Abstract
We describe nNPipe for the automated analysis of morphologically diverse catalyst materials. Automated imaging routines and direct-electron detectors have enabled the collection of large data stacks over a wide range of sample positions at high temporal resolution. Simultaneously, traditional image analysis approaches are slow and hence unsuitable for large data stacks and consequently, researchers have progressively turned towards machine learning and deep learning approaches. Previous studies often detail work on morphologically uniform material systems with clearly discernible features, limited workable image sizes and training data that may be biased due to manual labelling. The nNPipe data-processing method consists of two standalone convolutional neural networks that were exclusively trained on multislice image simulations and enables fast analysis of 2048 × 2048 pixel images. Inference performance compared between idealised and real industrial catalytic samples and insights derived from subsequent data analysis are placed into the context of an automated imaging scenario.
Similar content being viewed by others
Introduction
Materials analysis using (Scanning) transmission electron microscopy ((S)TEM) is a key characterisation tool which provides access to a variety of localised physical and chemical information with sub-100 pm resolution1. In the field of heterogeneous catalysis, it is commonly used for the analysis of nanoparticle size distributions and morphologies in various environments2,3,4,5,6, which are key parameters in understanding catalytic performance7,8. However, analysis post-image acquisition often involves manual system- and instrument-dependent post-processing, which is time-consuming for the large number of particles required to achieve statistically significant metrics for comparison with bulk measurements of properties.
One solution is to employ (semi-) automated approaches9,10,11,12,13,14,15. However, the applicability of these remains limited as experimental data with high contrast between the nanoparticles and the support material, and human intervention in the analysis, are often required. With the increasing use of direct-electron detectors16,17,18 and automated image acquisition19, high data throughputs are becoming common and thus, more robust methods with minimal manual intervention are needed for full analysis of large datasets20.
Artificial Intelligence (AI) approaches originally developed for computer vision have been introduced to nanoparticle analysis in electron microscopy. For example, machine learning (ML) methods have been used by ref. 21, who employed a genetic algorithm to analyse the morphological properties of more than 150,000 nanoparticles. Muneesawang et al.22. and Wang et al.23. alternatively used a K-means approach and noted that performance may be dependent on the image contrast and resolution. Prevalent architectures and their applications in the fields of Physical Sciences and Life Sciences, as well as error metrics, are reviewed elsewhere20,24,25,26,27,28,29.
Deep learning (DL) based solutions have been applied in various contexts for nanoparticle analysis in TEM imaging, including Convolutional Neural Networks (CNNs) for object detection and semantic segmentation at atomic30,31 and lower resolution32,33,34,35, analysis of the performance of the U-Net neural network36,37,38, as well as liquid-cell experiments39,40. The former are appealing to studies of heterogeneous catalysts as they allow a statistically significant determination of relevant material properties once the respective networks are trained. However, various aspects remain unaddressed, especially in light of automated and fast data acquisition and processing. Many studies32,37,38 are based on a topologically uniform support material where individual nanoparticles exhibit clear contrast with respect to the background. Moreover, the ground truth in training datasets is often provided using manual labelling, inevitably introducing a human bias. Finally, the realistic workable array size of individual images is linked to available GPU resources, limiting the range of possible working scenarios.
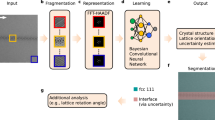
In this study, we use computationally generated models to form a training dataset of aberrated and augmented TEM image simulations (see ‘Methods’) to train a neural network pipeline; nNPipe. This passes fixed-sized raw TEM images (here 2048 × 2048 pixels) of various nanoparticle morphologies through the two standalone CNNs; YOLOv5 (https://ultralytics.com/yolov5) and SegNet41, and outputs semantically segmented maps of the same size (Fig. 1). We illustrate this approach using a diverse set of raw experimental images, taken from two datasets representing an idealised sample and a realistic catalytic material. The two nanoparticle systems used are as follows: A standard calibration sample of gold nanoparticles on ultrathin amorphous germanium (Au/Ge) film and an industrial model catalyst sample of nanoparticles on active carbon (Pd/C). In comparison to Au/Ge, the Pd/C system is morphologically much more diverse in terms of both particle size and local variations in substrate thickness. Thus, the latter provides valuable insight into how DL-based methods can be applied to realistic samples.
Training dataset generation is highlighted by green arrows: Au/Ge and Pd/C model computation, simulated training dataset generation46,47,48,49 and individual network training of YOLOv5x and SegNet41 for object detection and semantic segmentation, respectively. Red arrows highlight the inference stage, where an input image (in this case, an example of Au/Ge) is first inferred by object detection and instances of nanoparticles are subsequently semantically segmented as 256 × 256 sub-images by SegNet. The final post-processing stage reconstructs the label to its original size of 2048 × 2048 pixels.**.
While other CNN-based approaches, such as instance segmentation42 or pure semantic segmentation offer potential alternatives, we have opted for a two-step pipeline procedure where object detection and semantic segmentation are conducted in a serial fashion. Our reasons for this approach are as follows:
-
(i)
Speed
Object detection network architectures, YOLO43 and its more recent versions, such as YOLOv344 and YOLOv5 (https://ultralytics.com/yolov5), are distinguished by very high inference speed and competitive performance. Furthermore, the computational load is kept relatively low as all images are resized to 640 × 640 pixels for object detection, while only instances identified in bounding boxes of 256 × 256 pixels are passed to a subsequent SegNet41 network for semantic segmentation.
-
(ii)
Precision
Our Pd/C system exhibits nanoparticles that are often difficult to distinguish from the surrounding support film. nNPipe initially filters relevant instances from the full image and context before subsequently segmenting cropped image regions at full resolution.
-
(iii)
Modularity
Potential users can choose between using the whole pipeline or using the object detection output only. In addition, each module can be further developed and used with different image sizes.
-
(iv)
Feasibility
Our approach exploits established network architectures with minimal modification. The performance is entirely dependent on the training dataset instead of an optimised network architecture, enabling fast deployment.
