Abstract
Glioblastoma remains one of the deadliest brain malignancies. First-line therapy consists of maximal surgical tumor resection, accompanied by chemotherapy and radiotherapy. Malignant cells escape surgical resection by migrating into the surrounding healthy brain tissue, where they give rise to the recurrent tumor. Based on gene expression, tumor cores can be subtyped into mesenchymal, proneural, and classical tumors, each being associated with differences in genetic alterations and cellular composition. In contrast, the adjacent brain parenchyma where infiltrating malignant cells escape surgical resection is less characterized in patients. Using spatial transcriptomics (n = 11), we show that malignant cells within proneural or mesenchymal tumor cores display spatially organized differences in gene expression, although such differences decrease within the infiltrated brain tissue. Malignant cells residing in infiltrated brain tissue have increased expression of genes related to neurodevelopmental pathways and glial cell differentiation. Our findings provide an updated view of the spatial landscape of glioblastomas and further our understanding of the malignant cells that infiltrate the healthy brain, providing new avenues for the targeted therapy of these cells after surgical resection.
Similar content being viewed by others
Introduction
Glioblastoma (GBM) is the most lethal brain tumor, with a median survival rate of approximately 15 months1,2. First-line therapy consists of maximal surgical tumor resection, accompanied by concomitant and adjuvant temozolomide chemotherapy and radiotherapy3. The histological hallmarks of glioblastoma include pseudopalisading necrosis, microvascular proliferation (MVP), areas with high cellularity, and the diffuse infiltration of malignant cells into brain tissue4,5. Molecular profiling of patient tumors has identified 3 subtypes, namely the mesenchymal, proneural, and classical, which are associated with chromosomal alterations in NF1, CDK4, and EGFR, respectively6,7,8. Each subtype is composed of four main malignant cell states, namely (1) neural-progenitor-like (NPC-like), (2) oligodendrocyte-progenitor-like (OPC-like), (3) astrocyte-like (AC-like), and (4) mesenchymal-like (MES-like) cell states9,10,11,12. Intratumoral heterogeneity has been shown to be associated with treatment resistance13. Moreover, MES-like states are associated with hypoxia14, immune infiltration14,15, and have increased proportions at the time of tumor recurrence16. Moreover, the cellular composition or neighborhoods around malignant cells varies across tumor regions, with recruited monocytes and macrophages within the tumor17,18,19,20, and resident brain cells (neurons, astrocytes, oligodendrocytes, and microglia) interacting with infiltrating malignant cells5,21,22.
Malignant cells escape surgical resection by using brain vasculature and white matter tracts to diffusely infiltrate the surrounding brain parenchyma23,24. These cells give rise to the recurrent tumor within the resection margin in 90% of patients within one year after primary surgery25. Our understanding of the microenvironment in the infiltrated brain tissue is therefore crucial for understanding the progression from primary to recurrent tumors. The interactions between malignant cells and normal cells have been shown to facilitate disease progression in experimental models21,22,26, although how these cells interact within patient tumors is largely unknown. While a recent study found that brain tissue invasion is led by individual, unconnected cells in animal models27, the specific pathways that cause or are associated with malignant cell migration and their interactions with resident brain cells in patient tissue remain unknown. The missing information of infiltrating malignant cells that are left behind after primary treatment, hinders our ability to optimally treat GBM patients between the primary and recurrent tumors.
Recent spatial transcriptomics technologies28,29,30 allow the profiling of individual cells with hundreds or thousands of genes while retaining the coordinates of individual cells or specific cell populations within a region of interest (ROI). In this study, we hypothesized that profiling cells within the tumor core and infiltrated brain regions would reveal transcriptomic differences that would give insights into which biological processes are activated across the histological hallmarks of glioblastoma. We performed spatial transcriptomics with 2 complimentary methods (CosMx and GeoMx, NanoString Technologies) on a collection of primary grade 4 glioma tumors (IDH-wildtype GBM = 10, IDH-mutated astrocytoma = 1). MES-like states were primarily found within the tumor core and enriched in regions containing hypoxia and microvascular proliferation (MVP), where they, in addition to hypoxia-induced genes, expressed genes associated with vascular homeostasis. Both technologies showed malignant cells in infiltrated regions to increase expression of genes associated with AC -, NPC, and OPC-like malignant states, and this shift in malignant cell composition minimized transcriptional differences between mesenchymal and proneural GBM subtyped tumors in infiltrated brain tissue compared to the tumor core. We identified gene modules that were differentially expressed between the tumor core and infiltrated brain tissue, including genes associated with glial cell differentiation, synapses, and notch signaling. Only one module, enriched for gene ontologies suggesting cell differentiation was associated with poor survival in patients. Our results implicate multiple targetable pathways for disrupting the spread of glioblastoma tumors after tumor resection thus paving the way for developing new therapeutics against recurrent glioblastomas.
Results
Single-cell spatial profiling of the glioblastoma tumor core and infiltrated brain tissue shows different cellular composition compared to single-cell RNA sequencing datasets
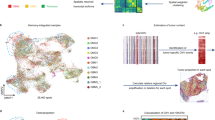
To profile cells within the tumor core and infiltrated brain tissue, we performed Spatial Molecular Imaging28 (CosMx, NanoString Technologies, Seattle, WA, USA) on formalin-fixed paraffin-embedded (FFPE) tissue from 5 primary GBM tumors using a panel of 960 genes (Supplementary Fig. S1A). Patient samples included areas with cellular tumor (n = 5), necrosis (n = 4), MVP (n = 5), and infiltrated brain tissue (n = 3) (Fig. 1A). Cell clusters were manually annotated as malignant or normal, using a combination of patient-specific clustering patterns and gene markers from publicly available databases from previous studies9,31,32 (Fig. 1B). The patient-specific clustering of malignant cells supported cell annotation, as normal cells clustered regardless of their sample of origin. Malignant cell states were then assigned using modified gene sets from previous studies9 (Fig. 1C). Differential expression analysis showed transcriptional markers were in alignment with previous studies32,33 (Fig. 1D). To investigate how cells were spatially associated, we correlated the cellular proportions from each field of view (FOV) (Fig. 1E, Supplementary Fig. S1C). Resident brain cells and vascular cells were, as expected, highly correlated. NPC-like states were strongly correlated with proliferating cells (G1/S, G2/M), while MES-like 1 cells were strongly correlated with TAMs as previously studied7,15 and vascular cells.
A Illustration of the spatial transcriptomics workflow. Archived formalin-fixed paraffin-embedded tissue sections from resected patient tumors (n = 5 patients) were screened for histological hallmarks of glioblastoma tumors. Spatial Molecular Imaging (CosMx, NanoString Technologies) was performed on all tumors using both gene transcripts and cell masks for cell profiling. B Uniform Manifold Approximation and Projection (UMAP) of all cells with individual plots for each patient sample using the same UMAP coordinates for each plot. C Mean expression of modified Neftel signatures across all malignant cells. D Dotplot of differential expression testing for all cell types. Color represents the average expression and dot size is the percentage of cells expressing the gene. E Correlation matrix of the distributions of cell types and malignant states across all patient regions of interest. F Composition of cells within our study and other single-cell RNA sequencing studies. G Heatmap of cell composition in each study, with values scaled and centered for each cell type. H Proportions of selected cell types within our data (red) (n = 1) and other studies (blue) (n = 16). Box plots show interquartile range (IQR), with the middle line indicating the median, and whiskers representing 1.5-fold IQR, and all individual points are shown. Source data are provided as a Source Data file. Panel A Created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
Spatial transcriptomics, in contrast to single-cell sequencing, does not require tissue dissociation and has been used to quantify cellular proportions of tumors34. We investigated how the composition of our dataset compares with other single-cell RNA sequencing studies annotated in a recently curated GBM atlas (GBMap)32 (Fig. 1F–H). Some of these studies sorted cells prior to single-cell RNA sequencing, making their cellular proportions not representative of the tumor (e.g. Darmanis et al.). However, the cellular composition of most studies were randomly sampled. Most malignant cell states and myeloid cells were of similar proportions in our data compared to the other studies. We found that astrocytes were found in high proportions in our study (with the highest being from Darmanis et al.11, where astrocytes were selected during sample collection), while oligodendrocytes were of average proportions. Lastly, our study showed remarkably high proportions of vascular cells and neurons compared to all other datasets.
