Abstract
Hereditary spastic paraplegias are a rare group of clinically and genetically heterogeneous neurodegenerative diseases, with upper motor neuron degeneration and progressive lower limb spasticity as their main phenotypic features. Despite that 76 distinct loci have been reported and some casual genes identified, most of the underlying causes still remain unidentified. Moreover, a wide range of clinical manifestations is present in most hereditary spastic paraplegias subtypes, adding further complexity to their differential clinical diagnoses. Here, we describe the first exon rearrangement reported in the SPG45/SPG65 (NT5C2) loci in a family featuring a complex hereditary spastic paraplegias phenotype. This study expands both the phenotypic and mutational spectra of the NT5C2-associated disease.
Similar content being viewed by others
Introduction
Hereditary spastic paraplegias (HSPs) are a rare group of genetically heterogeneous neurodegenerative disorders, characterized by a progressive lower limb spasticity and weakness that results from a loss of corticospinal motor tract function.1 Over 70 different HSP loci have already been reported, with several patterns of inheritance, including autosomal recessive (AR), autosomal dominant, and X-linked, being identified. Based on their accompanying clinical symptoms, HSPs are classified in two distinct categories: pure and complex. In complex HSP, intellectual disability (ID), epilepsy, ataxia, optic atrophy, deafness, peripheral neuropathy, and skin abnormalities might be observed. Most complicated forms are usually inherited in a recessive fashion.2
In this study, we report the identification of a novel mutation in the NT5C2 gene (MIM# 613162), also known as SPG45/SPG65, in a family presenting with a complex form of AR-HSP. The NT5C2 locus was first mapped to chromosome 10q24.3-q25.1 in a large AR family with five members featuring complex HSP. The disease was characterized by onset at birth, spastic gait, ID, optical atrophy, and visual defects.3 In 2014, the causal gene was cloned by the identification of pathogenic NT5C2 mutations in five different families.4 An additional family carrying a novel homozygous splice-site mutation has recently been reported.5
Results
Family report
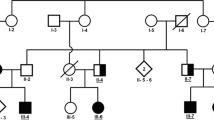
A consanguineous Iranian family with three affected siblings was referred to us (Fig. 1a). All three patients presented to our clinic with pronounced gait disturbances. Their mother had normal pregnancy with normal vaginal delivery, and all patients were normal at birth with no evidence of abnormality. The patients had delayed developmental milestones and ID became prominent with age. Walking began in about age 3 but gradually became spastic and problematic. There was no history of epileptic attacks, skeletal or facial deformity, failure to thrive, and no self-mutilation or aggression. There was no increased rate of infectious diseases or any skin abnormality. During examination, the patients showed ID but could perform visual contact and, to some extent, verbal contact. There was marked dysarthria and hypophonic speech, but other cranial exams were within normal limits. Spasticity of the limbs was observed during motor examination, with mild symmetric spastic paraparesis and increased deep tendon reflexes. Babinski sign was present and there was preserved abdominal-cutaneous reflex, but neither tremor nor dystonic features were observed. Coordination was normal and their gait was spastic. Although patients were not cooperative for sensory testing, no prominent sensory abnormality was detected. A summary of the clinical details can be found in Table 1.
a Pedigree structure of a family presenting with complex HSP due to a pathogenic NT5C2 mutation. Homozygous mutation carriers are represented as m/m and heterozygous carriers as wt/m. Dark squares (males) and circles (females) indicate HSP. b WGS reads of an affected sibling showing the 1.9 kb chromosome 10 deletion (c.771 + 573_814-298del) identified in all affected members are shown at the top, while Sanger chromatogram sequences corresponding to the deletion are shown at the bottom
Homozygosity mapping (HM)
Due to the AR inheritance pattern and the observed consanguinity in the affected pedigree (Fig. 1a), the HM method was used for disease loci identification. Eight different homozygous segments located on chromosomes 7, 8, 10, and 11 were identified to be shared by all affected individuals and to be present as heterozygous regions in non-affected family members (Supplementary Table 1).