We examine the ramifications for materials characterisation in differing imaging scenarios and sample morphologies that might be encountered in automated imaging procedures, i.e. images with varying nanoparticle load, discernibility of nanoparticles, support material thickness, etc. (compare ‘Methods’ and Tables 2, 3). Lastly, we compare the statistics obtained from the nNPipe approach against a manually labelled ground truth and provide an assessment metric for automated imaging. Information on the generation of sample models and training datasets and methods used for training are described in the ‘Methods’ section.
Results
Inference performance
During inference (Fig. 1), experimental images were first processed by the YOLOv5x object detection module that generated images with inferred classes and bounding boxes, together with corresponding lists containing vectors, di = [c,x,y,w,h] with a confidence of 0.5. Here, c refers to the inferred classes of ‘nanoparticle’ or ‘cluster’, while x,y,w,h denote the relative bounding box position, width and height, respectively. Clusters are instances of multiple joint nanoparticles that may also contribute to catalytic activity. However, a precise distinction between the nanoparticles’ shapes in projection is challenging and requires tailored data analysis approaches such as those discussed by Frei et al.45. In this work, we have included cluster instances in our training data (see ‘Methods’) such that their detection and segmentation remains optional. The results presented in this work focus on the analysis of individual nanoparticles, and consequently, centred 256 × 256-pixel image crops were only extracted from raw experimental images for instances with c = nanoparticle. These were subsequently passed as inputs to the SegNet module for semantic segmentation of individual nanoparticles from the background. Finally, 256 × 256-pixel images were subject to a post-processing routine to reconstruct a 2048 × 2048 label. Different routines (Supplementary Table 1) were used based on F1-scores for the Au/Ge and the Pd/C systems, respectively. A Meyer water-shedding step was included for the Pd/C system to enhance the separation of inferred nanoparticles from the varying contrast from the amorphous background. Full inference timings of respective datasets were separately measured for object detection and semantic segmentation with both neural networks running on the same GPU used for training. Table 1 compares full inference times for both material systems and stated timings include image writing processes. The resulting difference in the total inference time between the two material systems is attributed to the post-processing steps involved in the analysis of the Pd/C dataset.
Two experimental datasets respectively, containing 93 and 96 images for the Au/Ge and the Pd/C systems, were used for performance evaluation. Binary ground truth images (2048 × 2048 pixels) were obtained by manual segmentation of the raw image data by two experts working independently. A positive label was exclusively applied to nanoparticles that were clearly distinguishable. Clusters and nanoparticles that were cut by the image edges or exhibited an ambiguous contrast were assigned a negative ground truth label. These were allowed to be inferred by nNPipe, but were removed in the post-processing step (compare Supplementary Table 1). It is noted that both material systems contained a low number of indeterminate cases where an individual subjective judgement was unavoidable and examples of these are displayed in Supplementary Fig. 1.
Inference performance was determined by comparing pixel values of the binary ground truth images with the generated output as:
whereby TP refers to true positive, FP to false positive and FN to false negative pixels, respectively. For fully automated data acquisition, it is not possible to select suitable sample positions beforehand and it is, therefore, necessary to evaluate any method for different acquisition scenarios (see ‘Methods’ and above). Hence, TP, FN and FP pixels were colour coded into masks generated from overlapping pairs of ground truth images and inferred masks. Examples where nNPipe achieves particularly high and also lower performance were selected based on the F1-Score for further discussion. Figure 2 depicts exemplary cases of the Au/Ge system.
a, b Cases of high performance. c, d Cases with lower performance. The colour legend on the right of (b) is used for all colour-coded segmentation masks. ‘Nanoparticle’ instances are labelled as ‘AuNP’ while ‘cluster’ instances are labelled as ‘Cluster’ in the object detection outputs. Scale bar: 20 nm.
From an examination of mostly TP pixel contributions and high F1-scores in Fig. 2a, b, our method is clearly able to deliver a nearly identical segmentation mask (F1-scores 0.961 and 0.967) compared to the ground truth. However, closer inspection reveals FN pixel contributions at the edge of individual nanoparticle instances that can be attributed to a mismatch of the assigned particle border in the ground truth and the generated output (see Supplementary Fig. 2) with a border mismatch quantification given in the ‘Measurement performance’ section.
In contrast, Fig. 2c, d shows an example where the neural network pipeline achieves lower performance. Cut nanoparticles identified at the image border that were missed in the post-processing, as well as cluster instances mislabelled as nanoparticles in the object detection step, generate FP pixels in the resulting image segmentation. Conversely, nanoparticles mislabelled as clusters contribute to FN pixels (Fig. 2c). In addition, unknown image features (for example, lacy carbon at the top left in Fig. 2c, d) are identified as large clusters and some nanoparticles are unsegmented. This behaviour is attributed to comparable greyscale values of the lacy carbon with respect to clusters and with the instance size being significantly larger than a single nanoparticle. In Fig. 2d, it can also be observed that some nanoparticles which are in close proximity are mislabelled as clusters, although they are clearly separated. However, the semantic segmentation gives an overall high performance: particles cut by image edges (Fig. 2c, FP) and instances with shapes differing from those included in the training data are reliably separated from the background with high precision (Fig. 2).
In an analogous manner, the final segmentation performance was assessed for the Pd/C system, for which a differing performance was expected as this sample is morphologically more diverse and a more varied experimental imaging dataset was considered. Nonetheless, high-performance segmentation using the neural network pipeline was achieved, as highlighted in Fig. 3.
a, b Cases of high performance. c, d Cases with lower performance. The colour legend for this figure is shown only in (b). ‘Nanoparticle’ instances are labelled in orange as ‘PdNP’ while ‘cluster’ instances are labelled in yellow as ‘Cluster’ in the object detection outputs respectively. Scale bar: 20 nm.
