Abstract
There have been increasing efforts to develop prediction models supporting personalised detection, prediction, or treatment of ADHD. We overviewed the current status of prediction science in ADHD by: (1) systematically reviewing and appraising available prediction models; (2) quantitatively assessing factors impacting the performance of published models. We did a PRISMA/CHARMS/TRIPOD-compliant systematic review (PROSPERO: CRD42023387502), searching, until 20/12/2023, studies reporting internally and/or externally validated diagnostic/prognostic/treatment-response prediction models in ADHD. Using meta-regressions, we explored the impact of factors affecting the area under the curve (AUC) of the models. We assessed the study risk of bias with the Prediction Model Risk of Bias Assessment Tool (PROBAST). From 7764 identified records, 100 prediction models were included (88% diagnostic, 5% prognostic, and 7% treatment-response). Of these, 96% and 7% were internally and externally validated, respectively. None was implemented in clinical practice. Only 8% of the models were deemed at low risk of bias; 67% were considered at high risk of bias. Clinical, neuroimaging, and cognitive predictors were used in 35%, 31%, and 27% of the studies, respectively. The performance of ADHD prediction models was increased in those models including, compared to those models not including, clinical predictors (β = 6.54, p = 0.007). Type of validation, age range, type of model, number of predictors, study quality, and other type of predictors did not alter the AUC. Several prediction models have been developed to support the diagnosis of ADHD. However, efforts to predict outcomes or treatment response have been limited, and none of the available models is ready for implementation into clinical practice. The use of clinical predictors, which may be combined with other type of predictors, seems to improve the performance of the models. A new generation of research should address these gaps by conducting high quality, replicable, and externally validated models, followed by implementation research.
Similar content being viewed by others
Introduction
Attention-Deficit/Hyperactivity Disorder (ADHD) [1] is a neurodevelopmental condition which is characterized by age-inappropriate and impairing inattention and/or hyperactivity/impulsivity. Over the past decades, neurobiological research has resulted in a shift in the understanding of the pathophysiology of ADHD, from theoretical views of isolated brain dysfunctions to more complex models reflecting the heterogeneity of the clinical manifestations of ADHD [2]. However, neurobiological findings have not yet impacted clinical practice and, currently, the diagnosis of ADHD is exclusively based on a clinical assessment, with no established objective tests being available as standalone tools to diagnose ADHD [3]. The exact factors that predict the persistence of ADHD beyond adolescence are currently unclear. Furthermore, while effective (at least in the short-term) treatments are available [4], there are no established evidence-based prediction models to inform individualized treatment strategies based on the patient’s clinical, environmental, cognitive, genetic, or biological characteristics.
In the last decade, the new field of precision psychiatry has emerged, with the development of multivariable prediction models aimed at predicting the diagnosis, prognosis, or treatment response in relation to several mental health conditions [5, 6], considering individual variability in clinical characteristics, genes, environment, and lifestyle [7]. Advances in the field of prediction modelling have allowed the consolidation of an evidence-based science of precision medicine [8]. Prediction modelling studies investigate the development of such models, as well as their validation [9]. External validity is the extent to which predictions can be generalized to the data from other settings, while internal validity is the extent to which the predictions fit the derivation data [10].
Previous systematic reviews have identified a large number of prediction models across mental health conditions [10, 11]. Notably, in the last few years there has been a rapidly increasing interest in this field, and an emerging number of prediction models on ADHD have been rapidly published, making an updated evaluation of the status of the field essential. Furthermore, to our knowledge, no study has comprehensively and specifically reviewed the status of validated prediction models in ADHD, systematically assessing factors that can affect their predictive performance. Therefore, our primary aim was to systematically review and critically appraise available prediction models that might be considered for clinical use in the identification or management of ADHD. Our secondary aim was to test potential moderating factors that could affect the performance of available models as measured by their area under the curve (AUC), the most reliable and most reported metric across studies.
Methods
This study (pre-registered protocol: PROSPERO:CRD42023387502) was conducted and reported in accordance with the “Preferred Reporting Items for Systematic Reviews and Meta-analyses” (PRISMA) 2020 and the “Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis” (TRIPOD) statements and checklists (Tables S1–4, available online).
Search strategy and selection criteria
PubMed and Web of Science database including Web of Science Core Collection, BIOSIS Citation Index, KCI-Korean Journal Database, MEDLINE, Russian Science Citation Index, and SciELO Citation Index, Cochrane Central Register of Reviews, Embase and Ovid/PsycINFO databases, were searched from inception until 20/12/2023 with no language restrictions (search terms/syntax in Supplementary 1, available online). The references of the included articles and those in previous relevant reviews were manually searched to identify any possible additional relevant studies. Titles and abstracts were screened, and, after the exclusion of those not relevant, the full texts were assessed against the inclusion and exclusion criteria by a group of researchers who worked independently in pairs on one third of the hits each (GSdP, AB, AC, MD, AC, HS, VP).
