Abstract
Neuroimaging studies have uncovered the neural roots of individual differences in human general fluid intelligence (Gf). Gf is characterized by the function of specific neural circuits in brain gray-matter; however, the association between Gf and neural function in brain white-matter (WM) remains unclear. Given reliable detection of blood-oxygen-level-dependent functional magnetic resonance imaging (BOLD-fMRI) signals in WM, we used a functional, rather than an anatomical, neuromarker in WM to identify individual Gf. We collected longitudinal BOLD-fMRI data (in total three times, ~11 months between time 1 and time 2, and ~29 months between time 1 and time 3) in normal volunteers at rest, and identified WM functional connectomes that predicted the individual Gf at time 1 (n = 326). From internal validation analyses, we demonstrated that the constructed predictive model at time 1 predicted an individual’s Gf from WM functional connectomes at time 2 (time 1 ∩ time 2: n = 105) and further at time 3 (time 1 ∩ time 3: n = 83). From external validation analyses, we demonstrated that the predictive model from time 1 was generalized to unseen individuals from another center (n = 53). From anatomical aspects, WM functional connectivity showing high predictive power predominantly included the superior longitudinal fasciculus system, deep frontal WM, and ventral frontoparietal tracts. These results thus demonstrated that WM functional connectomes offer a novel applicable neuromarker of Gf and supplement the gray-matter connectomes to explore brain–behavior relationships.
Similar content being viewed by others
Introduction
Neuroimaging and psychological studies have investigated the neural basis of the cognitive processes that motivate novel insights about brain–behavior relationships1. An enduring aim of brain and cognitive sciences is to understand individual differences in human intelligence2. Human general fluid intelligence (Gf) refers to an ability to think logically and to solve novel problems that do not rely on previously acquired knowledge3,4. Gf has been broadly quantified using a series psychometric test, which further provides a foundation to process brain–behavior associations5,6. Given that individual differences are inherent to Gf, it is crucial to identify neural correlates of Gf and corresponding variations in brain structure and function.
The neural correlates of individual differences in Gf may be associated with variations in brain size and connections2. A larger brain size (volume) consistently indicates higher intelligence7; this concept of “the bigger brain, the better intelligence” may result from the efficiency of information flow among neurons5,8. Recently, the information flow among certain areas associated with Gf have been quantified by functional connectivity studies. These results showed that the variational relationships of regions engaging in common or related performance (even at rest) may be the basis of individual differences in Gf9,10,11, and measurements of the activity of the resting-state human brain might carry information about intelligence12. Furthermore, the rate of information flow among the distributed parietal and frontal areas composing the parieto-frontal integration theory (P-FIT) network are likely to play key roles in intelligence13. It is not surprising that a large number of these brain regions and brain functional networks are related to individual Gf12,14,15, due to the diverse abilities associated with Gf, including understanding of daily tasks and problem solving. Accordingly, whole-brain functional connectivity measures may provide more holistic insight to determine an individual’s Gf, rather than global brain size.
Gf is mainly rooted in the functional connectivity of specific neural circuits within gray-matter (GM); however, little is known regarding the neurofunctional substrates of individual differences in Gf in white-matter (WM)16,17,18. Recently, a small but increasing number of investigations have demonstrated a reliable detection of blood-oxygen-level-dependent functional magnetic resonance imaging (BOLD-fMRI) signals in WM. These studies indicate that neural activation elicits temporal and spectral profiles of hemodynamic responses in WM that are similar to those measured in GM during different functional tasks19,20,21,22,23,24. In parallel with the detection of task-related activations, BOLD-fMRI can also reflect the neural activity in WM at rest (i.e., absence of task requirement)25,26. Specifically, we found that, during the resting state, the power of low-frequency BOLD-fMRI fluctuations in WM exhibited a specific rather than a random distribution of noise26, and the WM functional connectome exhibited reliable and stable small-worldness and nonrandom modularity27. Abnormal small-worldness in the WM functional connectome was reported in patients with Parkinson’s disease28. Furthermore, the specific functional connectivity organization of the anatomical bundles was able to be identified by resting-state fMRI20,21,25,29. These investigations not only provided evidence of neural activity and connectivity, using fMRI, but also established cognitive biomarkers (i.e., for memory function) in WM30.
In the current study, using a cross-validation, data-driven analysis, we present novel findings that the whole-brain WM functional network predicted individual Gf and shed light on the possible neurofunctional correlates of Gf in WM. We first built a network-predicted model between connectivity strength and Gf scores of normal individuals following an initial examination (time 1, n = 326 participants). We demonstrated that the network-predicted model derived from these data could predict an individual’s Gf from his/her WM functional connectivity. This predictive model constructed at time 1 can be generalized to both the second (time 2, n = 105 participants) and the third examinations (time 3, n = 83 participants) using the overlapped individuals, thus, accurately predicting an individual’s Gf score from WM functional connectivity during the time 2 and time 3 scans for internal validation. Finally, to further test the generalizability of the predicted model, we showed that this model could also predict novel independent performance Gf (n = 53 participants) for external validation. These results suggested that the whole-brain functional connectivity of WM was a neuromarker of individual differences in Gf and would generalize to independent data to predict individual Gf.
Materials and methods
Participants
An overview of this study is shown in Fig. S1. Two independent cohorts were enrolled, namely, the internal and external validation groups. The interval validation group was further divided into the three subgroups. Internal validation I (time 1) was trained on time 1 data and also tested on time 1 data. Internal validation II (time 1 ∩ time 2) was trained on time 1 data (n − 1 participants) and tested on both time 1 and time 2 data (~11 months after time 1). Internal validation III (time 1 ∩ time 3) was trained on time 1 data (n − 1 participants) and tested on both time 1 and time 3 data (~29 months after time 1). All participants were normal college students from Southwest University of China, Chongqing, China. This study was approved by the Institutional Human Participants Review Board of the Southwest University Imaging Center. Written informed consent was obtained from all subjects. The data are available for research purposes through the International Data-sharing Initiative (http://fcon.1000.projects.nitrc.org/indi/retro/southwestuni/qiu/index.html). For detailed description about the participant information and data acquisition parameters, please see Liu et al.31.
