Abstract
Parkinson’s disease (PD) is the neurological disorder showing the greatest rise in prevalence from 1990 to 2016. Despite clinical definition criteria and a tremendous effort to develop objective biomarkers, precise diagnosis of PD is still unavailable at early stage. In recent years, an increasing number of studies have used omic methods to unveil the molecular basis of PD, providing a detailed characterization of potentially pathological alterations in various biological specimens. Metabolomics could provide useful insights to deepen our knowledge of PD aetiopathogenesis, to identify signatures that distinguish groups of patients and uncover responsive biomarkers of PD that may be significant in early detection and in tracking the disease progression and drug treatment efficacy. The present work is the first large metabolomic study based on nuclear magnetic resonance (NMR) with an independent validation cohort aiming at the serum characterization of de novo drug-naive PD patients. Here, NMR is applied to sera from large training and independent validation cohorts of German subjects. Multivariate and univariate approaches are used to infer metabolic differences that characterize the metabolite and the lipoprotein profiles of newly diagnosed de novo drug-naive PD patients also in relation to the biological sex of the subjects in the study, evidencing a more pronounced fingerprint of the pathology in male patients. The presence of a validation cohort allowed us to confirm altered levels of acetone and cholesterol in male PD patients. By comparing the metabolites and lipoproteins levels among de novo drug-naive PD patients, age- and sex-matched healthy controls, and a group of advanced PD patients, we detected several descriptors of stronger oxidative stress.
Similar content being viewed by others
Introduction
Parkinson’s disease (PD), after Alzheimer’s disease, is the second most prevalent neurodegenerative disease, affecting 1% of the population over the age of 601, and contributing significantly to the rise in the cost of public health. Despite many years of investigations on metabolic perturbances in PD2,3,4,5,6,7, the mechanisms of aetiopathogenesis, progression, and efficacy of drug treatment on the disease evolution need to be further explored. Currently, routine analytical tests have not yet provided sufficient information to identify reliable blood biomarkers to detect early signs of PD, to monitor the progression of the disease and to detect the effects of therapy intervention.
Metabolomics has become a powerful tool to characterize the biochemistry underlying the onset of different diseases and provides useful applications in the biomedical field.8,9,10,11,12,13. Nuclear Magnetic Resonance (NMR) is one of the most used analytical techniques for metabolomic investigations and it proved to be efficient in characterizing the metabolic composition of different biospecimens in the context of molecular medicine14,15,16,17,18,19,20,21,22,23. The NMR approach allows one to profile both small metabolites and lipoproteins.
To date, few metabolomic studies focused on de novo drug-naive PD patients (dn2PD). Most of them used limited number of subjects (<50)24,25 and the results are not always overlapping nor concordant, possibly because of the different analytical approaches or inclusion of PD patients with various phenotypes and stages of PD (genetic or idiopathic PD; de novo or advanced PD patients, etc.)11,26, or because of the limited sample size.
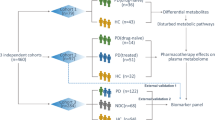
In this work, we look for the presence of a metabolomic fingerprint of Parkinson’s disease in the sera of dn2PD, and, if present, whether it is sex specific. To do so, we considered 307 subjects, both dn2PD (n = 228) and healthy controls (CTR, n = 79), partitioned between two large independent training and validation cohorts. The peculiar aspect of this collection is the presence of many dn2PD patients, to explore the serum biochemistry of the pathology independently of the effects of the antidopaminergic treatment. Additionally, a cohort of 22 advanced PD patients under dopaminergic treatment (advPD) was included to check and test for serum alterations in the disease progression. An overview of the present study design is illustrated in Fig. 1. The subjects in this study are a subset of the cohorts analyzed in the H2020 Project “PROPAG-AGEING” (www.propag-ageing.eu/project)27. To our best knowledge, this is the first large NMR-based study with an independent validation cohort aiming at the serum characterization of dn2PD patients.
Results
Exploratory analysis
As a first unsupervised approach, PCA was applied on all available samples (from both training and validation cohorts) to obtain an overview of the variation in the data. Figure 2 shows the PCA 3D score plot on bucketed 1D NOESY spectra color-coded by subject status proving the absence of outlier samples.
Predictive modeling: OPLS-DA analysis
Disease fingerprinting
OPLS-DA analysis was chosen as supervised approach to extrapolate the hidden variables that could be used to characterize the serum fingerprint28 (considering the whole NMR spectrum, therefore including the resonance signals arising from metabolites and lipoproteins) of dn2PD. First, the training cohort was used to explore differences between dn2PDs and CTRs and to derive a discriminant serum fingerprint to correctly assign the new samples according to the diagnosis (dn2PD or CTR). The OPLS-DA model built on bucketed serum 1D NOESY spectra of the training cohort showed an evident discrimination between subject groups, featuring an overall mean predictive accuracy of 75.3%, a specificity of 75.0%, and a sensitivity of 75.3% for the classification of dn2PD and CTR (Table 1, Supplementary Fig. 1a).
Sex-dependent fingerprint
Since it has been widely demonstrated that differences between the two sexes could affect manifestation, epidemiology, and pathophysiology of many diseases, and sex discrimination is apparent in metabolomics profiles29,30,31,32,33,34, two independent cross-validated models were created for male (Table 1, Supplementary Fig. 1b) and female training subjects (Table 1, Supplementary Fig. 1c). As shown in Table 1 the male model provides a better discrimination, with an overall predictive accuracy of 73.0% compared to the 61.6% predictive accuracy of the female model.
