Abstract
“NeoRAS WT” refers to the loss of RAS mutations (MTs) following first-line treatment in metastatic colorectal cancer (mCRC). We evaluate the incidence and clinicopathological characteristics of NeoRAS WT mCRC using next-generation sequencing of plasma circulating tumor DNA. Patients with mCRC enrolled in the GOZILA study initially diagnosed with tissue RAS MT mCRC and received subsequent systemic therapy are eligible. NeoRAS WT is defined as the absence of detectable RAS MT in plasma and assessed in all eligible patients (Group A) and in a subgroup with at least one somatic alteration detected in plasma (Group B). Overall, 478 patients are included. NeoRAS WT prevalence is 19.0% (91/478) in Group A and 9.8% (42/429) in Group B. Absence of liver or lymph node metastasis and tissue RAS MTs other than KRAS exon 2 MTs are significantly associated with NeoRAS WT emergence. Overall, 1/6 and 2/6 patients with NeoRAS WT treated with anti-EGFR monoclonal antibodies (mAbs) show partial response and stable disease for ≥6 months, respectively. NeoRAS WT mCRC is observed at a meaningful prevalence, and anti-EGFR mAb-based therapy may be effective.
Similar content being viewed by others
Introduction
RAS genes (KRAS, NRAS, and HRAS) are oncogenes that produce RAS proteins responsible for transmitting signals that promote cell growth1. RAS mutations (MTs) are associated with carcinogenesis in many cancers; however, effective therapies for these cancers are limited2. Overall, 17% of solid tumors involve KRAS MTs, including ~50% of metastatic colorectal cancer (mCRC) cases3,4. Anti-epidermal growth factor receptor (EGFR) monoclonal antibodies (mAbs), namely, cetuximab and panitumumab, are a mainstay of mCRC therapy, but they are ineffective against RAS MT mCRC5,6. The current clinical practice guidelines recommend RAS gene testing before administering anti-EGFR mAb-based therapy to patients with mCRC7,8,9. Given the lack of treatment options for these patients and their poor prognosis10, further therapeutic developments are urgently needed.
Understanding of the mechanisms by which the RAS mutational status evolves before and after treatment has recently advanced, primarily owing to advances in circulating tumor DNA (ctDNA)-based diagnostics that enable minimally invasive, simple, and repeatable testing11,12,13. RAS MTs in ctDNA have been identified in patients with anti-EGFR mAb-resistant RAS “WT” mCRC, many of whom acquire the condition after exposure to anti-EGFR mAbs14,15. Contrarily, initially diagnosed RAS MT patients have RAS WT after chemotherapy-based treatment. This phenomenon, called “NeoRAS WT” mCRC, has attracted research attention because anti-EGFR mAbs, conventionally ineffective against RAS MT mCRC, may be effective in NeoRAS WT mCRC16,17,18. Several clinical trials on the safety and efficacy of anti-EGFR mAbs for the treatment of NeoRAS WT mCRC are ongoing19,20.
The incidence of NeoRAS WT mCRC varied widely from 20% to 80% among previous studies17,21,22,23,24,25. However, these studies were limited by small sample sizes, varying lines of treatment, and lack of a consensus definition of the NeoRAS WT mCRC population. In particular, when only RAS MTs were measured, it was not possible to determine whether RAS MTs had disappeared or whether ctDNA was not detectable. Attempts have since been made to assess ctDNA using next-generation sequencing (NGS) or detect methylated ctDNA21,25.
The SCRUM-Japan GOZILA study is an ongoing Japanese nationwide screening project that analyzes cancer genomic information with NGS from the plasma ctDNA of patients with advanced gastrointestinal cancer using the Guardant360® assay26. Initial results demonstrated that ctDNA NGS had a faster turnaround time and greater diagnostic yield than tissue NGS26. In addition, other studies have validated ctDNA NGS for the assessment of RAS mutations, BRAF V600E mutation27, microsatellite instability28, and HER2 amplification29 with high concordance to tissue-based companion diagnostics in patients with mCRC. The present study aimed to use this platform to examine the incidence and clinicopathological characteristics of NeoRAS WT mCRC.
Results
Patient characteristics
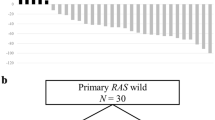
Among the 4991 patients with mCRC enrolled in GOZILA between March 2018 and February 2022, 647 patients were initially diagnosed with RAS MT mCRC using pretreatment tissue analysis. Among them, 169 patients in whom ctDNA was measured before the administration of first-line chemotherapy (N = 152) and in whom plasma RAS MTs were different from the tissue RAS MTs (N = 17) were excluded. Finally, 478 patients were evaluated. The CONSORT flow diagram is shown in Fig. 1. The median age at the time of blood sampling was 62.0 years (range, 25.0–85.0 years), and 249 (52.1%) patients were male. Among the 478 patients, 153 (32.0%) and 306 (64.0%) patients had right-sided tumors and multi-organ metastatic sites, respectively (Supplementary Data 1). The lungs were the most frequent site of metastasis (60.7%), followed by the liver (56.9%), lymph nodes (28.0%), and peritoneum (27.8%). Furthermore, 357 (74.7%) patients were treated with first-, second-, or third-line chemotherapy at the time of sampling; 129 (27.0%) patients received 5-fluorouracil, capecitabine, or S-1 and irinotecan combination therapy; and 334 (69.9%) patients received anti-vascular endothelial growth factor antibodies before blood collection for ctDNA analysis.
Tissue samples from 647 RAS MT mCRC patients are initially included. Among these, samples from 169 patients are excluded; finally, samples from 478 patients are included in the analysis. Somatic mutations in KRAS and NRAS are detected in 387 patients with mCRC in their plasma. Thus, the incidence of NeoRAS WT mCRC in Group A is 19%. Overall, 30 patients have no detectable genetic alteration, and 2 patients have germline mutations. The possibility of CH cannot be ruled out in 17 patients. Therefore, the incidence of NeoRAS WT mCRC in Group B is 9.8%. ctDNA: circulating tumor DNA, RAS: rat sarcoma viral oncogene homolog, WT wild-type, MT mutant, CH clonal hematopoiesis, mCRC metastatic colorectal cancer.
Association of clinicopathological characteristics with NeoRAS WT
Somatic MTs in KRAS and NRAS exons 2, 3, and 4 were detected in the plasma from 387 of the 478 (81.0%) patients with mCRC. Thus, the incidence of potential NeoRAS WT mCRC in Group A was 19.0% (91/478). Among the 91 patients, 30 patients had no detectable genetic alterations, and 2 had putative germline MTs (BRCA2 and ATM). The possibility of clonal hematopoiesis (CH) could not be ruled out in 17 patients (TP53, ATM, and GNAS) without any other somatic alternations. Therefore, the incidence of NeoRAS WT mCRC in Group B after excluding these potential confounding factors was 9.8% (42/429). Figure 2 shows the details of other somatic alterations in Group B. The median variant frequency was 0.23% (range: 0.08–29.05), and APC mutations were the most frequently detected mutations. When the limits of detection (i.e., minimum detectable mutant allele frequency [MAF]) were defined as 0.34% and 1%, the incidence rates of NeoRAS WT mCRC in Group B were 2.5% (10/397) and 1.5% (6/393), respectively.