For the external validation group, completely independent normal controls were recruited (see SI Materials and Methods for detailed information).
Assessment of fluid intelligence
For the internal validation group, the participants’ intellectual ability was assessed by the Combined Raven’s Test (CRT) (Chinese revised version) at time 132, which demonstrates a high degree of reliability and validity for intelligence testing32,33. This test gives an indication of the level of analogical thinking and abstract thought that a person has achieved; therefore, this test is known to be a good indicator of Gf5,11,34. The CRT scores (the number of correct answers given in 40 min) were used as a psychometric index of individual intelligence. In line with standard practice, the current study focused on the total score of the test4,34,35.
For the external validation group, the participants’ Gf abilities were assessed with performance general intelligence using the Chinese version of the Wechsler Adult Intelligence Scale (WAIS-RC). The Raven’s Test and WAIS-RC are clearly dominant in terms of interest, and are highly correlated between them in college students36.
Neuroimaging data acquisition
For the internal validation group, structural and functional MRI images were collected using a Siemens Trio 3.0T scanner (Siemens Medical, Erlangen, Germany) at the Southwest University China Center for Brain imaging31. The T1-weighted structural images (repetition time = 1900 ms, echo time = 2.52 ms, flip angle = 9°, field of view = 256 × 256 mm2, matrix size = 256 × 256, voxel size = 1 × 1 × 1 mm3, and slices = 176) were acquired. Subsequently, resting-state fMRI were acquired using a single-shot, gradient-recalled echo planar imaging sequence (repetition time = 2000 ms, echo time = 30 ms, flip angle = 90°, field of view = 220 × 220 mm2, matrix size = 64 × 64, voxel size = 3.4 × 3.4 × 3 mm3, and slices = 32). For each participant, a total of 242 functional volumes (484 s) were acquired. All participants were instructed to simply rest with their eyes closed, and not to think of anything in particular. For the external validation group, the parameters of data acquisition are list in “Materials and methods.”
Neuroimaging data preprocessing
Neuroimaging data were analyzed using the DPARSF (v4.3, www.restfmri.net) and SPM12 toolkits (www.fil.ion.ucl.ac.uk/spm/software/spm12). Slice-timing correction and realignment were applied to the remaining 235 functional images after excluding the first seven images. Structural images were then co-registered to the preprocessed functional images, and then segmented into GM, WM, and cerebrospinal fluid (CSF) by using DARTEL37. The mean signals from CSF (95% thresholded), 24 head motion parameters (six motion parameters, six temporal derivatives, and their respective squares) were regressed out from the data. To avoid elimination of important neural signals, we did not remove WM and brain global signals, as previous studies have suggested19,26,29.
To minimize mixing GM signals with WM signals, subsequent preprocessing for functional images was performed for WM alone. First, individual masks were obtained using a rigorous 90% threshold on the probability map of WM, which was produced by structural segmentation. Second, functional images were then spatially restricted into WM images using dot products between functional images and individual masks. Third, the WM functional images were then spatially normalized into the Montreal Neurologic Institute space by structural segmentation and were resampled into 3 × 3 × 3 mm3. Then, only voxels identified as WM across 80% of the participants were included into the group-level WM mask production. To exclude the impact of deep brain structures, the probability (25% threshold) Harvard-Oxford Atlas was used to remove subcortical nuclei (i.e., the bilateral thalamus, putamen, caudate, pallidum, and accumbens) from the group-level WM28,29,38. To minimize spurious local spatial correlations between voxels, spatial smoothing was not applied39,40. Subsequently, a band-pass filtering (0.01–0.10 Hz) was performed to minimize high-frequency physiological noise sources including the respiration rate. Finally, as functional connectivity is sensitive to the confounding factor of head motion, scrubbing was performed to reduce the negative influence41. If the framewise displacement (FD) exceeded 0.5 mm, the value of the signal at the point, as well as one forward point and two points previous to the signal point, were removed. Participants with 80% of their volumes remaining were included in further analyses.
Quality control
Exclusion criteria in selecting research participants are shown in Fig. S2. As head motion deteriorates the quality of fMRI data, we defined a set of standards to control for head motion. Specifically, if there was translational or rotational head movement > 2 mm or 2°, respectively, or head micromovements (mean FD) were >0.15 mm during resting-state fMRI scanning, these data were excluded. In addition, following scrubbing analyses, if a participant’s points on an image were <80%, their data were excluded. The quality of functional images (e.g., whether most of the temporal lobe was not visible) were then checked from the remaining participants. Quality control was applied to participants at time 1, time 2, and time 3. Finally, for each validation group, a prior outlier was defined as greater than mean + 2 standard deviation (SD) or less than mean − 2 SD for Gf scores, and were excluded. Finally, 326 participants (142 females, mean age for males and females = 20.03 years, SD = 0.072 years) were included in the internal validation I group (time 1); 105 participants were included in the internal validation II group (time 1 ∩ time 2); and 83 participants were included in the internal validation III group (time 1 ∩ time 3). The demographics for each time point/sample are shown in Table S1.