Model validation
An independent cohort was used to validate the existence of a serum metabolic fingerprint differentiating CTRs from dn2PD patients. Indeed, the efficacy of the global training model in discriminating dn2PD serum samples from CTR was tested using bucketed 1D NOESY spectra from the validation cohort, which were blindly projected onto the OPLS-DA discrimination space obtained from the previously described training model built on both sexes together (Supplementary Fig. 2). New test samples were predicted with an overall accuracy of 74.4%, a specificity of 65%, and a sensitivity of 75.6% (Table 2, Supplementary Fig. 2a, b), thus confirming the presence of a fingerprint discriminating CTRs from dn2PD patients. Then, training male and female models were used separately to blindly predict test samples (Table 2). As expected, the model built on the male group performs better in terms of accuracy, sensitivity, and specificity, which are around 70%, while the female model shown completely unbalanced performance values with a sensitivity as low as 41.7% (Table 2).
advPD vs dn2PD
Furthermore, OPLS-DA models were created with bucketed 1D NOESY to visualize the discrimination accuracies in the comparison of advPD patients with the dn2PD patients and then with CTR subjects. In the first comparison, we obtained an overall mean predictive accuracy of 87.4% (CI = 76.2–95.2%), a mean specificity of 85.1 (CI = 71.4–100%), and a mean sensitivity of 89.7 (CI = 76.2–100%), evidencing a clear discrimination between dn2PD and advPD. Optimal performances were obtained also by comparing the advPD patients’ spectra with those from CTRs, resulting in a mean accuracy of 86.9% (CI = 81.6–97.4%), a mean specificity of 87.5% (CI = 78.9–100%), and a mean sensitivity of 86.4% (CI = 78.9–94.7%). Further, projecting onto the OPLS-DA discrimination space built on the training models (dn2PD vs CTRs) the 22 bucketed serum 1D-NOESY spectra of advPD subjects, the presence of a serum metabolic signature characterizing healthy from diseased subjects at different stages of the pathology was highlighted. Indeed, all the advPD male and female patients resulted to be correctly classified as PD patients, with an accuracy of 100%. This result supports the idea of the presence of a specific metabolomic signature of PD in sera of affected patients, regardless of the dopaminergic treatment, as the use of PD medication by advPD patients does not influence the classification of those subjects in a model where only patients free from L-DOPA administration are included.
Metabolite and lipoprotein profiles
Profiling of dn2PD
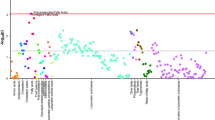
Based on multivariate results on NMR spectra, univariate statistics were applied on training cohort samples by keeping subjects separate according to the sex. Corroborating what previously demonstrated by the multivariate models, only male dn2PD patients showed a significantly different profile in terms of metabolites and lipoproteins compared to CTR males (Supplementary Table 1). A total of 26 compounds (metabolites and lipoproteins) resulted to be significantly different (FDR < 0.05) in the comparison of male dn2PD patients and CTR males: acetone, ornithine, and phenylalanine appear to be significantly higher in dn2PD patients with respect to CTRs, while all the 23 significant lipoproteins decrease in the dn2PD group (Fig. 3, Supplementary Table 1). Instead, no significant differences were detected when comparing female dn2PD patients with female CTRs of the training group, as reported in Supplementary Table 1. Binomial logistic regression models were built using the significant metabolites obtained by comparing the serum profile of male CTR and male dn2PD patients of the training cohort (see Fig. 3). The corresponding AUCs (area under the curve) were calculated (Table 3).
Looking at the discrimination between the two male groups (dn2PD and CTR), all the selected metabolites and lipoproteins have a probability between 68.4 and 80.2% to distinguish case (dn2PD) from the control (CTR) group. A model based on the concentrations of 27 metabolites and 111 lipoproteins was built to assess its performance in discriminating male dn2PD patients and male CTR subjects. In Supplementary Table 2 the performances obtained in the dn2PD vs CTR model (training cohort + validation cohort) are compared: (i) using the whole NMR spectrum; (ii) using the concentrations of the identified metabolites and lipoproteins. The former approach appears to perform slightly better (mean accuracy: 77.2%) than the latter (mean accuracy: 73.05%). The list of variable importance in projection (VIP) scores obtained with the latter approach is reported in Supplementary Fig. 3.
Profiling disease progression
To explore metabolic variations in more detail, serum metabolic profiles of all dn2PD, advPD patients, and CTRs were investigated combining training and validation cohort samples in a single heatmap (both males and females) showing the top 30 analytes identified by t-test (Fig. 4). In the two-way hierarchical clustering heatmap, groups and compounds are separated using hierarchical clustering (Ward’s algorithm), with the dendrogram being scaled to represent the distance between each branch (distance measure: Pearson’s correlation). In Fig. 4, a distinct profile signature emerges between PD patients (dn2PD and advPD) and CTRs as depicted by the column dendrogram. PD patients are characterized by higher levels of acetone compared to CTRs. Furthermore, the levels of acetone, formic acid, and histidine result to be higher in dn2PD than the advPD and CTR subjects. Instead, the PD profile seems to be characterized by lower levels of cholesterol (Chol), low-density lipoproteins (LDL), ApoB100, LDL-cholesterol (esterified and free), phospholipids, and ApoB concentrations in LDL and triglycerides content in LDL fractions (LDL, LDL3, LDL4, and LDL5). Additionally, we notice a marked decrease of ApoA1 lipoprotein and ApoA1 content in the HDL4 fraction, of free cholesterol in the HDL and HDL4 fractions, and of citric acid along the series CTRs>dn2PD > advPD. Likewise, a reduction is also observed in the concentration of LDL4, ApoB in LDL4, cholesterol (esterified and free), and phospholipids in LDL4 and LDL5 fractions. Finally, a lowering of ApoA2-HDL3, N,N-dimethylglycine, and methionine also occurs.