An inference very close to the ground truth is shown in Fig. 3a, where all clusters were correctly identified by YOLOv5x and nanoparticles were accurately segmented by the SegNet41 module (F1-score 0.889). Comparable to Fig. 2a, b, FN contributions in this segmentation mask arise from nanoparticle border mismatch of the generated mask and the original ground truth (see Supplementary Fig. 4). In addition, some darker carbon features were identified as nanoparticles, which were subsequently segmented as FP nanoparticles. It should, however, be noted, that the amorphous carbon background in Fig. 3a is uniform, which gives rise to a high contrast difference between the particles and background and, consequently, high segmentation performance. This is different for Fig. 3b–d, where the support material exhibits a differing morphology and thickness. However, nNPipe is still able to achieve high F1-scores, as shown in Fig. 3b. Nearly all nanoparticles are segmented correctly, whereby FP pixel contributions arise from a wrong classification in the object detection stage for a cluster, as well as for some nanoparticles that exhibit poorly defined borders. As already illustrated in Fig. 2, SegNet is also able to infer nanoparticle shapes correctly that were not included in the original training dataset for images in this sample system.
Higher rates of FN and FP pixels are observed for the cases shown in Fig. 3c, where the support material is thicker as compared to Fig. 3a. In this case, the contrast between individual nanoparticles and the support material is also lower. This is reflected in the object detection performance, where some nanoparticle instances were not identified, leading to FN nanoparticles in the segmentation mask. Contributions of FP pixels can also be identified around nanoparticle instances due to a border mismatch, as well as in darker areas of the support material, which are falsely classified as nanoparticles in the object detection stage due to these thickness variations.
To evaluate future automated acquisition strategies, an image in which nNPipe does not perform well has been included for completeness. Figure 3d shows the output from an image acquired at a sample position which contains a clear, flat carbon area and a thick area of support material. While nanoparticles located on the flat carbon film are segmented with high TP pixel contributions, the residual nanoparticles assigned in the ground truth are omitted, leading to numerous FN pixels. This is due to ‘cluster’ instances inferred in the thicker amorphous carbon area, which resembles the output generated by the YOLOv5x network for the lacy carbon in the Au/Ge system. Specifically, large, unknown image features exhibiting comparable greyscale values for both ‘nanoparticles’ and ‘clusters’ are inferred as ‘cluster’.
Established automated segmentation methods have also been used to put the performance of the method described into a broader context. Combined global and local Otsu thresholding has been used (with a rolling ball filter value of 600, a Gaussian kernel size of 7, a local filter size of 50 pixels and a minimum nanoparticle size of 50 pixels) to both datasets after contrast inversion and without water shedding using the python package ParticleSpy (https://zenodo.org/record/5094360). The resulting output was subject to the same performance measurements defined by Eqs. 1–3. Figure 4 graphically compares Kernel density estimation (KDE) plots for the F1-scores from these methods.
Graphs were generated using a Gaussian kernel and Scott’s rule53. Calculated F1-score means are illustrated by red dash-dotted lines, while maximum values of the KDE-plots are denoted with red crosses. F1-score values: Au GlobLocOtsu = 0.431, Au neural network = 0.883, Pd GlobLocOtsu = 0.260, Pd neural network = 0.598. KDE maxima at Au GlobLocOtsu = 0.578, Au neural network = 0.934, Pd GlobLocOtsu = 0.300, Pd neural network = 0.663.
Considering firstly, the performances of the various methods for the Au/Ge system (Fig. 4), both the achieved F1-score mean and F1-score at KDE density maximum are higher for nNPipe compared to combined global and local Otsu thresholding. Furthermore, the distribution for nNPipe is sharper, underlining the high F1-score probability around a maximum of 0.934. A similar trend holds true for the Pd/C system, where the F1-score mean and F1-score at the KDE density maximum achieve higher values for the neural network pipeline. However, there is a broader distribution of the F1-Scores in the Pd/C experimental dataset indicating a more varied analysis performance. In turn, this distribution highlights the morphological diversity and more variable imaging scenarios of the Pd/C dataset and hence represents a likelihood for inference errors.
Measurement performance
In addition to image inference, the measurement of selected material properties plays an important role in automated image acquisition and in the determination of the potential catalytic performance of a given material. An insight into this was achieved by applying the ‘particle_analysis’ function of the ParticleSpy (https://zenodo.org/record/5094360) package on ground truth images and neural network pipeline-generated masks. The following results are based on calculated nanoparticle areas, although we note that other properties such as major axis length, minor axis length, circularity, eccentricity and others can be equally used to address the full spectrum of features important to the functioning of the catalyst materials. Figure 5 shows histograms of inferred and ground truth nanoparticle areas alongside KDE-plots for the full Au/Ge dataset, as well as for datasets with F1-scores above a given threshold. The respective calculated mean areas and count instances are also displayed for comparison.
KDE-plots were generated using a Gaussian Kernel and Scott’s rule53. Tables containing the calculated mean nanoparticle area (nm2), nanoparticle counts in the dataset (|D|) and calculated errors with respect to the ground truth are also shown. A minimum nanoparticle size of 50 pixels was assumed for all images. a The full dataset, b Images with an F1-Score of >0.9, c Images with an F1-Score of >0.8, d Images with an F1-Score of >0.7 and e Images with an F1-Score of >0.6.
The overall observable errors of the mean nanoparticle area and nanoparticle counts are notably low across all the cases in Fig. 5. The lowest error is observed for Fig. 5b, which shows the evaluation for images that achieved an F1-score of 0.9 or higher. Although this sub-dataset only considers images that were inferred with the highest performance, an error of 7.37% remains for the mean nanoparticle area, although counts do not deviate. The inclusion of sub-datasets with lower F1-scores (Fig. 5c–e) does eventually lead to an increase in the error in both metrics but does not exceed 8.81% for the nanoparticle area and 2.73% for the counts. Supplementary Fig. 2 shows selected experimental HRTEM images that were included in specified F1-score intervals for overview.
The KDE-plots reveal a close similarity of the nanoparticle area distributions between the ground truth and inferred data. Supplementary Table 2 summarises estimated values for the most common nanoparticle areas (xymax) and full widths at half maximum of the KDE-plots (FWHM) for Fig. 5b–e. While the FWHM values remain similar in all cases, a nearly constant difference of ca. 7 nm2 is calculated for xymax values.