In Group A, there were significant differences in the frequency of NeoRAS between single-organ and multi-organ metastatic sites (30.2% vs. 12.7%, P < 0.001) and between the absence and presence of liver (34.0% vs. 7.7%, P < 0.001), lymph node (22.1% vs. 11.2%, P < 0.001), peritoneum (15.4% vs. 28.6%, P < 0.001), and bone (20.2% vs. 5.4%, P < 0.001) metastases. As for the treatment regimen and lines, there were significant differences in the frequency of NeoRAS between testing immediately prior to second-to-fourth-line treatment and to later-line treatment (21.8% vs. 10.7%, P = 0.007), prior use and no prior use of regorafenib (6.8% vs. 20.3%, P < 0.001), and history and no history of vascular endothelial growth factor inhibitor use (22.5% vs. 11.1%, P < 0.001). In Group B, there were significant differences in the frequency of NeoRAS between single-organ and multi-organ metastases (15.7% vs. 6.6%, P = 0.004), absence and presence of liver metastasis (16.6% vs. 5.6%, P < 0.001), and history and no history of vascular endothelial growth factor inhibitor use (11.9% vs 5.2%, P = 0.035).
We compared the clinical characteristics of tissue RAS WT (N = 1077), NeoRAS WT (Group A, N = 91; Group B, N = 42), and RAS MT/non-NeoRAS WT (N = 387) mCRC patients using the GOZILA database (Supplementary Data 2). NeoRAS WT mCRC patients had clinically similar characteristics to RAS MT mCRC patients. Regarding the primary site, NeoRAS WT patients had a higher incidence on the left side colon than RAS MT/non-NeoRAS WT patients, and the incidence was closer to that in RAS WT patients. There was no difference in incidence when the cutoff values for MAF were set at 0.34% and 1% compared with no cutoff.
Incidence of NeoRAS WT according to the tissue RAS variants
The identified RAS MTs are listed in Fig. 3. In Group A (N = 478), mutations in KRAS codons 12 and 13 were detected in 71.3% and 16.3% of patients, respectively. Mutations in KRAS codon 61 (3.3%), KRAS codon 117 (0.8%), KRAS codon 146 (3.6%), NRAS codon 12 (1.7%), NRAS codon 13 (0.6%), and NRAS codon 61 (2.3%) were less common (<10% of patients) than mutations in KRAS codons 12 and 13. The incidences of NeoRAS WT in KRAS codon 12, KRAS codon 13, KRAS codon 61, KRAS codon 146, NRAS codon 12, and NRAS codon 61 were 18.5%, 15.4%, 31.3%, 35.3%, 37.5%, and 18.2%, respectively. In Group A, the frequency of NeoRAS WT in KRAS exons 3 and 4 MT or NRAS MT mCRC tended to be higher than that in KRAS exons 2 and 3 MT (27.1% vs. 17.9%, P = 0.11)
The identified RAS MTs are listed. In both Group A and Group B, the frequency of NeoRAS WT in KRAS exons 3 and 4 MT or NRAS MT mCRC tends to be higher than that in KRAS exons 2 and 3 MT. Source data are provided as a Source Data file. RAS rat sarcoma viral oncogene homolog, MT mutant, mCRC metastatic colorectal cancer, WT wild-type.
In Group B (N = 429), mutations in KRAS codons 12 and 13 were detected in 70.9% and 17.0% of patients, respectively. Mutations in KRAS codon 61 (3.0%), KRAS codon 117 (0.9%), KRAS codon 146 (3.5%), NRAS codon 12 (1.4%), NRAS codon 13 (0.7%), and NRAS codon 61 (2.6%) were less common (<10% of patients) than mutations in KRAS codons 12 and 13. The incidences of NeoRAS WT in KRAS codon 12, KRAS codon 13, KRAS codon 61, KRAS codon 146, NRAS codon 12, and NRAS codon 61 were 8.6%, 9.6%, 15.4%, 26.7%, 16.7%, and 18.2%, respectively. The frequency of NeoRAS WT in KRAS exons 3 and 4 MT or NRAS MT mCRC also tended to be higher than that in KRAS exons 2 and 3 MT in Group B (17.3% vs. 8.8%, P = 0.076).
Multivariate analysis results
In the multivariate logistic regression analysis in Group A, absence of liver (odds ratio [OR]: 0.18, 95% confidence interval [CI]: 0.09–0.34, P < 0.00001) and lymph node (OR: 0.46, 95% CI: 0.22–0.94, P = 0.033) metastases was significantly related to the presence of NeoRAS WT mCRC (Fig. 4a). Similarly, in Group B, absence of liver metastasis and tissue RAS MTs other than KRAS exon 2 MTs were significantly related to the presence of NeoRAS WT mCRC (liver metastasis: OR: 0.29, 95% CI: 0.15–0.59, P = 0.0005; original RAS status: OR: 2.33; 95% CI: 1.0–5.42, P = 0.049; Fig. 4b).
Forest plot depicting the logistic regression multivariate analysis for the appearance of NeoRAS WT mCRC in Group A (4a) and Group B (4b). The adjusted odds ratio (OR: blue plot) and 95% confidence interval (CI: black horizontal lines) are shown for each factor. Various clinical factors and their association with the appearance of NeoRAS WT mCRC, as indicated by OR, are analyzed across the cohort using the two-sided Wald chi-squared test. The unadjusted OR of the appearance of NeoRAS WT mCRC is calculated as X/Y. For instance, about liver metastasis, X is defined as “the proportion of mCRC with liver metastases that became NeoRAS WT mCRC,” and Y is defined as “the proportion of mCRC without liver metastases that became NeoRAS WT mCRC.” An adjusted OR is an OR that has been adjusted to account for other predictor variables in the model (adjusted OR: 0.18, 95% CI: 0.09–0.34, P = 1.43 × 10−7). Vertical dotted line: the null hypothesis. Number of events: Group A = 91; Group B = 42. Source data are provided as a Source Data file. mCRC metastatic colorectal cancer, WT wild-type, RAS rat sarcoma viral oncogene homolog, VEGF vascular endothelial growth factor.
Clinical outcomes of NeoRAS WT mCRC patients treated with anti-EGFR therapy
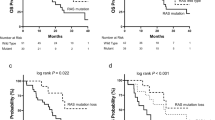
In this cohort, six NeoRAS WT patients were treated with anti-EGFR mAb-based therapy regimens. The treatment lines ranged from fourth to seventh, and the treatment regimens included anti-EGFR mAb monotherapy (N = 2) and mAb in combination with chemotherapy (N = 4). Among these 6 patients, 1 patient had partial response (PR), and another 2 patients had stable disease (SD) for at least 6 months. A summary of the treatment courses of the six patients is presented in Fig. 5.
Among the 6 NeoRAS WT patients who are treated with chemotherapy, including anti-EGFR therapy, 1 patient have partial response, and another 2 patients have stable disease for at least 6 months. EGFR: epidermal growth factor receptor, RAS: rat sarcoma viral oncogene homolog WT wild-type, FOLFIRI a combination of calcium folinate and fluorouracil with irinotecan hydrochloride hydrate, PFS progression-free survival, MAF mutant allele frequency, NA not available.