Construction of WM functional connectome
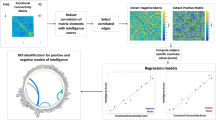
Nodes in WM functional networks were defined using group-wise voxel-based parcellation algorithms that produce roughly equal sizes within each node42. To obtain the 128 nodes used in this study, the parcellation algorithm was applied to the group-level WM mask from the remaining 326 participants at time 1. This parcellation scheme, which does not rely on a prior anatomical structure, may be better than the anatomical-based parcellation scheme, because of the uncertain match qualities between anatomical and functional images27, and is frequently used in GM mask parcellation40,43. In addition, the parcellation in the WM functional mask has been reported to have stable and reliable small-worldness27. For each participant, the WM functional connectivity matrix (128 × 128) was calculated by Pearson’s correlation between averaged BOLD signals of paired nodes. For the connectivity matrix, each cell represented the connection (i.e., edge) strength (Fisher’s r-to-Z normalized value) of each pair of nodes (step 1 in Fig. 1)44,45. Connectivity matrices were not thresholded or binarized46.
Step 1: We first obtained BOLD-fMRI signals in WM and then computed functional connectivity matrixes between each pair of nodes using Pearson’s correlation. Step 2: Each matrix values below or above the diagonal was reorganized to a row. Step 3: In each iteration, all rows were combined into a new matrix (m × n, where m represents the number of participants, and n represents the number of edges) across n − 1 participants (n is the number of participants). Step 4: Obtaining the correlation coefficients between each edge and an individual’s Gf abilities across n − 1 participants. Step 5: Significantly correlated edges (P < 0.01) with Gf was selected to construct features mask. These correlated edges were separated into a positive network (correlation coefficients were positive) or a negative network (correlation coefficients were negative). The dot product method was then applied between m × n matrix and feature masks. Step 6: Selected positive features and negative features were summed separately as two features into the predictive model (general linear model). Step 7: A general linear model was employed to construct the predictive model. Step 8: Test data matrixes were also performed using the dot product method with feature masks to obtain two features, which were added to the constructed predictive model to predict the Gf score. These steps were repeated N times, and Gf scores were obtained in all participants. Finally, correlation was calculated between observed and predicted Gf scores. BOLD-fMRI blood-oxygen-level-dependent functional magnetic resonance imaging, Gf general fluid intelligence.
Predictive model for internal validation
To investigate whether the strength of WM functional connectivity predicted individual Gf, we employed a leave-one-out cross-validation (LOOCV) method, which avoided overfitting by testing the strength of the relationship in an unseen participant1. Importantly, because head micromovements may cause spurious connectivity patterns, we first identified and excluded edges sensitive to head motion47. In each LOOCV, we randomly selected one participant as a testing set, and the remaining n − 1 participants were used as a training set (step 3 in Fig. 1)48. The predictive features were defined as the relevant edges to Gf scores at a significant threshold of P < 0.01 in the training set (n − 1 participants) (steps 4 and 5 in Fig. 1). The r values were then separated into a positive network (where r values were positive) and a negative network (where r values were negative)1,47,49. For each participant in the LOOCV, we summed the strength of edges from the WM functional connectivity matrix in the positive and negative sets, separately (step 6 in Fig. 1). Positive and negative sets have been reported to have different functional roles and might separately contribute to predictive power1,11,46. Hence, a simplified general linear model (GLM) protocol, which combined the positive and negative networks (step 7 in Fig. 1), was used to construct the training model in each LOOCV (the positive and negative models were also constructed for exploration). To predict the left-out participant’s (defined as testing data) Gf score, the training model was applied to the testing data, i.e., the strength of functional positive and negative networks (step 8 in Fig. 1). The flowchart of the predictive model is shown in Fig. 1. After repeating the abovementioned procedure n times, each participant’s Gf score was obtained. Finally, we computed the correlation between observed and predicted Gf scores using Pearson’s correlation to evaluate the power of the predictive model. K-fold cross-validation50 was also used to test the relationship between observed and predicted Gf scores. Participants were separated into 20-fold in line with previous work51, and ~16 participants were included in each fold. Similar to the LOOCV procedure, onefold was left out as a testing data, and the remaining 19-fold were used as training data to construct the predictive model in each cross-validation. After repeating this procedure 20 times, we obtained Gf scores. To minimize the random error on folds’ partition, we repeated the above predictive procedure 50 times52.
Permutation test
We first used parametric statistical analysis to obtain P values in the LOOCV procedure. However, the number of degrees of freedom are overestimated when LOOCV is performed within a single data set1,46. A permutation test is a commonly used statistical tool offering a simple way to compute the sampling distribution for any test statistic under the null hypothesis. We kept our WM functional connectivity matrices unchanged and randomly shuffled observed Gf scores 5000 times. The prediction procedure was then performed on each shuffled data. A distribution of the test statistic was obtained. The Ppermutation value was calculated by dividing shuffled times by the number that was greater or equal to the true prediction correlation. All P values estimated by the permutation test were under 0.05.
Because of nonoverlapping participants in the internal and external validation group, we evaluated the P value between the observed and predictive Gf scores using parametric statistical analysis only1.
Predictive model for external validation
To construct a predictive model of Gf to apply to a completely independent group, 326 participants at time 1 data were used to define consensus features53. Consensus features were defined as features that were selected in each LOOCV and were used to construct a predictive model of Gf. Next, the strength in the feature positions for each participant was computed in the external validation group. After performing LOOCV, we computed the correlations between observed and predicted Gf scores using Pearson’s correlation to evaluate the power of the predictive model.
Validation analysis of confounding factors
Validation analysis was performed to estimate the influence of confounding factors on predictive power. To better understand the variables in internal and external validation groups, the relationships between observed Gf scores and head motion, age, and sex were evaluated post-hoc using Pearson’s correlation analyses. To validate the specificity of the Gf predictive model, Pearson’s correlation analyses were also performed between age and head motion and predictive scores. In addition, the other influences of confounding factors on functional connectivity analysis was also considered in the Gf predictive model, including partial correlation analysis (controlling for age and sex), GM signals effect in data preprocessing, and group-level WM mask.
