap induces osteoblast differentiation by modulating Bmp signalling during zebrafish caudal fin regeneration. J. Cell Sci. 132, jcs231993 (2019).
Moro, A. et al. MicroRNA-dependent regulation of biomechanical genes establishes tissue stiffness homeostasis. Nat. Cell Biol. 21, 348–358 (2019).
Stewart, S. & Stankunas, K. Limited dedifferentiation provides replacement tissue during zebrafish fin regeneration. Dev. Biol. 365, 339–349 (2012).
Tu, S. & Johnson, S. L. Fate restriction in the growing and regenerating zebrafish fin. Dev. Cell 20, 725–732 (2011).
Knopf, F. et al. Bone regenerates via dedifferentiation of osteoblasts in the zebrafish fin. Dev. Cell 20, 713–724 (2011).
Sousa, S. et al. Differentiated skeletal cells contribute to blastema formation during zebrafish fin regeneration. Development 138, 3897–3905 (2011).
Stewart, S., Gomez, A. W., Armstrong, B. E., Henner, A. & Stankunas, K. Sequential and opposing activities of Wnt and BMP coordinate zebrafish bone regeneration. Cell Rep. 6, 482–498 (2014).
Smith, A., Avaron, F., Guay, D., Padhi, B. K. & Akimenko, M. A. Inhibition of BMP signaling during zebrafish fin regeneration disrupts fin growth and scleroblast differentiation and function. Dev. Biol. 299, 438–454 (2006).
Murciano, C. et al. Ray–interray interactions during fin regeneration of Danio rerio. Dev. Biol. 252, 214–224 (2002).
Halder, G., Dupont, S. & Piccolo, S. Transduction of mechanical and cytoskeletal cues by YAP and TAZ. Nat. Rev. Mol. Cell Biol. 13, 591–600 (2012).
Luo, Q. et al. Connective tissue growth factor (CTGF) is regulated by Wnt and bone morphogenetic proteins signaling in osteoblast differentiation of mesenchymal stem cells. J. Biol. Chem. 279, 55958–55968 (2004).
Kegelman, C. D. et al. Skeletal cell YAP and TAZ combinatorially promote bone development. FASEB J. 32, 2706–2721 (2018).
Pan, J.-X. et al. YAP promotes osteogenesis and suppresses adipogenic differentiation by regulating β-catenin signaling. Bone Res. 6, 18 (2018).
Seo, E. et al. SOX2 regulates YAP1 to maintain stemness and determine cell fate in the osteo-adipo lineage. Cell Rep. 3, 2075–2087 (2013).
Xiong, J., Almeida, M. & O’Brien, C. A. The YAP/TAZ transcriptional co-activators have opposing effects at different stages of osteoblast differentiation. Bone 112, 1–9 (2018).
Rinkevich, Y., Lindau, P., Ueno, H., Longaker, M. T. & Weissman, I. L. Germ-layer and lineage-restricted stem/progenitors regenerate the mouse digit tip. Nature 476, 409–414 (2011).
Grusche, F. A., Degoutin, J. L., Richardson, H. E. & Harvey, K. F. The Salvador/Warts/Hippo pathway controls regenerative tissue growth in Drosophila melanogaster. Dev. Biol. 350, 255–266 (2011).
Nuñez, V. A. et al. Postembryonic development of the posterior lateral line in the zebrafish. Evol. Dev. 11, 391–404 (2009).
Alexandre, D. & Ghysen, A. Somatotopy of the lateral line projection in larval zebrafish. Proc. Natl. Acad. Sci. USA 96, 7558–7562 (1999).
Ledent, V. Postembryonic development of the posterior lateral line in zebrafish. Development 129, 597–604 (2002).
Sarrazin, A. F., Nuñez, V. A., Sapède, D., Tassin, V. & Dambly-Chaudière, C. Origin and early development of the posterior lateral line system of zebrafish. J. Neurosci. 30, 8234–8244 (2010).
Lush, M. E. & Piotrowski, T. Sensory hair cell regeneration in the zebrafish lateral line. Dev. Dyn. 243, 1187–1202 (2014).