In order to understand the error origin in the inferred data, i.e. whether it mostly arises from a nanoparticle border mismatch or nanoparticle instances entirely consisting of FN and FP pixels (see results below Fig. 2), five inferred and ground truth image pairs were reconsidered for F1-score intervals of 0.1. In this case, only instances with mostly TP contributions were compared to isolate the error due to nanoparticle border mismatch. This was done for F1-score intervals of [0.9, 1], [0.8, 0.9], [0.7, 0.8] and [0.6, 0.7] to track changes with decreasing segmentation performance, the results of which are summarised in Supplementary Table 3 while representative example images are shown in Supplementary Fig. 5.
Across all sub-datasets, the nanoparticle border mismatch error remains constant in an interval between 8.16 and 10.07% with a characteristic appearance where the inner edge of the mismatch is considerably smoother than the outer one. This is consistent with the data shown in Fig. 2, where small contributions of FP and FN pixels were highlighted at the border of nanoparticles with high TP ratios. Therefore, we conclude that the method used slightly underestimates the individual borders of nanoparticles. As the inner edge of the mismatch is smoother, the underestimation can be understood by a bias of the SegNet network, originating from network training with synthetic images where individual nanoparticles have predominantly smooth surfaces. These arise due to approximations in the computational model building and the corresponding training data with respect to experimental images (see ‘Methods’). Within the error interval above, it is, therefore, reasonable to conclude that the error originated in the inferred data is predominantly a border mismatch leading to small histogram shifts towards smaller nanoparticle areas. This type of mismatch may be equally expected for manual ground truth labels prepared by human experts.
Following the analysis shown in Fig. 5, the measurement performance on the Pd/C system was evaluated by comparing histograms and KDE-plots of nanoparticle areas determined for the ground truth and inferred images (Fig. 6). As a consequence of a higher number of images with lower F1-Scores (see Fig. 3), there is a higher error in the histogram of the nanoparticle area measurement results for the full Pd/C dataset (Fig. 6a) compared to the Au/Ge dataset (Fig. 5a). This is particularly evident when comparing Fig. 6a with Fig. 6e, wherein the latter only considers the sub-dataset with images having an F1-score >0.5. While the error for the mean nanoparticle area is comparable, the count error is significantly higher for the full dataset, which can be explained by considering FN nanoparticle instances, as illustrated in Fig. 3d. Specifically, smaller nanoparticle areas are most affected with both histograms differing most at ca. 15 nm2 (Fig. 6a). This is consistent with local lower contrast affecting smaller particles more than larger ones, leading to undetected instances or errors in the segmentation stage. Nonetheless, errors calculated for sub-datasets with F1-scores >0.8 and >0.7 remain significantly lower than for the full dataset, particularly when the counting error is considered. Supplementary Fig. 6 displays examples of experimental images for indicated F1-score intervals.
KDE-plots were generated using a Gaussian Kernel and Scott’s rule53. Tables containing the calculated mean nanoparticle area (nm2), nanoparticle counts in the dataset (|D|) and calculated errors with respect to the ground truth are shown. Plots were generated for a The full dataset, b Images with an F1-Score of >0.8, c Images with an F1-Score of >0.7, d Images with an F1-Score of >0.6 and e Images with an F1-Score of >0.5.
High inference performance for selected sub-datasets is also reflected in the shape and peak positions of the respective KDE-plots when compared to the ground truth (Supp. Table 4). Both, the xymax prediction as well as the FWHM settle at xymax = 13.23, FWHM = 32.69 and xymax = 10.45, FWHM = 24.08 for the inference and ground truth respectively, once the sub-dataset F1-score >0.6 is included. Although the estimates in the KDE plot from the inference are higher, they remain close to the ground truth, indicating a high measurement performance for those sub-datasets, especially when considering the full shapes of both KDE-plots.
Both, nanoparticle border mismatch errors and entire FN and FP nanoparticle instance errors were separated in the same way as for the Au/Ge system with values provided in Supplementary Table 5, while image differences between the ground truth and inferred images in Supplementary Fig. 7. In contrast to the border mismatch edges highlighted in Supplementary Fig. 5, those generated for the Pd/C system have a more varied appearance. While smooth shapes indicating network bias are encountered for both the inner and outer edge, some mismatches also exhibit thick bulges. The latter are likely to arise due to contrast variations between the supporting amorphous carbon and individual nanoparticles, which was not included in the training of the segmentation network (Fig. 3). However, if Supplementary Table 5 is additionally considered, a low average error of 3.82% in the border mismatch for datasets with an F1-score >0.7 is obtained. Compared to the Au/Ge system, this indicates that a significantly larger part of the nanoparticle area error shown in Fig. 6 is likely to stem from ‘pure’ FP and FN instances originally passed from the object detection network for the corresponding sub-datasets. When sub-datasets with F1-scores >0.6 and >0.5 are considered, the border mismatch increases to an average of ca. 14.1% (Fig. 3b, c).
Discussion
As is evident from the high image inference performances shown in Figs. 2, 3, it is clearly possible to train neural networks for analysis tasks on large sample volumes using synthetic datasets generated using multislice image simulations46,47,48,49. This is particularly significant for the heterogeneous Pd/C sample system (Fig. 3), where the experimental morphological diversity was simplified in the sample modelling (see ‘Methods’). Nevertheless, it was possible for the neural network pipeline to achieve F1-scores higher than 0.85, which underlines the potential of deep learning methods to apply a learned task with high performance to images differing from those provided in training.
It follows that the advantages of this training approach are threefold:
(i) there is an opportunity to tailor sample models for individual purposes,
(ii) it is feasible to extract exact image-labels as pairs from image simulations and
(iii) there are virtually unlimited models and training images.
However, steps including model generation, image simulation and augmentation, as well as training of the individual networks, require considerable processing time before experimental data can be analysed. For cases where only small datasets are considered, it might therefore be advantageous to use shallow network segmentation or other machine learning techniques which have been shown to achieve high performance for uniform sample morphologies such as the Au/Ge system considered here without a significant training overhead37,50,51.
