Abstract
The essential G1-cyclin, CCND1, is frequently overexpressed in cancer, contributing to tumorigenesis by driving cell-cycle progression. D-type cyclins are rate-limiting regulators of G1-S progression in mammalian cells via their ability to bind and activate CDK4 and CDK6. In addition, cyclin D1 conveys kinase-independent transcriptional functions of cyclin D1. Here we report that cyclin D1 associates with H2BS14 via an intrinsically disordered domain (IDD). The same region of cyclin D1 was necessary for the induction of aneuploidy, induction of the DNA damage response, cyclin D1-mediated recruitment into chromatin, and CIN gene transcription. In response to DNA damage H2BS14 phosphorylation occurs, resulting in co-localization with γH2AX in DNA damage foci. Cyclin D1 ChIP seq and γH2AX ChIP seq revealed ~14% overlap. As the cyclin D1 IDD functioned independently of the CDK activity to drive CIN, the IDD domain may provide a rationale new target to complement CDK-extinction strategies.
Similar content being viewed by others
Introduction
Genome instability is a hallmark of cancer cells [1]. Chromosome instability (CIN) is often mutually exclusive from hypermutation genotypes and represents a distinct subtype of genome instability. CIN is an early event in tumor formation that causes either the inactivation of tumor suppressor genes, through defects in genome maintenance and destabilization of the nucleotide sequence or accelerates the gain of oncogenic loci. Aneuploidy, also known as somatic copy number alterations (SCNAs), is widespread in human cancers [2,3,4,5,6]. For most tumors, there is a positive correlation between SCNA levels and the total number of mutations. Although CIN is likely due to defects in a network of genes that regulate mitotic checkpoints, collaborative oncogenes, including c-Myc [7] or cyclin D1 [8,9,10] are sufficient for the induction of CIN. The induction of cellular growth and CIN by c-Myc involves dissociable domains [7], however, the domain of cyclin D1 required for the induction of CIN is unknown.
Cyclin D1 overexpression occurs in human breast, prostate, lung, and gastrointestinal malignancies and its abundance is induced at the level of transcription, translation and post-translational modifications [11, 12]. Phosphorylation and inactivation of the retinoblastoma (RB), the nuclear respiratory factor 1 (NRF1), and several other proteins by the cyclin D1/CDK4/6 complex is known to govern nuclear DNA synthesis and mitochondrial biogenesis [13,14,15,16]. Oncogenic signals induce cyclin D1 through multiple processes, including transcription, alterations in protein stability and miRNA which bind and regulate cyclin D1 via the 3’-UTR [17, 18]. In murine models, cyclin D1 was required for the growth of ErbB2-induced mammary tumors [19]. The murine cyclin D1 gene was shown to be required for Ras- or ErbB2-induced mammary tumorigenesis and APC-induced gastrointestinal tumorigenesis in cyclin D1 gene deletion mice [20, 21]. Although selective inhibitors of cyclin D1/CDK4/6 complexes have transitioned effectively to the clinic improving progression-free survival, therapy resistance subsequently arises [17, 22].
In addition to its well-documented function as the regulatory subunit of an RB kinase, cyclin D1 conducts important non-canonical activities, including the induction of cell migration [23], coordination of gene transcription [12, 14], the induction of chromosomal instability (CIN), and the induction of DNA damage repair (DDR) [24,25,26]. The DDR is associated with the induction of H2AX (Ser139P) and H2B (Ser14P) phosphorylation to establish a heterochromatin state which concentrates DNA repair factors [27]. Similar to several other proteins participating directly in the DDR [28], cyclin D1 can be recruited into chromatin and augments the DDR [25]. Cyclin D1 also regulates E2F-independent transcription through poorly understood mechanisms involving altering transcription factor (TF) recruitment into chromatin [20], binding in chromatin to induce gene transcription [29, 30], confirmed by genome-wide analysis [8, 31]. Genome-wide ChIP-seq studies identified regions of genes to which cyclin D1 was significantly recruited in chromatin with a preference for CTCF binding sites [8, 32], with similar levels of cyclin D1 enrichment found in other studies [8, 9, 31,32,33]. In vivo studies established that endogenous cyclin D1 is required for a substantial proportion of androgen-mediated gene expression in the prostate and estrogen-responsive gene expression in the mammary gland [34, 35]. Acute activation of a cyclin D1 cDNA, in either tissue culture or the murine mammary gland, induced gene expression within functional pathways governing, cell-cycle control, mitochondrial biogenesis, and the induction of chromosomal instability (CIN) and CIN gene expression [8]. The recruitment of cyclin D1 into chromatin at target gene promoters has been shown in fibroblasts [8, 9], neural [32] and lymphoma cells [33]. Furthermore, other components of the cell-cycle control apparatus have been identified in chromatin including CDKs and p27/p21 [36].
The molecular mechanisms by which a diverse array of genes may be coordinately regulated by cyclin D1 and the mechanisms by which cyclin D1 is recruited into chromatin to regulate gene transcription is unknown. Cyclin D1 does not contain transcriptional co-regulator domains which recognize histone motifs that have been modified by acetylation, phosphorylation, methylation, or citrullination, PHD (plant homeodomain) fingers and BAH (bromo-adjacent homology) domains, “Royal Family,” ADD (ATRX-DNMT3A-DNMT3L) domains or 14-3-3 histone interaction residues. In addition to reader domains, intrinsically disordered regions (IDR) govern protein-protein interactions. IDRs, constitute a substantial fraction of the eukaryotic proteome [37, 38]. Due to their large accessible surface area and inherent flexibility, IDRs are often involved in molecular recognition and protein-protein interaction events [39]. Intrinsically disordered proteins (IDPs) or regions of proteins (IDRs) abound in proteins with signaling functions and are found in TFs and proteins with chromatin-organizing function [40]. IDRs often mediate protein-protein interactions, undergoing induced folding that uncouples specificity from binding strength [41] and mediates distinct functions [42]. Furthermore, specific protein-protein interaction can occur even without full folding (“fuzzy” interactions [43]), as occurs in liquid-liquid phase separation [44]. Even in these cases, the interaction can carry a significant element of specificity, likely arising in an ensemble-encoded, “emergent” way [45]. IDPs/IDRs are frequently involved in dynamic interactions in signaling networks where fine-tuning of the cascade of interactions is exerted through multisite dependence. The current studies identified the cyclin D1 IDR as required for both DNA damage signaling and transcription, serving as a chromatin interaction interface binding H2BSer14P.
Results
The cyclin D1 carboxyl terminal region is required to selectively associate with Histone H2BSer14P
Prior studies have shown that cyclin D1 is recruited to sites of DNA damage with γH2AX [24,25,26]. Such sites are known to co-localize with H2BSer14P [27]. We, therefore, sort to determine whether cyclin D1 is directly associated with H2B or H2BSer14. GST pull-down experiments conducted with cyclin D1 and synthetic modified histone peptides showed cyclin D1 binds H2BS14P with increased affinity compared with GST alone and binds with higher affinity to a peptide encoding H2BS14P (Fig. 1C, D). Although cyclin D1 does not encode known histone protein interaction motifs (PHD fingers, Bromodomain tandem, Chromodomain, Tudor, MET, PWWP, WD40, YEATS, and the β propeller motifs), a glutamate-rich region (poly E region, aa 272-280), which have been identified in IDD, was considered as a candidate motif (Fig. 1A). A GST protein encoding a deletion of the glutamate rich region (cyclin D1ΔE) showed substantially decreased binding to the H2BS14P peptide (Fig. 1C, D). GST control had no signal (Fig. 1C, D). Consistent with prior studies, the GST-KDM4A fusion protein bound to an H3K4me3 peptide (Fig. 1E). We further investigated the strength of the interaction between cyclin D1 and modified histones using Surface Plasmon Resonance (SPR). This approach showed a strong and specific binding between cyclin D1 and H2BS14P when compared to GST control with significant binding to the H2BS14P peptide (Fig. 1F, G). The interaction with the H2BS14P peptide was abolished when the cyclin D1 ΔE mutant was introduced (Fig. 1G).
A Coomassie staining and (B). Western blot for cyclin D1 (CD1WT), cyclin D1 E-region deleted (CD1ΔE) and KDM4A GST fusion proteins. C GST pull-down, with (D). quantitation shown as mean ± SEM for multiplicate experiments for GST-CD1WT, GST-CD1ΔE or GST-ctrl fusion proteins with histone peptide (H2B) and S14-phosphorylated histone peptide (H2BS14P). In comparison, GST pull-down of histone H3K4me3 with KDM4A is shown in (E). F Schematic representation of GST-cyclin D1 wild type and ΔE mutant. G Surface plasmon resonance (SPR) analysis of histone peptide (H2BS14P) with GST-cyclin D1, GST-ΔE or GST-ctrl fusion proteins. (GST-Cyclin D1: blue:10 μM, light blue:7 μM, magenta:5 μM, red:3.5 μM, orange:1 μM).
A MST analysis with H2B (A, C) and H2BS14P (B, D), indicating Kd, or the lack of binding (Kd cannot be determined, n.d.). E Human cyclin D1 (CD1) (Alpha fold – AF-P24385-F1) is shown in electrostatic surface and H2B (PDB ID: 2RVQ) [102] is shown in the sticks (green or magenta). Cyclin D1 contains a large negatively charged cavity on its surface which is ideal for binding to the highly positively charged tail of H2B. It is worth noting that this pocket is formed by helices α2, α 6, and α9 and is away from the CDK4/6 interacting site of cyclin D1. F 180° rotation of the structure in (E). G Cartoon representation of cyclin D1-H2B structural model. Helices α2, α6, and α9 that form the H2B binding pocket of cyclin D1 are shown, as well as the pS14 and the glutamates that form the E-rich region. H Zoom in on the representation of the interaction between cyclin D1 and H2B same orientation as in (H). Cyclin D1 is shown in wheat color. The E-region which binds S14P is shown in blue. The phosphoserine S14P is shown in magenta on the green sticks of the histone peptide.
We further investigated the nature of the interaction between cyclin D1 and modified histones using microscale thermophoresis (MST). MST showed a strong and specific binding between cyclin D1 and H2BS14P when compared to the binding of unmodified H2B (Kd = 2.4 μM vs. 64 μM) (Fig. 2A, B). The interaction with both H2B and H2BS14P peptide was abolished when the cyclin D1 ΔE mutant was introduced (Fig. 2C, D).