As GBM cells can resemble neurodevelopmental cell states and differentiated brain cells such as neurons and astrocytes9,12, previous studies utilize statistical methods to quantify copy-number alterations to identify malignant populations9,10,11,33. We observe that the unique tumor genomes have altered transcriptional patterns that enable the annotation of malignant cells based on nearest-neighbor clustering. Moreover, we show that cellular compositions in spatial transcriptomics data differ from those in single-cell RNA sequencing studies and likely represent true cellular proportions of GBM tumors.
Mesenchymal malignant states are associated with areas of necrosis and microvascular proliferation and express genes associated with vascular growth and homeostasis
To gain a global overview of different cellular neighborhoods, defined by areas with different cellular compositions, we counted cell annotations of the 50 nearest neighbors to each cell, creating an annotation by cell count matrix. Dimension reduction and clustering were performed to identify cellular neighborhoods (Fig. 2A) and results were visually assessed using local transcriptional neighborhoods (Supplementary Fig. S2A–C) (see methods) and adjacent sections stained with H&E. To characterize each neighborhood, we performed Fisher’s exact test to identify which cell types were enriched (Fig. 2B). Neighborhoods 7, 8 and 9 showed enrichment for resident brain cells and were annotated as infiltrated brain tissue. Multiple neighborhoods were associated with tumor tissue (5, 1, 0, and 6), and differed in the proportions of malignant states. Neighborhoods 2, 3, and 4 were highly correlated within the same FOV (Fig. 2C) and located in areas with high levels of microvascular proliferation2,3 and/or necrosis2,4 on adjacent H&E stained sections. Neighborhood 2 was enriched for the MES-like 1 state and TAMs, neighborhood 3 was enriched for vascular cells, and neighborhood 4 was enriched for MES-like 2 and MES-like 1 states. These findings support previous hypotheses of pseudopalisades surrounding necrosis to be composed of the MES-like 2 state9,14 (Fig. 2D).
A Cellular neighborhoods in CosMx data visualized by Uniform Manifold Approximation and Projection. B Cell type or state enrichment across neighborhoods by Fisher’s exact test. C Pearson’s correlation matrix of cellular neighborhood fraction across field of views. D Histology of Hematoxylin and Eosin stained sections from adjacent tissue sections (top), with cell polygons colored by cell annotation (middle) and pixels colored by cellular neighborhoods (bottom) on example field of views with necrosis and microvascular proliferation. Clusters are the same as in (A). E Differential expression using a two-sided Wilcoxon Rank Sum test between cells in hypoxic/necrotic areas (cells = 3768, 5 samples) (neighborhood 4) compared to other neighborhoods (cells = 44251, 5 samples), with top genes being shown in Ivy-gap data (n = 122 across 10 patient samples) (F). G Differential expression using a two-sided Wilcoxon Rank Sum test between cells surrounding microvascular proliferation (cells = 6360, 5 samples) (neighborhood 3) compared to other neighborhoods (cells = 41,659, 5 samples), with top genes being shown in Ivy-gap data (n = 122 across 10 patient samples) (H). I Sections with the highest expression of MGP and TIMP1 in a recently published spatial transcriptomics dataset (Ravi et al.). J Differentially expressed genes for each patient in transcriptional clusters with the highest expression of MGP and TIMP1. K Expression of MGP, TIMP1, and all collagen genes in the dataset aggregated for each cluster, showing high expression of many, but not all collagens within these clusters. F, H Box plots show interquartile range (IQR), with the middle line indicating the median, and whiskers representing 1.5-fold IQR, and all individual points are shown. B, E, G, J All p-values were adjusted for using Benjamini–Hochberg correction. Source data are provided as a Source Data file.
Pseudopalisading cells surrounding necrosis and MVP are histological hallmarks of GBM. We tested for differential expression between malignant cells in neighborhood 4 and malignant cells in other neighborhoods to determine which genes were expressed by malignant cells surrounding necrotic tissue (Fig. 2E). This identified multiple genes induced by hypoxia (VEGFA, HILPDA) and also associated with the mesenchymal state (NDRG1, VIM, CHI3L1) to be upregulated, while stemness-associated markers (SOX2, EGFR) were most downregulated. Similar patterns of expression were found using bulk RNA-sequencing data from the Ivy-gap consortium5. When repeating the analysis for neighborhood 3 (MVP), we found a downregulation of EGFR, but not SOX2 (Fig. 2G) and upregulation of genes associated with mesenchymal states (e.g. CHI3L1 and VIM). While many of these genes were not specifically upregulated in areas of microvascular proliferation in Ivy-gap data, both Matrix Gla protein (MGP) and TIMP Metallopeptidase Inhibitor 1 (TIMP1) were most expressed in these samples (Fig. 2H). Both TIMP1 and MGP are found in the MES-like 1 signature from Neftel et al.9, although our results indicate that this upregulation is increased in only a subset of MES-like 1 cells. MGP, a calcium-binding protein, plays a crucial role in vascular homeostasis through the regulation of tissue mineralization35,36, and was found to be expressed in both MES-like malignant cells and perivascular fibroblasts in data from GBMap32. To further investigate the roles of these genes, we investigated the expression of MGP in a spatial transcriptomics dataset with multicellular resolution14, and found its expression to be highest in 3 patient samples, with spatial colocalization with TIMP1 in two samples (Fig. 2I). MGPhigh and TIMP1high clusters (Supplementary Fig. S2E) were tested for differential expression (Fig. 2J), and showed increased expression of MGP and TIMP1, pericyte/fibroblast markers (DCN, FN1) and collagens (e.g. COL1A1, COL1A2). Further analysis showed both patient MGPhigh clusters had higher expression of genes encoding collagen proteins, compared to the other transcriptional clusters (Fig. 2K).
Our findings show that genes associated with the MES-like 2 state were most expressed in necrotic areas with stemlike-related genes being downregulated in these cells. Necrotic and MVP neighborhoods were spatially correlated, and malignant cells within MVP neighborhoods overexpressed genes associated with the MES-like 1 state, and genes associated with vascular homeostasis. Further investigation of these patterns in an external dataset identified spatial co-expression with areas of high collagen expression. While it is well-established that hypoxia induces the expression of angiogenic genes, such as VEGFA, the association between MGP-expressing malignant cells and MVP indicates that MES-like cells may both initiate and continue to facilitate vascular development within and outside hypoxic areas.