Top features ranked by t-test to retain the most contrasting patterns. Heatmap displays average features concentrations for each group (CTR, dn2PD, and advPD). The horizontal axis represents the groups, and the vertical axis represents 30 selected features concentrations in which features with similar trends cluster in rows. The magnitude of abundance change (“red” increased or “blue” decreased) is shown in accordance with the color scale on the right.
Discussion
To our knowledge, this is the first large NMR-based metabolomics study dealing with the characterization of serum profiles of de novo Parkinson’s disease patients free from dopaminergic treatment compared to healthy sex/age-matched volunteers. In this context, advanced PD patients under dopaminergic treatment are also examined. The use of many well-characterized dn2PD and the presence of an external validation cohort represent major strengths of this study. Fingerprinting of the full NMR spectra discriminates dn2PD and CTR with a predictive accuracy of 75.3%. The NMR spectra of advPD classify them in the dn2PD group with 100% accuracy, proving the existence of a PD serum signature which is clearly recognizable even in the presence of the confounding effect of the dopaminergic therapy.
A more pronounced fingerprint and profile of PD is found in men with respect to women. Increasing evidence points to sex as a crucial determining factor in the development and phenotypical expression of PD. The risk of developing PD is a factor of two higher in men than in women35,36. Our observation could support the idea that disease development, at least in dn2PD patients, might involve different pathogenetic mechanisms in male and female subjects.
Multivariate analysis detects significant alterations of several metabolites and lipoproteins in PD patients, some of which also survive the scrutiny of univariate analysis. The observation of acetone being higher in dn2PD patients reinforces previous studies8 and is in line with a general condition of oxidative stress and of increased oxidative stress defense37,38. Acetone is a waste product formed from acetoacetic acid. Accumulation of acetone in blood and in breath is common in diabetes, in patients during anesthesia or surgical stress. Other factors such as body weight, age, and especially alcoholism influence the resting blood concentration of acetone39. Higher acetone, 3-hydroxybutyrate, and acetate levels have been described also in blood samples from patients with other neurodegenerative diseases, such as multiple sclerosis40 and amyotrophic lateral sclerosis41. To date, the biological reason for serum acetone to be associated with the onset of PD remains not completely elucidated.
Formic acid and histidine are significantly higher in dn2PD when compared with advPD patients and CTRs. Formic acid has been classically identified as mitochondrial toxin and could be also potentially implied in the dopaminergic pathway, explaining the lower level of this molecule in patients under L-DOPA treatment. Instead, literature data report a controversial role of histidine regarding its pro-/anti-oxidant activity8,42. Increased concentrations of histidine and phenylalanine have been recently observed also in saliva of PD patients43, thus suggesting alterations in neurotransmitters, especially dopamine44. Other detected metabolites, such as citric acid, methionine, and N,N-dimethylglycine, playing important roles against oxidative stress damage45,46,47, have been found downregulated in our PD groups. Citric acid alteration could explain the interplay between oxidants and energy metabolism in PD48. Methionine and N,N-dimethylglycine are produced by the betaine-homocysteine methyltransferase from homocysteine and betaine. Other studies suggest accumulation of homocysteine in blood and CSF as a risk factor for PD and dementia49,50, and this could explain lower levels of the end products in PD patients.
Increased levels of ornithine have been previously detected in serum samples of de novo PD and advanced PD patients6. Here, we found higher level of ornithine only in the male dn2PD group compared to CTR subjects in the training cohort. However, this metabolite is not significantly altered in the male validation cohort, nor was it found to be altered in the dn2PD female population.
Several studies pose lipids and lipoproteins as central players in Parkinson’s disease51,52,53,54,55,56,57,58. Various subclasses of fatty acids, glycerolipids, phospholipids, sterol, and lipoproteins may contribute to PD pathogenesis. However, controversial, fragmented and not always reproducible data are available in the literature. Mollenhauer et al.59 reported lower serum cholesterol levels in PD patients, independent of nutritional status and body mass index (BMI); dysregulation of cholesterol trafficking was shown to be involved in the pathogenesis of neurodegeneration in PD. Higher cholesterol and LDL-Chol levels might attenuate neurodegenerative process in PD-affected males60. Consistently, we found lower cholesterol levels in male patients at early PD stage. For the first time, we also characterize the lipoprotein subclasses profile of serum samples of PD patients. Our results point to a statistically significant decrease of the concentration of small low-density lipoproteins in dn2PD patients. Lower LDL-related parameters, especially LDL-cholesterol levels, have been associated with higher PD risk52,53,61 and generally, small dense LDL particles (sdLDL) are more susceptible to oxidation than larger LDLs. Therefore, we suggest that sdLDL particles may provide an optimal substrate for ROS oxidative action, which appears to be increased in dn2PD patients54,55,58.
An interesting result of the present research is that females have a much weaker PD fingerprint than males. Other investigations have found that there are differences in the serum profiles of PD males and females. Because of the scarcity of sex-specific analyses in PD research, the complicated connection between sex and metabolism in relation to PD remains unknown. Further research should consider the modulatory effect of sex hormones, blood metabolites, and PD.