We conducted a structural analysis to define the mechanism by which cyclin D1 interacts with H2B. Because the available X-ray and cryo-EM structures of human cyclin D1 lack the E-region we used the AlphaFold generated human cyclin D1 (AF-P25322-F1) structural model [46] with the NMR H2B coordinates PDB ID: 2VQ and deployed the CABS-dock server. Our approach allowed us to independently identify the cyclin D1 binding site of the unstructured tail of histone H2B residues containing S14P. Since the length of the histone tail that interacts with cyclin D1 is unknown, we performed docking using two different length peptides of the H2B unstructured tail, both of which contain S14P. The CABS-dock server placed either of the 2 longer peptides in precisely the same location on the surface of cyclin D1 (Fig. 2E, F). The cyclin D1 binding pocket predicted for H2B is formed by helices α2, α6, and α9, located away from the CDK4/6 interacting site, spanning the entire length of the cyclin D1 molecule (Fig. 2G). The pocket is highly negatively charged, ideal for binding to the positively charged H2B tail. The cyclin D1 residues E74, E76, and E162 are predicted to be likely involved in binding to the K25, R26, and R30 of the H2B C-terminal part of the tail. The cyclin D1 residues E66 and E69 most likely engage K17 and K21, which form part of the center of the H2B tail. The N-terminal portion of the H2B tail forms a hairpin-like structure with Ser14P at the tip of this hairpin. The H2B tail hairpin inserts itself into a cavity formed by helix α9, the E-region, and the following connecting loop (Fig. 2G). In this cavity, H2B residues K8, K9, and K13 most likely engage cyclin D1 residues Q261, Q264, and D282. These interactions place S14P directly in coordinating distance from the E-region glutamates E274 and E278 (Fig. 2H).
Combinatorial modifications of histones govern cyclin D1 binding
To assess the binding of cyclin D1 to post-translationally modified histones in an unbiased manner, the cyclin D1 and KDM4A fusion proteins were assessed for interaction with histone arrays. The arrays consist of 384 unique histone modification combinations in duplicate, including up to four different modifications on the same 19mer peptide (Fig. 3A). As each spot is arrayed in duplicate, binding intensity analysis was conducted for each of the duplicate spots to compare relative binding intensity (left vs. right) which demonstrated a highly significant correlation for binding of each duplicate spot (R2 = 0.93) as one indicator of binding specificity (the red dotted line in Fig. 3B). The GST-cyclin D1 protein bound modified histones (H2B > H4 > H3), and the GST-cyclin D1ΔE protein showed substantially reduced binding (Fig. 3Ca–c). The pattern of binding by KDM4A resembled previously described preferences (Fig. 3Cd) [47]. To further interrogate the repertoire of post-translational histone modifications that may determine binding to cyclin D1, we quantitated interactions from multiplicate analysis of the 384 unique histone modification arrays on the 19mer peptides (Fig. 3D–I). H2B binding to cyclin D1 was enhanced by the additional modification of H2BS14 (Fig. 3D). This enhancement of binding was observed for H2BK5Ac, H2BK12Ac and for H2BK15Ac (Fig. 3F). Cyclin D1 recognized and showed enhanced binding for H2B that had been modified by multiple clusters of modifications (H2BK5Ac vs. H2BK5Ac-S14P, H2BK12Ac vs. H2BK12Ac-S14P, H2BK15Ac vs. H2BK15Ac-S14P) (Fig. 3F).
A Representative example of histone array consisting of 384 unique histone modification combinations in duplicate. The array includes up to four different modifications on the same 19mer peptide. For the individual modifications, please see Supplemental Table 3. The Y axis is labeled alphabetically for a given histone in each row, with modifications labeled numerically on the X axis. As each spot is arrayed in duplicate, representative examples of cyclin D1 binding to modified histones as tested by histone-modification arrays and analyzed by active Motif software are shown. B Binding is representative of the average positive intensity over the average negative intensity of each modification. The correlation between densitometric analysis of duplicate interactions for each interaction from two separate arrays is shown as intensity left vs. intensity right. The intensity is highly reproducible between duplicate binding assessments (R2 = 0.927). (P refers to the designation of the column on the array (H2B), and the number refers to the numerical labeling of the row). C Representative examples of histone arrays (384 unique histone modifications on the same 19mer peptide), with binding shown to GST-cyclin D1 (GST-CD1WT), GST-CD1ΔE, GST-KDM4A or GST-ctrl fusion proteins. D Representative and (E) quantitated binding, representing average positive intensity over the average negative intensity of each modification for H2B 19mer peptide. F Graphical quantification of cyclin D1 binding to post-translationally modified H2BS14 19mer peptide.
An intrinsically disordered domain (IDD) in cyclin D1
The strongly acidic nature of the cyclin D1 carboxyl-terminus, suggests this region may be intrinsically disordered. To assess the disorder status of the cyclin D1 carboxyl-terminal region, we applied a range of disorder predictors, including PrDOS, IUPred, and PONDR (Fig. 4A). The prediction results obtained showed a strong correlation with the consensus prediction taken from MobiDB 4.0 (Fig. 4A). A broad range of prediction models derived “Disorder consensus” for the region (Fig. S1A). The predictors rely on different principles, and their consensus is a highly dependent indication of the fundamental structural status of the protein. The predictors agree that most of the cyclin D1 sequence encodes for a structured protein (aa1-aa255, corresponding to a carboxyl-terminal and an amino-terminal domain), whereas the carboxyl-terminal 40 residues (aa256-aa295) have a strong tendency to be intrinsically disordered. Considering the training sets of the predictors relying on examples of IDPs/IDRs, we can be sure that this region does not fluctuate around an equilibrium conformation (that is, it is not “flexible”). Rather it is disordered and can assume a broad ensemble of conformations most appropriate for potentially adapting to diverse binding partners. This is particularly true for the E-rich region. The last carboxyl-terminal 15 residues appear as a down-peak on the disordered pattern (Fig. 4A) with alternating acidic and aliphatic residues, and its behavior of predicted disorder and local hydrophobicity is characteristic of IDP binding motifs. In agreement, this region gives a very strong signal by the ANCHOR predictor (Fig. 4A) developed to recognize specific protein recognition sites embedded in IDPs/IDRs [48].
A Disorder tendency and disordered binding site prediction. PRDos (blue), IUPred long (maroon), PONDR (orange) predictions show that the carboxy-terminal 40 residues (aa 256 - aa295) have a strong tendency to be intrinsically disordered, whereas ANCHOR (black) suggests that last 15 amino acids contain a protein-protein interaction site. B Multiple sequence alignment of the last (carboxyl-terminal) 100 amino acids of cyclin D1 from various species and of human cyclin A1, B1 and E1. The E-box motif was found in 148 human proteins by BLASTP, and their enrichment (C) and ratio (D) in GO Biological Process (BP) categories (“regulation of…”, no. of hits in the given category/148 polyE proteins) is shown.
Within the carboxyl-terminal 100 amino acids, the carboxyl-terminal hydrophobic stretch of the last 15 residues (VDLACTPTDVRDVDI) is the most conserved element in cyclin D1, which, along with the poly E region, is partially present in the remotely related zebrafish cyclin D1 (Fig. 4B). The E-rich region is specific to cyclin D1. It is not found in the closest homologs cyclin D2 and D3 (Fig. S1B). Otherwise, little conservation with distant species is seen in the rest of its C-terminal 100 residues, arguing strongly for the functional importance of the polyE region and the last 15 residues of cyclin D1. Interestingly, the full polyE repeat appears to have arisen in mammals, which fits into the general evolutionary trend that repeat expansion generating homopolymeric amino acid runs is a vehicle of late evolutionary innovation, as their level is much higher in eukaryotes than in prokaryotes and much higher in mammals than in lower vertebrates [49]. The observed conservation pattern of the polyE region in the CTD of cyclin D1 is consistent with this finding. It suggests that the functionality we report is a sign of an evolutionary functionalization event that occurred at the dawn of mammals.
Using the protein BLAST (BLASTP) data mining approach, the cyclin D1 E-rich motif, at least the minimum 9 glutamate residues, can be found in 148 unrelated proteins (Table S1), many of which have chromatin-related functions (Fig. 4C, D). By scrambling the sequences of these hits, we determined the expected distribution of poly E segments in them. The observed 9 residue-long or longer stretches occur highly significantly more frequently (Fig. S2, U-test p-value < 1E-70) than expected by chance (see Materials and Methods for details), which points to the functionality of this motif. In accord, we found that ~50% of these proteins containing E-rich regions participate in chromatin binding, transcriptional regulation, or chromosomal organization. This conclusion is supported by a detailed analysis for GO Biological Process (BP), Molecular Function (MF), and Cellular Component (CC) ontologies, determining enrichments with reference to the whole human genome (Fig. 4C), as shown by hypergeometric test to be highly significant (Fig. S3).
Cyclin D1 governs Top2A abundance, transcription, and co-activator recruitment to the Top2A promoter
In order to examine the potential role of the E domain in gene transcription we conducted ChIP analysis to assess recruitment of cyclin D1 to transcriptional targets. The lipoprotein lipase (LPL) gene is repressed by cyclin D1, as a component of cyclin D1-mediated restraint of lipogenesis [50], via transcriptional mechanisms associated with the recruitment of cyclin D1 in ChIP assays [29, 30]. As a form of positive control, ChIP assays of the Lpl promoter in cyclin D1+/+ 3T3 cells were conducted, showing the previously described presence of cyclin D1 in the context of local chromatin, an increased abundance of HDAC3, reduced H3K9 acetylation, the presence of heterochromatin protein 1 alpha (HP1α) and SUV39H1. Furthermore, associated with histone methylase SUV39H1, we identified increased di- and tri-methylation of H3K9 (Fig. 5A).