Results
Internal validation I: prediction from WM functional connectivity at time 1
We first verified that the observed Gf scores did not correlate with head micromovements (mean FD) during scanning (internal validation group, time 1: r(324) = −0.051, P = 0.358; time 2: r(103) = 0.095, P = 0.468; time 3: r(81) = 0.027, P = 0.812; external validation group: r(51) = −0.028, P = 0.841) and age (internal validation group, time 1: r(324) = −0.026, P = 0.635; time 2: r(103) = 0.126, P = 0.200; time 3: r(81) = 0.315, P = 0.112; external validation group: r(51) = 0.025, P = 0.861) (Fig. S3). We also confirmed no differences in the observed Gf scores between females and males using two tailed t-tests (internal validation group, time 1: t(324) = 1.337, P = 0.182; time 2: t(103) = 1.661, P = 0.100; time 3: t(81) = 1.004, P = 0.318; external validation group: t(51) = 1.124, P = 0.266) (Fig. S4).
After performing the predictive model analyses, we found that WM functional connectivity could be used as a feature to predict individual Gf (r(324) = 0.238, P = 1.44 × 10−5) (Fig. 2a). In addition, the Ppermutation was 0.004, reliably suggesting the significant correlation. The power of the predictive model was not affected by head motion, as the features sensitive to head motion were excluded and the predicted Gf scores were not correlated with mean FD values (r(324) = −0.082, P = 0.141) (Fig. S5A). As an observation of the predictive power of negative and positive networks, we also performed the correlation analysis between predicted and observed Gf values for the positive-feature and negative-feature models. The negative-feature model also generated significant prediction (r(324) = 0.160, Ppermutation = 0.009, Fig. 2b), and the positive-feature model exhibited marginal prediction (r(324) = 0.100, P = 0.071,, Fig. 2c). The predictions of positive-feature and negative-feature models were not significantly different (Steiger’s z value = 0.681, P = 0.516; Steiger’s z value was obtained from https://www.psychometrica.de/correlation.html), whereas the combined negative and positive features (GLM model) were more accurate than the prediction of the positive-feature model (Steiger’s z value = 2.84, P = 0.004), and marginally more accurate than the prediction of the negative-feature model (Steiger’s z value = 1.38, P = 0.084), suggesting, to some extent, that the negative and positive networks provided some complementary information. Thus, the GLM model was applied to further internal validations and external validation.
Results from a LOOCV comparing predicted and observed individuals’ Gf scores (n = 326). Scatter plots showed the predictions of the a GLM model (combined the strength of negative and positive features), b negative-feature model, and c positive-feature model at feature selection of P < 0.01 thresholded. Each dot represented each participant. The GLM model and negative-feature model showed significant predictions, and the positive-feature model was in marginal prediction. The negative-feature and positive-feature may provide nonoverlapped information on predicting individuals’ Gf abilities. WM white-matter, Gf general fluid intelligence, LOOCV leave-one-out cross-validation, GLM general linear model.
Furthermore, the correlation coefficient between observed and predicted Gf scores was 0.204 ± 0.013 (mean ± SD) in k-fold cross-validation. The significant correlation in the 20-fold cross-validation protocol indicated the rigorous presence of WM functional connectivity in an individual’s Gf abilities.
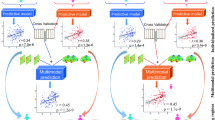
Internal validation II: prediction from WM functional connectivity at time 2
We next demonstrated that the constructed predictive model of Gf at time 1 data were generalized to time 2. To this end, we selected the participants who participated both in time 1 and time 2 scans (n = 105). We reconstructed the predictive model of Gf described in the above section “Internal validation I: prediction from WM functional connectivity at time 1” at time 1 (n = 105). As expected, models trained on 105 participants’ data also significantly predicted Gf values at time 1 (r(103) = 0.385, Ppermutation = 0.004) (Fig. 3a). To determine the generalization performance of the predictive model of Gf in the same group, we constructed the WM functional connectivity at time 2. We used the same procedure to predict an individual’s Gf scores as was done at time 1, except that the test data was from the WM functional connectivity matrix at time 2.
Scatter plots show correlations between observed and predicted Gf abilities. a, b WM functional network models were iteratively trained on time 1 data from n − 1 participants (n = 105) and tested on time 1 and time 2 data from the left-out individual. c, d Similarly, the WM functional network model was also iteratively trained on time 1 data from n − 1 participants (n = 83) and tested on time 1 and time 3 data from the left-out individual. There was no difference between the two powers of the predictive models from WM connectivity matrixes at time 1 and time 2 (Steiger’s z value = 1.773, P = 0.076) and time 1 and time 3 (Steiger’s z value = 1.163, P = 0.244). Gf general fluid intelligence, WM white-matter.
We demonstrated that the training model, based on WM functional connectivity at time 1, predicted an individual’s Gf scores from the WM connectivity matrix at time 2. The significant correlation parameters between the observed Gf scores and predicted Gf scores were r(103) = 0.204, Ppermutation = 0.018 (Fig. 3b). Although the correlation coefficient was numerically attenuated at time 2, the powers of the predictive model, from WM connectivity matrices at time 1 and time 2, showed no difference (Steiger’s z value = 1.773, P = 0.077). These results indicated, within WM functional connectivity, that there was a basis for neural correlates of individual Gf abilities and also suggested reliability of the overall model. This effect also could not be explained by head motion, as the predicted Gf scores from time 2 were not correlated with mean FD values (r(103) = −0.009, P = 0.930) (Fig. S5B).