Whole genome sequencing (WGS)
To reduce the number of disease-associated loci and assist with the disease gene identification, WGS was carried out in two affected siblings. After excluding splice-site, coding non-synonymous, small insertions and deletions, as well as stop gained/loss mutations as the disease-causing mutations, copy-number analysis of the WGS data was carried out. A novel, 1954-bp homozygous deletion at the NT5C2 locus involving the entire coding exon 11 was identified to be present in both sequenced, affected siblings. This deletion (NG_042272.1: g.101589_103554del), found to be located in a previously identified, disease-associated locus at chromosome 10 (Supplementary Table 1), resulted in c.771 + 573_814-298del nucleotide change and p.(Lys258_Lys271del) amino-acid change (Fig. 1b, Table 2). Subsequently, the examination of the deletion’s flanking regions in the remaining family members through Sanger sequencing revealed that this deletion was present in homozygous state in all affected members, while the three available unaffected cases were found to be heterozygous deletion carriers. This was later verified by the amplification of the NT5C2 exon 11 in all family members, which certified the absence of the exon 11 in all affected members and its presence in the healthy family members (Fig. 1a), further confirming its segregation with the disease status. The NT5C2 exon 11 was found to be present in the 50 different NT5C2 transcripts reported in NCBI website (https://www.ncbi.nlm.nih.gov/gene/22978?report=full_report), further supporting its presence in all reported human transcripts. No additional mutation was identified in the remaining disease-associated loci.
Discussion
We describe the identification of a novel NT5C2 mutation in a family with complex AR-HSP. By combining both linkage and WGS data, we were able to identify a large homozygous 1954-bp NT5C2 deletion as the causative genetic variation for a complex form of HSP. The clinical phenotype, which resembles previous reported cases, is characterized by an infantile onset, ID, delayed walking, increased deep tendon reflexes, and pronounced gait disturbances. Unlike other reported cases, no skeletal deformities were observed. While marked dysarthria and hypophonic speech, not previously reported in NT5C2-associated disease,3,4,5 were present in all three patients (Table 1).
We here report the first NT5C2 exon rearrangement in a family with complex HSP. Six other NT5C2 mutations, including slice-site, nonsense, and frameshift mutations, were previously reported in complex HSP (Table 2). We here report the first exon rearrangement described at the NT5C2 locus. Given the high frequency of exon rearrangements in other complex AR-HSP genes, such as SPG11,6 it is not surprising to identify these types of mutations in this recently identified HSP gene. However, copy number variations (CNVs) are more likely to be captured through WGS, rather than whole exome sequencing, as WGS captures both coding and non-coding genetic variation, allowing us to map the CNV breakpoints, and leads to improved detection of CNV and de novo variations due to its read coverage uniformity and allele reduced bias.7, 8 Therefore, with the growth and availability of WGS technology, we are confident that more CNVs would be mapped and identified at the NT5C2 and other HSP loci, which will certainly aid in the differential diagnosis of this heterogeneous group of neurodegenerative diseases.
This report expands both the phenotypic and mutational spectra of the NT5C2-associated disease.
Material and methods
Subjects
A consanguineous family with AR-HSP was clinically examined (Fig. 1a). The local ethics committee at the Shahid Beheshti University of Medical Sciences approved this study, and informed consent according to the Declaration of Helsinki from all participants was obtained. DNA samples from all members were isolated from whole blood using standard procedures.
Homozygosity mapping
High-throughput single nucleotide polymorphism genotyping was carried out in all available family members (n = 6) (Fig. 1a) using the HumanOmniExpress Exome arrays v1.3 and HiScanSQ system (Illumina Inc., San Diego, CA, USA). The GenomeStudio program (GS; Illumina) was used to undertake quality assessments and generate PLINK input reports9 for HM, and HM analyses were carried out as previously described.10, 11
Whole genome sequencing
Two affected family members were subjected to WGS analyses. WGS was carried out at the New York Genome Center. Sequencing libraries were constructed with the TruSeq PCR-free library kit (Illumina) following the manufacturer’s recommended protocol. Libraries were sequenced on the Illumina HiSeq X instruments, with 2 × 150 bp paired reads, to a minimum coverage of >30×. Sequencing data was processed as previously described.11
Validation and disease-segregation
The NT5C2 deletion identified through WGS was validated through direct Sanger sequencing by using primers flanking the deletion breakpoints and primers amplifying the NT5C2 exon 11. Primer sequences were designed by using a public primer design website (http://ihg.gsf.de/ihg/ExonPrimer.html) (sequences available upon requested). All purified PCR products were sequenced in both forward and reverse directions with Applied Biosystems BigDye Terminator v3.1 sequencing chemistry as per the manufacturer’s instructions, and resolved and analyzed as described elsewhere.10
Data availability
The DNA variation identified in this study is deposited in Open source databases, such as The Leiden Open Variation Database (http://databases.lovd.nl/shared/phenotypes/0000073477) and the ClinVar NCBI database (https://submit.ncbi.nlm.nih.gov/subs/clinvar_wizard/SUB2299204/overview). The raw data that support the findings of this study are now available from the corresponding author upon reasonable request, and will be deposited in dbGAP upon completion of the grant proposal.