Endogenous murine LPL ChIP assay (A) and Top2A ChIP assays (B−I) with antibodies directed to target proteins, as indicated in the figure. C−I Recruitment of the indicated proteins to Top2A promoter was determined by ChIP and real-time quantitative PCR (qPCR). All data are mean ± SEM and represent n = 3 experiments. P-values were determined by the student’s t-test. J Western blot for Top2α and β tubulin abundance in cyclin D1-/- 3T3 cells rescued with a cyclin D1 expression vector with quantitation of multiplicate experiments. K Top2a intensity assessed by immunofluorescent staining in cyclin D1-/- cyclin D1 rescue 3T3 cells with data shown as mean ± SEM for N = 5. L Relative Top2A mRNA abundance in the mammary gland of Tet-inducible cyclin D1 mammary epithelial cell targeted transgenic mice (7 days of transgenic induction) or MMTV-cyclin D1 transgenic induced mammary gland tumors.
Prior genome-wide ChIP seq studies identified regions of genes to which cyclin D1 was significantly recruited into chromatin [8]. Cyclin D1 bound to the Top2A gene at −262 to −180, a site distinct from the previously characterized cell-cycle control element (−51 to +90) [51]. The Top2A gene is a representative example of the chromosomal instability gene (CIN) signature induced by cyclin D1 [8]. ChIP analysis was conducted to characterize the cyclin D1-mediated chromatin-associated complexes recruited to the endogenous Top2A gene regulatory region (Fig. 5B). In cyclin D1-/- 3T3 cells rescued with cyclin D1WT, recruitment of cyclin D1 to the murine Top2A promoter was associated with increased HP1α, p300 and with reduced abundance of HDAC3 (Fig. 5C–F). A quantitative analysis of multiple experiments demonstrated that cyclin D1 recruitment to the Top2A regulatory region (Fig. 5F, P = 0.035) was associated with the co-recruitment of p300 and HP1α (Fig. 5D, E, P = 0.011 and P = 0.029, respectively), reduced recruitment of HDAC3 (Fig. 5F, P = 0.009), correlating with substantially reduced H3K9me2 (Fig. 5H, P = 0.005) and unaltered H3K9 acetylation (Fig. 5I).
In cyclin D1+/+ (cyclin D1-/-cyclin D1) vs cyclin D1-/- 3T3 cells, Top2A abundance was increased by Western blot (Fig. 5J, P = 0.038) and by IHC (Fig. 5K). To define the impact of cyclin D1 on Top2A abundance in vivo, inducible mammary gland-targeted transgenic mice were deployed (Fig. 5L). Transient (7 day) expression of transgenic cyclin D1 in the mammary epithelium is sufficient for the induction of CIN in vivo [8]. The abundance of Top2A was induced 7-fold by cyclin D1 expression in the mammary gland of transgenic mice and Top2A expression was induced ~19-fold in MMTV-cyclin D1 tumors (Fig. 5L). The rescue of cyclin D1 into cyclin D1-/- cells in this system provides cyclin D1 within the physiological range [8], and the tet-inducible cyclin D1 transgenic mice express cyclin D1 at levels observed in mammary tumors [8, 9].
Analysis of Top2A, Top2B, and cyclin D1 expression in 2254 human breast cancer samples [8] showed cyclin D1, and TOP2A co-expression was associated with ERα+ tumors (Fig. S4A, B). Furthermore, scatter plot of cyclin D1 and Top2A abundance by breast cancer subtype depicts the significant correlation between cyclin D1 and Top2A in luminal B (r = 0.08. p = 0.24) and normal-like (r = 0.29, p = 1.2e-05) breast cancers (Fig. S4D, F).
A cyclin D1 carboxyl terminal glutamate-rich motif is required for recruitment to the Top2A promoter and DNA damage repair signaling, but not to induce cellular polarity and migration
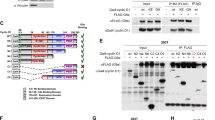
Because cyclin D1 is recruited to the Top2A promoter in ChIP-Seq [8] and ChIP, we deployed the Top2A promoter to define the domain(s) of cyclin D1 required for recruitment into the context of local chromatin using expression vectors encoding FLAG-tagged cyclin D1WT, and cyclin D1 mutants (Fig. 6A). Western blot demonstrated the greater abundance of the cyclin D1ΔE compared with the cyclin D1WT mutant in cultured cells (Fig. 6B, C). The abundance of the cyclin D1 ΔE mutant was 3-fold greater than cyclin D1WT when normalized to vinculin (Fig. 6B, C, P < 0.010). Immunofluorescence analysis of the transduced cells demonstrated similar levels of GFP expression from the retrovirus internal ribosome entry site in transfected cells (Fig. 6D). Cyclin D1 induces cellular polarity and cellular migration [23]. Herein, the reintroduction of cyclin D1WT or the cyclin D1ΔE mutant into cyclin D1-/- 3T3 cells restored cellular polarity (Fig. S5) and transwell migration (Fig. S6), whereas the mutant of cyclin D1 that is defective in CDK4/6 binding due to a point substitution of K114 (cyclin D1KE) and a mutant of cyclin D1 encoding amino acids 179−295 (cyclin D1C4 mutant) were defective (Fig. S5, 6).
A Schematic representation of cyclin D1wt and cyclin D1ΔE mutant with internal deletion of the E-rich region (red), FLAG tag (gray), C terminal region (pink), and remaining (orange) region. B Western blot detection of the FLAG-tagged expression vectors and (C) densitometric analysis of cyclin D1 proteins shown as mean ± SEM for N = 5 separate experiments (P < 0.05). D Immunofluorescence staining of the FLAG epitope. GFP (Green) from the vector IRES, DAPI (Blue), and FLAG (Red). E Mammalian 2-hybrid interaction of cyclin D1wt and cyclin D1ΔE. shown as mean ± SEM for N > 5 separate transfections conducted in MCF7 cells. F ChIP assays of cyclin D1-/- 3T3 cells rescued with the indicated FLAG-tagged cyclin D1 proteins conducted with the Top2A promoter with oligonucleotide pairs that were either specific (S) to the region identified in ChIP-seq or non-specific (NS) targeted to a region not identified in ChIP-seq as binding to cyclin D1. PCR and (G) detected occupancy. Quantitation of PCR products was shown as mean ± SEM for N = 3. H, I Representative ChIP analysis and quantitation of the PCR products derived from cyclin D1 proteins binding Zw10 or (J, K) Mlf1 regulatory region.
Cyclin D1WT bound CDK4 in mammalian two-hybrid assays in MCF7 breast cancer cells (Fig. 6E, P < 0.001) and 293 T cells (Fig. S7). The cyclin D1ΔE mutant showed enhanced CDK4 binding in MCF7 (Fig. 6E, P < 0.001) and 293 T cells (Fig. S7C). An expression vector encoding three distinct point mutations within the E region (E278A, E280A, E278/280 A) also maintained enhanced binding to CDK4 (Fig. S7C). Mutation of the CDK4/6 binding site of cyclin D1 (cyclin D1KE) abolished binding to CDK4 (Fig. S7A–C). Amino-terminal deletion mutants of cyclin D1 that removed the CDK binding site (C1-C5), abolished interaction with CDK4 (Fig. S7C). The reintroduction of cyclin D1 into cyclin D1-/- 3T3 cells increased the proportion of cells in the S phase and G2/M. Induction of G2/M was abolished by the cyclin D1ΔE mutant whereas induction of S phase was maintained (Fig. S8). The cyclin D1 C4 mutant was sufficient to induce the G2/M phase but did not induce S phase. These findings are consistent with our model in which the CDK binding and thereby S phase function are distinct and uncoupled from the transcriptional activities through the E domain to induce the expression of genes governing chromosomal instability [8] in a CDK-independent manner [9].
Cyclin D1 enhanced doxorubicin-induced phosphorylation of H2A at Ser139 and increased comet tail formation, as previously shown [25], with a reduction upon deletion of the E rich region (Fig. S9A, B). Phosphorylated H2AX (γ-H2AX) is a well-accepted marker of DNA double strand breaks [52]. Upon doxorubicin treatment Western blot analysis showed increased γ-H2AX abundance in cyclin D1WT rescued cyclin D1-/- MEFs at passage 20 (p20) (Fig. S9C) and passage 21 (p21) (Fig S9D) but not in cyclin D1ΔE rescue cells. These findings suggest that the E region is required for the cyclin D1-mediated augmentation of the DNA damage response. To determine the functional significance of cyclin D1 in Top2A activity, we assessed the role of cyclin D1 in regulating sensitivity to the Top2A inhibitor etoposide, which forms a ternary complex with DNA and the topoisomerase II enzyme (which aids in DNA unwinding), preventing re-ligation of the DNA strands, and by doing so causes DNA strands to break. Because Top2A levels vary throughout the cell cycle, increasing after DNA replication [51], cyclin D1-/- cells rescued with either cyclin D1WT or the cyclin D1ΔE mutant were treated with mimosine (200 μM, 24 h) to arrest the cells prior to DNA replication [53], and Western blot conducted for Top2A (Fig. S10A). Top2A induction by cyclin D1 was reduced by the cyclin D1ΔE mutant (Fig. S10A). Consistent with the increased abundance of Top2A, the cyclin D1 rescued cells were more sensitive to etoposide (Fig. S10B), with a modest but significant reduction in sensitivity upon deletion of the E domain.
Analysis was conducted of cyclin D1 binding in chromatin using cyclin D1-/- 3T3 cells rescued with cyclin D1WT or cyclin D1 mutants. Using two distinct pairs of oligonucleotide probes encompassing the cyclin D1 binding region identified through ChIP-Seq. Top2A promoter ChIP analysis demonstrated the recruitment of cyclin D1WT and cyclin D1KE (P < 0.010) (Fig. 6F, G). In contrast, the cyclin D1ΔE mutant was not recruited to the Top2A promoter (P < 0.010) (Fig. 6F, G). Probes directed to regions of the Top2A gene, that did not bind cyclin D1 in ChIP-seq did not show binding in ChIP (Fig. 7F, NS). The FLAG antibody that immunoprecipitated cyclin D1 in ChIP from cyclin D1-/-Rescue cells, did not ChIP the negative control GFP transduced cyclin D1-/-GFP 3T3 cells (Fig. 6F). Two additional target genes, Zw10 and Mlf1, which like Top2A participate in chromosomal stability, and were identified in ChIP-Seq as cyclin D1 target genes [8, 9], bound cyclin D1WT and cyclin D1KE with two distinct sets of oligonucleotide primers but did not bind cyclin D1ΔE in ChIP assays (Fig. 6H–K).