Representative cases of NeoRAS WT mCRC
We describe below two representative cases of mCRC patients undergoing standard-of-care regimens for RAS MT mCRC and reverting from RAS MT to NeoRAS WT.
Case 1 was a 57-year-old woman with metachronous hepatic and lung-metastasized sigmoid cancer (Fig. 6) Initially, this patient was diagnosed with localized sigmoid colon adenocarcinoma. Primary resection was performed followed by adjuvant chemotherapy. The primary tumor and lymph node lesions demonstrated the presence of NRAS G12C MT, although the MAF was close to the test’s cutoff value (primary average MAF: 5.0%, lymph node MAF: 1%) according to whole-exome tissue sequencing (Supplementary Table 1). On genomic profiling of tissue samples, using both whole-exome sequencing and FoundationOne CDx®, NRAS G12C MT, APC MT (R1114*, K1310fa), and TP53 MT (R175H) were detected. Shortly after adjuvant chemotherapy, liver and lung metastases were detected. Genomic profiling by plasma NGS (Guardant 360®) before chemotherapy initiation revealed APC MT (R1114*, K1310fa) and TP53 MT (R175H), but NRAS G12C MT was not detected.
mCRC metastatic colorectal cancer, WT wild-type, RAS rat sarcoma viral oncogene homolog, MAF mutant allele frequency, PCR-rSSO polymerase chain reaction-reverse sequence-specific oligonucleotide, CapeOX combination of capecitabine and oxaliplatin, FOLFIRI combination of calcium folinate, fluorouracil, and irinotecan hydrochloride hydrate, AFL aflibercept, Rego regorafenib, FTD/TPI trifluridine tipiracil hydrochloride, BV bevacizumab.
Case 2 was a 46-year-old woman with synchronous hepatic-, lung-, and ovary-metastasized rectal cancer (Fig. 7). Primary resection and metastasectomy of the ovary were performed before chemotherapy initiation because of rectal stenosis. On genomic profiling using whole-exome sequencing, an NRAS MT (Q61H) in both primary (average MAF: 55.7%) and metastatic (average MAF: lymph node, 13.7%; ovary, 33.2%) tumors was detected (Supplementary Table 2). This patient had received a first-line standard-of-care regimen (capecitabine, oxaliplatin and bevacizumab [BV]) that led to PR. Therefore, surgery for liver metastasis was performed. NRAS MT (Q61H) was detected, but MAF was very low at 0.95%, and relative clonality of NRAS was also low at 26.8% (Supplementary Table 2). The number of lung metastases increased after liver metastasectomy, and radical resection was considered difficult; hence, we decided to continue chemotherapy. The chemotherapy regimen was switched to folinic acid, 5-fluorouracil, and irinotecan (FOLFIRI) + BV owing to chemotherapy-induced peripheral neuropathy. After 1 year and 6 months of chemotherapy, the number of lung metastases did not increase and continued to decrease, and the patient underwent lung metastasectomy. Repeat whole-exome sequencing was performed using a lung metastasis specimen, and NRAS MT (Q61H) was detected (average MAF: 33.4%) (Supplementary Table 3). Approximately 1 year after lung metastasectomy, peritoneal metastases occurred, and FOLFIRI + BV was re-started. After disease progression following FOLFIRI + BV treatment, alterations including NRAS Q61H were not detected using plasma NGS (Guardant 360®).
mCRC metastatic colorectal cancer, WT wild-type, RAS rat sarcoma viral oncogene homolog, MAF mutant allele frequency, PCR-rSSO polymerase chain reaction-reverse sequence-specific oligonucleotide, CapeOX combination of capecitabine and oxaliplatin, BV bevacizumab, FOLFIRI combination of calcium folinate, fluorouracil, and irinotecan hydrochloride hydrate, FTD/TPI trifluridine tipiracil hydrochloride, Rego regorafenib.
Except the two cases for which exome sequencing was performed, there were 18 cases for which tissue NGS data (SCRUM-JAPAN) were available. Supplementary Data 3 shows a comparison between tissue and ctDNA genomic data.
Discussion
Patients with RAS MT mCRC who have failed chemotherapy have a poor prognosis and few effective treatment options. In the current study, we defined NeoRAS WT as a meaningful subset of RAS MT mCRC, with a prevalence of nearly 10%, that might benefit from available anti-EGFR mAb-based therapies. Previous studies in this area reported highly variable results17,21,22,23,24,30,31 (Supplementary Table 3) owing, in varying degrees, to small sample sizes, divergent definitions of the indication, and limited diagnostic methods. Critically, previous reports examining the incidence of NeoRAS WT mCRC have measured only RAS MTs to determine the RAS mutational status. For example, Sato et al. reported conversion from RAS MT to RAS WT after first-line chemotherapy in 27 out of 62 (43.5%) patients, with a higher incidence in patients with lung metastases only, those who underwent primary tumor resection, and those who achieved first-line treatment response30. In addition, Sunakawa et al. reported that RAS MTs were undetected during disease progression in 62% of the patients based on the ctDNA data of 29 patients who were treated with modified FOLFOXIRI + BV as first-line chemotherapy (JACCRO CC-11 trial)24. However, the measurement of RAS MTs is influenced by the characteristics of the ctDNA assay, and this may lead to a high false-negative rate, especially when the tumor volume is low. Moreover, some ctDNA assays use cutoff values rather than limit of detection of ctDNA to determine wild-type and mutant clones; thus, even if a mutant clone is detected, it may still be reported as wild-type31.
To overcome this limitation, the presence of ctDNA should be confirmed using NGS or profiling of methylated genes. However, when methylated genes are used as the basis for the presence of ctDNA, the difference in sensitivity to detect somatic alteration should be taken into consideration. Henry et al. used NGS to assess the loss of RAS MT in patients with mCRC who had not previously been treated with anti-EGFR mAb. They reported that the conversion rate of RAS status ranged from 2% to 8% and that changes in RAS MT status from MT to WT in mCRC were relatively rare21. In addition, Nicolazzo et al. reported that using either NGS or methylation assay was a reliable method for confirming the presence of ctDNA in patients with NeoRAS WT mCRC25. The incidence of NeoRAS WT was 37.5% in patients with RAS MT mCRC diagnosed using both primary tumor tissue and ctDNA prior to treatment and treated with at least first-line chemotherapy25. These results suggest that when evaluating the incidence of NeoRAS WT mCRC, the results obtained from ctDNA should be interpreted carefully. In addition, the results would be more reliable if MTs other than RAS, such as APC MTs or methylated genes, could be confirmed.