Our data indicate serum alterations which are consistent with increase oxidative stress in PD, suggesting that the early stage of PD may be characterized by increased oxidative defenses and a worsening of the oxidative stress status. The signature of such process is detectable in circulating metabolites and lipoproteins at the very beginning of the clinical onset of the disease. Oxidative stress imbalance bridges the disease with the aging process, the major risk factor for PD. Thus, this result can be somehow interpreted as a peculiar sign of accelerated PD ageing. Of course, further biological investigations are needed to explore in more detail the peripheral signature of ROS stress and PD pathogenesis in Central Nervous System. Our results also support the presence of a complex interaction between sex and metabolism in relation to PD.
Methods
Patient cohorts
The study population consists of a total of 329 German subjects, including dn2PD patients, advPD patients, and healthy CTR. In detail (Fig. 1), patient cohorts included: (1) a training cohort, consisting of 72 dn2PD and 59 CTR, for a total of 131 subjects from the baseline visit of the Kassel cohort, as previously published27,59,62,63; (2) an independent validation cohort consisting of samples from 156 dn2PD, 20 CTR, and 22 advPD patients, for a total of 198 subjects, as part of the cross-sectional Kassel cohort27. Patients enrolled in this study were clinically phenotyped before sample collection. Phenotyping included 1.5 Tesla magnetic resonance imaging (MRI) to determine structural abnormalities, quantitative levodopa testing as published64, smell identification test (Sniffin’ sticks, Burghardt Messtechnik GmbH, Wedel, Germany), Mini Mental Status Examination (MMSE) followed by further cognitive testing and video-supported polysomnography to determine REM sleep behavior disorder in a subset of patients. The phenotyping was done based on these results and in accordance to established criteria for PD (UK Brain Bank Criteria)65, multiple system atrophy (MSA)66, dementia with Lewy bodies (DLB)67, progressive supranuclear palsy (PSP)68, corticobasal degeneration (CBD)69, Alzheimer’s disease and frontotemporal dementia (FTD)70. Subjects with marked vascular lesions in MRI indicative of a vascular comorbidity and subjects with normal pressure hydrocephalus by MRI were excluded.
A complete overview of the demographic characteristics of analyzed patients is reported in Table 4. The study was conducted according to the Declaration of Helsinki and with informed written consent provided by all subjects. The study was approved by the ethics committee of the Physician’s Board Hesse, Germany (Approval No. FF89/2008 for DeNoPa) and the University Medical Center Goettingen, Germany (Approval No. 9/7/04 and 36/7/02 for Kassel cohort).
NMR sample preparation and analysis
All blood samples from training and validation cohorts were collected between 8 AM and 9 AM under fasting condition using Sarstedt tubes for serum collection by venous puncture. Tubes with blood samples were centrifuged at 2500 × g for 10 min; serum was aliquoted and frozen within 30 min after collection. The samples were stored at −80 °C, transported in dry ice, until analysis following standard procedures for NMR metabolomic studies28,71,72. Samples were then prepared at the Magnetic Resonance Center (CERM/CIRMMP) University of Florence, Italy73. The analytical preparation of serum samples and their NMR spectra acquisition followed the protocols detailed elsewhere28.
For each serum sample, the one-dimensional (1D) NOESY pulse sequence was applied to acquire 1H-NMR spectra, using a Bruker 600 MHz spectrometer, with a proton Larmor frequency of 600.13 MHz and equipped with a 5 mm PATXI 1H-13C-15N and 2H decoupling probe. The spectrometer includes a z axis gradient coil, an automatic tuning-matching (ATM), and an automatic and refrigerated sample changer (SampleJet). To stabilize at the level of ±0.1 K the sample temperature, a BTO 2000 thermocouple was employed, and each NMR tube was kept for about 5 min inside the NMR probe head to equilibrate at the acquisition temperature of 310 K.
Spectral processing
Before applying Fourier transform, raw data were multiplied by an exponential line-broadening function of 0.3 Hz. Transformed spectra were automatically corrected for phase and baseline distortions and calibrated to a reference signal (anomeric glucose proton signal at 5.24 ppm), using Topspin 3.2 software (Bruker BioSpin).
Each 1D serum spectrum, in the range of 0.2–10.0 ppm, was bucketed into 0.02 ppm chemical shift segments using AssureNMR (version 2.2) software (Bruker BioSpin). Regions containing the residual water signal (between 4.68 and 4.84 ppm) were removed.
Serum and lipoprotein identification and quantification
Twenty-seven metabolites and 111 lipoprotein components (Supplementary Table 1) were identified and quantified from 1D 1H-NOESY NMR spectra using the AVANCE Bruker IVDr (Clinical Screening and In Vitro Diagnostics research, Bruker BioSpin)74 software. For all serum samples, different lipoproteins (VLDL, LDL, IDL, HDL) and different lipoprotein subclasses, classified according to density and size, for a total of 15 subclasses (VLDL-1 to VLDL-5, LDL-1 to LDL-6, and HDL-1 to HDL-4), were detected. For each main class and subclass, reported data consist of concentrations of lipids (total cholesterol, free cholesterol, phospholipids, and triglycerides) contained in each fraction. Concentrations of apolipoproteins ApoA1 and ApoA2 were estimated for HDL class and each relative subclass, while Apo-B concentrations were calculated for VLDL, IDL classes, and all LDL subclasses.