Certain limitations of our training approach have been highlighted in the results presented. Very thick support material (which included buried nanoparticles with unclear borders) posed a challenge to the object detection network, resulting in high FN particle rates in the subsequent segmentation. This can be understood as adequate image simulations of this scenario were not included in the training dataset and were therefore not addressed appropriately during inference. This also holds true for nanoparticle shapes that showed a larger variety in appearance in experimental images than in image simulations. The identified border mismatch highlighted in Figs. 2, 3 as well as in Supplementary Tables 3, 5 is hence likely to stem from a bias of the neural network expecting shapes present in the training data.
A potential expedient to this would be to retrain. Sub-sets of experimental images and corresponding manually labelled ground truths could be exploited for image augmentation to form a sufficiently large training dataset. The disadvantage of this strategy would be a potential expert bias, where cases with subjective labelling play a particular role (Supplementary Fig. 1). Investigation of whether retraining increases the image inference performance, and the nanoparticle area measurement is, however, important and will be the subject of future work.
The very high similarity of nanoparticle area distributions with respect to the ground truth was achieved for the whole Au/Ge dataset (Fig. 5a), and the F1-scores >0.8 are especially significant. For the sub-datasets with F1-scores >0.9 and >0.8, the counting error remains below 2% while the determined mean nanoparticle area differs by only 7.37 and 8.60%, respectively (Fig. 5b, c). For the analysis summarised in Supplementary Tables 3, 4, it can be concluded that nNPipe can achieve a virtually close-to-human-like performance for sample systems with a uniform sample morphology on the full dataset acquired during automated image acquisition.
High performance has also been demonstrated for the morphologically more diverse Pd/C system. However, for this system, it is necessary to record data from sample positions where very thick amorphous carbon is avoided. While the measurement performance remains close to the one shown for the Au/Ge system for datasets with F1-scores >0.7, higher deviations in the inferred mean nanoparticle area and count become more likely for images at F1-scores <0.7. This can be understood by examining images such as those in Fig. 3c, d, in which the image fields of view include thicker areas of the support material, which cause blurring of the nanoparticle borders. A close-to-human performance can, therefore, only be expected for images with more uniform background contrast (Fig. 3a, b).
In the context of a practical automated imaging scenario, selecting acquired images based on F1-scores is not possible due to the lack of ground truth. However, a reasonable estimate can be obtained by comparing the number of instances generated at the YOLOv5x object detection stage with the ones generated at the SegNet segmentation stage: If the difference is below a dataset-dependent threshold, a higher chance for TP nanoparticle instances should follow. To illustrate this, Fig. 7 shows a point distribution plot of the F1-score in each image as a function of the introduced instance difference (ID).
As the overall F1-score for the Au/Ge system was high, selecting data according to the ID is not a strict requirement. For instance, setting a boundary criterion of ID <10% leads to a mean F1-score of 0.91. However, this only holds true for 52.63% of the acquired images of the full dataset to give only an increase of 0.03 in the F1-score.
This contrasts with the Pd/C system, for which the ID was originally implemented. As shown in Fig. 8, the likelihood of a high F1-score is significantly higher at low IDs. In this case, a boundary criterion of ID <7.5% leads to an F1-score of 0.70, which is a more significant improvement compared to the overall F1-Score of 0.598 (Fig. 4). For this dataset, the suggested boundary condition would be applicable to 40.82% of the data. However, it would ensure a close-to-human performance measurement (Fig. 8)
As highlighted in the results section, nNPipe also gives access to a range of nanoparticle properties of relevance to catalytic performance. Figure 9 shows examples in which the nanoparticle area is plotted against the nanoparticle eccentricity via bivariate KDE-plots, while sub-datasets can be obtained using the ID (Figs. 7, 8).
Histograms were binned at 1 nm2 ∙ bin−1 for the nanoparticle area and at 0.01∙bin−1 for eccentricity. All KDE-plots were obtained using Gaussian kernels and Scott’s rule53 a Au/Ge sub-datasets with F1-scores >0.8. b Pd/C sub-datasets with F1-scores >0.7.
In the method described, nNPipe implements a deep learning approach for the analysis of large-scale morphologically diverse catalyst systems. Only a few computationally generated models are required to generate large synthetic datasets approximating experimental EM images, which are suitable for successful training of a two-step neural network pipeline to high performance (F1-score up to 0.883). Given that manual segmentation of a single experimental image takes ~30 min, this method provides a significant advance in efficiency. nNPipe requires only 125.25 s (2.08 min) and 300.34 s (5.01 min) on an NVIDIA GeForce RTX 2080 Ti GPU, while manual segmentation would require typically 1100x and 580x more time for the analysis of the Au/Ge or Pd/C data consisting of 93 and 96 images, respectively.
Once suitable datasets are selected and analysed, relevant material properties such as nanoparticle area distributions and outliers or eccentricity could be extracted with marginal accuracy losses when compared to respective ground truth plots. This is of relevance for industrial applications where access to statistically significant metrics of morphologically diverse heterogeneous catalyst systems, such as the nanoparticle size distribution, nanoparticle aspect ratios, etc., can be directly linked to the system’s catalytic activity and offer a fast pathway for materials optimisation. Whilst feasible and performance-enhancing retraining methods are currently under investigation to ensure applicability to other nanoparticle catalyst systems, it can be foreseen that nNPipe is equally suitable in other imaging scenarios, for example, in situ heating or gas-cell experiments. Those are part of continuing research and will be published subsequently. Finally, further optimisations of the neural network architectures, an extension of training datasets or post-processing routines may enable new analytical modes, such as a determination of the z-height of individual nanoparticles for 3D distribution on the support material.
Methods
Sample preparation
A standard calibration sample of gold nanoparticles supported on ultrathin amorphous germanium (Au/Ge) film (ca. 4 nm) was prepared by JEOL Ltd. and provided on a standard copper TEM grid. Samples of palladium nanoparticles (1 wt%) on active carbon (Pd/C) were prepared as powder supplied by Johnson Matthey plc. The powder was derived from alginate and fired at 450 °C. TEM grids of this sample were prepared by a dry deposition method, where the Pd/C powder was mechanically crushed using two clean glass slides. A TEM copper grid with a lacy carbon film was subsequently sandwiched between the two exposed sides of the glass without additional mechanical force. This as-prepared TEM grid was used for image acquisition.