Spatial trajectories identify differentially expressed genes associated with malignant states across the tumor core and adjacent infiltrated brain tissue
Multiple GBM states have been associated with brain invasion in vitro12 and in animal models27, although the transcriptomic profiling at single-cell resolution in infiltrated brain tissue remains underexplored. Therefore, to identify gene modules that are upregulated in malignant cells within infiltrated tissue, we developed an algorithm to identify genes with high variance across a spatial trajectory and used this on a trajectory leading from the tumor core to infiltrated brain tissue (Fig. 3A). Briefly, the designated trajectory is binned into individual segments, and gene expression for cells of the same type is aggregated for each bin. Genes that have a high spatial variance are retained for downstream analysis and clustered into gene modules.
A Overview of the developed algorithm for detecting gene modules within a spatial trajectory in CosMx data. B Hematoxylin and eosin stains and selected field of views chosen for spatial trajectory analysis, along with the visualization of the spatial trajectory and identified gene modules (C). D Shared genes up- and downregulated in malignant cells and tumor-associated macrophages and microglia (TAMs) between the two patient samples. E Transcriptional clusters and predicted chromosome values (7 and 10) in a 10x Visium GBM dataset. F Volcano plot of differentially expressed genes between the infiltrated area and tumor core using a two-sided Wilcoxon Rank Sum test. G Gene set enrichment analysis of Neftel malignant state signatures (G) and PangloDB cell type signatures (H) using a two-sided Fisher’s exact test. I Non-negative matrix factorization (NMF) programmes identified in this dataset. J Odds ratio of gene weights in the invasive NMF11 compared to peripheral and core NMFs for previously identified genes for malignant cells, and aggregated for each Neftel cell state (K). F–H All p-values were adjusted for using Benjamini–Hochberg correction. Source data are provided as a Source Data file.
Two patient samples had intact trajectories between the core and infiltrated brain tissue (Fig. 3B, Supplementary Fig. S3A, B). To enable the comparison of trajectories between samples, we identified spatially variant gene clusters with a low clustering resolution for the purpose of between-patient comparison (Fig. 3C). In malignant cells and myeloid cell populations, we identified genes with spatial variance that were unique and common between samples (Fig. 3D, Supplementary Fig. S3C). To assess which cells and malignant states expressed these genes, we aggregated gene expression for each cell type or state in GBMap32 (Fig. 3D). Genes commonly expressed in the tumor core were highly expressed in MES-like GBM states. Amongst these genes was CHI3L1, a gene recently associated with connected and non-invasive malignant cells37. The few genes upregulated in invasive cells were most associated with the AC-like state and growth factor receptors (EGFR, FGFR3), or NPC-like/OPC-like states, indicating their increased presence in malignant cell composition outside the tumor. TAM genes upregulated in the core were expressed by both blood-derived macrophages and microglia, while those in the infiltrated area were, as expected, most expressed in microglia.
Our findings indicated that MES-like signatures decreased in invasive GBM cells, while the remaining GBM states were associated with the infiltrated brain. However, these findings are limited to two patient samples, and a panel covering 960 genes. To further investigate infiltrated areas containing tumor cells, we identified infiltrative sections from a whole-transcriptome spatial transcriptomics dataset with multicellular resolution14. We identified one patient with 3 distinct regions (core = high CNV alterations, invasive = intermediate CNV alterations, periphery = low CNV alterations) (Fig. 3E). When testing between the infiltrated and tumor core areas, we identified a large number of differentially expressed genes, although no malignant state signatures were significantly enriched in either area (Fig. 3F, G). Conversely, neuronal cell signatures were upregulated in infiltrated tissue (Fig. 3H). However, as GBM cells share transcriptional programs with resident brain cells, it is difficult to distinguish between which differentially expressed genes are derived from malignant and non-malignant populations.
To circumvent the limitations arising with multicellular resolution, we used non-negative matrix factorization (NMF) to deconvolute the count matrix to 15 individual factors (Supplementary Fig. S3D). This identified NMF factors that were primarily expressed within the tumor core, the infiltrated area, and the tumor periphery (Fig. 3I). To identify genes that were more expressed in the infiltrating malignant cells, but not the cells of the healthy brain, we calculated the odds ratio of a gene being expressed in the factor associated with the infiltrated area compared with it being expressed in the peripheral factor (normal brain). We repeated this comparison with the factors found within the tumor core and were able to identify which genes were highly expressed in the infiltrated brain tissue compared to the tumor periphery (Fig. 3J, right vs left), and the infiltrated brain tissue compared to the tumor core (Fig. 3J, top vs bottom). This showed similar patterns to our previous findings with Delta Like Canonical Notch Ligand 1 (DLL1) being the gene most upregulated in the infiltrated area. DLL1 is also used as a NPC-like 1 marker gene, and to further investigate how genes associated with malignant states were associated with this invasive signature, we averaged the odds ratio of each malignant state from Neftel et al., and found the NPC-like 1 to be most associated with infiltrated brain tissue, with mesenchymal states being most present in the tumor core (Fig. 3K).
Our results show how transcription varies in malignant and immune cells spatially located along a trajectory from the tumor core to infiltrated brain tissue. Malignant cells within the tumor core have increased expression of genes associated with MES-like states, while cells in the infiltrated tissue increase expression of AC-, OPC-, and NPC-like states. These genes consist of growth factor receptors and notch signaling ligands, indicating their potential role in brain invasion. However, as these analyses were performed in only three patient samples, any results and interpretations may not be representative across GBM tumors.
Mesenchymal tumors become more proneural within infiltrated brain tissue
To extend our spatial analysis of the tumor core and infiltrated brain tissue, we performed Digital Spatial Profiling (GeoMx, NanoString Technologies, Seattle, WA, USA) on FFPE archived resected tissue from seven primary GBM tumors and one grade 4 primary IDH-mutated astrocytoma (Supplementary Fig. S4A). Using our cohort of TP53 mutated tumors with overexpression of Tumor Protein P53 (p53) in the majority of cells within the tumor core, we were able to identify regions of diffuse brain invasion and selected regions of interest (ROIs) from the tumor core, infiltrated brain tissue, periphery (few malignant cells), and brain tissue void of malignant cells (normal) (Fig. 4A). In each region, we used antibodies targeting p53 for malignant cells and Ionized calcium-binding adaptor molecule 1 (iba1) for TAMs to segment cells and extract gene transcripts from each cell population across ROIs (Fig. 4B). After quality control, we retained 103 iba1 and 96 p53 segments, spread across 37 core, 31 infiltrated, 28 periphery, and 10 normal ROIs. Principal component analysis showed a clear difference between p53 and Iba1 segments (Fig. 4C). To ensure that the p53 segmentation captured the transcriptomic profile of malignant cells, we performed copy-number variation (CNV) analysis to approximate the malignant cell purity of each segment (Fig. 3D). p53 segments showed the characteristic gain of chromosome 7 and loss of chromosome 10, although the tumor periphery had the highest proportion of segments with the lowest magnitude of CNVs (Supplementary Fig. S4B). As each patient’s genome is unique (Supplementary Fig. S4B), a unified threshold for low-purity cannot be made. Therefore, we removed p53 segments that were within 2 standard deviations from the same patient’s Iba1 segment chromosome 7 and 10 combined values.