A Schematic representation of cyclin D1wt and mutants with internal deletion of the E-rich region (red), FLAG tag (gray), C terminal region (pink), and remaining (orange) region. B Karyotype determined through SKY analysis for cyclin D1-/- cells rescued with either cyclin D1wt or cyclin D1 mutants with (C) mean polyploidy shown. D ChIP assays of cyclin D1-/- 3T3 cells rescued with the indicated FLAG-tagged cyclin D1 proteins were conducted with the Top2A promoter. E Top2A promoter luciferase reporter assays are shown as mean ± SEM for N > 5 separate transfections. F Cellular proliferation assays were conducted in cyclin D1-/- 3T3 cells, rescued either with cyclin D1WT, mutant or control vector. G Summary of cyclin D1 mutant properties. The items labeled with * are based on ref. [99].
The C-terminal E domain of cyclin D1 is required to induce chromosomal instability
Previous studies using SKY analysis and gene expression profiling had demonstrated that cyclin D1 re-expression results in CIN, and this action is kinase-independent [8, 9]. To define the domain of cyclin D1 involved in the induction of CIN, we transduced cyclin D1-/- MEFs with expression vectors encoding cyclin D1WT, C4, ΔE, or ctrl and conducted karyotyping (Fig. 7A). SKY was conducted comparing the cyclin D1WT, C4 and ΔE mutants vs. vector control (GFP) cells. Expression of cyclin D1WT, or cyclin D1C4 was sufficient for the induction of aneuploidy. Aneuploidy was induced at 72 h by cyclin D1WT (90%, P < 0.03) and by cyclin D1C4 (100%, P < 0.01). In contrast, the cyclin D1ΔE mutant failed to induce aneuploidy compared with control cells (Fig. 7B, C).
We next assessed the transcriptional activation by cyclin D1 using the Top2A promoter linked to a luciferase reporter. The site of cyclin D1 binding to Top2A in 3T3 cells (−262 to −180), is distinct from the previously characterized cell-cycle control element (−51 to +90) [51]. Cyclin D1WT and cyclin D1C4 bound to the Top2A promoter in ChIP (Fig. 3D) and induced activity of the Top2A promoter 3.5-fold (Fig. 7D). The cyclin D1ΔE mutant that failed to bind the Top2A promoter in ChIP assays also failed to induce the Top2A promoter luciferase reporter gene (Fig. 7E). Cyclin D1WT increased proliferation 2-fold compared with the empty vector (Fig. 7F). The cyclin D1KE, cyclin D1C4, and cyclin D1ΔE mutant failed to promote the growth of 3T3 cyclin D1-/- rescued cells compared to cyclin D1WT (Fig. 7F). Thus, induction of aneuploidy, chromatin recruitment, Top2A promoter induction and the induction of cellular proliferation by cyclin D1 requires the poly E region (Fig. 7G). γH2AX is recruited into chromatin in response to signaling stimuli including NMDA [54,55,56,57]. Consistent with a model in which cyclin D1 is recruited into chromatin at sites of active DNA damage, ChIP-Seq analysis showed coincident binding for cyclin D1 at ~14% of γH2AX binding sites (Fig. S11A), with several gene targets involved in chromatin structure function and chromosomal instability (Fig. S11B).
These studies suggest a model in which cyclin D1 initiates a DNA damage response, characterized by the induction of γH2AX. Cyclin D1 is known to be recruited to sites of DNA damage with γH2AX [24,25,26], and such sites co-localize with H2BSer14 [27]. Cyclin D1 associated with high affinity to H2BS14P via the E domain. Collectively these studies identify the E region as an intrinsically disordered domain (IDD) that participates in both the induction of the DNA damage response and transcriptional regulation.
Discussion
The current studies extend our current understanding of the molecular mechanisms by which cyclin D1 mediates the transcriptional activity. First, these studies identified the minimal E region as required for cyclin D1 binding in chromatin to target genes identified in ChIP-seq (Top2a, Zw10, Mlf1). The region of cyclin D1 identified herein is distinct from the region binding directly to TFs, the basal transcription apparatus, or co-activators (AIB1, p300) [32, 58]. Second, we show that the E region is necessary for transcriptional induction by cyclin D1, aneuploidy induction, and the DNA damage response. Deletion of the E region did not affect cyclin D1-mediated induction of polarity, cellular migration, or binding to CDK4 in mammalian two-hybrid analysis in multiple cell types. Third, we show that the carboxyl-terminus is an IDD, particularly at the E region, with features of a protein binding interface seen in other IDD. Fourth, the E region was required for interaction with histone (H2BS14P), and in MST, the affinity to H2BSer14P was similar to that seen with other IDR interactions. Finally, we showed that the E region is present in 148 proteins, many of which are involved in chromosomal function. Cyclin D1 binds in chromatin to induce the transcription of genes governing CIN. As the deletion of the cyclin D1 E domain abolished both the recruitment of cyclin D1 into chromatin and CIN, we propose that the loss of transcriptional function is responsible for the loss of CIN.
Herein the cyclin D1 E region participated in both the DNA damage response and gene transcription, consistent with findings documenting the co-dependence of these processes for particular target genes [54, 56, 59, 60] (reviewed in [61]). Our analysis of cyclin D1 target genes identified in ChIP-Seq [8, 9] showed cyclin D1WT but not cyclin D1ΔE bound in ChIP assays to a variety of target promoters and augmented the DNA damage response. Double-stranded breaks facilitate gene induction mediated by nuclear receptors [60, 62, 63], heat shock, serum stimulation [54], and behavioral responses [55, 57]. TOPIIA (Top2α and Top2β) induce transient double-strand breaks and dissipate torsional stress by relaxing DNA supercoiling [61]. In cyclin D1 ChIP seq the most enriched sites for binding were CTCF sites [8, 9, 32]. CTCF sites are highly enriched for both γH2AX and Top2β within common genomic regions [56]. γH2AX is mainly confined to actively transcribed genes and their downstream regions [57]. Although the precise mechanisms remain to be determined, the induction of gene expression upon DNA damage may involve Top2β-mediated dissolution of CTCF-mediated restraint of enhancer-promoter interactions [56].
Cyclin D1 is recruited to sites of DNA damage [23,24,25, 64] and previous studies comparing the cyclin D1a and cyclin D1b isoform [25, 65], implicated the unique C-terminal sequence of cyclin D1a in the induction of the DNA damage response. Herein, cyclin D1 was recruited to H2BS14P via the E domain. H2BS14P co-localizes with γH2AX as the primary double-strand break marker in DNA damage foci [27]. Cyclin D1 bound the H2BS14P peptide with significantly higher affinity than H2BS14 (Kd = 2.4 μM vs. 64 μM) and deletion of the E-rich region (cyclin D1ΔE) abrogated binding to either of the peptides. The Kd of cyclin D1 binding to H2BS14P is similar to that of the 14-3-3 protein for H3S10P which ranged from 25-to 92 μM in two different studies [66, 67]. The strength of the cyclin D1/phospho-H2B interaction is comparable to those between other chromatin binding modules and their respective histone modifications, as the binding affinity of methylated H3 tail peptides to chromodomains of HP1 [68] and chromomethylase [69], ranges from 10 to 100 μM. In previous studies of IDD and fuzzy interaction motifs, multiple low-affinity interactions in the pM to μM range [70, 71], are necessary and contribute to the binding specificity of these motifs [71,72,73]. In our MST experimental protocol, the protein is labeled, therefore the mobility changes, induced by the binding of a small peptide assessed by amplitude of the changes in Fnorm, are small [74, 75]. The altered directionality of the Fnorm changes with H2BS14P (Fig. 6G) likely reflects a different mode of binding, as previously described [76]. The peptide concentrations used in MST with cyclin D1ΔE were considerably higher than with cyclin D1WT, resulting in slightly higher instability of the MST signal and higher scattering.
The C-terminal region was predicted herein to be intrinsically disordered, with predicted disorder scores dropping toward ordered values in the last 10−15 residues (VDLACTPTDVRDVDI), and strong interaction potential was predicted by ANCHOR [48]. Such patterns are often seen at short (linear) motifs that mediate protein-protein interactions in IDPs/IDRs [77]. It is highly plausible that these prediction tools capture the recently identified degron motif [78]. IDRs are often involved in molecular recognition and protein-protein interaction events due to their large accessible surface area and inherent flexibility [39]. IDPs/IDRs abound in homopolymeric runs of amino acids, and amino acid repeats often mediate their functional interactions [79]. The absolute sequence conservation of the cyclin D1 carboxyl-terminal hydrophobic region among different species underscores the functional importance of this region. Such short motifs of IDPs/IDRs often undergo induced folding (disorder-to-order transition) upon partner binding [41], which might be the case with the binding motif of cyclin D1. The disorder pattern, its hydrophobic character and evolutionary conservation corroborate the experimental observations that the carboxyl-terminal acidic region is involved in mediating the interaction functions of cyclin D1.
IDD interactions can be mediated by post-translational modifications including phosphorylation and acetylation [80]. For example, the IDD identified within CBP and p53 involve transient post-translational modifications of their substrates [81] and the Zinc finger protein 106, upon acetylation by CBP, interacts with an IDD within CBP [82]. Based on the precedent of other IDD behaviors, the impact of histone phosphorylation is predicted to govern folding to a structured state promoting protein-protein interactions [83,84,85], rather than charge change, which is a relatively trivial difference (pI Cyclin D1: 4.97, pI Cyclin D1 CTD: 3.79, vs. pI H3.1: 11.13), as a single phosphorylation event adds 1.5 negative charges to H3.1 (pI H3.1-phospho: 10.87). The Kd values for the association of multiply acetylated H4 tails and hTAFII250 double bromodomain are between 1 and 50 μM [86]. For example, the binding of folded domains, such as SH2 domains, to phosphorylated residues (pTyr), the added negative charge(s) of the phosphate group, results in binding in a dedicated groove, as opposed to the rather unspecific interaction of the unmodified peptide.