Lecaudey, V., Cakan-Akdogan, G., Norton, W. H. J. & Gilmour, D. Dynamic Fgf signaling couples morphogenesis and migration in the zebrafish lateral line primordium. Development 135, 2695–2705 (2008).
Atkinson, P. J., Huarcaya Najarro, E., Sayyid, Z. N. & Cheng, A. G. Sensory hair cell development and regeneration: similarities and differences. Development 142, 1561–1571 (2015).
Kim, N.-G., Koh, E., Chen, X. & Gumbiner, B. M. E-cadherin mediates contact inhibition of proliferation through Hippo signaling-pathway components. Proc. Natl. Acad. Sci. USA 108, 11930–11935 (2011).
Benham-Pyle, B. W., Pruitt, B. L. & Nelson, W. J. Mechanical strain induces E-cadherin-dependent Yap1 and β-catenin activation to drive cell cycle entry. Science 348, 1024–1027 (2015).
Kozlovskaja-Gumbriene, A. et al. Proliferation-independent regulation of organ size by Fgf/Notch signaling. Elife 6, e21049 (2017).
Loh, S. L. et al. Zebrafish yap1 plays a role in differentiation of hair cells in posterior lateral line. Sci. Rep. 4, 1–9 (2014).
Agarwala, S. et al. Amotl2a interacts with the Hippo effector Yap1 and the Wnt/β-catenin effector Lef1 to control tissue size in zebrafish. Elife 4, e08201 (2015).
Wang, X. et al. Lpar2b controls lateral line tissue size by regulating Yap1 activity in zebrafish. Front. Mol. Neurosci. 11, 34 (2018).
Pistocchi, A. et al. The zebrafish prospero homolog prox1 is required for mechanosensory hair cell differentiation and functionality in the lateral line. BMC Dev. Biol. 9, 58 (2009).
Yung, Y. C., Stoddard, N. C. & Chun, J. LPA receptor signaling: pharmacology, physiology, and pathophysiology. J. Lipid Res. 55, 1192–1214 (2014).
Yu, F.-X. et al. Regulation of the Hippo-YAP pathway by G-protein-coupled receptor signaling. Cell 150, 780–791 (2012).
Okudaira, S., Yukiura, H. & Aoki, J. Biological roles of lysophosphatidic acid signaling through its production by autotaxin. Biochimie 92, 698–706 (2010).
Lai, S.-L. et al. Autotaxin/Lpar3 signaling regulates Kupffer’s vesicle formation and left-right asymmetry in zebrafish. Development 139, 4439–4448 (2012).
Frisca, F. et al. Role of ectonucleotide pyrophosphatase/phosphodiesterase 2 in the midline axis formation of zebrafish. Nature 6, 37678 (2016).
Yukiura, H. et al. Autotaxin regulates vascular development via multiple Lysophosphatidic Acid (LPA) receptors in zebrafish. J. Biol. Chem. 286, 43972–43983 (2011).
Lee, S.-J. et al. LPA1 is essential for lymphatic vessel development in zebrafish. FASEB J. 22, 3706–3715 (2008).
Stone, L. S. The development of lateral-line sense organs in amphibians observed in living and vital-stained preparations. J. Comp. Neurol. 57, 507–540 (1933).
Stone, L. S. Further experimental studies of the development of lateral-line sense organs in amphibians observed in living preparations. J. Comp. Neurol. 68, 83–115 (1937).
Dijkgraaf, S. The functional and significance of the lateral-line organs. Biol. Rev. 38, 51–105 (1962).
Ma, E. Y., Rubel, E. W. & Raible, D. W. Notch signaling regulates the extent of hair cell regeneration in the zebrafish lateral line. J. Neurosci. 28, 2261–2273 (2008).
Namdaran, P., Reinhart, K. E., Owens, K. N., Raible, D. W. & Rubel, E. W. Identification of modulators of hair cell regeneration in the zebrafish lateral line. J. Neurosci. 32, 3516–3528 (2012).