Statistical analyses
All data analyses were performed using R (version 3.6.1), an open source software for the statistical management of data75.
Multivariate data analysis was conducted on bucketed 1D NMR spectra of all available samples to visualize if spectral features contribute to the separation between groups. Principal component analysis (PCA) was used as a first exploratory approach76 to visualize the presence of outliers and to investigate, in an unsupervised manner, the data structure both in the training and in the validation cohorts. PCA analysis was performed on data scaled to unit variance.
Orthogonal projections to latent structures discriminant analysis (OPLS-DA) was employed as supervised technique to check for the presence of a serum metabolomic signature of the disease that distinguishes dn2PD patients from CTR77. The OPLS-DA approach was applied because it is an improvement of the PLS-DA method. Compared to PLS-DA, the advantage of OPLS-DA is that just one component is employed as a class predictor, while the remaining components reflect variations orthogonal to the first predictive component. A regression model is created in OPLS-DA using multivariate data and a response variable that simply contains class information.
When an unbalanced number of subjects was compared, OPLS-DA models were built by reducing groups to the same size by random sampling, thus including in the model an equal number of subjects from each group. The procedure was repeated 100 times and the results averaged over the 100 models. All discriminant and predictive analyses were performed on bucketed 1D-NOESY spectra without prior normalization. The OPLS-DA models were validated by repeated twofold cross validation (2CV) method. The 2CV scheme (detailed in ref. 78), briefly consists of two nested loops CV1 and CV2, where CV1 optimize the number of components to be used in the OPLS-DA model, while the CV2 is to assess the final model performance. In the outer loop (CV2) the complete dataset is split into a test set and a training set: the test set is set aside, and the training set is used in the loop CV1. Within CV1 the training set is split into an internal validation set and another training set (this operation is repeated 50 times). This last training set is used to develop a series of OPLS-DA models with different number of latent variables from which the samples in the internal validation set are predicted. In the CV2 the 90% of the data are randomly chosen at each iteration as a training set to build the model. Then, the remaining 10% are tested. The full procedure is repeated 100 times to derive average discrimination accuracy, sensitivity, and specificity and their confidence intervals (95% CI). Overall, for different classifications, accuracy, sensitivity, and specificity were calculated according to standard definitions, by means of a Monte-Carlo (MC) 2CV scheme (R script written in-house).
Moreover, to externally test the efficacy of the training model in discriminating dn2PD from CTR subjects, bucketed 1D NOESY spectra of the validation cohort serum samples (dn2PD, CTR, and advPD subjects of the validation cohort) were blindly projected onto the OPLS-DA score plot resulting from the training model.
Univariate Wilcoxon test79 was employed to compare metabolite and lipoprotein concentrations between patient groups. The Benjamini & Hochberg method80 was applied to correct for multiple testing, and adjusted P-values (FDR) < 0.05 were considered statistically significant. Log2 fold change (FC) ratios of the median intensities were also calculated for all analyses performed. Effect sizes were estimated for group comparisons using Cliff’s delta formulation81 (Cd), which contributes to the characterization of the meaningful signals by giving an estimation of the magnitude of the separation in the different comparisons. Magnitude is evaluated using the thresholds provided in Romano et al.82, where Cd values < 0.147 are “negligible”, (Cd) < 0.33 are “small”, (Cd) < 0.474 are “medium”, and (Cd) > 0.474 are “large”.
Association between each statistically significant analyte of the training cohort and the disease was assessed and validated using logistic regression in combination with ROC curve analysis. Before performing any analysis, continuous values of the analytes were standardized by centering and dividing by two standard deviations83 using the “rescale” function of the R package “arm”.
First, a binomial logistic regression model was built, for each statistically significant analyte found in the training cohort (i.e., a metabolite or a lipoprotein concentration), using the “glm” function in the R package “stats”. These analytes (metabolites and lipoproteins) were used as the predictors (x), while the dichotomic variable indicating the status (i.e., CTR or dn2PD) was used as the dependent variable (y) to be predicted.
The fitted values obtained for each analyte and for each subject were used to estimate areas under the ROC curves (AUC values, using the “colAUC” function of the R package “caTools”). Subsequently, the fitted regression models built on the training set were used to predict probabilities of samples in the validation cohort (values between 0 and 1) to be classified as CTR or dn2PD (“predict.glm” function in the R package “stats”). These predicted probabilities were used to calculate AUC values for the validation cohort that were further compared with AUC values reported for the training cohort.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The metabolite and lipoprotein concentrations dataset used and/or analyzed during the current study are available as Supplementary Data Table.
Code availability
Statistical codes used will be shared by request from any qualified investigator.
References
Tysnes, O.-B. & Storstein, A. Epidemiology of Parkinson’s disease. J. Neural Transm. Vienna Austria 1996(124), 901–905 (2017).
Agarwal, P. A. & Stoessl, A. J. Biomarkers for trials of neuroprotection in Parkinson’s disease. Mov. Disord. J. Mov. Disord. Soc. 28, 71–85 (2013).
Trezzi, J.-P. et al. Distinct metabolomic signature in cerebrospinal fluid in early Parkinson’s disease. Mov. Disord. J. Mov. Disord. Soc. 32, 1401–1408 (2017).
Chang, K.-H. et al. Alternations of metabolic profile and kynurenine metabolism in the plasma of Parkinson’s disease. Mol. Neurobiol. 55, 6319–6328 (2018).