The emerging view is that nucleosomes can cluster into a dynamic, fuzzy phase [87] mediated by promiscuous, dynamic interactions of the disordered appendages of core and linker histones and other proteins. Some IDPs can exhibit relatively compact structures such as polyglutamines [88]. These interactions are referred to as “fuzzy” as the weak and transient complexes confer specificity without a structured state [43, 89]. Such an organization has also been observed in the heterochromatin, mediated by the interaction of methylated H3K9 and Heterochromatin protein 1 (HP1) [90]. Our results on the specific and modification-dependent interaction of the disordered region of cyclin D1 and histone tails is also of relevance in light of recent observations that interactions mediated by IDPs/IDRs in chromatin underlie the formation of a liquid-like state, in which nucleosomes are organized in a dynamic, condensed fashion [91]. Such condensates, in general, can form by liquid-liquid phase separation mediated by multivalent interactions of disordered proteins [44]. The E-rich motif is conserved with 148 proteins, many of which are involved in chromatin reorganization. Future studies will be directed to the role of these IDD-containing proteins in chromatin reorganization.
Materials and methods
Cell Culture, plasmids, chemicals, and luciferase reporter assays
The HEK293T cell line was obtained from ATCC (Manassas, VA) in the early 2000s. Ccnd1+/+, Ccnd1-/- or Ccnd1-/- cyclin D1 rescued mouse 3T3 fibroblasts were generated from the mouse in this lab [8]. The cells were maintained in Dulbecco’s modified Eagle’s media (DMEM) supplemented with 10% FBS, 2 mmol/L glutamine, 100 units/mL penicillin, and 100 μg/mL streptomycin in a humidified atmosphere containing 5% CO2 at 37 °C. The early passages of the cells were stored. Mycoplasma contamination was tested using the Universal Mycoplasma Detection Kit from ATCC to ensure the absence of mycoplasma contamination. The cells thawed from low passage stocks were used within one month of the initial thaw. During the experiments, the morphology of all cell lines was routinely checked under a phase contrast microscope. All of the newly revived cells were treated with Plasmocin (InvivoGen), and any mycoplasma contamination was determined with Hoechst 33258 staining under a high magnification fluorescent microscope routinely.
The mouse stem cell virus-internal ribosome entry site-green fluorescent protein retrovirus vectors, pMSCV-IRES-GFP, and the ecotropic, packaging vector, pSV-ψ-E-MLV and infection methods were as described previously [50]. The cDNA encoding cyclin D1 wt, N1 (aa1-aa287), C4 (aa179-aa295), and deletion of the E-rich motif (ΔE) (aa272-aa280) were generated by PCR. EcoR1 and Bam HI restriction sites were introduced via PCR primers. The PCR products were cloned into pMSCV-IRES-GFP vector. The GST expression plasmids were generated using either full length cyclin D1 cDNA, or cyclin D1 ΔE mutant. These fragments were subcloned into the BamHI and EcoR1 sites of pGEX-4T-3 vector (Amersham). For the cloning of the VP16-cyclin D1 constructs the cDNA encoding D1 wt, N1 (aa1-aa287), C4 (aa179-aa295) and deletion of E-rich motif (aa272-aa280) were generated by PCR. BamHI and XbaI restriction sites were introduced via the primers. The PCR products were cloned into pACT vector (Promega). The human TOP2A promoter linked to a luciferase reporter gene was previously described [92] and the data were normalized through co-transfection of a renilla luciferase reporter plasmid (1 ng). Mimosine was from Santa Cruz Biotechnology, and doxorubicin was from Sigma.
MSCV retroviruses were prepared by transient co-transfection with helper virus into 293 T cells, using calcium phosphate precipitation. The retroviral supernatants were harvested 48 h after transfection and filtered through a 0.45-μm filter. Ccnd1-/- 3T3 cells were incubated with fresh retroviral supernatants in the presence of 8 μg/ml Polybrene for 24 h, cultured for an additional 4 days, and then subjected to fluorescence-activated cell sorting (FACS) (FACStar Plus; BD Biosciences, San Jose, CA) to select for cells expressing GFP. GFP+ cells were subsequently analyzed. Ccnd1-/- 3T3 cells were rescued with cyclin D1 protein expression vectors with FLAG-tagged cyclin D1 wild type (WT) or the mutants; amino- and carboxyl-terminal deletions (C4 and N1, respectively) and E-rich motif internal deletion (ΔE) and compared with cells transduced with an empty vector expressing IRES-GFP as a control. Wortmannin was used at a final concentration of 200 nM for 6 h.
Animal model
Animal experiments were approved by Thomas Jefferson University’s Institutional Animal Care and Use Committee (IACUC). Eight to ten weeks-old female mice were used for in vivo experiments. All mouse experiments using the MMTV-cyclin D1 transgenic mice and the tetracycline-inducible mammary epithelial cell targeted cyclin D1a transgenic mice (MMTV-rtTA-cyclin D1) in the FVB strain background were conducted following the guidelines for the care and use of laboratory animals at Thomas Jefferson University.
Spectral karyotyping analysis
For spectral karyotyping (SKY) analysis, fluorescence color images of chromosomes stained by Rhodamine, Texas Red, Cy5, FITC, and Cy5.5 were captured under a Nikon microscope equipped with a spectral cube and Interferometer module. SKY View software (version 1.62), was used to analyze the chromosomal number and structural alterations of chromosomes, including simple balanced translocations, unbalanced (or nonreciprocal) translocations, deletions, and duplications. At least 10 metaphases were analyzed per sample. Statistical significance was calculated using the chi-square test of association (Pearson).
Western blotting and Immunofluorescence
Whole-cell lysates (50 μg) were separated by 10% SDS-PAGE gel, and the proteins were transferred to nitrocellulose membrane for Western blotting using the following antibodies: anti-FLAG antibody (M2 Aldrich, #F1804, St. Louis, MO), anti-GST (Cell Signaling, #2624, Danvers, MA), anti-β-tubulin (Santa Cruz Biotechnology, #sc-55529, Santa Cruz, CA), anti-Vinculin (Sigma-Aldrich, #V9131 St. Louis, MO), Topoisomerase 2 A (Santa Cruz Biotechnology, #sc-365916, Santa Cruz, CA), Histone 2B Serine 14 P (Santa Cruz Biotechnology, #sc-31671, Santa Cruz, CA), Histone 2B (Santa Cruz Biotechnology, #sc-10808, Santa Cruz, CA), γH2A.X (Cell Signaling, #9718, Danvers, MA) and H2A.X (Cell Signaling, #2595, Danvers, MA).
Immunofluorescence was performed on Ccnd1-/- cells transduced with retrovirus expressing cyclin D1 WT, C4, N1, ΔE, or vector control as previously described [8]. The subcellular localization was determined using the anti-FLAG M2-antibody.
Chromatin immunoprecipitation assay
Chromatin immunoprecipitation (ChIP) analysis was conducted as described [8] following the protocol provided by Upstate Biotechnology (Charlottesville, VA). Chromatin solutions were precipitated overnight at 4 °C with rotation, using 8 μg of anti-FLAG, 4 μg of anti-Cyclin D1 (HD-11, Santa Cruz Biotechnology, Santa Cruz, CA), H3(Ac-K9) (Millipore, #07-352, Billerica, MA), p300, (Santa Cruz Biotechnology, #sc-585X, Santa Cruz, CA), HDAC3 (Abcam, #ab7030, Cambridge, UK), H3(Dimethyl-K9) (Millipore, #1220, Billerica, MA), SUV39H1 (Millipore, #05-615, Billerica, MA) and HP1α (Millipore, #05-685, Billerica, MA). For negative control, rabbit IgG and mouse IgG were immunoprecipitated. ChIP analysis for each immunoprecipitated protein was conducted on the endogenous murine Top2A promoter (35 cycles of PCR). ChIP-DNA quantitation was conducted in an Agilent 2100 bioanalyzer (Agilent Technologies, Palo Alto, CA), using Power SYBR Green (AB Biosciences) according to the manufacturer’s guidelines. Equal quantities of ChIP-DNA were used for the real-time PCR quantitation. Ct values were used to calculate the relative fold enrichment (2-ΔCt, ΔCt = Ctinput - CtIgG). A one-way ANOVA followed by Student’s t-test comparison was performed to compare the relative fold enrichment (n = 3).
Comet assays
Neutral pH comet assays were conducted using the Comet- Assay Kit (Trevigen) as previously described [25]. After treatment with doxorubicin or control, cells were harvested and mixed with low-melting temperature agarose. After lysis, electrophoresis was conducted at 1 V/cm for 20 min. Slides were stained with SYBG green dye for 10 min and visualized on a Zeiss LSM 510 META confocal microscope with a Å~20 objective. The relative length and intensity of SYBG green–stained DNA tails to heads was proportional to the amount of DNA damage present in the individual nuclei and was measured by Olive tail moment using OpenComet software (http://www.cometbio.org/index.html) [93].
Protein-protein interaction by mammalian two-hybrid reporter assay
Mammalian two-hybrid was conducted following the manufacturer’s instruction (Promega, Madison, WI). The human CDK4 cDNA was cloned into pBIND vector, and the human cyclin D1 cDNA (wild type or mutants) was cloned into pACT vector to generate fusion proteins with the DNA-binding domain of Gal4 (Gal4-CDK4), and the activation domain of VP16 (VP16-cyclin D1), respectively. Gal4-CDK4, VP16-cyclin D1 and pG5-Luc reporter containing five copies of Gal4 binding sites upstream of a minimal TATA box and the firefly luciferase gene (Promega) were co-transfected into MCF-7 or HEK-293T cells with Superfect reagent 48 h after transfection, the cells were lysed, and the amount of luciferase activity was quantitated. Transfection efficiency was normalized using renilla luciferase.
Peptide and GST pull-down assays
The Peptide pull-down assays were performed following established protocols. Biotin peptides Histone H2B and Histone H2BS14P were customized and purchased from Anaspec. Pull-down assays were performed using bacterial-purified GST-Cyclin D1 WT, Cyclin D1ΔE deletion, GST-KMD4A, and GST control. Equal molarity of GST-tagged proteins was used. 25 μg of the histone peptides were bound to Streptavidin agarose beads in PBS for 30 min at room temperature, and then the beads were washed 3 times with PBS. Cyclin D1WT, Cyclin D1ΔE, and GST were incubated with the beads for 1 h at room temperature, with constant mixing, followed by 3 washing steps with PBS (https://www.sigmaaldrich.com/US/en/technical-documents/protocol/protein-biology/enzyme-activity-assays/pull-down-assays).