A p53 mutated patient cohort was screened for high immunohistochemical p53 expressing tumor cells. Tumors with areas of the transition zone (tumor border) and tumor periphery (scarcity of tumor cells) were included in the study. Scale bars in smaller images are representative for all images of the same type. B GeoMx experimental overview. Immunofluorescence multiplexing of Iba1, p53, Glial Fibrillary Acidic Protein (GFAP) and DAPI. Iba1 and p53 were used to segment cells. C Principle component analysis (PCA) of areas of interest (AOIs) colored by segmentation marker. D InferCNV predicted copy number alterations in p53 and iba1 AOIs. E PCA of p53 AOIs only, with arrows illustrating the shift between core and infiltrated regions for each patient. F Heatmap of Neftel et al. signatures ranked by the PC1 axis in E. Volcano plots for differential expression analysis (DESeq2) using two-sided test between core (G) and infiltrated (H) p53 segments, divided into mesenchymal (n = 4) and proneural (n = 3) subtypes (top) followed by gene set enrichment analysis (bottom). I Gene markers found by Neftel et al. and the significance of a gene being expressed in the core or infiltrated regions of a tumor, split into mesenchymal (n = 4) and proneural (n = 3) tumors. -log10(P.adjusted) values are shown on the y-axis. To visualize whether a gene is differentially expressed across malignant programs, we assign a value equal to the sign of the LogFC (1 or −1) multiplied by -log10(p.adj). This approach allows us to distinguish between genes upregulated in the core or infiltrated regions. Genes upregulated in the core tissue appear on the bottom half of the plot, while genes upregulated in the infiltrated tissue appear on the top half. Box plots show interquartile range (IQR), with the middle line indicating the median, and whiskers representing 1.5-fold IQR, and all individual points are shown. G, H All p-values were adjusted for using Benjamini–Hochberg correction. Source data are provided as a Source Data file. Panel B Created with BioRender.com released under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International license.
While the GeoMx technology can identify single populations of cells, it is not at single-cell resolution, making each area of interest (AOI) a mix of multiple malignant states. Using principal component analysis on p53 AOIs, we found the first component to identify a mesenchymal to proneural axis (Supplementary Fig. S4C, D), a pattern commonly observed in single-cell RNA sequencing experiments16,38. We found most tumors (n = 5) transitioned towards a proneural composition in infiltrated brain tissue (Fig. 4E, F), while those that were most proneural (n = 2) stayed proneural but with an increase in genes associated with neuronal markers, neurotransmitter transportation, and presynaptic and axonal genes (Supplementary Fig. S4D). As GBM is currently classified into 3 main subtypes (Mesenchymal, Classical, and Proneural) based on the characterization of tumor core samples, little is known about how these subtypes differ in infiltrated brain tissue. We divided our samples into the three most proneural (RH11, RH8, and RH7), and four most mesenchymal tumors (RH9, RH1, RH3, and RH10) and tested for differential expression in tumor core segments (Fig. 4G), revealing as expected the most significant gene ontologies being either the proneural or mesenchymal Verhaak signature. However, when repeating the analysis for infiltrated regions, the most significant genes were those located on chromosome 12 next to CDK4, a chromosomal amplification associated with proneural tumors and amplified in 2 of our proneural samples6 (Fig. 4H, Supplementary Fig. S1B). This shows that while we were able to detect transcriptional heterogeneity, these differences were largely attributed to genomic alterations and not derived from the microenvironment—possibly indicating that these genomic alterations may be causal in shaping the tumor microenvironment. Amongst the upregulated genes in proneural tumors in infiltrated regions was OS9, which encodes for a protein that participates in the degradation of hypoxia-inducible factor 1 subunit and regulates its abundance in the cytoplasm, effectively reducing cellular response to the hypoxic environment39.
To compare how Neftel et al. malignant state signatures9 varied in expression across the tumor core and infiltrated brain tissue for each tumor subtype, we tested for differential expression between these areas in p53 AOIs (Fig. 4I). Mesenchymal tumors showed large differences, where MES-like 1 marker genes within the tumor core were replaced with marker genes of the AC-, OPC- and NPC-like 1 states in the infiltrated brain tissue. We observed minimal differences for proneural tumors across regions.
Our findings validate the use of p53 as a malignant cell marker in TP53-mutated tumors with high p53 expression. As expected, we found many genes differentially expressed between mesenchymal and proneural tumor cores, which did not persist outside of the tumor core. This was found to be largely due to the downregulation of genes associated with MES-like states and the upregulation of AC-, NPC- and OPC-like states in the tumor periphery.
Genes associated with cell differentiation, notch signaling and synapses are upregulated in malignant cells residing outside of the tumor core
Principal component analysis showed two main trajectories in how p53 segments changed from the tumor core to infiltrated brain tissue. Most tumors (n = 5) followed a mesenchymal to proneural transition, while the two most proneural tumors became more neural-like (Supplementary Fig. S4D). These results illustrate that GBM invasion is, again, heterogeneous across tumors and likely is influenced by genomics and the microenvironments inside and outside of the tumor. As five of the tumors followed a similar pattern, we continued to analyze these tumors to test which gene modules become upregulated in this infiltrative pattern. As we sought to identify regulatory gene modules associated with brain invasion, we identified co-expressed gene modules within our dataset and tested whether they were differentially expressed between p53 core and brain infiltrated AOIs (Fig. 5A, Supplementary Fig. S5A). To annotate these gene modules based on previous literature, we performed gene set enrichment analysis to test which ontologies were overrepresented in each gene module (Fig. 5B). While most gene modules upregulated in the infiltrated brain tissue were not significantly enriched with a known ontology, three modules were enriched with ontologies associated with synaptic vesicles, glial cell differentiation, oligodendrocyte specification and differentiation, notch signaling and neural crest differentiation.
A All gene modules were tested for being differentially expressed between core and infiltrated tissue areas of interest (AOI) (n = 5) using a linear mixed-effects model and ANOVA to estimate the effect on location. Box plots show interquartile range (IQR), with the middle line indicating the median, and whiskers representing 1.5-fold IQR. B Gene set enrichment analysis (GSEA) for all gene modules using Fisher’s exact test. Only significant ontologies were retained in the figure. C Correlation matrix (left) of all significantly up- or downregulated gene modules between the tumor core and infiltrated brain tissue in TCGA (The Cancer Genome Atalas) data. Modules are clustered into module clusters using hierarchical clustering. Expression heatmap of gene modules across all annotated GBMap cells (right). Rows are the same in both heatmaps. D GSEA (using fgsea) for all significant gene modules for differential expression analysis results between tumor and infiltrating cells from Darmanis et al. E Module score expression in TCGA data (left) and forest plots using multivariable Cox regression including each of the the three infiltrated tissue module clusters and covariates (MGMT- and IDH-status and age) (right). Box plots show interquartile range (IQR), with the middle line indicating the median, and whiskers representing 1.5-fold IQR, and all individual points are shown. Error bars for forest plot represent the 95% CI. *p < 0.05 after multiple hypothesis correction (A, D). All p-values were adjusted for using Benjamini–Hochberg correction. *p < 0.05, **p < 0.01. Source data are provided as a Source Data file.
To address how our gene modules are expressed and co-expressed across many patient samples, we correlated each module’s expression score in TCGA data (Fig. 5C). Hierarchical clustering identified 6 module clusters which consisted exclusively of either gene modules that were upregulated in either the tumor core or infiltrated brain tissue. To inspect which cell types expressed these modules, we aggregated the expression of GBMap across all cell types or cell states. Tumor core modules were mostly expressed by the mesenchymal malignant states, endothelial cells and pericytes (annotated mural cells in GBMap), and myeloid cells (monocytes, dendritic cells, TAMs). Conversely, modules significantly expressed in the infiltrated tissue regions were most expressed in NPC- and OPC-like malignant states, and resident brain cells (astrocytes, neurons, oligodendrocytes, OPCs). However, the GBMap atlas is generated primarily from resected patient tumors with little tissue derived from infiltrated brain tissue. To compare our findings with those from a previous study using single-cell RNA sequencing on biopsies from the core and infiltrated brain tissue11, we downloaded and processed their data using standard processing pipelines40. Using copy-number analysis to identify malignant cells, we found two patient samples containing malignant cells from both the core and infiltrated brain tissue, and differential expression analysis was performed for each patient for malignant cells residing in either the tumor or infiltrated brain tissue (Fig. 5D, Supplementary Fig. S5B). The direction of regulation of each significant gene module in our dataset was almost perfectly in alignment with those found in the single-cell dataset, supporting that the upregulated genes in the infiltrated brain tissue were not only expressed by non-malignant resident brain cells.