Phelan, M. M., Caamaño-Gutiérrez, E., Gant, M. S., Grosman, R. X. & Madine, J. Using an NMR metabolomics approach to investigate the pathogenicity of amyloid-beta and alpha-synuclein. Metabolomics 13, 151 (2017).
Hatano, T., Saiki, S., Okuzumi, A., Mohney, R. P. & Hattori, N. Identification of novel biomarkers for Parkinson’s disease by metabolomic technologies. J. Neurol. Neurosurg. Psychiatry 87, 295–301 (2016).
Emamzadeh, F. N. & Surguchov, A. Parkinson’s disease: biomarkers, treatment, and risk factors. Front. Neurosci. 12, 612 (2018).
Nagesh Babu, G. et al. Serum metabolomics study in a group of Parkinson’s disease patients from northern India. Clin. Chim. Acta Int. J. Clin. Chem. 480, 214–219 (2018).
Shao, Y. & Le, W. Recent advances and perspectives of metabolomics-based investigations in Parkinson’s disease. Mol. Neurodegener. 14, 3 (2019).
Lei, S. & Powers, R. NMR metabolomics analysis of Parkinson’s disease. Curr. Metabolomics 1, 191–209 (2013).
Troisi, J. et al. A metabolomic signature of treated and drug-naïve patients with Parkinson’s disease: a pilot study. Metabolomics 15, 90 (2019).
Takis, P. G., Ghini, V., Tenori, L., Turano, P. & Luchinat, C. Uniqueness of the NMR approach to metabolomics. TrAC Trends Anal. Chem. 120, 115300 (2019).
Dani, C. et al. Metabolomic profile of term infants of gestational diabetic mothers. J. Matern. Fetal Neonatal Med. 27, 537–542 (2014).
Meoni, G. et al. The metabolic fingerprints of HCV and HBV infections studied by Nuclear Magnetic Resonance Spectroscopy. Sci. Rep. 9, 4128 (2019).
Vignoli, A. et al. Metabolic signature of primary biliary cholangitis and its comparison with celiac disease. J. Proteome Res. 18, 1228–1236 (2019).
Vignoli, A. et al. Fingerprinting Alzheimer’s disease by 1H nuclear magnetic resonance spectroscopy of cerebrospinal fluid. J. Proteome Res. 19, 1696–1705 (2020).
Caracausi, M. et al. Plasma and urinary metabolomic profiles of Down syndrome correlate with alteration of mitochondrial metabolism. Sci. Rep. 8, 2977 (2018).
Ghini, V. et al. Metabolomics profiling of pre-and post-anesthesia plasma samples of colorectal patients obtained via Ficoll separation. Metabolomics 11, 1769–1778 (2015).
Romano, F. et al. Effect of non-surgical periodontal therapy on salivary metabolic fingerprint of generalized chronic periodontitis using nuclear magnetic resonance spectroscopy. Arch. Oral. Biol. 97, 208–214 (2018).
Citterio, F. et al. Changes in the salivary metabolic profile of generalized periodontitis patients after non-surgical periodontal therapy: a metabolomic analysis using nuclear magnetic resonance spectroscopy. J. Clin. Med. 9, 3977 (2020).
Vignoli, A. et al. NMR-based metabolomics identifies patients at high risk of death within two years after acute myocardial infarction in the AMI-Florence II cohort. BMC Med. 17, 3 (2019).
Di Donato, S. et al. Serum metabolomic as biomarkers to differentiate early from metastatic disease in elderly colorectal cancer (crc) patients. Ann. Oncol. 27, 2762 (2016).
Meoni, G. et al. Metabolomic/lipidomic profiling of COVID-19 and individual response to tocilizumab. PLOS Pathog. 17, e1009243 (2021).
D’Andrea, G. et al. Different circulating trace amine profiles in de novo and treated Parkinson’s disease patients. Sci. Rep. 9, 6151 (2019).
Shao, Y. et al. Comprehensive metabolic profiling of Parkinson’s disease by liquid chromatography-mass spectrometry. Mol. Neurodegener. 16, 4 (2021).
Trupp, M. et al. Metabolite and peptide levels in plasma and CSF differentiating healthy controls from patients with newly diagnosed Parkinson’s disease. J. Park. Dis. 4, 549–560 (2014).
Pirazzini, C. et al. A geroscience approach for Parkinson’s disease: conceptual framework and design of PROPAG-AGEING project. Mech. Ageing Dev. 194, 111426 (2021).
Vignoli, A. et al. High-throughput metabolomics by 1D NMR. Angew. Chem. Int. Ed. Engl. 58, 968–994 (2019).
Vignoli, A., Tenori, L., Luchinat, C. & Saccenti, E. Age and sex effects on plasma metabolite association networks in healthy subjects. J. Proteome Res. 17, 97–107 (2018).
Wallner-Liebmann, S. et al. The impact of free or standardized lifestyle and urine sampling protocol on metabolome recognition accuracy. Genes Nutr. 10, 441 (2015).
Wallner-Liebmann, S. et al. Individual human metabolic phenotype analyzed by (1)H NMR of saliva samples. J. Proteome Res. 15, 1787–1793 (2016).
Assfalg, M. et al. Evidence of different metabolic phenotypes in humans. Proc. Natl Acad. Sci. USA 105, 1420–1424 (2008).
Bernini, P. et al. Individual human phenotypes in metabolic space and time. J. Proteome Res. 8, 4264–4271 (2009).
Ghini, V., Saccenti, E., Tenori, L., Assfalg, M. & Luchinat, C. Allostasis and resilience of the human individual metabolic phenotype. J. Proteome Res. 14, 2951–2962 (2015).