López-Schier, H. & Hudspeth, A. J. A two-step mechanism underlies the planar polarization of regenerating sensory hair cells. Proc. Natl. Acad. Sci. USA 103, 18615–18620 (2006).
Ye, Z. et al. Yap-lin28a axis targets let7-Wnt pathway to restore progenitors for initiating regeneration. Elife 9, e55771 (2020).
Aman, A. & Piotrowski, T. Wnt/β-catenin and Fgf signaling control collective cell migration by restricting chemokine receptor expression. Dev. Cell 15, 749–761 (2008).
Jiang, L., Romero-Carvajal, A., Haug, J. S., Seidel, C. W. & Piotrowski, T. Gene-expression analysis of hair cell regeneration in the zebrafish lateral line. Proc. Natl. Acad. Sci. USA 111, E1383–E1392 (2014). https://doi.org/10.1073/pnas.1402898111
Giffen, K. P., Liu, H., Kramer, K. L. & He, D. Z. Expression of protein-coding gene orthologs in zebrafish and mouse inner ear non-sensory supporting cells. Front. Neurosci. 13, 1117 (2019).
Jin, J. et al. Evidence that Lin28 stimulates translation by recruiting RNA helicase A to polysomes. Nucleic Acids Res. 39, 3724–3734 (2011).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Sci. (80-.) 318, 1917–1920 (2007).
Zhang, J. et al. LIN28 regulates stem cell metabolism and conversion to primed pluripotency. Cell Stem Cell 19, 66–80 (2016).
Golden, E. J., Benito-Gonzalez, A. & Doetzlhofer, A. The RNA-binding protein LIN28B regulates developmental timing in the mammalian cochlea. Proc. Natl. Acad. Sci. USA 112, E3864–E3873 (2015).
Ramachandran, R., Fausett, B. V. & Goldman, D. Ascl1a regulates Müller glia dedifferentiation and retinal regeneration through a Lin-28-dependent, let-7 microRNA signalling pathway. Nat. Cell Biol. 12, 1101–1107 (2010).
Corey, D. P. et al. TRPA1 is a candidate for the mechanosensitive transduction channel of vertebrate hair cells. Nature 432, 723–730 (2004).
Siemens, J. et al. Cadherin 23 is a component of the tip link in hair-cell stereocilia. Nature 428, 950–955 (2004).
Söllner, C. et al. Mutations in cadherin 23 affect tip links in zebrafish sensory hair cells. Nature 428, 955–959 (2004).
Santos, F., Macdonald, G., Rubel, E. W. & Raible, D. W. Lateral line hair cell maturation is a determinant of aminoglycoside susceptibility in zebrafish (Danio rerio). Hear. Res. 213, 25–33 (2006).
Buck, L. M. J., Winter, M. J., Redfern, W. S. & Whit, T. T. Ototoxin-induced cellular damage in neuromasts disrupts lateral line function in larval zebra fish. Hear. Res. 284, 67–81 (2012).
Forbes, S. J. & Newsome, P. N. Liver regeneration—mechanisms and models to clinical application. Nat. Rev. Gastroenterol. Hepatol. 13, 473–485 (2016).
Wang, S., Miller, S. R., Ober, E. A. & Sadler, K. C. Making it new again: insight into liver development, regeneration, and disease from zebrafish. Curr. Top. Dev. Biol. 124, 161–195 (2017).
Wang, C. et al. Differences in Yes-associated protein and mRNA levels in regenerating liver and hepatocellular carcinoma. Mol. Med. Rep. 5, 410–414 (2012).
Grijalva, J. L. et al. Dynamic alterations in Hippo signaling pathway and YAP activation during liver regeneration. Am. J. Physiol. Gastrointest. Liver Physiol. 307, G196–G204 (2014).
Bai, H. et al. Yes-associated protein regulates the hepatic response after bile duct ligation. Hepatology 56, 1097–1107 (2012).
Aloia, L. et al. Epigenetic remodelling licences adult cholangiocytes for organoid formation and liver regeneration. Nat. Cell Biol. 21, 1321–1333 (2019).