Finally, to investigate whether our invasive gene module clusters were associated with patient outcomes, we used a Cox regression model to test whether they were significantly associated with survival. Using gender, age, MGMT- and IDH-status as covariates (Fig. 5E), we found module cluster 6 was significantly associated with poor survival. Interestingly, this module cluster was most expressed in healthy brain tissue. Similar findings of a neural phenotype have recently been shown to be associated with poor patient outcomes41, and our findings indicate that this gene module is found to a higher degree in infiltrating malignant cells.
Our findings identify multiple gene modules associated with the tumor core and infiltrated brain tissue. Many modules were not enriched in known pathways, although few were significantly associated with cell fate, synapses, and notch signaling. Our gene modules were similarly correlated across TCGA tumors, and one gene module cluster had highest expression in the healthy brain and was also associated with poor patient survival.
Discussion
In this study, we present a multimodal spatial transcriptomics approach to profile cells from the tumor core and infiltrated brain tissue. Using imaging approaches with subcellular resolution, we identify multiple cellular neighborhoods within the glioblastoma tumor and compare how malignant cells influence and are influenced by their local tumor environment. Using our unique cohort of TP53 mutated patients with high protein expression of p53, we were able to screen resected tumor sections to identify regions with infiltrated brain tissue and profile the malignant cells using the GeoMx platform. While our study is limited in its number of patients, we find similar trends in a growing cohort of datasets characterizing primary GBM tumor cores. Our work investigates the patterns of malignant brain infiltration and identifies multiple transcriptional patterns which are both shared and not shared with resident brain cells. As infiltrating malignant cells escape tumor resection, the characterization of these cells provides insights into potential therapeutic targets to improve patient outcomes.
MES-like states have been described as injury-response, hypoxic, and immune-reactive states9,14,33, with our findings supporting these descriptions and additionally identifying its association with MVP where it expresses genes associated with vascular homeostasis. While the mesenchymal phenotype is associated with metastasis in other cancers42, our findings suggest this state to be exclusive to the tumor core. Conversely, when transitioning outside of mesenchymal subtyped tumors, we found the reduction of MES-like 1 markers to be replaced with increased expression of genes associated with AC-, NPC-, and OPC-like states. Moreover, we found increased expression of notch signaling consistently across all analyses. Two ligands of the notch signaling pathway (DLL1 and DLL3) are markers of NPC-like 1 cells9 and are known to play a role in maintaining quiescence and regulating asymmetric mitosis in adult neural stem cells43, as well as mediating neuronal migration. However, more research is needed to determine whether these ligands directly influence brain infiltration. A recent study using multiple animal models to study GBM infiltration identified unconnected cells to be more invasive than connected cells27. The authors observed a tendency for unconnected cells to be more NPC/OPC-like, while connected cells were more AC/MES-like. However, without having both spatial and transcriptomic information, it is difficult to quantify which states are most invasive, as any subset of a cell population may become one of the few unconnected cells that drive brain invasion. Our findings suggest that both AC-, OPC- and NPC-like states are associated with the infiltration of adjacent brain tissue.
We identify multiple gene modules that are upregulated in AOIs in the infiltrated brain tissue. Of these modules, we identified highly correlated module clusters in a larger cohort. One module cluster contained genes most expressed in differentiated non-malignant cells (e.g. neurons, oligodendrocytes, astrocytes), was most expressed in control brain tissue, and was prognostic of poor patient survival. Others have identified a neuronal-like GBM subtype to be prognostic of poor patient survival41, malignant cells to hijack neuronal features during brain invasion27, and that malignant cells become more neuronal at the time of recurrence measured by bulk RNA-sequencing44,45. Conversely, single-nuclei RNA sequencing shows an increased abundance of neurons at recurrence45,46 and the largest single-nuclei RNA sequencing dataset shows a proneural to mesenchymal transition between the primary and recurrent tumor16. Moreover, while our CosMx panel was limited to 960 genes, we did not find any indication of neuronal differentiation in the malignant cells of the infiltrated regions in CosMx data, although this was shown using the GeoMx platform. The discrepancy in these findings may be caused by the limited overlap of neuronal genes within the 960-plex CosMx panel. However, while it is plausible that malignant cells become more neuronal-like at recurrence or during cell invasion, it is also plausible that some of these findings lie in difficulties in distinguishing between malignant neuronal cells and non-malignant neuronal cells.
The amplification of CDK4 has been shown to predispose tumors to the proneural subtype6,9, with the main hypothesis deriving from a study showing CDK4 and CDK2 to increase differentiation of neural stem cells47. While CDK4 plays a critical role in cell proliferation, its effect on malignant cell composition is less clear. Alternatively, OS9, a neighboring gene to CDK4 on chromosome 12 and therefore equally amplified in proneural tumors, has been shown to regulate the partial degradation of hypoxia-inducible factor 139, which induces many genes associated with the mesenchymal subtype6,7,8,9. We speculate whether the amplification of the oncogene CDK4 partakes in driving tumorigenicity, although the parallel amplification of OS9 may result in decreased hypoxic signaling, due to increased degradation of hypoxia-inducible factor 1. As the mesenchymal subtype and MES-like states are associated with hypoxia-induced genes, the resulting dysfunction in hypoxic signaling associated with OS9 could explain the major differences between proneural and mesenchymal tumors, and why such differences are limited to the tumor core. Future studies are required for properly clarifying differences between tumor subtypes, although it is intriguing to speculate whether OS9 is causal for the major differences between mesenchymal and proneural subtypes and whether it can be exploited therapeutically.
In conclusion, our study has mapped the transcriptional profiles of malignant and normal cells within and surrounding GBM tumors. By highlighting how cells within GBM primary tumor cores and infiltrated brain tissue differ in gene expression, we pinpoint which biological pathways are activated in each region. Our findings show how malignant mesenchymal states are associated with necrosis and microvascular proliferation, while the relative proportion of AC-, OPC, and NPC-like states increase in the infiltrated brain tissue. We identify multiple gene modules upregulated within infiltrated brain tissue, with some being exclusively expressed by malignant cells, including one enriched for genes involved in notch signaling. Other modules, including one that is prognostic of patient survival, was found to be expressed in both malignant and non-malignant cells, and future studies are needed to further understand which cells are responsible for influencing patient outcomes. As the treatment of GBM relies on the surgical removal of tumors, further characterization of the residual cells in the brain is crucial for understanding and designing future therapeutic strategies to prevent the development of recurrent tumors.
Methods
Ethical approval
This study was approved by the Regional Committee on Health Research Ethics for the Capital Region of Denmark (Project- ID H-20027055) as well as the Danish Data Protection Agency (file number: P-2020-695). Permission to use patient tissue is automatically granted from The Danish Tissue Application Register in the absence of expressed prohibition from patients. The use of tissue was not prohibited by any patient according to the Danish Tissue Application Register.