Although the mechanism of conversion to NeoRAS WT mCRC remains unclear, one hypothesis is that mCRC with low-MAF RAS mutations may represent evidence of a mixture of RAS MT and WT clones within a tumor and that conversion to RAS WT after treatment occurs owing to a decrease in the proportion of RAS MT clones relative to RAS WT32. Similarly, Henry et al. reported that the loss of RAS MT is associated with a low pretreatment MAF21. In this study, the incidence of NeoRAS WT in tumors with KRAS exon 2 MT status tended to be lower than that in tumors with other RAS MTs. Furthermore, multivariate analysis showed that tissue RAS MTs other than KRAS exon 2 MTs were significantly related to the emergence of NeoRAS WT. This finding may reflect an underlying tendency of MTs other than KRAS exon 2 MTs toward increased clonal diversity, with both RAS MT and WT clones coexisting prior to therapy. This would explain the observed increase in NeoRAS WT conversion. Previous reports support our findings. Sato et al. reported that all patients (N = 4) with tissue RAS Q61H mutations had NeoRAS WT30. Yoshinami et al. also showed a high incidence of NeoRAS among patients with tissue RAS MTs in exons other than the KRAS exon 2 (5/8, 62.5%), and this finding was related to the emergence of NeoRAS WT in their multivariate analysis33,34. Loree et al.35 similarly described that the allelic fractions of MTs in KRAS exons 3 and 4 and NRAS were lower than those of KRAS exon 2 MTs.
mCRC with low MAF of RAS may reflect part of tumor heterogeneity. The two main types of heterogeneity in mCRC are spatial and temporal heterogeneity. Spatial heterogeneity includes inter- or intratumor heterogeneity and inter-metastatic heterogeneity. Regarding metastatic sites, mCRC patients with liver and lymph node metastases had a low frequency of NeoRAS WT in this study. In a previous meta-analysis, the pooled discordance proportion of KRAS between the primary tumor and metastatic site was 8% (95% CI: 5–10%)36, with a relatively high clonality. In addition, there have been several studies on liver metastasis and KRAS exon 2 MTs. Therefore, RAS clonality may differ among metastatic sites and RAS variants36,37. In temporal heterogeneity, clonal selection develops, and a series of secondary resistance mechanisms is established during systemic therapies. The representative cases of NeoRAS WT mCRC in this study are shown in Figs. 6 and 7. In Case 1, the genetic alterations were different between the primary tumor and lymph node metastasis, with intratumor and inter-metastatic heterogeneity. Treatment (adjuvant chemotherapy) caused clonal selection, which could have eliminated the RAS mutant clones. Therefore, the original tumor had a subclonal NRAS MT, which was unlikely to be driving the cancer and was simply lost during treatment.
In Case 2, the gene alterations were also different among the primary tumor, lymph node metastasis, and synchronous ovarian metastasis, with intratumor and inter-metastatic heterogeneity. Treatment (palliative chemotherapy) caused clonal selection and decreased both variant allele frequency and clonality of RAS. Therefore, clonal NRAS MT, which was outcompeted by a NRAS WT subclone, could result in a reversion from RAS MT to a phenotypically distinct RAS WT. One of the central roles of ctDNA is to overcome the difficulty of obtaining comprehensive genetic information owing to heterogeneity. ctDNA results should be interpreted carefully because ctDNA reflects comprehensive genomic information of the tumor that is easier to detect in mCRC patients with liver and lymph node metastases than in those with lung or peritoneal metastases38. To confirm this hypothesis, future analyses should compare the MAF and concordance of RAS status between liver and lymph node metastasis and other metastatic sites in patients with KRAS exon 2 and in those with other RAS MTs using samples of both tissue and ctDNA before and after treatment.
Furthermore, the NeoRAS WT phenomenon mainly occurs in bevacizumab-treated patients, confirming previous reports that bevacizumab as a first-line therapy is an independent predictor of RAS mutation clearance in ctDNA at the time of progressive disease39. However, the biological rationale for the association of bevacizumab with RAS mutation clearance is currently unknown. The present study observed a significant difference in univariate analysis, but it disappeared in multivariate analysis. Therefore, further research is needed to determine if bevacizumab history increases the frequency of NeoRAS WT.
The major concern related to NeoRAS WT mCRC is whether anti-EGFR mAbs have chemotherapeutic effects. In the present study, three of the six patients with NeoRAS WT mCRC treated with anti-EGFR mAb as a later-line treatment showed treatment response or long progression-free survival (PFS). Before considering the efficacy of anti-EGFR mAbs against NeoRAS WT mCRC, the clinical outcomes of anti-EGFR mAbs against low-MAF RAS mCRC should be reviewed. In the ad-hoc biomarker analysis of the CO.17 trial, 1 patient with a KRAS A59T MT (MAF = 2%) responded to cetuximab35. Furthermore, in the post-hoc analysis of the CRYSTAL trial, patients with low extended RAS MT levels, with signals between 0.1% and 5%, benefited from the addition of cetuximab. Similar findings were reported by Laurent-Puig et al., who found that patients with 1% mutated KRAS alleles had a similar response to patients with KRAS WT40, suggesting a potential gradient of efficacy based on the MAF of mutant RAS in the tumor6. The status of low-RAS MAF mCRC may change from MT to WT after treatment32. Therefore, anti-EGFR mAb is expected to have a chemotherapeutic effect on NeoRAS WT mCRC.
Several prospective and retrospective studies on the chemotherapeutic effects of anti-EGFR mAbs on NeoRAS WT mCRC have been conducted16,17,18,22,30,41. Mohamed et al. conducted a proof-of-concept study of anti-EGFR mAbs in patients with NeoRAS WT mCRC. The main results were an objective response rate of 55.6% and a PFS of 9 months in patients with Neo RAS WT mCRC who were treated with a combination of FOLFIRI and cetuximab41. Most patients received two or more chemotherapy regimens prior to FOLFIRI and cetuximab41. In addition, Nicolazzo et al. reported the efficacy of anti-EGFR mAb in combination with standard chemotherapy for NeoRAS WT mCRC identified by NGS and/or methylation analysis when patients were resistant to first-line therapy25. Among the 20 patients enrolled at the time of first-line treatment resistance, 6 patients received anti-EGFR mAbs and had a median PFS of 6.5 months25. Among the 20 patients enrolled at the time of second/third-line treatment resistance, 4 patients received anti-EGFR mAbs and had a median PFS of 6.0 months25. This result suggests that anti-EGFR mAbs may be effective for patients with NeoRAS WT mCRC. Therefore, in clinical practice, the RAS status of mCRC patients with tissue RAS MT other than in KRAS exon 2 and without liver or lymph node metastases should be re-assessed using ctDNA at the time of progressive disease, and the indication for treatment with anti-EGFR mAb should be explored. Several clinical trials are ongoing to evaluate the therapeutic efficacy of anti-EGFR mAbs against NeoRAS WT mCRC19,20.
Our study has some limitations. It lacked information on ctDNA prior to treatment. Furthermore, the number of EGFR antibody-treated individuals was too small to make definitive conclusions. In addition, details of genetic alterations in pretreatment tissues of NeoRAS WT patients who responded to treatment with anti-EGFR mAb were unclear.