Surface plasmon resonance
To analyze further the specific interactions between cyclin D1 and histone modifications, we deployed Surface Plasmon Resonance (SPR). SPR between the proteins was measured using surface plasmon resonance measurements on a Bio-Rad ProteOnTM XPR 36 Protein Interaction Array System. Biotinylated histone peptides were immobilized on the surface of a ProteOn™ NLC Sensor Chip (# 1765021). The chip was initialized and equilibrated as suggested by the manufacturer. The buffer used for ligand capture and partner binding was: 20 mM TRIS, 150 mM NaCl, 2 mM EDTA, 0.5% Tween-20, and pH8.0. The peptides were loaded on the chip surface at a concentration of 5 μg/ml with a flow rate of 30 μl/min to a level of 300 RU (response units). Ligand capture and partner binding experiments were carried out at 25 oC. Binding of different proteins was detected using a concentration range between 0.1 and 2 μM (cyclin D1ΔE, GST, and Tudor domain) or 1 and 10 μM (cyclin D1wt). Flowrate used for binding was 40 μl/min, the association time was 240 s and the dissociation time varied between 560 and 960 s. A lane without ligand added was used as negative control and the resulting baseline curves were subtracted before binding analysis.
Microscale Thermophoresis (MST) analysis
Binding assays were carried out on a Microscale Thermophoresis (MST) system (Monolith NT. 115 from NanoTemper Technologies, München, Germany) in reaction mixtures containing fluorescently labeled cyclin D1 variants and unlabeled H2B peptides in PBS. Standard-treated capillaries (Cat. No.: MO-K002) were used for measurements. Instrument settings were as follows: First round: LED Power: 10%, MST Power 20%, Before MST: 5 s, MST on: 30 s, After MST: 5 s, Delay: 25 s. The second round had the same settings, but the MST Power was 40%. For the measurements, GST-tagged Cyclin D1WT and GST-tagged Cyclin D1ΔE were fluorescently labeled with an amine-reactive fluorescent dye (Monolith Protein Labeling Kit RED-NHS 2nd Generation, Cat. No.: MO-L011) according to the protocol provided by the manufacturer. Labeled protein concentrations were set to give an initial raw fluorescence between 800 and 1000 counts and were varied between 3.5 μM and 12.5 μM. All experiments were conducted at room temperature. Labeling and MST measurements were carried out in phosphate-buffered saline (PBS, Cat. No.: 524650, Merck KGaA, Darmstadt, Germany). The exact composition was: 140 mM NaCl, 10 mM phosphate buffer, and 3 mM KCl, pH 7.4. Anaspec (Fremont CA, USA) synthesized histone peptides with C-terminal biotin labels. The sequences of the peptides were: Unmodified H2B: PEPAKSAPAPKKGSKKAVTKAQ, Phosphorylated H2B: PEPAKSAPAPKKGpSKKAVTKAQ. Starting peptide concentrations were 15 times higher than the Cyclin D1 concentration in the reaction mixture. Curve fitting and Kd calculations were carried out with the software provided with the instrument (MO Affinity Analysis, Version 2.3). A detailed description of the MST method and the interpretation of the results can be found in Jerabek-Willemsen et al. [76, 94].
Histone peptide array
The modified histone peptide arrays were purchased from Active Motif (Active Motif, # 13001, Carlsbad, CA) and analyzed as described by the manufacturer’s protocol using the buffer (100 mM KCl, 20 mM HEPES pH 7.5, 1 mM EDTA, 0.1 mM DTT and 10% glycerol). The membranes were incubated with purified protein either from full-length or mutant-expressed GST-Cyclin D1WT or GST-Cyclin D1ΔE.
Protein data mining analysis and prediction of disorder tendency and multiple sequence alignment
BLASTP (National Library of Medicine) was used to retrieve all proteins that contain a minimum of 9 tandem glutamine repeats. The search retrieved 149 hits, including cyclin D1, of which there were 148 unique proteins (and one pseudogene). The 148 unique proteins were inputted into the DAVID functional annotation clustering algorithm to display and cluster similar annotation terms with an enrichment score of >1.5.
Four different disorder prediction algorithms assessed structural disorder: PrDOS [95], IUPred [96], RONN [97] and DISOPRED2 [98]. The results obtained showed a high correlation with the consensus prediction from MobiDB 4.0 (https://mobidb.bio.unipd.it). These predictors rely on different principles and represent different stringencies. Thus their consensus offers exquisite reliability in the disorder status of a (segment) of a polypeptide chain. Briefly, PrDOS is composed of two predictors, one based on local amino acid sequence and another based on template proteins, and is most appropriate for predicting short, disordered segments. IUPred relies on assessing the total pairwise interaction energy encoded in a polypeptide chain and has options for predicting long and short disorder. The “long” option can detect context-independent global disorder of a protein that encompasses at least 30 consecutive residues of predicted disorder. The “short” option is suitable for predicting short, probably context-dependent disordered regions. RONN (Regional Order Neural Network) is a bio-basis function neural network trained on disordered proteins that predicts regions that lack a well-defined 3D structure under native conditions. DISOPRED2 is based on cascaded support vector machine classifiers trained on PSI-BLAST profiles.
Multiple sequence alignments were conducted with the ClustalW server at the EMBL-EBI website (http://www.ebi.ac.uk/Tools/msa/clustalw2/) on sequences downloaded from the UniProt site (http://www.uniprot.org/). A graphic representation of the data was prepared with the ClustalX software.
To evaluate the significance of the enrichment of E-rich region sequences, the length distribution of poly E segments was extracted and compared to random sequences. The 148 protein hits (Table S2) were scrambled 100 times, and the average length distribution of poly E segments was recorded. Mann-Whitney’s U-test indicated that the E-rich region sequences and their scrambled variants have extremely significantly different poly E length distributions, with a p-value < 10−70, suggesting that the observed distribution (and the presence of a poly E region of 9 consecutive Glu-structure) could not have occurred by chance.
Cyclin D1-H2B model generation
The human cyclin D1-H2B model was generated using the CABS-dock server (http://biocomp.chem.uw.edu.pl/CABSdock/) [99, 100] without defining a specific H2B binding site of cyclin D1 thus allowing for the software to identify the best fit independently. H2B docking into cyclin D1 structure was carried out in three steps [99, 100]. Briefly, the server first converts the cyclin D1 into a coarse‐grained representation thus simulating the behavior of complex systems. The CABS-dock server uses a Replica Exchange Monte Carlo pseudo‐dynamics with annealing for protein dynamic simulation. The next step involves the placement of H2B in various possible protein binding pockets approximately 20 Å from the cyclin D1 surface. H2B peptides used in docking were randomly selected from a library of 10 generic H2B starting conformations. We selected the default CABS-dock mode, which uses 10 replicas and 20 annealing cycles. The available x-ray or cryo-EM structures of human cyclin D1 lack the E-region so we used the AlphaFold generated CD1 (AF-P25322-F1) and the NMR H2B coordinates PDB ID: 2VQ in our modeling. The docking was carried out with two different length peptides of the H2B tail: PEPAKSAPAPKKGSKKAVTKAQKKDGKKRKRSR, and PEPAKSAPAPKKGSKKAVTKAQKKDGKKRKR, as it is not clear what is the precise length of the H2B tail that binds cyclin D1. Both peptides contain the residue S14 but either lack or include residues C- and N-terminal to it. The CABS-dock server placed both longer peptides in precisely the same location on the surface of cyclin D1. In this study, we selected the best binding mode of H2B from the 10-top scores. The figures were made in PYMOL (The PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC) and the electrostatic surface charge generated with APBS [101].
ChIP-Seq analysis
Cyclin D1 CHIP-Seq data in mouse embryonic fibroblasts is from our previous work (GSE207361) [8]. γH2A.X Chip-Seq data is from Dr. Madabhushi’s published work (GSE61887) [56]. The track of cyclin D1 and γH2A.X in the chip-seq were viewed and captured with Integrated Genome Browser.
Statistical analysis
The statistical analysis was performed using Prism 4 software (GraphPad Software, San Diego, CA). Differences were considered statistically significant with P-values < 0.05.
Data availability
All data generated or analyzed during this study are included in this published article and its supplementary information files.
References
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Cahill DP, Kinzler KW, Vogelstein B, Lengauer C. Genetic instability and darwinian selection in tumours. Trends Cell Biol. 1999;9:M57–60.
Davoli T, Uno H, Wooten EC, Elledge SJ. Tumor aneuploidy correlates with markers of immune evasion and with reduced response to immunotherapy. Science. 2017;355:eaaf8399
Fujiwara T, Bandi M, Nitta M, Ivanova EV, Bronson RT, Pellman D. Cytokinesis failure generating tetraploids promotes tumorigenesis in p53-null cells. Nature. 2005;437:1043–7.
Sack LM, Davoli T, Li MZ, Li Y, Xu Q, Naxerova K, et al. Profound tissue specificity in proliferation control underlies cancer drivers and aneuploidy patterns. Cell. 2018;173:499–514 e423.
Sotillo R, Schvartzman JM, Socci ND, Benezra R. Mad2-induced chromosome instability leads to lung tumour relapse after oncogene withdrawal. Nature. 2010;464:436–40.
Kuzyk A, Mai S. c-MYC-induced genomic instability. Cold Spring Harb Perspect Med. 2014;4:a014373.
Casimiro MC, Crosariol M, Loro E, Ertel A, Yu Z, Dampier W, et al. ChIP sequencing of cyclin D1 reveals a transcriptional role in chromosomal instability in mice. J Clin Investig. 2012;122:833–43.
Casimiro MC, Di Sante G, Crosariol M, Loro E, Dampier W, Ertel A, et al. Kinase-independent role of cyclin D1 in chromosomal instability and mammary tumorigenesis. Oncotarget. 2015;6:8525–38.
Duffy S, Fam HK, Wang YK, Styles EB, Kim JH, Ang JS, et al. Overexpression screens identify conserved dosage chromosome instability genes in yeast and human cancer. Proc Natl Acad Sci USA. 2016;113:9967–76.