Internal validation III: prediction from WM functional connectivity at time 3
To further explore whether the predictive model was still stable after a longer fMRI scanning (~29 months interval), we reconstructed the predictive model of Gf based on 83 participants’ fMRI data at time 1 (time 1 ∩ time 3: n = 83). The strength of WM functional connectivity also significantly predicted Gf values (r(81) = 0.363, Ppermutation = 0.012, Fig. 3c). This trained model was applied to the WM connectivity matrix at time 3. Significant correlations emerged between predicted and observed Gf scores (r(81) = 0.217, Ppermutation = 0.020, Fig. 3d). Although the correlation coefficient between predicted and observed Gf values at time 3 was numerically smaller than at time 1, the predictive power showed no difference (Steiger’s z value = 1.163, P = 0.244). The predictions further indicated the reliable neuromarker of Gf from WM functional connectivity. The power of this predictive model was not affected by head motion, thus excluding the head motion confounding factor (time 3: r(81) = −0.019, P = 0.866) (Fig. S5C). In addition, we have also constructed a full correlation matrix across times 1, 2, and 3 for observed and predicted Gf scores. We also found that the observed Gf score correlated with predicted Gf scores in GLM (Table S2). Collectively, these results suggested that although, to a certain degree, brain functional connectivity patterns were metastable or unstable during short or long-time development, the Gf predictive model from WM functional connectivity was relatively stable, and did not change across longitudinal scanning times (time 1, time 2, and time 3).
External validation: prediction at an independent sample from another center
In order to generalize the predictive model of Gf to a completely independent data set, we entered positive and negative values into the predictive model to predict individual Gf scores. The observed Gf scores were marginally correlated with predicted Gf scores in the independent sample group (r(51) = 0.276, P = 0.045, Fig. 4). The correlation coefficient (r) was relatively small, but this value was typically between 0.2 and 0.5, according to the previous studies1,54. Prediction results will often have lower within-sample effect sized than results generated with simple correlation analysis1. Similarly, predicted Gf scores were not correlated with mean FD values, ruling out the head motion confounding factor (r(51) = −0.169, P = 0.228, Fig. S5D).
Validations of the effects of confounding factors
To validate the specificity of the Gf predictive model, we investigated the correlation between predicted Gf scores and confounding factors. There were no significant correlations between predicted Gf scores and age in internal and external validation groups (time 1: r(324) = −0.008, P = 0.884; time 2: r(103) = 0.086, P = 0.384; time 3: r(81) = −0.086, P = 0.441; independent sample group: r(51) = −0.116, P = 0.409) (Fig. S5E–H). Otherwise, we performed partial correlation analyses controlling for age and sex in the Gf predictive model1,46,54, which resulted in significant correlations between observed and predicted Gf scores at time 1 (r(324) = 0.211, P = 1 × 10−4, Fig. S6). To further examine the GM signals effect on our results27, we regressed out the global brain signals (including GM, CSF, and WM signals) in the neuroimaging data preprocessing and maintained all other processes. We observed that the predicted Gf scores were also correlated with the observed Gf scores (r(324) = 0.217, P < 1 × 10−4, Fig. S7). Finally, we created a new WM mask across all participants to validate the parcellation effect on our results. We used time 1 data as the mask validation sample, and found that the predictive power still remained at time 1 data (r(324) = 0.207, P = 2 × 10−4, Fig. S8). The two predictive models based on different masks showed no difference on predictive power (Steiger’s z value = 0.581, P = 0.561). These results suggested that the Gf predictive model based on WM functional connectivity could identify relevant Gf attributes rather than other confounding factors.
Functional locations of consensus features on WM connectome
To illustrate the biological substrates underlying the neural correlates of individual Gf and WM functional connectivity, we described the most specific edges (i.e., consensus features) contributing to the predictive model1,53. We first divided 128 nodes into 11 specific WM networks (considered as mask) based on a previous study on human WM functional networks (Fig. 5a). We then examined the within- and between-WM functional connectivity consensus features. Although these features were generated from data-driven analyses that were not dependent on prior acknowledge, the features exhibited specific distributions, and were primarily located between the superior longitudinal fasciculus and deep frontal WM and ventral frontoparietal tracts; the number of connections is shown in Fig. 5b. The involvement of the superior longitudinal fasciculus in WM functional networks suggested that the connections did not provide structural information but provided functional information corresponding to Gf scores6,13.
a The location of consensus features. We divided the 128-ROI in WM into 11 WM functional networks. Macroscale regions were mainly located in the superior longitudinal fasciculus, deep frontal white-matter, and ventral frontoparietal tracts. The gray column in each node represented the degree of this node. b The differences in the number of edges in the WM functional networks. Gf fluid intelligence, ROI region of interest; WM white-matter.
Discussion
We showed that the supplementary neuromarker-WM functional connectivity was both reliable and generalizable to establish brain–behavior relationships. We demonstrated that the WM functional profiles could be used as features to predict the fundamental cognitive traits of individuals’ Gf abilities. We used a cross-validation, data-driven analysis to identify the connectivity strength of WM functional networks in predicting Gf. This predictive model was generalizable to predict individual Gf abilities in the same group at the time of scanning (time 1), 11 months later (time 2), and 29 months later (time 3), providing evidence for stable Gf features from WM functional connectivity. To demonstrate more generalization and robustness of the predictive model, we used a completely independent sample group as a test cohort. The features that predicted individual Gf abilities at time 1 (n = 326 participants) also predicted Gf scores in an independent sample. These results that the predictive model from WM functional networks predicted an individual’s Gf abilities in disparate groups and with difference assessment scales, underscored the potential to investigate the neuromarkers of cognitive trait from whole-brain WM functional connectivity.