Patient tissue selection
Archived histology slides from glioblastoma patients diagnosed between January 1st, 2016, and December 31st, 2019, at Rigshospitalet, Copenhagen University Hospital, Copenhagen, were acquired. We evaluated the slides using a BX51 microscope (Olympus Denmark A/S, Ballerup, Denmark) and subsequently reviewed them with a neuropathologist (BWK or DS). The inclusion criteria for the patients used in the GeoMx experiments were as follows: I) Presence of tumor core, infiltrated brain tissue, and tumor periphery on the same slide; II) High immunohistochemical p53 tumor cell expression to enable identification of malignant cells in the periphery; and III) Sufficient tissue in the block for analysis.
Histology stains
The selected formalin-fixed, paraffin-embedded (FFPE) tissue were sectioned to a thickness of 5 µm using a microtome and mounted on glass slides. To ensure our selected patients still complied with the aforementioned inclusion criteria, two different staining methods were performed:
-
1.
Hematoxylin and eosin: The tissue was deparaffinized, rehydrated and stained with Hematoxylin for 4 min and eosin 0,1% for 5 min. Slides were then subject to dehydration and coverslip mounting.
-
2.
Immunohistochemistry was performed on an automated stainer (Ventana Discovery). The slides underwent deparaffinization and heat-induced-epitope retrieval (HIER) using Ultra Cell Conditioning Solution (Tris Based Buffer) for 32 minutes at 99 °C. After blocking endogenous peroxidase activity, tissue sections were incubated with the p53 antibody (Clone: DO-7, BioRad) for 32 minutes at 37 °C. Antibody detection was performed using a DAB detection kit, nuclei were counterstained with Hematoxylin II and Bluing Reagent to increase nuclei contrast.
GeoMx RNA slide preparation
We followed the GeoMx® DSP Manual Slide Preparation User Manual (MAN−10150), RNA Slide Preparation Protocol. The selected FFPE sections were freshly cut to a thickness of 5 µm using a microtome and mounted on TOMO slides, spaced approximately 2-3 mm apart, allowing for mounting of two tissue sections on one slide. The tissue sections were less than one week old when subjected to GeoMx DSP slide preparation.
Slides were baked in a 60 °C drying oven for 2 h and subsequently subjected to deparaffinization and rehydration. Heat-induced epitope retrieval (HIER) was performed with Tris-Based Buffer for 20 min at 100 °C. To expose RNA targets, slides were incubated in Proteinase K solution (Thermo Fisher, AM2548) at a concentration of 0.1 µg/ml for 15 min. The tissue was post-fixed according to the User Manual. Slides were labeled with an RNA probe mix containing the GeoMX Human Whole Transcriptome RNA assay probe set (Nanostring Technologies, Seattle, WA, USA) and subjected to in situ hybridization overnight in the HybEZ II Oven (ACDBIO) for 20 h. A morphology marker solution was added, including p53 (clone: DO-7, Bio-Rad, 1:50), IBA1 (clone: GA5, Millipore, 1:25), GFAP (Clone: 20A12.1, Novus, 1:2000), and nuclei were labeled with Syto13.
The prepared slides were loaded into the GeoMx DSP instrument, following the GeoMx® DSP Instrument User Manual (MAN−10152-01). Slide scanning, ROI selection, segmentation, and barcode collection were performed using the GeoMx Digital Spatial Profiler (NanoString Technologies). For ROI selection, 24 ROIs of approximately 600 µm in diameter were chosen per slide (12 ROIs per patient). ROI selection was based on p53, IBA1, and GFAP morphology marker staining, as well as a comparison with consecutive chromogenic slides stained with H&E and p53.
The GeoMx® DSP NGS Readout User Manual (MAN−10153) was followed, and DNA barcode libraries were prepared according to the manufacturer’s instructions. The libraries were subsequently sequenced on the Illumina NovaSeq 6000 platform and the resulting BCL files converted to FASTQ via Illumina bcl2fastq v2.20.0, using FastQC v0.11.8 for quality control. Sequencing reads were then processed into digital count conversion (DCC) files using NanoString’s GeoMx NGS Pipeline v2.3.3.10. These DCC files were then uploaded to the GeoMx DSP system.
CosMx RNA slide preparation
A 5μm section from 6 glioblastoma samples were profiled using the CosMx spatial molecular imager as described in He et al.28. In brief, each sample was mounted to a histological slide, and a flow cell for reagent administration was affixed to the slide. The panel was profiled with the 1000-plex Universal Cell Characterization panel and was imaged with stains for Histone (blue channel), 18S rRNA (yellow channel), GFAP (red channel), and DAPI (UV). Reagents were flowed across the slide by the CosMx machine to cyclically image each of the targets in the panel, which were decoded based on their assigned barcode sequences and localization.
Cell segmentation
4’,6-diamidino-2-phenylindole (DAPI) was used for nuclei segmentation using CellPose 2.048, using default settings apart from the pretrained model (CP) and diameter size (60 = ~10 microns). RNA transcripts located within a nuclei mask were marked in a new column with a unique cell identifier and was used for the Baysor algorithm49. We optimized the Baysor algorithm for our dataset by iterating through different parameters. Each parameter was visually inspected by creating images with the transcripts of known nuclear genes (MALAT1), cytoplasmic genes (HSP90AA1), DAPI immunohistochemistry, and cell masks. Highly expressed nuclear and cytoplasmic genes were identified and used as priors for segmentation (Nuclear: MALAT1, MZT2A, NEAT1; Cytoplasmic: CLU, VIM, CD74, GLUL, SOD2, B2M, SPARCL1, CD81, HSP90AB1, GPX1). Each patient sample was run individually.
CosMx counts normalization, dimension reduction, clustering and cell annotation
Cell segmentation results from Baysor was pooled across samples and was used to create cell metadata and a count matrix. Cells containing less than 30 RNA counts were removed. The count matrix and metadata was used to create a Seurat object40, of which we ran the standard pipeline of running ScTransform50, RunPCA, RunUMAP, Findneighbors, and FindClusters. All cell clusters were manually annotated using differentially expressed markers from a recently published GBM atlas32 to annotate cell types, PanglaoDB31, as well as previously curated signatures for glioblastoma states9. After annotation, clusters were split into groups of broader cell types (immune, vessels, brain and tumor), and starting from RunPCA we reran the Seurat pipeline on each group to identify higher resolution of cell types and states. Harmony batch correction51 was used only on the Tumor group, to force cells into clustering with malignant cells from other patient samples.
Cell neighborhood identification
The dbscan R package52 was used to identify the 50 nearest neighbors of each cell. Neighboring cell annotations were counted for each cell, creating a cell annotation x cell matrix, of which we computed the shared nearest neighbors for each cell’s neighborhood using the Seurat function FindNeighbors. Neighborhoods were identified by graph-based clustering using the Seurat function FindClusters with parameters; k.param = 50 resolution = 0.02.
Local transcriptional neighborhoods
To visualize, quantify, and annotate local gene neighborhoods withing our spatial data, we used a subset of the mRNA molecules to identify reoccurring patterns of spatial adjacency. The nearest 50 molecules for this subset were identified, weighted by distance, and summed to a gene by molecule matrix. This matrix was scaled and centered, and principal component loadings from the single-cell analysis were projected onto this matrix. We attempted to perform principal component on the molecules themselves, although this identified nuclear/cytoplasmic differences and despite being more informative, were complicated to use. The projected embeddings were used for downstream cluster analysis and to determine the odds ratio of a gene being found in a given cluster (Supplementary Fig. S2A, B). While these findings were not used in our study, this approach facilitated the exploratory analysis to identify pitfalls and interesting differences across samples.