Cerri, S., Mus, L. & Blandini, F. Parkinson’s disease in women and men: what’s the difference? J. Park. Dis. 9, 501–515 (2019).
Baldereschi, M. et al. Parkinson’s disease and parkinsonism in a longitudinal study: two-fold higher incidence in men. ILSA Working Group. Italian Longitudinal Study on Aging. Neurology 55, 1358–1363 (2000).
McPherson, P. A. C. & McEneny, J. The biochemistry of ketogenesis and its role in weight management, neurological disease and oxidative stress. J. Physiol. Biochem. 68, 141–151 (2012).
Jimenez-Moreno, N. & Lane, J. D. Autophagy and redox homeostasis in Parkinson’s: a crucial balancing act. Oxid. Med. Cell. Longev. 2020, e8865611 (2020).
Straub, J. M. & Hausdörfer, J. Accumlation of acetone in blood during long-term anaesthesia with closed systems. Br. J. Anaesth. 70, 363–364 (1993).
Cocco, E. et al. 1H-NMR analysis provides a metabolomic profile of patients with multiple sclerosis. Neurol. Neuroimmunol. Neuroinflammation 3, e185 (2015).
Kumar, A. et al. Metabolomic analysis of serum by (1) H NMR spectroscopy in amyotrophic lateral sclerosis. Clin. Chim. Acta Int. J. Clin. Chem. 411, 563–567 (2010).
Paul, V. N., Chopra, K. & Kulkarni, S. K. Prooxidant role of histidine in hypoxic stressed mice and Fe(3+)-induced lipid peroxidation. Methods Find. Exp. Clin. Pharmacol. 22, 551–555 (2000).
Kumari, S. et al. Quantitative metabolomics of saliva using proton NMR spectroscopy in patients with Parkinson’s disease and healthy controls. Neurol. Sci. 41, 1201–1210 (2020).
Figura, M. et al. Serum amino acid profile in patients with Parkinson’s disease. PLoS ONE 13, e0191670 (2018).
Abdel-Salam, O. M. E. et al. Citric acid effects on brain and liver oxidative stress in lipopolysaccharide-treated mice. J. Med. Food 17, 588–598 (2014).
Takahashi, T. et al. N, N-Dimethylglycine decreases oxidative stress and improves in vitro development of bovine embryos. J. Reprod. Dev. 62, 209–212 (2016).
Campbell, K., Vowinckel, J., Keller, M. A. & Ralser, M. Methionine metabolism alters oxidative stress resistance via the pentose phosphate pathway. Antioxid. Redox Signal. 24, 543–547 (2016).
Quijano, C., Trujillo, M., Castro, L. & Trostchansky, A. Interplay between oxidant species and energy metabolism. Redox Biol. 8, 28–42 (2015).
Postuma, R. B. & Lang, A. E. Homocysteine and levodopa: should Parkinson disease patients receive preventative therapy? Neurology 63, 886–891 (2004).
Rozycka, A., Jagodzinski, P. P., Kozubski, W., Lianeri, M. & Dorszewska, J. Homocysteine level and mechanisms of injury in Parkinson’s disease as related to MTHFR, MTR, and MTHFD1 genes polymorphisms and L-dopa treatment. Curr. Genomics 14, 534–542 (2013).
Pizarro, C., Esteban-Díez, I., Espinosa, M., Rodríguez-Royo, F. & González-Sáiz, J.-M. An NMR-based lipidomic approach to identify Parkinson’s disease-stage specific lipoprotein–lipid signatures in plasma. Analyst 144, 1334–1344 (2019).
Zhang, L. et al. Circulating cholesterol levels may link to the factors influencing Parkinson’s risk. Front. Neurol. 8, 501 (2017).
Huang, X. et al. Lower low density lipid cholesterol levels are associated with Parkinson’s disease. Mov. Disord. J. Mov. Disord. Soc. 22, 377–381 (2007).
Lau, D., L, L. M., Koudstaal, P. J., Hofman, A. & Breteler, M. M. B. Serum cholesterol levels and the risk of Parkinson’s disease. Am. J. Epidemiol. 164, 998–1002 (2006).
Gudala, K., Bansal, D. & Muthyala, H. Role of serum cholesterol in Parkinson’s disease: a meta-analysis of evidence. J. Park. Dis. 3, 363–370 (2013).
Hu, L. et al. Integrated metabolomics and proteomics analysis reveals plasma lipid metabolic disturbance in patients with Parkinson’s disease. Front. Mol. Neurosci. 13, 80 (2020).
Xicoy, H., Wieringa, B. & Martens, G. J. M. The role of lipids in Parkinson’s disease. Cells 8, 27 (2019).
Hu, G. Total cholesterol and the risk of Parkinson’s disease: a review for some new findings. Park. Dis. 2010, 836962 (2010).
Mollenhauer, B. et al. Nonmotor and diagnostic findings in subjects with de novo Parkinson disease of the DeNoPa cohort. Neurology 81, 1226–1234 (2013).
Rozani, V. et al. Higher serum cholesterol and decreased Parkinson’s disease risk: a statin-free cohort study. Mov. Disord. 33, 1298–1305 (2018).
Huang, X., Abbott, R. D., Petrovitch, H., Mailman, R. B. & Ross, G. W. Low LDL cholesterol and increased risk of Parkinson’s disease: prospective results from Honolulu-Asia Aging Study. Mov. Disord. 23, 1013–1018 (2008).