Experimental HRTEM imaging
TEM images were acquired using a double-corrected JEOL-ARM300CF instrument operated at 300 kV on a Gatan OneView Model 1095 4096 × 4096 pixel CCD-camera, at magnifications between 300,000–400,000X. Images were binned by 2 × 2 pixels during acquisition, resulting in a final resolution of 2048 × 2048 pixels with a sampling size of 0.0824–0.0618 nm∙pixel−1. Images of a variety of scenarios, i.e. a low and high particle load, in- and out-of-focus images, favourable and unfavourable contrast from the support material as examples, were included to mimic a more realistic automated imaging procedure and to explore the working limits of the method.
Sample model generation
In order to obtain a sufficiently large training dataset containing different sample configurations, sample models composed of {x,y,z}-coordinate lists were generated in customised Mathematica 12.1 notebooks and later used as input for TEM multislice simulations. Relevant system properties such as nanoparticle size and aspect ratio distributions were manually measured on ca. 200 individual nanoparticles from initial experimental images using an elliptic approximation and included in the sample model generation process.
Morphologies of the support material were approximated by qualitative comparisons of multislice simulations (see ‘Training Data’) and experimental images. While a flat amorphous Ge film was assumed for the Au/Ge system, a different strategy was applied for the Pd/C system, as the amorphous carbon support material was morphologically more diverse. 2D country outlines from a polyconic map projection were processed using a random set of erosion, blur and rotations to generate a diverse set of shapes mimicking the support material observed in experimental images. Subsequently, multiple country-shaped polygons—‘Countrygons’—were merged and linearly extended along the z direction to create a 3D structure, within which carbon atom positions were assigned. Up to five layers were shifted in the x–y-plane and stacked to better model the experimental support (see Supplementary Fig. 8). For both systems, different thicknesses and widths of the support material were considered. Atom coordinates inside the layers were assigned randomly, whereby the minimum average interatomic distance was calculated from respective densities (Table 2) and set as the sole boundary condition.
A randomised number of nanoparticles were placed on the generated substrates with icosahedral, cuboctahedral, decahedral and ellipsoidal morphologies considered. In addition, nanoparticle clusters consisting of multiple joined nanoparticles were incorporated to account for the presence of aggregates of nanoparticles in the experimental images. Atom coordinates were assigned with a cubic F lattice symmetry with lattice parameters of aAu =0.40782 nm and apd = 0.38907 nm, respectively. Table 2 summarises the properties assigned to the respective sample systems, while Supplementary Fig. 9 shows two examples of 3D hull models taken from the notebooks used. Note that for the Pd/C system, nanoparticles were also placed on the support materials’ sides, but such cases did not occur for the Au/Ge system. The models were not subject to molecular dynamics (MD) or density functional theory (DFT) calculations for energy-state relaxation.
Training data
The as-generated sample models were used as input for simulation using the multislice algorithm implemented in MULTEM code46,47,48,49. For augmentation, each of the 3500 sample models was subject to five randomised combinations of aberration values that matched typical experimental conditions (Table 3). In addition, the defocus spread modelling of the partial temporal coherence was randomised to introduce blurring into the image simulations. Image histograms of the resulting outputs were adjusted to histograms of a selection of experimental images and noise functions, including Gaussian- and Poisson-noise, beam-shape dependent brightness variations and different signal-to-noise ratio (SNR) were applied. The sampling of the image simulations was adjusted to 0.0824 nm∙pixel−1, to account for the sampling in most experimental images acquired. As the generated sample models varied in size, this adjustment naturally led to varying image sizes.
The as-generated models contained a number of nanoparticles exhibiting significantly lower contrast with respect to the background than those observed experimentally. Initial CNN training attempts (see ‘Architectures, Training and Inference’) concluded that such training examples would diminish the quality of the training dataset as the computer vision tasks became unclear, resulting in the inference of nanoparticles where none should be visible. Energetic relaxation of the models through MD calculations was not pursued due to the significant computational resources required for models of this size. As an alternative, nanoparticles were removed from the model coordinate list based on an empirically set minimum Grey-Level Difference (GLD) criterion (Table 3) that accounts for the contrast between individual nanoparticle instances and their surroundings. Pruned lists were then simulated for a second time with identical aberration and noise values, as applied previously. A comparison before and after Grey-Level Difference (GLD) pruning is provided in the Supplementary Information (Supplementary Fig. 10) and Supplementary Fig. 11 shows a side-by-side comparison of simulated and experimental images for both systems. Ultimately, the training examples also included images in which no nanoparticles were present. Labels for object detection and segmentation were obtained by correlating the coordinate lists with as-obtained image simulations, whereby a positive label was applied to pixels referring to nanoparticles and clusters, respectively. In the final step, the image simulations were subject to rotations and flipping operators for further augmentation. Table 3 contains all relevant information for the training dataset generation. Note that as for Table 2, ranges refer to limits from which a random number was drawn and applied, while aberration values use the notation described in ref. 52.
Architectures, training and inference
YOLOv5x (https://ultralytics.com/yolov5) was downloaded from github (https://github.com/ultralytics/yolov5) while SegNet was rebuilt in Mathematica 12.1 as outlined in ref. 41. A logistic sigmoid function was exploited instead of a softmax function for the output layer to adjust to binary classification. In all training cases, the individual networks were trained ab initio on a single NVIDIA GeForce RTX 2080 Ti GPU with a 90/10 training/validation data split and ADAM optimisation with a learning rate of 0.001. For YOLOv5x, the training was stopped after 100 epochs whereby the weights obtained at best mean Average Precision (mAP) were saved for the image inference. In the case of SegNet, training was stopped when the validation error did not improve over multiple epochs, which occured after 82 epochs for the Au/Ge and after 71 epochs for the Pd/C system. Weights with the lowest training and validation set errors were chosen for image inference. Further relevant hyperparameters, as well as respective training curves for both systems, are provided in the Supplementary Information (Supplementary Tables 6, 7 and Supplementary Figs. 12, 13).