Spatial trajectory analysis
In this section, we outline the methods and procedures used for trajectory analysis in the context of spatial gene expression data. Taking two points and the width of the trajectory, we calculate the positions of cells within a tissue sample, and filter those that are outside the width or length or the trajectory. For each bin on the trajectory (n = 20), the mean expression of each cell type is calculated from the scaled gene expression across all cell types. As spatial data has a lot of noise due to e.g. poor segmentation, we scaled the matrix for all cell types prior to performing the trajectory analysis on individual cell types, which could then be used to cluster gene expression patterns into those that were inherently lowly or highly expressed in a cell type compared to other cell types. The variance of each gene expression across the trajectory is then calculated, and the genes with over 1.5 standard deviations over the mean of the variance across all genes were kept for downstream clustering and visualization. Clustering techniques were graph-based, using the Seurat algorithms FindNeighbors (knn = 5) and FindClusters (resolution = 1).
10x Visium GBM atlas data
Publicly available data from a recently published GBM atlas was downloaded at Datadryad (https://doi.org/10.5061/dryad.h70rxwdmj) and each sample was processed using ScTransform, RunPCA, RunUMAP, FindNeighbors, and FindClusters. Ivy-Gap annotation was performed by using SingleR to predict the histology of each spot in Visium data. InferCNV was used to infer chromosomal alterations.
GeoMx data processing
Data normalization was performed using quantile normalization as recently proposed by others53. The whole-transcriptome gene panel used for this analysis contains negative probes that do not bind to any known human gene. We used these genes to identify which genes had a variance or mean expression 2 standard deviations higher than negative probes. InferCNV was used to infer chromosomal alterations and these were used to discard samples with low malignant cell purity.
Differential expression analysis
Differential expression analysis was performed using the DESeq2 R package54 with GeoMx data, or the Seurat function FindMarkers with CosMx data. DESeq2 tests were designed with ~ Patient + Group for all tests with multiple observations coming from one patient. When testing across patients (Tumor group vs tumor group) segments were aggregated by mean and compared using the design ~ group.
Gene set enrichment analysis
Gene set enrichment analysis for Fig. 4 was performed using R packages fgsea55, org.hs.eg.db and msigdbr56 on the -log10(p.adjusted) *sign(log2FC) values of all genes tested. Gene set enrichment analysis for Fig. 5 was performed on gene clusters using the enrichR API57 and the following ontologies: KEGG_2021_Human and Human_Phenotype_Ontology58,59,60,61,62,63.
Statistics
Multiple statistical methods are described above. P values < 0.05 were considered significant when testing for differential expression, and p values were adjusted using false discovery rate correction.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The CosMx and GeoMx data generated in this study have been deposited in the Zenodo database under the https://doi.org/10.5281/zenodo.10521687 [https://zenodo.org/records/10521687]. Any additional information regarding the data is available from the corresponding authors upon request. Source data are provided with this paper.
Code availability
The code used for data processing and analysis in this study is available upon request.
References
Wen, P. Y. et al. Glioblastoma in adults: a Society for Neuro-Oncology (SNO) and European Society of Neuro-Oncology (EANO) consensus review on current management and future directions. Neuro Oncol. 22, 1073–1113 (2020).
Ostrom, Q. T. et al. CBTRUS Statistical Report: Primary brain and other central nervous system tumors diagnosed in the United States in 2010-2014. Neuro Oncol. 19, v1–v88 (2017).
Stupp, R. et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 352, 987–996 (2005).
Hambardzumyan, D. & Bergers, G. Glioblastoma: Defining Tumor Niches. Trends Cancer 1, 252–265 (2015).
Puchalski, R. B. et al. An anatomic transcriptional atlas of human glioblastoma. Science 360, 660–663 (2018).
Brennan, C. W. et al. The somatic genomic landscape of glioblastoma. Cell 155, 462–477 (2013).
Wang, Q. et al. Tumor Evolution of Glioma-Intrinsic Gene Expression Subtypes Associates with Immunological Changes in the Microenvironment. Cancer Cell. 32, 42–56.e6 (2017).
Verhaak, R. G. et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell. 17, 98–110 (2010).
Neftel, C. et al. An Integrative Model of Cellular States, Plasticity, and Genetics for Glioblastoma. Cell 178, 835–49.e21 (2019).
Patel, A. P. et al. Single-cell RNA-seq highlights intratumoral heterogeneity in primary glioblastoma. Science 344, 1396–1401 (2014).
Darmanis, S. et al. Single-Cell RNA-Seq Analysis of Infiltrating Neoplastic Cells at the Migrating Front of Human Glioblastoma. Cell Rep. 21, 1399–1410 (2017).
Bhaduri, A. et al. Outer Radial Glia-like Cancer Stem Cells Contribute to Heterogeneity of Glioblastoma. Cell. Stem Cell. 26, 48–63.e6 (2020).
Gimple, R. C., Yang, K., Halbert, M. E., Agnihotri, S. & Rich, J. N. Brain cancer stem cells: resilience through adaptive plasticity and hierarchical heterogeneity. Nat. Rev. Cancer 22, 497–514 (2022).
Ravi, V. M. et al. Spatially resolved multi-omics deciphers bidirectional tumor-host interdependence in glioblastoma. Cancer Cell. 40, 639–55.e13 (2022).
Hara, T. et al. Interactions between cancer cells and immune cells drive transitions to mesenchymal-like states in glioblastoma. Cancer Cell. 39, 779–92.e11 (2021).
Wang, L. et al. A single-cell atlas of glioblastoma evolution under therapy reveals cell-intrinsic and cell-extrinsic therapeutic targets. Nat. Cancer 3, 1534–1552 (2022).
Pombo Antunes, A. R. et al. Single-cell profiling of myeloid cells in glioblastoma across species and disease stage reveals macrophage competition and specialization. Nat. Neurosci. 24, 595–610 (2021).
Jain, S. et al. Single-cell RNA sequencing and spatial transcriptomics reveal cancer-associated fibroblasts in glioblastoma with protumoral effects. J. Clin. Invest. 133, e147087 (2023).
Al-Dalahmah, O. et al. Re-convolving the compositional landscape of primary and recurrent glioblastoma reveals prognostic and targetable tissue states. Nat. Commun. 14, 2586 (2023).
Ravi, V. M. et al. T-cell dysfunction in the glioblastoma microenvironment is mediated by myeloid cells releasing interleukin−10. Nat. Commun. 13, 925 (2022).
Venkatesh, H. S. et al. Electrical and synaptic integration of glioma into neural circuits. Nature 573, 539–545 (2019).
Venkatesh, H. S. et al. Targeting neuronal activity-regulated neuroligin-3 dependency in high-grade glioma. Nature 549, 533–537 (2017).
Scherer, H. J. Structural Development in Gliomas. Am. J. Cancer 34, 333–351 (1938).
Cuddapah, V. A., Robel, S., Watkins, S. & Sontheimer, H. A neurocentric perspective on glioma invasion. Nat. Rev. Neurosci. 15, 455–465 (2014).
Petrecca, K., Guiot, M. C., Panet-Raymond, V. & Souhami, L. Failure pattern following complete resection plus radiotherapy and temozolomide is at the resection margin in patients with glioblastoma. J. Neurooncol. 111, 19–23 (2013).
Venkataramani, V. et al. Glutamatergic synaptic input to glioma cells drives brain tumour progression. Nature 573, 532–538 (2019).
Venkataramani, V. et al. Glioblastoma hijacks neuronal mechanisms for brain invasion. Cell 185, 2899–917.e31 (2022).