Moreover, there was no consensus on the definition of CH in mCRC. In this study, CH was identified only on a reported basis, and DNA from white blood cells was not examined. Finally, the sensitivity of ctDNA detection may have affected the study results. We did not set a detection limit for MAF, and thus, the incidence of NeoRAS WT in Group B was defined as the absence of detectable RAS MT, but detectable for other alterations regardless of MAF, excluding possible CH alterations. As reported in a previous study, Gardant 360 had predictive sensitivities of >98%, 84.0%, and 50% when MAF was set as ≥1%, >0.34%–<1%, and ≤0.34%, respectively42 Future technology that can accurately determine the presence of ctDNA with a cutoff neighborhood is desirable, and further large-scale analyses using both tissue and ctDNA information before and after treatment are needed to accurately clarify the incidence and clinicopathological characteristics of NeoRAS WT mCRC.
In conclusion, NeoRAS WT mCRC is associated with the absence of liver and lymph node metastases, as well as RAS MTs other than KRAS exon 2 MTs. Therefore, treatment with anti-EGFR mAbs may be effective in patients with NeoRAS WT mCRC.
Methods
Patients
This study was approved by the Institutional Review Board of the Cancer Institute Hospital of the Japanese Foundation of Cancer Research (IRB number: 2021-GB-009) and was conducted in accordance with the principles of the Declaration of Helsinki. The protocol was described on the hospital’s website, and the participants were given the opportunity to opt out of the study.
NGS data (Guardant360®, Guardant Health, Inc, Redwood City, CA, USA) were used to confirm the incidence of NeoRAS WT mCRC. Patients with mCRC enrolled in the GOZILA study between March 2018 and February 2022 and who exhibited tissue RAS MTs (KRAS or NRAS exon 2, 3, and 4 mutations) before chemotherapy initiation were eligible, regardless of the treatment line. Briefly, the GOZILA study is a nationwide plasma genomic profiling study involving 31 core cancer institutions in Japan. Patients with metastatic gastrointestinal cancers are eligible for enrollment26.
Blood samples, circulating tumor DNA isolation, and sequencing
NGS of ctDNA was performed using the Guardant360® system at Guardant Health. Guardant360 detects single-nucleotide variants, indels, fusions, copy number alterations, and microsatellite instability in 74 genes26. In the GOZILA study, 2 × 10 mL whole blood was collected in a cell-free DNA BCT® (Streck, Inc., La Vista, NE, USA) and sent to Guardant Health. Then, 5–30 ng of cell-free DNA (cfDNA) isolated from plasma was labeled with non-redundant oligonucleotides (“molecular barcoding”), enriched using targeted hybridization capture, and sequenced on the NextSeq 550 platform (Illumina, San Diego, CA, USA)26. Base call files generated using RTA software version 2.12 (Illumina, San Diego, CA, USA) were demultiplexed using bcl2fastq (version 2.19) and processed using a custom pipeline for molecular barcode detection, sequencing adapter trimming, and base quality trimming. Processed reads were aligned to hg19 using the Burrows–Wheeler aligner-MEM algorithm (arXiv:1303.3997v2). The ctDNA fraction was determined using the maximum variant allelic fraction26.
Tumor tissue DNA sequencing
All patients were diagnosed with mCRC based on the analysis of tissue RAS MTs. The mutation profiles of the tissue samples were determined using standard-of-care procedures validated by each hospital. For example, the RASKET-B KIT (MBL, Nagoya, Japan), which applied the PCR-reverse sequence-specific oligonucleotide (PCR-rSSO) method, was used according to the manufacturer’s protocol43. The PCR-rSSO and Luminex MAP technologies allow multiplex molecular testing in a single well. We examined 12 types of RAS exon 2 mutations (G12S, G12C, G12R, G12D, G12V, G12A, G13S, G13C, G13R, G13D, G13V, and G13A), 8 types of RAS exon 3 mutations (A59T, A59G, Q61K, Q61E, Q61L, Q61P, Q61R, and Q61H), and 4 types of RAS exon 4 mutations (K117N, A146T, A146P, and A146V), as previously described43. The procedure of whole-exome sequencing is shown in the Supplementary Note44,45. Relative clonality was defined as “subclonal” if the MAF was less than 30% of the highest MAF in the sample and was defined as “clonal” if it was above this threshold29.
Endpoints
The primary endpoint was confirmation of the incidence of NeoRAS WT mCRC using NGS. The patients were divided into two groups based on genetic alterations, as follows: Group A, with tissue-confirmed pretreatment RAS MTs and no post-treatment RAS MTs detected on ctDNA analysis; Group B, with tissue-confirmed pretreatment RAS MTs, no post-treatment RAS MTs detected on ctDNA analysis, and other gene MTs confirmed post-treatment without CH. The following genes reported to be potentially CH genes in CRC patients46,47 were excluded: DNMT3A, TET2, TP53, CEBPA, ETV6, HRAS, PDGFRA, KMT2A, EZH2, GATA2, GNAS, PPM1D, ASLX1, SF3B1, SRSF2, CHEK2, and KMT2D. The secondary endpoints were differences in the incidence of NeoRAS WT mCRC according to the clinicopathological factors and chemotherapeutic efficacy of anti-EGFR mAbs against NeoRAS WT mCRC. Whole-exome multiregion spatial sequencing was also performed on DNA to confirm intratumor heterogeneity in representative NeoRAS WT mCRC (N = 2).
Data collection
Data on the following clinicopathological factors were collected: age, sex, primary tumor location, metastatic site, timing of sample collection, treatment regimen, treatment lines at the time of sampling, and therapeutic effects depending on the tissue RAS variant. Complete response (CR), PR, SD, and progressive disease were defined following the Response Evaluation Criteria in Solid Tumours guidelines, v1.148. The response rate indicated the proportion of patients who achieved CR or PR to chemotherapy, and the disease control rate indicated the proportion of patients who had a CR, PR, or SD response to therapy. PFS was defined as the time from the first day of anti-EGFR therapy to the first objective evidence of disease progression or death from any cause.
Statistical analysis
Clinicopathological factors were compared between patients with NeoRAS WT and with RAS MT mCRC. Continuous variables were compared with the Mann–Whitney U test, and between-group comparisons were performed with the χ2 test and Fisher’s exact test. Univariate and multivariate logistic regression analyses were performed to examine the factors related to NeoRAS WT mCRC. All factors that showed significance in the univariate analysis were included in the multivariate analysis. All statistical analyses were performed using the statistical software “EZR” (Saitama Medical Center, Jichi Medical University, Saitama, Japan) in R and R commander49. All tests were two-sided, and P < 0.05 was considered significant. We have uploaded the raw data used in this study as a Source Data file.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The authors declare that all variant data used in the conduct of the analyses are available within the article and its supplementary information. The datasets, including individual participant data supporting the results reported in this article, will be made available within 3 months from initial request to researchers depending on methodological considerations. The initial contact for the request will be made with the corresponding authors Eiji Shinozaki (eiji.shinozaki@jfcr.or.jp) and Takayuki Yoshino (tyoshino@east.ncc.go.jp). In this cohort, raw ctDNA sequence data was not provided by Guardant Health, other than the source data used for the analysis because of data privacy regulations and restrictions on their use in patient consent forms. Source data are provided as a source data file. Source data are provided with this paper.
References
Barbacid, M. ras genes. Annu. Rev. Biochem. 56, 779–827 (1987).