Ethical approval
Ethical approval was obtained from the Institutional Review Board of the Shahid Beheshti University of Medical Sciences and all methods were performed in accordance with relevance guidances and regulations.
References
Blackstone, C. Cellular pathways of hereditary spastic paraplegia. Annu. Rev. Neurosci. 35, 25–47 (2012).
Klebe, S., Stevanin, G. & Depienne, C. Clinical and genetic heterogeneity in hereditary spastic paraplegias: from SPG1 to SPG72 and still counting. Rev. Neurol. (Paris) 171, 505–530 (2015).
Dursun, U., Koroglu, C., Kocasoy Orhan, E., Ugur, S. A. & Tolun, A. Autosomal recessive spastic paraplegia (SPG45) with mental retardation maps to 10q24.3-q25.1. Neurogenetics 10, 325–331 (2009).
Novarino, G. et al. Exome sequencing links corticospinal motor neuron disease to common neurodegenerative disorders. Science 343, 506–511 (2014).
Elsaid, M. F. et al. NT5C2 novel splicing variant expands the phenotypic spectrum of spastic paraplegia (SPG45): case report of a new member of thin corpus callosum SPG-Subgroup. BMC Med. Genet. 18, 33 (2017).
Gunther, S. et al. High frequency of pathogenic rearrangements in SPG11 and extensive contribution of mutational hotspots and founder alleles. Hum. Mutat. 37, 703–709 (2016).
Gilissen, C. et al. Genome sequencing identifies major causes of severe intellectual disability. Nature 511, 344–347 (2014).
Ritter, D. I. et al. Identifying gene disruptions in novel balanced de novo constitutional translocations in childhood cancer patients by whole-genome sequencing. Genet. Med. doi:10.1038/gim.2014.189 (2015).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Krebs, C. E. et al. The Sac1 domain of SYNJ1 identified mutated in a family with early-onset progressive parkinsonism with generalized seizures. Hum. Mutat. 34, 1200–1207 (2013).
Sanchez, E. et al. Identification of a large DNAJB2 deletion in a family with spinal muscular atrophy and parkinsonism. Hum. Mutat. 37, 1180–1189 (2016).
Acknowledgements
We thank the patients and relatives for their cooperation in this study. This work was in part supported by the Shahid Beheshti University of Medical Sciences and the National Institute of Neurological Disorders and Stroke of the National Institutes of Health (R01NS079388; CPR).
Author information
Authors and Affiliations
Contributions
H.D. and C.P-R. conceived and designed the study. A.T. and M.A. performed clinical assessments. L.J.A., A.A. and C.P-R. performed experiments, and contributed to data acquisition, analysis and interpretation. C.P-R. drafted the manuscript. All authors contributed to critical revision of the manuscript for intellectual content and final approval of the manuscript.
Corresponding author
Ethics declarations
Competing interests
Authors declare that they have no competing financial interests.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Darvish, H., Azcona, L.J., Tafakhori, A. et al. Whole genome sequencing identifies a novel homozygous exon deletion in the NT5C2 gene in a family with intellectual disability and spastic paraplegia. npj Genomic Med 2, 20 (2017). https://doi.org/10.1038/s41525-017-0022-7
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41525-017-0022-7
- Springer Nature Limited
This article is cited by
-
Expanding the clinical relevance of the 5′-nucleotidase cN-II/NT5C2
Purinergic Signalling (2018)