Data availability
Resources comprising of image and ground truth datasets, as well as relevant code, a descriptive manual and other data required to replicate the results presented herein analysis, are available on https://zenodo.org/record/7024893#.YwjMkHbMJPZ.
References
Morishita, S. et al. Attainment of 40.5 pm spatial resolution using 300 kV scanning transmission electron microscope equipped with fifth-order aberration corrector. Microscopy 67, 46–50 (2018).
Jones, L., MacArthur, K. E., Fauske, V. T., van Helvoort, A. T. J. & Nellist, P. D. Rapid estimation of catalyst nanoparticle morphology and atomic-coordination by high-resolution Z-contrast electron microscopy. Nano Lett. 14, 6336–6341 (2014).
Michen, B. et al. Avoiding drying-artifacts in transmission electron microscopy: characterizing the size and colloidal state of nanoparticles. Sci. Rep. 5, 9793 (2015).
Zhao, Y., Tang, R. & Huang, R. Palladium supported on graphitic carbon nitride: an efficient and recyclable heterogeneous catalyst for reduction of nitroarenes and suzuki coupling reaction. Catal. Lett. 145, 1961–1971 (2015).
Ward, M. R., Theobald, B., Sharman, J., Boyes, E. D. & Gai, P. L. Direct observations of dynamic PtCo interactions in fuel cell catalyst precursors at the atomic level using E(S)TEM. J. Microsc. 269, 143–150 (2017).
He, L. et al. Morphology-dependent catalytic activity of Ru/CeO2 in dry reforming of methane. Molecules 24, 526 (2019).
Crozier, P. A. & Hansen, T. W. In situ and operando transmission electron microscopy of catalytic materials. MRS Bull. 40, 38–45 (2015).
Kumar, S., Varma, R. S., Zboril, R. & Gawande, M. B. Nanoparticle Design and Characterization for Catalytic Applications in Sustainable Chemistry (eds Prinsen, P. & Luque, R.) Ch. 3 (Royal Society of Chemistry, 2019).
Gontard, L. C., Ozkaya, D. & Dunin-Borkowski, R. E. A simple algorithm for measuring particle size distributions on an uneven background from TEM images. Ultramicroscopy 111, 101–106 (2010).
Park, C. et al. A multistage, semi-automated procedure for analyzing the morphology of nanoparticles. IIE Trans. 44, 507–522 (2011).
Mondini, S., Ferretti, A. M., Puglisi, A. & Ponti, A. Pebbles and PebbleJuggler: software for accurate, unbiased, and fast measurement and analysis of nanoparticle morphology from transmission electron microscopy (TEM) micrographs. Nanoscale 4, 5356–5372 (2012).
Park, C., Huang, J. Z., Ji, J. X. & Ding, Y. Segmentation, inference and classification of partially overlapping nanoparticles. IEEE Trans. Pattern Anal. Mach. Intell. 35, 1–1 (2013).
Epicier, T., Sato, K., Tournus, F. & Konno, T. Chemical composition dispersion in bi-metallic nanoparticles: semi-automated analysis using HAADF-STEM. J. Nanopart. Res. 14, 1106 (2012).
Groom, D. J. et al. Automatic segmentation of inorganic nanoparticles in BF TEM micrographs. Ultramicroscopy 194, 25–34 (2018).
Uusimaeki, T., Wagner, T., Lipinski, H.-G. & Kaegi, R. AutoEM: a software for automated acquisition and analysis of nanoparticles. J. Nanopart. Res. 21, 122 (2019).
Clough, R. & Kirkland, A. I. In Advances in Imaging and Electron Physics (Ed. Hawkes, P. W.) Ch. 1 (Academic Press, 2016).
Faruqi, A. R. & McMullan, G. Direct imaging detectors for electron microscopy. Nucl. Instrum. Methods Phys. Res. A 878, 180–190 (2018).
MacLaren, I., Macgregor, T. A., Allen, C. S. & Kirkland, A. I. Detectors—The ongoing revolution in scanning transmission electron microscopy and why this important to material characterization. APL Mater. 8, 110901 (2020).
Schorb, M., Haberbosch, I., Hagen, W. J. H., Schwab, Y. & Mastronarde, D. N. Software tools for automated transmission electron microscopy. Nat. Methods 16, 471–477 (2019).
Spurgeon, S. R. et al. Towards data-driven next-generation transmission electron microscopy. Nat. Mater. 20, 274–279 (2021).
Lee, B. et al. Statistical characterization of the morphologies of nanoparticles through machine learning based electron microscopy image analysis. ACS Nano 14, 17125–17133 (2020).
Muneesawang, P. & Sirisathitkul, C. Size measurement of nanoparticle assembly using multilevel segmented TEM images. J. Nanomater. 2015, 1–8 (2015).
Wang, X. et al. AutoDetect-mNP: an unsupervised machine learning algorithm for automated analysis of transmission electron microscope images of metal nanoparticles. JACS Au 1, 316–327 (2021).
Kalinin, S. V., Sumpter, B. G. & Archibald, R. K. Big–deep–smart data in imaging for guiding materials design. Nat. Mater. 14, 973–980 (2015).
Kalinin, S. V. et al. Lab on a beam—Big data and artificial intelligence in scanning transmission electron microscopy. MRS Bull. 44, 565–575 (2019).
Dan, J., Zhao, X. & Pennycook, S. J. A machine perspective of atomic defects in scanning transmission electron microscopy. InfoMat 1, 359–375 (2019).
Ge, M., Su, F., Zhao, Z. & Su, D. Deep learning analysis on microscopic imaging in materials science. Mater. Today Nano 11, 100087 (2020).
Ede, J. M. Deep learning in electron microscopy. Mach. Learn. Sci. Technol. 2, 011004 (2021).
Treder, K. P., Huang, C., Kim, J. S. & Kirkland, A. I. Applications of deep learning in electron microscopy. Microscopy 71, i100–i115 (2022).
Madsen, J. et al. A deep learning approach to identify local structures in atomic‐resolution transmission electron microscopy images. Adv. Theory Simul. 1, 1800037 (2018).
Groschner, C. K., Choi, C. & Scott, M. C. Machine learning pipeline for segmentation and defect identification from high-resolution transmission electron microscopy data. Microsc. Microanal. 27, 549–556 (2021).