Our WM functional connectome-based predictive model, which was generalized to a completely independent cohort, provides a new insight into investigating neuromarkers related to Gf abilities. It is noteworthy that this predictive power accounts for ~7.62% of the explained variance. According to previous protocols of connectome-based predictive modeling1, this correlation will typically be between 0.2 and 0.5; because, the predictive power will often have lower within-sample effect sizes than results generated with simple correlation analysis. In addition, the cross independent data validation could provide a more conservative estimate of the strength of the brain–behavior relationship and is more likely to generalize independent data1. The model showed that human intelligence was unlikely supported by one brain area or one system, but rather was related to several cognitive components (such as the cognitive control network)55. This powerful model was successfully applied to illustrate the relationships between functional connectivity in GM and Gf scores11 and sustained attention46. But these predictive models did not focus on WM functional connectivity in part because of the longstanding controversy regarding the functional role of WM BOLD-fMRI signals.
The first debate relates to the way in which WM, relative to GM, contains lower cerebral blood flow and volume. Cerebral blood flow and volumes are the basis of the BOLD signal in neural activity56,57,58. Another debate centers on how BOLD signals recognize local field action potentials in GM, but not in WM16,59. However, in comparison with GM, and regardless of large discrepancies in respect to the physiological factors observed between GM and WM, WM maintains a higher ratio of glial cells to neurons60, and has an approximately equal oxygen extraction fraction61. Many studies identifying WM signals in resting-state fMRI have been reported16,25,26,29, suggesting that there are no fundamental barriers or direct sources of evidence against the possibility of detecting WM neural activities using BOLD-fMRI16. Furthermore, our previous study has proven that several confounding factors (i.e., CSF signals, global brain signals, and physical distance) did not influence the small-worldness property of WM functional networks27.
In addition to showing that the WM connectivity-based predictive model was able to predict Gf abilities, the current results also suggested that the distributed networks relevant to Gf support partial classical P-FIT characterizing human intelligence. P-FIT describes the interactions of distributed networks in frontal and parietal regions in order to predict the differences in an individual’s Gf abilities13. The P-FIT model summarizes five steps for information processing of Gf13. First, salient information is mainly collected by auditory and visual methods, thus, occipital and temporal regions are critical to sensory information processing. Second, parietal regions mainly in the supramarginal, angular, and superior parietal gyrus extract perceptual information. Third, the assumption of interactions between parietal and frontal regions is used to test the hypotheses. Next, the anterior cingulate cortex is used to select and inhibit responses. Finally, the speedy and error-free information processing from the parietal to frontal regions is dependent on WM structures, such as the superior longitudinal fasciculus6. Our results have added to the P-FIT model in that functional information in the superior longitudinal fasciculus system is a basis for neural correlates with Gf. Other regions related to Gf mainly included the visual superficial WM system; deep frontal WM (including the cingulum); the ventral frontoparietal tracts; and the forceps minor system29. Several of these WM functional systems are correlated with GM networks, such as the visual superficial WM system, which itself is highly correlated with visual GM networks. Our finding regarding a direct relationship between these WM functional networks and Gf is in agreement with steps 1–4 in P-FIT information processing. However, there were other WM functional networks, including the sensorimotor and ventral and dorsal attention GM networks, that showed high correlation with Gf, which were not included in the P-FIT model. Thus, our results demonstrated the importance of whole-brain imaging to investigate an individual’s Gf abilities.
The specific distributions of relevant edges associated with Gf may indicated that BOLD-fMRI signals in WM were nonartifactual and nonrandom noises. Otherwise, the excluded possibility of observed WM functional connectivity from random noise can be validated by the fact that the predictive model is generalized to an independent data set. In this study, we also applied several methods to ensure that WM BOLD-fMRI signals were not affected by GM signals, by strictly controlling the boundary between WM and GM (i.e., we employed a 90% threshold on the WM probability map); by separating WM and GM functional images in preprocessing26,29; and by identifying participants’ voxels only in WM to create WM masks. Moreover, from the architecture of the brain venous systems, the possibility that deoxygenated blood was drained from cortical GM to deep WM was rather small. In fact, there are two venous systems in normal neuroanatomy: one is the superficial venous system, which drains deoxygenated blood in GM and superficial WM in the cortex into the pial veins; the other is deep system draining deoxygenated blood in deep WM into the subependymal veins19,62. The venous architecture does not overlap in brain regions. Deoxygenated blood drainage from GM cortex to the deep venous system through the WM does exist, but the probability of draining is less than 3%19,62. Collectively, these methods, and the brain venous system architecture, ensured that BOLD-fMRI signals analyzed herein were in fact from WM.
Although our findings provided a novel WM connectivity-based predictive model to predict an individual’s Gf abilities, several limitations in this study remain. First, the sample in this study was comprised of college students, limiting the application of WM functional connectome-based predictive model for other populations. Future studies should employ a wider population base to investigate the relationship between Gf and WM functional connectomes. Second, individuals’ Gf abilities were assessed by the CRT in this study. Although the completely independent sample from another center was assessed by WAIS-RC, given the various types of psychometric tests for human intelligence, future studies should further test the specificity and generalization of the current predictive model using a wide variety of intelligence measures to detect the consensus neuromarker of intelligence in WM. Third, the current work used WM functional connectivity as a neuromarker, ignoring the causality connectivity (effective connectivity) among the brain regions. Future effective connectivity studies on WM functions are therefore needed. Finally, the physiological basis of BOLD-fMRI signals in WM remains unknown. However, a previous study suggested that the impact of artifacts (such as physiological noise and head motion) in WM activity is relatively small16,26. In this study, the possible effects of artifacts were strictly controlled by exclusion of excessive head motion, regression of CBF signals, regression of global brain signals, and exclusive preprocessing of WM. Future neurophysiological studies on BOLD-fMRI signals in WM are needed to more accurately interpret the current predictive model.
In conclusion, we demonstrated that functional connectivity in WM was a neuromarker to predict an individual’s Gf abilities and that the distributed networks supported the P-FIT model. Beyond the current findings, the predictive model from whole-brain functional connectivity in WM can be used to investigate other cognitive abilities and clinical symptoms in psychiatric diseases.