Hong, D. S. et al. KRASG12C inhibition with sotorasib in advanced solid tumors. N. Engl. J. Med. 383, 1207–1217 (2020).
AACR Project GENIE Consortium AACR Project GENIE: powering precision medicine through an international consortium. Cancer Discov. 7, 818–831 (2017).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Douillard, J. Y. et al. Panitumumab-FOLFOX4 treatment and RAS mutations in colorectal cancer. N. Engl. J. Med. 369, 1023–1034 (2013).
Van Cutsem, E. et al. Fluorouracil, leucovorin, and irinotecan plus cetuximab treatment and RAS mutations in colorectal cancer. J. Clin. Oncol. 33, 692–700 (2015).
Hashiguchi, Y. et al. Japanese Society for Cancer of the Colon and Rectum (JSCCR) guidelines 2019 for the treatment of colorectal cancer. Int. J. Clin. Oncol. 25, 1–42 (2020).
Morris, V. K. et al. Treatment of metastatic colorectal cancer: ASCO Guideline. J. Clin. Oncol. 41, 678–700 (2023).
Cervantes, A. et al. Metastatic colorectal cancer: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up. Ann. Oncol. 34, 10–32 (2023).
Fakih, M. et al. Real-world study of characteristics and treatment outcomes among patients with KRAS p.G12C-mutated or other KRAS mutated metastatic colorectal cancer. Oncologist 27, 663–674 (2022).
Siravegna, G., Marsoni, S., Siena, S. & Bardelli, A. Integrating liquid biopsies into the management of cancer. Nat. Rev. Clin. Oncol. 14, 531–548 (2017).
Wan, J. C. M. et al. Liquid biopsies come of age: towards implementation of circulating tumour DNA. Nat. Rev. Cancer 17, 223–238 (2017).
Osumi, H., Shinozaki, E., Yamaguchi, K. & Zembutsu, H. Clinical utility of circulating tumor DNA for colorectal cancer. Cancer Sci. 110, 1148–1155 (2019).
Misale, S. et al. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature 486, 532–536 (2012).
Siravegna, G. et al. Clonal evolution and resistance to EGFR blockade in the blood of colorectal cancer patients. Nat. Med. 21, 795–801 (2015).
Gazzaniga, P., Raimondi, C., Urbano, F. & Cortesi, E. EGFR inhibitor as second-line therapy in a patient with mutant RAS metastatic colorectal cancer: circulating tumor DNA to personalize treatment. JCO Precis. Oncol. 2, 1–6 (2018).
Raimondi, C. et al. Transient disappearance of RAS mutant clones in plasma: a counterintuitive clinical use of EGFR inhibitors in RAS mutant metastatic colorectal cancer. Cancers 11, 42 (2019).
Osumi, H. et al. NeoRAS wild-type in metastatic colorectal cancer: myth or truth?-Case series and review of the literature. Eur. J. Cancer 153, 86–95 (2021).
Fernández Montes, A. et al. FOLFIRI plus panitumumab as second-line treatment in mutated RAS metastatic colorectal cancer patients who converted to wild type RAS after receiving first-line FOLFOX/CAPOX plus bevacizumab-based treatment: phase II CONVERTIX trial. Ann. Oncol. 30, iv23–iv24 (2019).
Osumi, H. et al. Multicentre single-arm phase II trial evaluating the safety and effiCacy of panitumumab and iRinOtecan in NeoRAS Wild-type mEtaStatic colorectal cancer patientS (C-PROWESS trial): study protocol. BMJ Open 12, e063071 (2022).
Henry, J. et al. NeoRAS: incidence of RAS reversion from RAS mutated to RAS wild type. J. Clin. Oncol. 38, 180 (2020).
Nicolazzo, C. et al. Circulating methylated DNA to monitor the dynamics of RAS mutation clearance in plasma from metastatic colorectal cancer patients. Cancers 12, 3633 (2020).
Moati, E. et al. Plasma clearance of RAS mutation under therapeutic pressure is a rare event in metastatic colorectal cancer. Int. J. Cancer 147, 1185–1189 (2020).
Sunakawa, Y. et al. Dynamic changes in ras gene status in circulating tumour DNA: a phase II trial of first-line FOLFOXIRI plus bevacizumab for RAS-mutant metastatic colorectal cancer (JACCRO CC-11). ESMO Open 7, 100512 (2022).
Nicolazzo, C. et al. True conversions from RAS mutant to RAS wild-type in circulating tumor DNA from metastatic colorectal cancer patients as assessed by methylation and mutational signature. Cancer Lett. 507, 89–96 (2021).
Nakamura, Y. et al. Clinical utility of circulating tumor DNA sequencing in advanced gastrointestinal cancer: SCRUM-Japan GI-SCREEN and GOZILA studies. Nat. Med. 26, 1859–1864 (2020).
Aoki, Y. et al. Clinical validation of plasma-based genotyping for RAS and BRAF V600E mutation in metastatic colorectal cancer: SCRUM-Japan GOZILA substudy. JCO Precis. Oncol. 7, e2200688 (2023).
Nakamura, Y. et al. Clinical validity of plasma-based genotyping for microsatellite instability assessment in advanced GI cancers: SCRUM-Japan GOZILA substudy. JCO Precis. Oncol. 6, e2100383 (2022).
Nakamura, Y. et al. Circulating tumor DNA-guided treatment with pertuzumab plus trastuzumab for HER2-amplified metastatic colorectal cancer: a phase 2 trial. Nat. Med. 27, 1899–1903 (2021).
Sato, S. et al. Chemotherapy-induced reversion of mutant RAS to wild-type RAS in metastatic colorectal cancer. Anticancer Res. 42, 2625–2635 (2022).
Bando, H. et al. A multicentre, prospective study of plasma circulating tumour DNA test for detecting RAS mutation in patients with metastatic colorectal cancer. Br. J. Cancer 120, 982–986 (2019).
Udagawa, S. et al. Circulating tumor DNA: the dawn of a new era in the optimization of chemotherapeutic strategies for metastatic colo-rectal cancer focusing on RAS mutation. Cancers 15, 1473 (2023).
Yoshinami, Y. et al. A multi-institutional observational study evaluating the incidence and the clinicopathological characteristics of neo RAS wild-type metastatic colorectal cancer. J. Clin. Oncol. 41, 206–211 (2023).
Osumi, H. et al. A multi-institutional observational study evaluating the incidence and the clinicopathological characteristics of NeoRAS wild-type metastatic colorectal cancer. Transl. Oncol. 35, 101718 (2023).
Loree, J. M. et al. Expanded low allele frequency RAS and BRAF V600E testing in metastatic colorectal cancer as predictive biomarkers for cetuximab in the randomized CO.17 Trial. Clin. Cancer Res. 27, 52–59 (2021).
Bhullar, D. S. et al. Biomarker concordance between primary colorectal cancer and its metastases. EBiomedicine 40, 363–374 (2019).
Kim, M. J. et al. Different metastatic pattern according to the KRAS mutational status and site-specific discordance of KRAS status in patients with colorectal cancer. BMC Cancer 12, 347 (2012).