Li, W. et al. A homeostatic arid1a-dependent permissive chromatin state licenses hepatocyte responsiveness to liver-injury-associated YAP signaling. Cell Stem Cell 25, 54–68 (2019).
Dong, J. et al. Elucidation of a universal size-control mechanism in Drosophila and mammals. Cell 130, 1120–1133 (2007).
Sadler, K. C., Amsterdam, A., Soroka, C., Boyer, J. & Hopkins, N. A genetic screen in zebrafish identifies the mutants vps18, nf2 and foie gras as models of liver disease. Development 132, 3561–3572 (2005).
Cox, A. G. et al. Yap reprograms glutamine metabolism to increase nucleotide biosynthesis and enable liver growth. Nat. Cell Biol. 18, 886–896 (2016).
Brandt, Z. J., Echert, A. E., Bostrom, J. R., North, P. N. & Link, B. A. Core Hippo pathway components act as a brake on Yap/Taz in the development and maintenance of the biliary network. Development 4, dev.184242 (2020).
Driskill, J. H. & Pan, D. The Hippo pathway in liver homeostasis and pathophysiology. Annu. Rev. Pathol. Mech. Dis. 16, 299–322 (2021).
Yimlamai, D. et al. Hippo pathway activity influences liver cell fate. Cell 157, 1324–1338 (2014).
Molina, L. M. et al. Compensatory hepatic adaptation accompanies permanent absence of intrahepatic biliary network due to YAP1 loss in liver progenitors. Cell Rep. 36, 109310 (2021).
Verboven, E. et al. Regeneration defects in Yap and Taz mutant mouse livers are caused by bile duct disruption and cholestasis. Gastroenterology 160, 847–862 (2021).
MacCarthy-Morrogh, L. & Martin, P. The hallmarks of cancer are also the hallmarks of wound healing. Sci. Signal. 13, eaay8690 (2020).
Moroishi, T., Gram Hansen, C. & Guan, K.-L. The emerging roles of YAP and TAZ in cancer. Nat. Rev. Cancer 15, 73–79 (2015).
Johnson, R. & Halder, G. The two faces of Hippo: targeting the Hippo pathway for regenerative medicine and cancer treatment. Nat. Rev. Drug Discov. 13, 63–79 (2014).
Merkes, C. et al. Ewing sarcoma Ewsa protein regulates chondrogenesis of Meckel’s Cartilage through modulation of Sox9 in zebrafish. PLoS One 10, e0116627 (2015).
Mayrhofer, M. et al. A novel brain tumour model in zebrafish reveals the role of YAP activation in MAPK- and PI3K-induced malignant growth. Dis. Model. Mech. 10, 15–28 (2017).
Brandt, Z. J., North, P. N. & Link, B. A. Somatic mutations of lats2 cause peripheral nerve sheath tumors in zebrafish. Cells 8, 972 (2019).
Acknowledgements
Work ongoing in the Gram Hansen lab is supported by a University of Edinburgh Chancellor’s Fellowship as well as by Worldwide Cancer Research (19-0238) and LifeArc-CSO. Additional funding has been obtained from the Bone Cancer Research Trust (BCRT), Sarcoma UK (SUK202.2016), the Wellcome Trust-University of Edinburgh Institutional Strategic Support Fund (ISSF3). Y.F. received a Wellcome Trust Sir Henry Dale Fellowship (100104/Z/12/Z); Cancer Research UK early detection project award (C38363/A26931) and a Cancer Research UK pioneering award (C38363/A25107). S.E.R. is funded by a Wellcome Trust PhD studentship [108906/Z/15/Z]. Figures, besides Fig. 1, created in Adobe Illustrator.
Author information
Authors and Affiliations
Contributions
S.E.R. and C.G.H. drafted the manuscript. S.E.R., Y.F., and C.G.H. revised the manuscript. C.G.H. and Y.F. supervised the project. S.E.R. prepared the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Riley, S.E., Feng, Y. & Hansen, C.G. Hippo-Yap/Taz signalling in zebrafish regeneration. npj Regen Med 7, 9 (2022). https://doi.org/10.1038/s41536-022-00209-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41536-022-00209-8
- Springer Nature Limited