Code availability
The code of this predictive model was provided by Shen et al. at https://www.nitrc.org/projects/bioimagesuite/.
References
Shen, X. et al. Using connectome-based predictive modeling to predict individual behavior from brain connectivity. Nat. Protoc. 12, 506–518 (2017).
Barbey, A. K. Network neuroscience theory of human intelligence. Trends Cogn. Sci. 22, 8–20 (2018).
Deary, I. J. Intelligence. Annu. Rev. Psychol. 63, 453–482 (2012).
Jaeggi, S. M., Buschkuehl, M., Jonides, J. & Perrig, W. J. Improving fluid intelligence with training on working memory. Proc. Natl Acad. Sci. USA 105, 6829–6833 (2008).
Haier, R. What does a smart brain look like? Sci. Am. Mind 23, 26–33 (2009).
Penke, L. et al. Brain white matter tract integrity as a neural foundation for general intelligence. Mol. Psychiatry 17, 1026–1030 (2012).
Pietschnig, J. et al. Meta-analysis of associations between human brain volume and intelligence differences: how strong are they and what do they mean? Neurosci. Biobehav. Rev. 57, 411–432 (2015).
Roth, G. & Dicke, U. Evolution of the brain and intelligence. Trends Cogn. Sci. 9, 250–257 (2005).
Song, M. et al. Brain spontaneous functional connectivity and intelligence. Neuroimage 41, 1168–1176 (2008).
Basten., U., Hilger, K. & Fiebach, C. J. Where smart brains are different: a quantitative meta-analysis of functional and structural brain imaging studies on intelligence. Intelligence 51, 10–27 (2015).
Finn, E. S. et al. Functional connectome fingerprinting: identifying individuals using patterns of brain connectivity. Nat. Neurosci. 18, 1664–1671 (2015).
Dubois, J., Galdi, P., Paul, L. K. & Adolphs, R. A distributed brain network predicts general intelligence from resting-state human neuroimaging data. Philos. Trans. R. Soc. Lond. B Biol. Sci. 373, 20170284 (2018).
Jung, R. E. & Haier, R. J. The parieto-frontal integration theory (P-FIT) of intelligence: converging neuroimaging evidence. Behav. Brain. Sci. 30, 135–154 (2007). Discussion 154-187.
Santarnecchi., E. et al. Network connectivity correlates of variability in fluid intelligence performance. Intelligence 65, 35–47 (2017).
Ferguson, M. A., Anderson, J. S. & Spreng, R. N. Fluid and flexible minds: Intelligence reflects synchrony in the brain’s intrinsic network architecture. Netw. Neurosci. 1, 192–207 (2017).
Gawryluk, J. R., Mazerolle, E. L. & D’Arcy, R. C. Does functional MRI detect activation in white matter? A review of emerging evidence, issues, and future directions. Front. Neurosci. 8, 239 (2014).
Courtemanche, M. J. et al. Detecting white matter activity using conventional 3 Tesla fMRI: an evaluation of standard field strength and hemodynamic response function. Neuroimage 169, 145–150 (2018).
Gawryluk, J. R., Mazerolle, E. L., Beyea, S. D. & D’Arcy, R. C. Functional MRI activation in white matter during the Symbol Digit Modalities Test. Front. Hum. Neurosci. 8, 589 (2014).
Huang, Y. et al. Voxel-wise detection of functional networks in white matter. Neuroimage 183, 544–552 (2018).
Marussich, L., Lu, K. H., Wen, H. & Liu, Z. Mapping white-matter functional organization at rest and during naturalistic visual perception. Neuroimage 146, 1128–1141 (2017).
Wu, X. et al. Functional connectivity and activity of white matter in somatosensory pathways under tactile stimulations. Neuroimage 152, 371–380 (2017).
Ding, Z. et al. Spatio-temporal correlation tensors reveal functional structure in human brain. PLoS ONE 8, e82107 (2013).
Ding, Z. et al. Visualizing functional pathways in the human brain using correlation tensors and magnetic resonance imaging. Magn. Reson. Imaging 34, 8–17 (2016).
Li, M. et al. Characterization of the hemodynamic response function in white matter tracts for event-related fMRI. Nat. Commun. 10, 1140 (2019).
Ding, Z. et al. Detection of synchronous brain activity in white matter tracts at rest and under functional loading. Proc. Natl Acad. Sci. USA 115, 595–600 (2018).
Ji, G. J. et al. Low-frequency blood oxygen level-dependent fluctuations in the brain white matter: more than just noise. Sci. Bull. 62, 656–657 (2017).
Li, J. et al. Exploring the functional connectome in white matter. Hum. Brain Mapp. 40, 4331–4344 (2019).
Ji, G. J. et al. Regional and network properties of white matter function in Parkinson’s disease. Hum. Brain Mapp. 40, 1253–1263 (2019).
Peer, M. et al. Evidence for functional networks within the human brain’s white matter. J. Neurosci. 37, 6394–6407 (2017).
Makedonov, I., Chen, J. J., Masellis, M. & MacIntosh, B. J. Physiological fluctuations in white matter are increased in Alzheimer’s disease and correlate with neuroimaging and cognitive biomarkers. Neurobiol. Aging 37, 12–18 (2016).
Liu, W. et al. Longitudinal test-retest neuroimaging data from healthy young adults in southwest China. Sci. Data 4, 170017 (2017).
Li., D. et al. The testing results report on the Combined Raven’s Test in Shanghai. Psychol. Sci. 4, 27–31 (1988).
Wang., D. et al. Revision on the combined Raven’s Test for the rural in China. Psychol. Sci. 5, 23–27 (1989).
Sun, J. et al. Verbal creativity correlates with the temporal variability of brain networks during the resting state. Cereb. Cortex 29, 1047–1058 (2018).