Mollenhauer, B. et al. Monitoring of 30 marker candidates in early Parkinson disease as progression markers. Neurology 87, 168–177 (2016).
Sixel-Döring, F., Zimmermann, J., Wegener, A., Mollenhauer, B. & Trenkwalder, C. The evolution of REM sleep behavior disorder in early Parkinson disease. Sleep 39, 1737–1742 (2016).
Schade, S. et al. Acute levodopa challenge test in patients with de novo Parkinson’s disease: data from the DeNoPa cohort. Mov. Disord. Clin. Pract. 4, 755–762 (2017).
Clarke, C. E. et al. Clinical effectiveness and cost-effectiveness of physiotherapy and occupational therapy versus no therapy in mild to moderate Parkinson’s disease: a large pragmatic randomised controlled trial (PD REHAB). Health Technol. Assess. Winch. Engl. 20, 1–96 (2016).
Wenning, G. K., Tison, F., Ben Shlomo, Y., Daniel, S. E. & Quinn, N. P. Multiple system atrophy: a review of 203 pathologically proven cases. Mov. Disord. 12, 133–147 (1997).
McKeith, I. Dementia with Lewy bodies. Dialogues Clin. Neurosci. 6, 333–341 (2004).
Collins, S. J., Ahlskog, J. E., Parisi, J. E. & Maraganore, D. M. Progressive supranuclear palsy: neuropathologically based diagnostic clinical criteria. J. Neurol. Neurosurg. Psychiatry 58, 167–173 (1995).
Reich, S. G. & Grill, S. E. Corticobasal degeneration. Curr. Treat. Options Neurol. 11, 179–185 (2009).
Warren, J. D., Rohrer, J. D. & Rossor, M. N. Frontotemporal dementia. BMJ 347, f4827 (2013).
ISO/DIS 23118 Molecular in vitro diagnostic examinations—specifications for pre-examination processes in metabolomics in urine, venous blood serum and plasma. ISO, https://www.iso.org/obp/ui/#iso:std:iso:23118:ed-1:v1:en (2021).
Bernini, P. et al. Standard operating procedures for pre-analytical handling of blood and urine for metabolomic studies and biobanks. J. Biomol. NMR 49, 231–243 (2011).
Ghini, V., Quaglio, D., Luchinat, C. & Turano, P. NMR for sample quality assessment in metabolomics. N. Biotechnol. 52, 25–34 (2019).
Lipoprotein Subclass Analysis Enabling Tools on the IVDr Platform. Bruker.com https://www.bruker.com/products/mr/nmr-preclinical-screening/lipoprotein-subclass-analysis.html.
Ihaka, R. & Gentleman, R. R: a language for data analysis and graphics. J. Comput Stat. Graph 5, 299–314 (1996).
Serneels, S. & Verdonck, T. Principal component analysis for data containing outliers and missing elements. Comput. Stat. Data Anal. 52, 1712–1727 (2008).
Ebbels, T. M. D. in The Handbook of Metabonomics and Metabolomics 201–226 (Elsevier Science B.V., 2007).
Szymańska, E., Saccenti, E., Smilde, A. K. & Westerhuis, J. A. Double-check: validation of diagnostic statistics for PLS-DA models in metabolomics studies. Metabolomics 8, 3–16 (2012).
Neuhäuser, M. in International Encyclopedia of Statistical Science 1656–1658 (Springer, 2011).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B Methodol. 57, 289–300 (1995).
Cliff, N. Ordinal Methods for Behavioral Data Analysis (Psychology Press, 1996).
Romano, J., Kromrey, J., Coraggio, J. & Skowronek, J. Appropriate statistics for ordinal level data: should we really be using t-test and Cohen’sd for evaluating group differences on the NSSE and other surveys? In: Annual meeting of the Florida Association of Institutional Research, BibSonomy (2006).
Gelman, A. Scaling regression inputs by dividing by two standard deviations. Stat. Med. 27, 2865–2873 (2008).
Acknowledgements
G.M., L.T., Cristina L., P.T., and Claudio L. acknowledge the support and the use of resources of Instruct-ERIC, a Landmark ESFRI project, and specifically the CERM/CIRMMP Italy Centre. This work was supported by the Horizon 2020 Framework Programme (grant number 634821, PROPAG-AGEING).
Author information
Authors and Affiliations
Consortia
Contributions
Claudio L., B.M., C.F., C.T., and P.T. conceived and designed the study. C.T., B.M., and S.S. collected laboratory data and patients’ medical records. B.M. and C.T. provided serum samples. G.M. obtained NMR data. G.M., Cristina L., and L.T. performed metabolomic analyses. G.M. and Cristina L. wrote the paper. M.G.B., P.G., P.T., C.T., C.F., B.M., and Claudio L. critically revised the manuscript. All authors reviewed and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Meoni, G., Tenori, L., Schade, S. et al. Metabolite and lipoprotein profiles reveal sex-related oxidative stress imbalance in de novo drug-naive Parkinson’s disease patients. npj Parkinsons Dis. 8, 14 (2022). https://doi.org/10.1038/s41531-021-00274-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41531-021-00274-8
- Springer Nature Limited
This article is cited by
-
Identification of metabolites reproducibly associated with Parkinson’s Disease via meta-analysis and computational modelling
npj Parkinson's Disease (2024)
-
Transcriptome-based biomarker prediction for Parkinson’s disease using genome-scale metabolic modeling
Scientific Reports (2024)