Diehl JA, Cheng M, Roussel MF, Sherr CJ. Glycogen synthase kinase-3beta regulates cyclin D1 proteolysis and subcellular localization. Genes Dev. 1998;12:3499–511.
Pestell RG. New roles of cyclin D1. Am J Pathol. 2013;183:3–9.
Ewen ME, Sluss HK, Sherr CJ, Matsushime H, Kato J-Y, Livingston DM. Functional interactions of the retinoblastoma protein with mammalian D-type cyclins. Cell. 1993;73:487–97.
Hydbring P, Malumbres M, Sicinski P. Non-canonical functions of cell cycle cyclins and cyclin-dependent kinases. Nat Rev Mol Cell Biol. 2016;17:280–92.
Kato J-Y, Matsushime H, Hiebert SW, Ewen ME, Sherr CJ. Direct binding of cyclin D to the retinoblastoma gene product (pRb) and pRb phosphorylation by the cyclin D-dependent kinase CDK4. Genes Dev. 1993;7:331–42.
Wang C, Li Z, Lu Y, Du R, Katiyar S, Yang J, et al. Cyclin D1 repression of nuclear respiratory factor 1 integrates nuclear DNA synthesis and mitochondrial function. Proc Natl Acad Sci USA. 2006;103:11567–72.
Di Sante G, Page J, Jiao X, Nawab O, Cristofanilli M, Skordalakes E, et al. Recent advances with cyclin-dependent kinase inhibitors: therapeutic agents for breast cancer and their role in immuno-oncology. Expert Rev Anticancer Ther. 2019;19:569–87.
Yu Z, Wang C, Wang M, Li Z, Casimiro MC, Liu M, et al. A cyclin D1/microRNA 17/20 regulatory feedback loop in control of breast cancer cell proliferation. J Cell Biol. 2008;182:509–17.
Lee RJ, Albanese C, Fu M, D’Amico M, Lin B, Watanabe G, et al. Cyclin D1 is required for transformation by activated Neu and is induced through an E2F-dependent signaling pathway. Mol Cell Biol. 2000;20:672–83.
Hulit J, Wang C, Li Z, Albanese C, Rao M, Di Vizio D, et al. Cyclin D1 genetic heterozygosity regulates colonic epithelial cell differentiation and tumor number in ApcMin mice. Mol Cell Biol. 2004;24:7598–611.
Yu Q, Geng Y, Sicinski P. Specific protection against breast cancers by cyclin D1 ablation. Nature. 2001;411:1017–21.
Sherr CJ, Beach D, Shapiro GI. Targeting CDK4 and CDK6: from discovery to therapy. Cancer Discov. 2016;6:353–67.
Li Z, Wang C, Jiao X, Lu Y, Fu M, Quong AA, et al. Cyclin D1 regulates cellular migration through the inhibition of thrombospondin 1 and ROCK signaling. Mol Cell Biol. 2006;26:4240–56.
Jirawatnotai S, Hu Y, Michowski W, Elias JE, Becks L, Bienvenu F, et al. A function for cyclin D1 in DNA repair uncovered by protein interactome analyses in human cancers. Nature. 2011;474:230–4.
Li Z, Jiao X, Wang C, Shirley LA, Elsaleh H, Dahl O, et al. Alternative cyclin D1 splice forms differentially regulate the DNA damage response. Cancer Res. 2010;70:8802–11.
Li Z, Chen K, Jiao X, Wang C, Willmarth NE, Casimiro MC, et al. Cyclin D1 integrates estrogen-mediated DNA damage repair signaling. Cancer Res. 2014;74:3959–70.
Fernandez-Capetillo O, Allis CD, Nussenzweig A. Phosphorylation of histone H2B at DNA double-strand breaks. J Exp Med. 2004;199:1671–7.
Soutoglou E, Misteli T. Activation of the cellular DNA damage response in the absence of DNA lesions. Science. 2008;320:1507–10.
Bienvenu F, Barre B, Giraud S, Avril S, Coqueret O. Transcriptional regulation by a DNA-associated form of cyclin D1. Mol Biol Cell. 2005;16:1850–8.
Fu M, Rao M, Bouras T, Wang C, Wu K, Zhang X, et al. Cyclin D1 inhibits peroxisome proliferator-activated receptor gamma-mediated adipogenesis through histone deacetylase recruitment. J Biol Chem. 2005;280:16934–41.
Bienvenu F, Jirawatnotai S, Elias JE, Meyer CA, Mizeracka K, Marson A, et al. Transcriptional role of cyclin D1 in development revealed by a genetic-proteomic screen. Nature. 2010;463:374–8.
Pauklin S, Madrigal P, Bertero A, Vallier L. Initiation of stem cell differentiation involves cell cycle-dependent regulation of developmental genes by Cyclin D. Genes Dev. 2016;30:421–33.
Albero R, Enjuanes A, Demajo S, Castellano G, Pinyol M, Garcia N, et al. Cyclin D1 overexpression induces global transcriptional downregulation in lymphoid neoplasms. J Clin Investig. 2018;128:4132–47.
Casimiro MC, Wang C, Li Z, Di Sante G, Willmart NE, Addya S, et al. Cyclin D1 determines estrogen signaling in the mammary gland in vivo. Mol Endocrinol. 2013;27:1415–28.
Ju X, Casimiro MC, Gormley M, Meng H, Jiao X, Katiyar S, et al. Identification of a cyclin D1 network in prostate cancer that antagonizes epithelial-mesenchymal restraint. Cancer Res. 2014;74:508–19.
Kollmann K, Heller G, Schneckenleithner C, Warsch W, Scheicher R, Ott RG, et al. A kinase-independent function of CDK6 links the cell cycle to tumor angiogenesis. Cancer Cell. 2013;24:167–81.
Berlow RB, Dyson HJ, Wright PE. Expanding the paradigm: intrinsically disordered proteins and allosteric regulation. J Mol Biol. 2018;430:2309–20.
Csizmok V, Follis AV, Kriwacki RW, Forman-Kay JD. Dynamic protein interaction networks and new structural paradigms in signaling. Chem Rev. 2016;116:6424–62.
Tompa P, Schad E, Tantos A, Kalmar L. Intrinsically disordered proteins: emerging interaction specialists. Curr Opin Struct Biol. 2015;35:49–59.
Tompa P. Intrinsically disordered proteins: a 10-year recap. Trends Biochem Sci. 2012;37:509–16.
Wright PE, Dyson HJ. Linking folding and binding. Curr Opin Struct Biol. 2009;19:31–38.
Tompa P, Szasz C, Buday L. Structural disorder throws new light on moonlighting. Trends Biochem Sci. 2005;30:484–9.
Tompa P, Fuxreiter M. Fuzzy complexes: polymorphism and structural disorder in protein-protein interactions. Trends Biochem Sci. 2008;33:2–8.
Shin Y, Brangwynne CP (2017). Liquid phase condensation in cell physiology and disease. Science. 2017;357.
Pancsa R, Schad E, Tantos A, Tompa P. Emergent functions of proteins in non-stoichiometric supramolecular assemblies. Biochim Biophys Acta Proteins Proteom. 2019;1867:970–9.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–9.
Su Z, Wang F, Lee JH, Stephens KE, Papazyan R, Voronina E, et al. Reader domain specificity and lysine demethylase-4 family function. Nat Commun. 2016;7:13387.
Dosztanyi Z, Meszaros B, Simon I. ANCHOR: web server for predicting protein binding regions in disordered proteins. Bioinformatics. 2009;25:2745–6.
Alba MM, Tompa P, Veitia RA. Amino acid repeats and the structure and evolution of proteins. Genome Dyn. 2007;3:119–30.
Wang C, Pattabiraman N, Zhou JN, Fu M, Sakamaki T, Albanese C, et al. Cyclin D1 repression of peroxisome proliferator-activated receptor gamma expression and transactivation. Mol Cell Biol. 2003;23:6159–73.
Falck J, Jensen PB, Sehested M. Evidence for repressional role of an inverted CCAAT box in cell cycle-dependent transcription of the human DNA topoisomerase IIalpha gene. J Biol Chem. 1999;274:18753–8.
Kuo LJ, Yang LX. Gamma-H2AX - a novel biomarker for DNA double-strand breaks. Vivo. 2008;22:305–9.
Park SY, Im JS, Park SR, Kim SE, Wang HJ, Lee JK. Mimosine arrests the cell cycle prior to the onset of DNA replication by preventing the binding of human Ctf4/And-1 to chromatin via Hif-1alpha activation in HeLa cells. Cell Cycle. 2012;11:761–6.
Bunch H, Lawney BP, Lin YF, Asaithamby A, Murshid A, Wang YE, et al. Transcriptional elongation requires DNA break-induced signalling. Nat Commun. 2015;6:10191.
Delint-Ramirez I, Konada L, Heady L, Rueda R, Jacome ASV, Marlin E, et al. Calcineurin dephosphorylates topoisomerase IIbeta and regulates the formation of neuronal-activity-induced DNA breaks. Mol Cell. 2022;82:3794–3809.e3798.
Madabhushi R, Gao F, Pfenning AR, Pan L, Yamakawa S, Seo J, et al. Activity-Induced DNA breaks govern the expression of neuronal early-response genes. Cell. 2015;161:1592–605.
Stott RT, Kritsky O, Tsai LH. Profiling DNA break sites and transcriptional changes in response to contextual fear learning. PLoS One. 2021;16:e0249691.
Fu M, Wang C, Rao M, Wu X, Bouras T, Zhang X, et al. Cyclin D1 represses p300 transactivation through a cyclin-dependent kinase-independent mechanism. J Biol Chem. 2005;280:29728–42.
Gomez-Cabello D, Pappas G, Aguilar-Morante D, Dinant C, Bartek J. CtIP-dependent nascent RNA expression flanking DNA breaks guides the choice of DNA repair pathway. Nat Commun. 2022;13:5303.
Ju BG, Lunyak VV, Perissi V, Garcia-Bassets I, Rose DW, Glass CK, et al. A topoisomerase IIbeta-mediated dsDNA break required for regulated transcription. Science. 2006;312:1798–802.