Takeuchi, H. et al. White matter structures associated with creativity: evidence from diffusion tensor imaging. Neuroimage 51, 11–18 (2010).
Paul, S. M. The advanced Raven’s progressive matrices. J. Exp. Educ. 54, 95–100 (1986).
Ashburner, J. A fast diffeomorphic image registration algorithm. Neuroimage 38, 95–113 (2007).
Desikan, R. S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31, 968–980 (2006).
Ji, G. J. et al. Decreased network efficiency in benign epilepsy with centrotemporal spikes. Radiology 283, 186–194 (2017).
Zhang, Z. et al. Altered functional-structural coupling of large-scale brain networks in idiopathic generalized epilepsy. Brain 134, 2912–2928 (2011).
Power, J. D. et al. Spurious but systematic correlations in functional connectivity MRI networks arise from subject motion. Neuroimage 59, 2142–2154 (2012).
Zalesky, A. et al. Connectivity differences in brain networks. Neuroimage 60, 1055–1062 (2012).
Wang, J. et al. Disrupted functional brain connectome in individuals at risk for Alzheimer’s disease. Biol. Psychiatry 73, 472–481 (2013).
Wang, J. H. et al. Graph theoretical analysis of functional brain networks: test-retest evaluation on short- and long-term resting-state functional MRI data. PLoS ONE 6, e21976 (2011).
Achard, S. et al. Hubs of brain functional networks are radically reorganized in comatose patients. Proc. Natl Acad. Sci. USA 109, 20608–20613 (2012).
Rosenberg, M. D. et al. A neuromarker of sustained attention from whole-brain functional connectivity. Nat. Neurosci. 19, 165–171 (2016).
Wang, D. et al. Individual-specific functional connectivity markers track dimensional and categorical features of psychotic illness. Mol. Psychiatry https://doi.org/10.1038/s41380-41018-40276-41381 (2018).
Li, J. et al. More than just statics: temporal dynamics of intrinsic brain activity predicts the suicidal ideation in depressed patients. Psychol. Med. 49, 852–860 (2019).
Beaty, R. E. et al. Robust prediction of individual creative ability from brain functional connectivity. Proc. Natl Acad. Sci. USA 115, 1087–1092 (2018).
Poldrack, R. A., Huckins, G. & Varoquaux, G. Establishment of best practices for evidence for prediction: a review. JAMA Psychiatry https://doi.org/10.1001/jamapsychiatry.2019.3671 (2019).
Kashyap, R. et al. Individual-specific fMRI-subspaces improve functional connectivity prediction of behavior. Neuroimage 189, 804–812 (2019).
Liao, W. et al. Static and dynamic connectomics differentiate between depressed patients with and without suicidal ideation. Hum. Brain Mapp. 39, 4105–4118 (2018).
Dosenbach, N. U. et al. Prediction of individual brain maturity using fMRI. Science 329, 1358–1361 (2010).
Scheinost, D. et al. Ten simple rules for predictive modeling of individual differences in neuroimaging. Neuroimage 193, 35–45 (2019).
Hampshire, A., Highfield, R. R., Parkin, B. L. & Owen, A. M. Fractionating human intelligence. Neuron 76, 1225–1237 (2012).
Rostrup, E. et al. Regional differences in the CBF and BOLD responses to hypercapnia: a combined PET and fMRI study. Neuroimage 11, 87–97 (2000).
Preibisch, C. & Haase, A. Perfusion imaging using spin-labeling methods: contrast-to-noise comparison in functional MRI applications. Magn. Reson. Med. 46, 172–182 (2001).
Helenius, J. et al. Cerebral hemodynamics in a healthy population measured by dynamic susceptibility contrast MR imaging. Acta Radiol. 44, 538–546 (2003).
Logothetis, N. K. et al. Neurophysiological investigation of the basis of the fMRI signal. Nature 412, 150–157 (2001).
Azevedo, F. A. et al. Equal numbers of neuronal and nonneuronal cells make the human brain an isometrically scaled-up primate brain. J. Comp. Neurol. 513, 532–541 (2009).
Raichle, M. E. et al. A default mode of brain function. Proc. Natl Acad. Sci. USA 98, 676–682 (2001).
Ruiz, D. S., Yilmaz, H. & Gailloud, P. Cerebral developmental venous anomalies: current concepts. Ann. Neurol. 66, 271–283 (2009).
Acknowledgements
We are grateful to all the participants in this study. We thank International Science Editing (http://www.internationalscienceediting.com) for editing this manuscript. We thank Dr Alex Fornito (Turner Institute for Brain and Mental Health, Monash University) for his assistance in Fig. 5a. This work was supported by the National Key Project of Research and Development of Ministry of Science and Technology (2018AAA0100705), the National Natural Science Foundation of China (61871077, 61533006, U1808204, 81771919, and 61673089), and Sichuan Science and Technology Program (2018TJPT0016).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Li, J., Biswal, B.B., Meng, Y. et al. A neuromarker of individual general fluid intelligence from the white-matter functional connectome. Transl Psychiatry 10, 147 (2020). https://doi.org/10.1038/s41398-020-0829-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-020-0829-3
- Springer Nature Limited
This article is cited by
-
White matter dysfunction in psychiatric disorders is associated with neurotransmitter and genetic profiles
Nature Mental Health (2023)
-
Longitudinal development of the human white matter structural connectome and its association with brain transcriptomic and cellular architecture
Communications Biology (2023)
-
The challenges and prospects of brain-based prediction of behaviour
Nature Human Behaviour (2023)
-
Less reduced gray matter volume in the subregions of superior temporal gyrus predicts better treatment efficacy in drug-naive, first-episode schizophrenia
Brain Imaging and Behavior (2021)
-
Abnormal connectivity of anterior-insular subdivisions and relationship with somatic symptom in depressive patients
Brain Imaging and Behavior (2021)