Bando, H. et al. Effects of metastatic sites on circulating tumor DNA in patients with metastatic colorectal cancer. JCO Precis. Oncol. 6, e2100535 (2022).
Nicolazzo, C. et al. RAS mutation conversion in bevacizumab-treated metastatic colorectal cancer patients: a liquid biopsy based study. Cancers 14, 802 (2022).
Laurent-Puig, P. et al. Clinical relevance of KRAS-mutated subclones detected with picodroplet digital PCR in advanced colorectal cancer treated with anti-EGFR therapy. Clin. Cancer Res. 21, 1087–1097 (2015).
Bouchahda, M. et al. Undetectable RAS-mutant clones in plasma: possible implication for anti-EGFR therapy and prognosis in patients with RAS-mutant metastatic colorectal cancer. JCO Precis. Oncol. 4, 1070–1079 (2020).
Caughey, B. A. et al. Identification of an optimal mutant allele frequency to detect activating KRAS, NRAS, and BRAF mutations in a commercial cell-free DNA next-generation sequencing assay in colorectal and pancreatic adenocarcinomas. J. Gastrointest. Oncol. 14, 2083–2096 (2023).
Taniguchi, H. et al. Clinical validation of newly developed multiplex kit using Luminex xMAP technology for detecting simultaneous RAS and BRAF mutations in colorectal cancer: results of the RASKET-B study. Neoplasia 20, 1219–1226 (2018).
Ichimiya, S. et al. Contribution of pre-existing neoantigen-specific T cells to a durable complete response after tumor-pulsed dendritic cell vaccine plus nivolumab therapy in a patient with metastatic salivary duct carcinoma. Immunol. Invest. 51, 1498–1514 (2022).
Morisaki, T. et al. Neoantigens elicit T cell responses in breast cancer. Sci. Rep. 11, 13590 (2021).
Ptashkin, R. N. et al. Prevalence of clonal hematopoiesis mutations in tumor-only clinical genomic profiling of solid tumors. JAMA Oncol. 4, 1589–1593 (2018).
Wu, H.-T. et al. Characterization of clonal hematopoiesis of indeterminate potential mutations from germline whole exome sequencing data. J. Clin. Oncol. 38, 1525 (2020).
Eisenhauer, E. A. et al. New response evaluation criteria in solid tumours: revised RECIST guideline. version 1.1. Eur. J. Cancer 45, 228–247 (2009).
Kanda, Y. Investigation of the freely available easy-to-use software “EZR” for medical statistics. Bone Marrow Transpl. 48, 452–458 (2013).
Acknowledgements
The authors would like to thank all the patients and their families, all the investigators, research nurses, and study coordinators who participated in this study. Collaborators with GI-SCREEN and GOZILA study Aichi Cancer Center Hospital: Yukiya Narita and Toshiki Masuishi; Gifu University Hospital: Kazuhiro Yoshida; Kagawa University Hospital: Akihito Tsuji; Kanazawa University: Yuta Adachi and Koushiro Ohtsubo; Kyorin University: Junji Furuse; Kyushu University: Eiji Oki; National Cancer Center Hospital: Ken Kato; Saitama Medical University: Yosuke Horita; Shimane Prefectural Central Hospital: Akiyoshi Kanazawa; Shizuoka Cancer Center: Kentaro Yamazaki; St. Marianna University: Takako Nakajima; University of Tsukuba Hospital: Toshikazu Moriwaki. This work was supported by the National Cancer Center Research and Development Fund (31-A-5 to A. Ohtsu) and SCRUM-Japan Funds (http://www.scrum-japan.ncc.go.jp/index.html).
Author information
Authors and Affiliations
Contributions
Hiroki Osumi (conceptualization: lead; data curation: lead; formal analysis: lead; investigation: lead; methodology: lead; project administration: lead; and writing – original draft: lead). Eiji Shinozaki (conceptualization: equal; project administration: lead; supervision: lead; and writing – review & editing: lead). Yoshiaki Nakamura (funding acquisition: lead; methodology: lead; resources: lead; and visualization: lead; writing – review & editing: lead) Taito Esaki (writing – review & editing: supporting). Hiroya Taniguchi (writing – review & editing: supporting). Hironaga Satake (writing – review & editing: supporting). Yu Sunakawa (writing – review & editing: supporting). Yoshito Komatsu (writing – review & editing: supporting). Yoshinori Kagawa (writing – review & editing: supporting). Tadamichi Denda (writing – review & editing: supporting). Manabu Shiozawa (writing – review & editing: supporting). Taroh Satoh (writing – review & editing: supporting). Tomohiro Nishina (writing – review & editing: supporting). Masahiro Goto (writing – review & editing: supporting). Naoki Takahashi (writing – review & editing: supporting). Takeshi Kato (writing – review & editing: supporting). Hideaki Bando (writing – review & editing: supporting). Kensei Yamaguchi (supervision: supporting; writing – review & editing: supporting). Takayuki Yoshino (funding acquisition: lead; methodology: lead; project administration: lead; resources: lead; supervision: equal; writing – review & editing: supporting).