Puc J, Aggarwal AK, Rosenfeld MG. Physiological functions of programmed DNA breaks in signal-induced transcription. Nat Rev Mol Cell Biol. 2017;18:471–6.
Haffner MC, Aryee MJ, Toubaji A, Esopi DM, Albadine R, Gurel B, et al. Androgen-induced TOP2B-mediated double-strand breaks and prostate cancer gene rearrangements. Nat Genet. 2010;42:668–75.
Lin C, Yang L, Tanasa B, Hutt K, Ju BG, Ohgi K, et al. Nuclear receptor-induced chromosomal proximity and DNA breaks underlie specific translocations in cancer. Cell. 2009;139:1069–83.
Casimiro MC, Di Sante G, Ju X, Li Z, Chen K, Crosariol M, et al. Cyclin D1 promotes androgen-dependent DNA damage repair in prostate cancer cells. Cancer Res. 2016;76:329–38.
Solomon DA, Wang Y, Fox SR, Lambeck TC, Giesting S, Lan Z, et al. Cyclin D1 splice variants. Differential effects on localization, RB phosphorylation, and cellular transformation. J Biol Chem. 2003;278:30339–47.
Macdonald N, Welburn JP, Noble ME, Nguyen A, Yaffe MB, Clynes D, et al. Molecular basis for the recognition of phosphorylated and phosphoacetylated histone h3 by 14-3-3. Mol Cell. 2005;20:199–211.
Walter W, Clynes D, Tang Y, Marmorstein R, Mellor J, Berger SL. 14-3-3 interaction with histone H3 involves a dual modification pattern of phosphoacetylation. Mol Cell Biol. 2008;28:2840–9.
Jacobs SA, Taverna SD, Zhang Y, Briggs SD, Li J, Eissenberg JC, et al. Specificity of the HP1 chromo domain for the methylated N-terminus of histone H3. EMBO J. 2001;20:5232–41.
Lindroth AM, Shultis D, Jasencakova Z, Fuchs J, Johnson L, Schubert D, et al. Dual histone H3 methylation marks at lysines 9 and 27 required for interaction with CHROMOMETHYLASE3. EMBO J. 2004;23:4286–96.
Dogan J, Gianni S, Jemth P. The binding mechanisms of intrinsically disordered proteins. Phys Chem Chem Phys. 2014;16:6323–31.
Shammas SL, Rogers JM, Hill SA, Clarke J. Slow, reversible, coupled folding and binding of the spectrin tetramerization domain. Biophys J. 2012;103:2203–14.
Davey NE, Van Roey K, Weatheritt RJ, Toedt G, Uyar B, Altenberg B, et al. Attributes of short linear motifs. Mol Biosyst. 2012;8:268–81.
Hoepffner HJ, Dreyer M, Reimers U, Schmidt-Preuss U, Koepp HP, Rudiger HW. A new familial syndrome with impaired function of three related peptide growth factors. Hum Genet. 1989;83:209–16.
Bartoschik T, Galinec S, Kleusch C, Walkiewicz K, Breitsprecher D, Weigert S, et al. Near-native, site-specific and purification-free protein labeling for quantitative protein interaction analysis by MicroScale Thermophoresis. Sci Rep. 2018;8:4977.
Huang L, Zhang C. Microscale Thermophoresis (MST) to detect the interaction between purified protein and small molecule. Methods Mol Biol. 2021;2213:187–93.
Jerabek-Willemsen M, Andre T, Wanner R, Roth HM, Duhr S, Baaske P, et al. MicroScale thermophoresis: interaction analysis and beyond. J Mol Struct. 2014;1077:101–13.
Oldfield CJ, Cheng Y, Cortese MS, Romero P, Uversky VN, Dunker AK. Coupled folding and binding with alpha-helix-forming molecular recognition elements. Biochemistry. 2005;44:12454–70.
Li Y, Jin K, Bunker E, Zhang X, Luo X, Liu X, et al. Structural basis of the phosphorylation-independent recognition of cyclin D1 by the SCF(FBXO31) ubiquitin ligase. Proc Natl Acad Sci USA. 2018;115:319–24.
Tompa P. Intrinsically unstructured proteins evolve by repeat expansion. Bioessays. 2003;25:847–55.
Bah A, Forman-Kay JD. Modulation of intrinsically disordered protein function by Post-translational Modifications. J Biol Chem. 2016;291:6696–705.
Krois AS, Dyson HJ, Wright PE. Long-range regulation of p53 DNA binding by its intrinsically disordered N-terminal transactivation domain. Proc Natl Acad Sci USA. 2018;115:E11302–10.
Contreras-Martos S, Piai A, Kosol S, Varadi M, Bekesi A, Lebrun P, et al. Linking functions: an additional role for an intrinsically disordered linker domain in the transcriptional coactivator CBP. Sci Rep. 2017;7:4676.
Bah A, Vernon RM, Siddiqui Z, Krzeminski M, Muhandiram R, Zhao C, et al. Folding of an intrinsically disordered protein by phosphorylation as a regulatory switch. Nature. 2015;519:106–9.
Thapar R. Structural basis for regulation of RNA-binding proteins by phosphorylation. ACS Chem Biol. 2015;10:652–66.
Wagner MJ, Stacey MM, Liu BA, Pawson T. Molecular mechanisms of SH2- and PTB-domain-containing proteins in receptor tyrosine kinase signaling. Cold Spring Harb Perspect Biol. 2013;5:a008987.
Jacobson RH, Ladurner AG, King DS, Tjian R. Structure and function of a human TAFII250 double bromodomain module. Science. 2000;288:1422–5.
Watson M, Stott K. Disordered domains in chromatin-binding proteins. Essays Biochem. 2019;63:147–56.
Crick SL, Jayaraman M, Frieden C, Wetzel R, Pappu RV. Fluorescence correlation spectroscopy shows that monomeric polyglutamine molecules form collapsed structures in aqueous solutions. Proc Natl Acad Sci USA. 2006;103:16764–9.
Sharma R, Raduly Z, Miskei M, Fuxreiter M. Fuzzy complexes: specific binding without complete folding. FEBS Lett. 2015;589:2533–42.
Strom AR, Emelyanov AV, Mir M, Fyodorov DV, Darzacq X, Karpen GH. Phase separation drives heterochromatin domain formation. Nature. 2017;547:241–5.
Maeshima K, Ide S, Hibino K, Sasai M. Liquid-like behavior of chromatin. Curr Opin Genet Dev. 2016;37:36–45.
Lok CN, Lang AJ, Mirski SE, Cole SP. Characterization of the human topoisomerase IIbeta (TOP2B) promoter activity: essential roles of the nuclear factor-Y (NF-Y)- and specificity protein-1 (Sp1)-binding sites. Biochem J. 2002;368:741–51.
Gyori BM, Venkatachalam G, Thiagarajan PS, Hsu D, Clement MV. OpenComet: an automated tool for comet assay image analysis. Redox Biol. 2014;2:457–65.
Jerabek-Willemsen M, Wienken CJ, Braun D, Baaske P, Duhr S. Molecular interaction studies using microscale thermophoresis. Assay Drug Dev Technol. 2011;9:342–53.
Ishida T, Kinoshita K. PrDOS: prediction of disordered protein regions from amino acid sequence. Nucleic Acids Res. 2007;35:W460–464.
Dosztanyi Z, Csizmok V, Tompa P, Simon I. IUPred: web server for the prediction of intrinsically unstructured regions of proteins based on estimated energy content. Bioinformatics. 2005;21:3433–4.
Yang ZR, Thomson R, McNeil P, Esnouf RM. RONN: the bio-basis function neural network technique applied to the detection of natively disordered regions in proteins. Bioinformatics. 2005;21:3369–76.
Ward JJ, Sodhi JS, McGuffin LJ, Buxton BF, Jones DT. Prediction and functional analysis of native disorder in proteins from the three kingdoms of life. J Mol Biol. 2004;337:635–45.
Blaszczyk M, Kurcinski M, Kouza M, Wieteska L, Debinski A, Kolinski A, et al. Modeling of protein-peptide interactions using the CABS-dock web server for binding site search and flexible docking. Methods. 2016;93:72–83.
Kurcinski M, Jamroz M, Blaszczyk M, Kolinski A, Kmiecik S. CABS-dock web server for the flexible docking of peptides to proteins without prior knowledge of the binding site. Nucleic Acids Res. 2015;43:W419–424.
Jurrus E, Engel D, Star K, Monson K, Brandi J, Felberg LE, et al. Improvements to the APBS biomolecular solvation software suite. Protein Sci. 2018;27:112–28.
Moriwaki Y, Yamane T, Ohtomo H, Ikeguchi M, Kurita J, Sato M, et al. Solution structure of the isolated histone H2A-H2B heterodimer. Sci Rep. 2016;6:24999.
Acknowledgements
This work was supported in part by an award from the National Institutes of Health (R01 CA 132115, R21 CA235139-01 to RGP), the Breast Cancer Research Breakthrough Program (#W81XWH1810605) (RGP), a grant from the Ralph and Marian C Falk Medical Research Trust (RGP) and grants from K125340 (National Research and Development Office Hungary) to AT, grant K124670 (National Research and Development Office Hungary) and the Odysseus grant G.0029.12 (F.W.O, Research Foundation Flanders) to PT.
Author information
Authors and Affiliations
Contributions
RGP designed the study; RGP, XJ, GDS and YD wrote the manuscript; RGP and XJ revised manuscripts; GDS, XJ, MCC, AWA, ZL. YQ, ADR, CP, ZZ and KA performed experiments, acquired and analyzed data; AK conducted statistical analyses., CW performed protein-protein interaction by mammalian two-hybrid reporter assay; ZY generated the microarray data set; MC generated 3T3 cells cyclin D1-/-; RV and MM performed histone peptide array; AT performed the Microscale Thermophoresis (MST) analysis; PT and AT generated the prediction of disorder tendency and multiple protein alignment; TL carried out statistical analysis of poly E regions; DKB and XJ performed densitometry analysis of histone peptide array.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiao, X., Di Sante, G., Casimiro, M.C. et al. A cyclin D1 intrinsically disordered domain accesses modified histone motifs to govern gene transcription. Oncogenesis 13, 4 (2024). https://doi.org/10.1038/s41389-023-00502-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41389-023-00502-1
- Springer Nature Limited