Oktay, A. B. & Gurses, A. Automatic detection, localization and segmentation of nano-particles with deep learning in microscopy images. Micron 120, 113–119 (2019).
Zafari, S., Eerola, T., Ferreira, P., Kälviäinen, H. & Bovik A. Automated segmentation of nanoparticles in BF TEM images by U-Net binarization and branch and bound. Computer Analysis of Images and Patterns—CAIP 2019 113–125 (Springer, 2019).
Colliard-Granero, A. et al. Deep learning for the automation of particle analysis in catalyst layers for polymer electrolyte fuel cells. Nanoscale 14, 10–18 (2021).
Zhang, F. et al. Rodlike nanoparticle parameter measurement method based on improved mask R-CNN segmentation. SIViP 15, 579–587 (2021).
Ronneberger, O., Fischer, P. & Brox, T. U-Net: convolutional networks for biomedical image segmentation. Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015 234–241 (Springer, 2015).
Horwath, J. P., Zakharov, D. N., Mégret, R. & Stach, E. A. Understanding important features of deep learning models for segmentation of high-resolution transmission electron microscopy images. NPJ Comput. Mater. 6, 108 (2020).
Saaim, K. M., Afridi, S. K., Nisar, M. & Islam, S. In search of best automated model: explaining nanoparticle TEM image segmentation. Ultramicroscopy 233, 113437 (2021).
Yao, L., Ou, Z., Luo, B., Xu, C. & Chen, Q. Machine learning to reveal nanoparticle dynamics from liquid- phase TEM videos. ACS Cent. Sci. 6, 1421–1430 (2020).
Jamali, V. et al. Anomalous nanoparticle surface diffusion in LCTEM is revealed by deep learning-assisted analysis. Proc. Natl Acad. Sci. USA 118, e2017616118 (2021).
Badrinarayanan, V., Kendall, A. & Cipolla, R. SegNet: A Deep Convolutional Encoder-Decoder Architecture for Image Segmentation. IEEE Transactions on Pattern Analysis and Machine Intelligence 39, 2481–2495 (2017).
He, K., Gkioxari, G., Dollár, P. & Girshick, R. Mask R-CNN. Proceedings of the IEEE International Conference on Computer Vision (ICCV) 2961–2969 (IEEE, 2017).
Redmon, J., Divvala, S., Girshick, R. & Farhadi, A. You only look once: unified, real-time object detection. IEEE Conference on Computer Vision and Pattern Recognition (CVPR) 779–788 (2016).
Redmon, J. & Farhadi, A. YOLOv3: an incremental improvement. Preprint at https://arxiv.org/abs/1804.02767 (2018).
Frei, M. & Kruis, F. E. Image-based size analysis of agglomerated and partially sintered particles via convolutional neural networks. Powder Technol. 360, 324–336 (2020).
Cowley, J. M. & Moodie, A. F. The scattering of electrons by atoms and crystals. I. A new theoretical approach. Acta Cryst. 10, 609–619 (1957).
Goodman, P. & Moodie, A. F. Numerical evaluations of N-beam wave functions in electron scattering by the multi-slice method. Acta Cryst. A30, 280–290 (1974).
Lobato, I. & van Dyck, D. MULTEM: a new multislice program to perform accurate and fast electron diffraction and imaging simulations using graphics processing units with CUDA. Ultramicroscopy 156, 9–17 (2015).
Lobato, I., Aert, S. V. & Verbeeck, J. Progress and new advances in simulating electron microscopy datasets using MULTEM. Ultramicroscopy 168, 17–27 (2016).
Ito, E., Sato, T., Sano, D., Utagawa, E. & Kato, T. Virus particle detection by convolutional neural network in transmission electron microscopy images. Food Environ. Virol. 10, 201–208 (2018).
Bell, C. G. et al. Trainable segmentation for transmission electron microscope images of inorganic nanoparticles, J. Microsc. https://doi.org/10.1111/jmi.13110. (2022).
Typke, D. & Dierksen, K. Determination of image aberrations in high-resolution electron microscopy using diffractogram and cross-correlation methods. Optik 4, 155–166 (1995).
Scott, D. W. in Kernel Density Estimators Ch. 6 (John Wiley & Sons, Incorporated, 2015).
Shiojiri, M., Saito, Y., Okada, H. & Sasaki, H. Densities of amorphous thin films. Jpn J. Appl. Phys. 18, 1931 (1979).
Wypych, G. Databook of Antistatics (ChemTec Publishing, 2014).
Acknowledgements
We acknowledge funding from the EPSRC and Johnson Matthey plc for an iCASE award (2113841, K.P.T.), EPSRC Grant number EP/S001999/1 (A.I.K.) and the Rosalind Franklin Institute and EPSRC Grant Number EP/T033452/1 (A.I.K.). We thank Diamond Light Source for access and support in the use of the electron Physical Science Imaging Centre (Instrument E02, MG25427-41,-47,-57, -58, -81 and MG23814-25) that contributed to the results presented here.
Author information
Authors and Affiliations
Contributions
K.P.T.: Conceptualisation, code development, TEM-grid preparation, HRTEM imaging, preparation of manual ground truth data, deep-learning strategy, data analysis and interpretation and preparation of the manuscript. C.H.: Conceptualisation, coding guidance, creation and provision of SegNet in Mathematica 12.1 training environment, HRTEM imaging, deep learning strategy and user-support at ePSIC. C.G.B.: Preparation of ground truth data. T.J.A.S.: Data interpretation, advice on data analysis and related coding and user-support at ePSIC. M.E.S. and D.Ö.: provided Pd/C catalyst powder sample. M.E.S., D.Ö., J.S.K. and A.I.K.: Conceptualisation and data and project guidance. All authors reviewed and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Treder, K.P., Huang, C., Bell, C.G. et al. nNPipe: a neural network pipeline for automated analysis of morphologically diverse catalyst systems. npj Comput Mater 9, 18 (2023). https://doi.org/10.1038/s41524-022-00949-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-022-00949-7
- Springer Nature Limited