Corresponding authors
Ethics declarations
Competing interests
Eiji Shinozaki: Takeda Pharma, Merck, Lily, and Chugai Pharma. Yoshiaki Nakamura: Chugai Pharma, Guardant Health AMEA, and Merck Research Funding: Taiho Pharmaceutical (Inst), Guardant Health (Inst), Genomedia (Inst), Chugai Pharma (Inst), Guardant Health (Inst), Seattle Genetics (Inst), and Roche (Inst) Taito Esaki: Lilly, Taiho Pharmaceutical, Daiichi Sankyo, and Chugai Pharma Research Funding: Daiichi Sankyo (Inst), MSD (Inst), Novartis (Inst), Ono Pharmaceutical (Inst), Astellas Pharma (Inst), Lilly (Inst), Bayer (Inst), Nihon Kayaku (Inst), Amgen Astellas BioPharma (Inst), Parexel (Inst), IQVIA (Inst), Quintiles (Inst), Eisai (Inst), Pfizer (Inst), Chugai Pharma (Inst), Syneos Health (Inst), Asahi Kasei Pharma (Inst), Amgen (Inst), Dainippon Sumitomo (Inst), and Dainippon Sumitomo (Inst) Hisateru Yasui: Taiho Pharmaceutical, Chugai Pharma, Bristol-Myers Squibb Japan, Daiichi Sankyo, Lilly, Yakult Honsha, Bayer Yakuhin, Takeda, and Ono Pharmaceutical Research Funding: MSD (Inst), Ono Pharmaceutical (Inst), Astellas Pharma (Inst), AstraZeneca (Inst), and Daiichi Sankyo (Inst) Hiroya Taniguchi: Bayer, Sanofi, Takeda, Chugai Pharma, Taiho Pharmaceutical, Lilly, Merck Serono, Yakult Honsha, Medical & Biological Laboratories Co., Ltd, Bristol-Myers Squibb Japan, MSD K.K, Novartis, Daiichi Sankyo, Mitsubishi Tanabe Pharma, Nippon Kayaku, and Ono Yakuhin Research Funding: Dainippon Sumitomo Pharma (Inst), Array BioPharma (Inst), MSD Oncology (Inst), Ono Pharmaceutical (Inst), Daiichi Sankyo (Inst), Sysmex (Inst), Novartis (Inst), and Takeda (Inst) Satake Hironaga: Bristol-Myers Squibb Co., Ltd., Bayer Co., Ltd., Chugai Pharmaceutical Co., Ltd, Daiichi Sankyo Co., Ltd., Eli Lilly Japan Co., Ltd., Merck Bio Pharma Co., Ltd., MSD Co., Ltd., Ono Pharmaceutical Co., Ltd., Sanofi Co., Ltd., Taiho Pharmaceutical Co., Ltd., Takeda Co., Ltd. and Yakult Honsha Co., Ltd. Research funding: Ono Pharmaceutical Co Ltd, Daiichi Sankyo, Taiho Pharmaceutical Co Ltd, and Takeda Pharmaceutical Co., Ltd. Yu Sunakawa: Taiho Pharmaceutical, Chugai Pharma, Takeda, Bayer Yakuhin, Bristol-Myers Squibb Japan, Lilly, Merck, Sysmex, MSD K.K, Ono Pharmaceutical, Daiichi Sankyo, Guardant Health, and Incyte Consulting or Advisory Role: Bristol-Myers Squibb Japan, MSD K.K, Daiichi Sankyo, and Merck Research Funding: Taiho Pharmaceutical, Takeda, Chugai Pharma, Lilly, Sanofi, and Otsuka Yoshito Komatsu: Lilly Japan, Taiho Pharmaceutical, Chugai Pharma, Takeda, Bayer Yakuhin, Bristol-Myers Squibb Co, Sanofi/Aventis, Merck, Yakult Honsha, Ono Pharmaceutical, Nipro Corporation, Moroo Co, Asahi Kasei, Mitsubishi Tanabe Pharma, Otsuka, Medical Review Co., Ltd, and Daiichi Sankyo Research Funding: MSD K.K, Taiho Pharmaceutical, Yakult Honsha, Bayer Yakuhin, DAIICHI SANKYO CO., Ltd, Ono Pharmaceutical, NanoCarrier, Eisai, Sanofi/Aventis, Sysmex, Shionogi, IQVIA, Parexel International Corporation, Astellas Pharma, Mediscience Planning, Sumitomo Dainippon Pharma Co., Ltd, A2 Healthcare, Incyte, Lilly (Inst), Nipro Corporation (Inst), and BeiGene (Inst) Yoshinori Kagawa. Consulting or Advisory Role: Taiho Pharmaceutical, Merck, and Lilly Speakers’ Bureau: Lilly, Sanofi, Takeda, Merck, Taiho Pharmaceutical, MSD, Chugai Pharma, Yakult Pharmaceutical, Bayer, and Ono Pharmaceutical. Research Funding: Ono Pharmaceutical Tadamichi Denda: Daiichi Sankyo and Ono Pharmaceutical, Sysmex. Research Funding: Ono Pharmaceutical (Inst), Amgen (Inst), MSD, Pfizer, and Bristol-Myers Squibb Foundation. Shiozawa Manabu: Lilly, Takeda, Taiho, Ono, Yakult, and Merck. Taroh Satoh: Chugai Pharma, Merck Serono, Bristol-Myers Squibb, Takeda, Yakult Honsha, Lilly, Bayer Yakuhin, Ono Pharmaceutical, Merck, Astellas Pharma, Taiho Pharmaceutical, Nihon Kayaku, and Daiichi Sankyo Consulting or Advisory Role: Bayer Yakuhin, Lilly, Ono Pharmaceutical, Takara Bio, Merck Serono, and Nihon Kayaku Research Funding: Yakult Honsha (Inst), Chugai Pharma (Inst), Ono Pharmaceutical (Inst), Sanofi (Inst), Lilly (Inst), Daiichi Sankyo (Inst), Merck (Inst), Merck Serono (Inst), Gilead Sciences (Inst), Dainippon Sumitomo Pharma (Inst), and IQVIA (Inst) Tomohiro Nishina: Taiho Pharmaceutical, Ono Pharmaceutical, Bristol-Myers Squibb Japan, Daiichi Sankyo, Chugai Pharma Research Funding: MSD (Inst), Ono Pharmaceutical (Inst), Astellas Pharma (Inst), Daiichi Sankyo/UCB Japan (Inst), Bristol-Myers Squibb Japan (Inst), Chugai Pharma (Inst), Taiho Pharmaceutical (Inst), and AstraZeneca (Inst). Masahiro Goto: Daiichi Sankyo Company, Limited, Ono Pharmaceutical, Taiho Pharmaceutical, MSD K.K, Takeda, Sumitomo Dainippon Pharma Co., Ltd, Yakult Pharmaceutical, and Lilly Research Funding: Chugai Pharma, Taiho Pharmaceutical, and Nippon Kayaku Naoki Takahashi: Ono Pharmaceutical, Bristol-Myers Squibb Japan, and Taiho Pharmaceutical Takeshi Kato: Chugai Pharma, Ono Pharmaceutical, Takeda, Lilly, and Asahi Kasei Research Funding: Chugai Pharma Hideaki Bando: Taiho, Lilly, and Ono. M.W. Nihon Medi-Physics. Research grant: Ono Pharmaceutical. Kensei Yamaguchi. Consulting or Advisory Role: Bristol-Myers Squibb Japan, and Daiichi Sankyo. Speakers’ Bureau: Chugai Pharma, Bristol-Myers Squibb Japan, Takeda, Taiho Pharmaceutical, Lilly, Ono Pharmaceutical, Daiichi Sankyo, and Merck Research Funding: Ono Pharmaceutical (Inst), Taiho Pharmaceutical (Inst), Daiichi Sankyo (Inst), Lilly (Inst), Gilead Sciences (Inst), Yakult Honsha (Inst), Chugai Pharma (Inst), Boehringer Ingelheim (Inst), Eisai (Inst), MSD Oncology (Inst), Sanofi (Inst), and Bristol-Myers Squibb (Inst) Takayuki Yoshino: Chugai Pharma, Takeda Pharma, Merck, Bayer Yakuhin, Ono Pharmaceutical, and MSD K.K. Consulting Fee: Sumitomo Corp. Research Funding: MSD (Inst), Daiichi Sankyo Company, Limited (Inst), Ono Pharmaceutical (Inst), Taiho Pharmaceutical (Inst), Amgen (Inst), Sanofi (Inst), Pfizer (Inst), Genomedia (Inst), Sysmex (Inst), Nippon Boehringer Ingelheim (Inst), Chugai Pharma (Inst), and Eisai (Inst). No other potential conflicts of interest were reported.
Peer review
Peer review information
Nature Communications thanks Sebastian Stintzing and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Osumi, H., Shinozaki, E., Nakamura, Y. et al. Clinical features associated with NeoRAS wild-type metastatic colorectal cancer A SCRUM-Japan GOZILA substudy. Nat Commun 15, 5885 (2024). https://doi.org/10.1038/s41467-024-50026-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-50026-4
- Springer Nature Limited