He, S. et al. High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat. Biotechnol. 40, 1794–1806 (2022).
Chen, K. H., Boettiger, A. N., Moffitt, J. R., Wang, S. & Zhuang, X. RNA imaging. Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348, aaa6090 (2015).
Beechem, J. M. High-Plex Spatially Resolved RNA and Protein Detection Using Digital Spatial Profiling: A Technology Designed for Immuno-oncology Biomarker Discovery and Translational Research. Methods Mol. Biol. 2055, 563–583 (2020).
Franzen, O., Gan, L. M. & Bjorkegren, J. L. M. PanglaoDB: a web server for exploration of mouse and human single-cell RNA sequencing data. Database 2019, baz046 (2019).
Ruiz-Moreno, C. et al. Harmonized single-cell landscape, intercellular crosstalk and tumor architecture of glioblastoma. Preprint at https://www.biorxiv.org/content/10.1101/2022.08.27.505439v1 (2022).
Richards, L. M. et al. Gradient of Developmental and Injury Response transcriptional states defines functional vulnerabilities underpinning glioblastoma heterogeneity. Nat. Cancer 2, 157–173 (2021).
Liu, I. et al. The landscape of tumor cell states and spatial organization in H3-K27M mutant diffuse midline glioma across age and location. Nat. Genet. 54, 1881–1894 (2022).
Luo, G. et al. Spontaneous calcification of arteries and cartilage in mice lacking matrix GLA protein. Nature 386, 78–81 (1997).
Cancela, M. L., Laize, V. & Conceicao, N. Matrix Gla protein and osteocalcin: from gene duplication to neofunctionalization. Arch. Biochem. Biophys. 561, 56–63 (2014).
Hai, L. et al. A clinically applicable connectivity signature for glioblastoma includes the tumor network driver CHI3L1. Nat. Commun. 15, 968 (2024).
Wang, L. et al. The Phenotypes of Proliferating Glioblastoma Cells Reside on a Single Axis of Variation. Cancer Discov. 9, 1708–1719 (2019).
Baek, J. H. et al. OS-9 interacts with hypoxia-inducible factor 1alpha and prolyl hydroxylases to promote oxygen-dependent degradation of HIF-1alpha. Mol. Cell. 17, 503–512 (2005).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573–87 e29 (2021).
Drexler, R. et al. A prognostic neural epigenetic signature in high-grade glioma. Nat. Med., 30, 1622–1635 (2023).
Thiery, J. P. Epithelial-mesenchymal transitions in tumour progression. Nat. Rev. Cancer 2, 442–454 (2002).
Kawaguchi, D., Furutachi, S., Kawai, H., Hozumi, K. & Gotoh, Y. Dll1 maintains quiescence of adult neural stem cells and segregates asymmetrically during mitosis. Nat. Commun. 4, 1880 (2013).
Varn, F. S. et al. Glioma progression is shaped by genetic evolution and microenvironment interactions. Cell 185, 2184–99.e16 (2022).
Kim, K. H. et al. Integrated proteogenomic characterization of glioblastoma evolution. Cancer Cell. 42, 358–77.e8 (2024).
Hoogstrate, Y. et al. Transcriptome analysis reveals tumor microenvironment changes in glioblastoma. Cancer Cell. 41, 678–92.e7 (2023).
Lim, S. & Kaldis, P. Loss of Cdk2 and Cdk4 induces a switch from proliferation to differentiation in neural stem cells. Stem Cells 30, 1509–1520 (2012).
Pachitariu, M. & Stringer, C. Cellpose 2.0: how to train your own model. Nat. Methods 19, 1634–1641 (2022).
Petukhov, V. et al. Cell segmentation in imaging-based spatial transcriptomics. Nat. Biotechnol. 40, 345–354 (2022).
Hafemeister, C. & Satija, R. Normalization and variance stabilization of single-cell RNA-seq data using regularized negative binomial regression. Genome Biol. 20, 296 (2019).
Korsunsky, I. et al. Fast, sensitive and accurate integration of single-cell data with Harmony. Nat. Methods 16, 1289–1296 (2019).
Hahsler, M., Piekenbrock, M. & Doran, D. dbscan: Fast Density-Based Clustering with R. J. Stat. Softw. 91, 1–30 (2019).
van Hijfte, L. et al. Alternative normalization and analysis pipeline to address systematic bias in NanoString GeoMx Digital Spatial Profiling data. iScience 26, 105760 (2023).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Sergushichev, A. A. An algorithm for fast preranked gene set enrichment analysis using cumulative statistic calculation. Preprint at https://www.biorxiv.org/content/10.1101/060012v1 (2016).
Liberzon, A. et al. The Molecular Signatures Database (MSigDB) hallmark gene set collection. Cell Syst. 1, 417–425 (2015).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Gillespie, M. et al. The reactome pathway knowledgebase 2022. Nucleic Acids Res. 50, D687–D692 (2022).
Kanehisa, M., Furumichi, M., Sato, Y., Kawashima, M. & Ishiguro-Watanabe, M. KEGG for taxonomy-based analysis of pathways and genomes. Nucleic Acids Res. 51, D587–D592 (2023).
Miller, R. A. et al. Understanding signaling and metabolic paths using semantified and harmonized information about biological interactions. PLOS One. 17, e0263057 (2022).
Koopmans, F. et al. SynGO: An Evidence-Based, Expert-Curated Knowledge Base for the Synapse. Neuron 103, 217–34.e4 (2019).
Kohler, S. et al. The Human Phenotype Ontology in 2021. Nucleic Acids Res. 49, D1207–D1217 (2021).
Gene Ontology, C. et al. The Gene Ontology knowledgebase in 2023. Genetics 224, iyad031 (2023).
Acknowledgements
This project is supported by grants from Neye-Foundation, DCCC Brain Tumor Center funded by the Danish Cancer Society, Folketingsmand Jens Christensen og hustru Korna Christensens Fond, Arvid Nilssons Fond, Fabrikant Einar Willumsens Mindelegat, and Aase og Ejnar Danielsens Fond and Tømrermester Jørgen Holm og Hustru Elisa f. Hansens Mindelegat. Hannah Rostalski received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie grant agreement no 945322 (LEAD).
Author information
Authors and Affiliations
Contributions
D.S.L.H., V.P., S.R.M., and B.W.K. conceived the project. V.P., S.R.M., and D.S.L.H. screened and selected archived patient tumor tissue for experiments. B.W.K. and D.S. supervised the pathological assessment of tissue. D.S.N., J.W., M.J.S.L., A.L., and U.L. curated clinical and genomic data. V.P., S.R.M., N.S.B., A.A., K.J. and S.R.M. optimized and performed GeoMx experiments and multiplexed immunofluorescence. A.Y.S., T.O.S., and F.O.B., performed sequencing on GeoMx libraries. NanoString Technologies performed CosMx experiments through a technology access program. D.S.L.H. planned and performed all computational analyses under the supervision of K.V.S. Results were interpreted by D.S.L.H, K.V.S., D.H.H., V.P., S.R.M., H.R., and B.W.K. D.S.L.H. wrote the manuscript with input to methods from V.P., N.S.B., and T.O.S. and feedback from all authors. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Yuan Wang, Ankur Sharma and Frederick Varn for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Harwood, D.S.L., Pedersen, V., Bager, N.S. et al. Glioblastoma cells increase expression of notch signaling and synaptic genes within infiltrated brain tissue. Nat Commun 15, 7857 (2024). https://doi.org/10.1038/s41467-024-52167-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-52167-y
- Springer Nature Limited