The inclusion criteria were: (a) original individual studies; (b) conducted in children and/or adults with ADHD according to established diagnostic criteria (DSM or ICD—any version); (c) reporting on multivariable internally and/or externally [12] validated prediction models; (d) providing diagnostic, prognostic, or treatment-response estimates at the individual subject level or in subgroups; (e) providing at least discrimination as per the AUC (i.e., the ability of the model to separate individuals who develop events from those who do not), accuracy (i.e., the degree of closeness of the measured value), or classification measures (sensitivity, specificity, or predictive values) (definitions in Table 1). The exclusion criteria were: (a) abstracts, conference proceedings, reviews, or meta-analyses; (b) prediction model studies that did not evaluate or report their internal or external validation; (c) predictor-finding studies that included one predictor only.
Descriptive measures and data extraction
Data extraction items (Supplementary 2, available online) were based on the “Checklist for critical Appraisal and data extraction for systematic Reviews of prediction Modelling Studies” (CHARMS) and the “Transparent Reporting of a multivariable prediction model for Individual Prognosis Or Diagnosis” (TRIPOD) statements. The model’s ability to separate individuals with and without the outcome, e.g., AUC, was selected as the main outcome. Discriminative validity is usually considered ‘acceptable’ when AUC scores are between 0.7–0.8, ‘good’ between 0.8–0.9, and ‘excellent’ when >0.9 [13]. We extracted information on the performance of each model assessed by other measures when reported. When more than one outcome per study was found in the same category, we extracted the information for the primary outcome, as defined in each article, unless the study reported multiple primary co-outcomes. We relied on what the individual authors reported as their primary outcome.
Quality assessment
Risk of bias was assessed for each of the included studies with a validated version - previously used in mental health research- of the Prediction Model Risk of Bias Assessment Tool (PROBAST v5/05/2019) [9] (Supplementary 3, available online).
Strategy for data synthesis
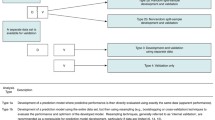
Data from the included studies were first summarized in descriptive tables. The top 10% of the most commonly employed predictor types were shown in a bar chart. We then conducted meta-regressions to estimate the association, when data were available, between AUC and: (i) the type of validation (internal vs external); (ii) the age range (children and adolescents vs adults vs combined/not reported); (iii) the type of model (diagnostic vs prognostic vs treatment-response model); (iv) the number of predictors; (v) the type of predictors [clinical/sociodemographic vs any biomarker (neuroimaging, electroencephalography, magnetoencephalography, proteomic, genetic, cognitive, or a combination of modalities)] [10]; (vi) the modality of predictors [unimodal, using only one type of predictor (e.g., clinical only) vs multimodal, using more than one type of predictor (e.g., clinical and biomarker)] and (vii) the quality of the studies (low risk vs unclear risk vs high risk). We used a random-effects model to allow for heterogeneity in underlying associations across studies. Number of studies permitting, we also planned sensitivity analyses to assess the impact of studies being at low risk of bias and without suboptimal validation. Suboptimal validation was appraised by two statisticians (RI and MHI) with a focus on: (1) double dipping (i.e., performing feature selection or selection of tuning -or penalty- parameters on data samples from both the training and the test set) [14]; (2) reporting apparent/non-validated predictive performance instead of the validated predictive performance; (3) reporting the size and significance of apparent regression coefficients rather than the cross-validated performance measure, and (4) re-estimating regression coefficients in the test set, instead of applying the apparently validated model. The meta-regression was performed with Comprehensive Meta-Analysis Version 3. Statistical significance was set at p < 0.05.
Results
After removing duplicates, from an initial pool of 7764 references, we retained 100 eligible studies (Fig. 1). None of the models reported in the included studies was implemented into clinical practice. 96 (96.0%) and seven (7.0%) models were internally and externally validated, respectively. Among the eligible studies, 88.0% reported on diagnostic prediction models, 5.0% on prognostic models (with outcomes such as symptom change or development of substance use disorders), and 7.0% on treatment-response models. The retained studies most frequently used clinical (35.0%), neuroimaging (31.0%) and cognitive (27.0%), predictors (Fig. 2). The total sample size was 323,554 individuals, ranging from 10 to 238,696 individuals per study. The average age was 15.7 years. The source of data encompassed case-control studies (73 studies, 73.0%), cohort studies (23 studies, 23.0%), and clinical trials (4 studies, 4.0%). AUC was the most commonly reported measure of model performance (61.0%), followed by accuracy (36.0%). Eight studies (8.0%) only reported the sensitivity and specificity of the models.
Predictors in prediction models
In the 88 diagnostic prediction models, studies used cognitive (K = 5 studies) [15,16,17,18,19], clinical (K = 13) [20,21,22,23,24,25,26,27,28,29,30,31]287, neuroimaging (K = 19) [32,33,34,35,36,37,38,39,40,41,42,43,44,45,46,47,48,49,50], EEG (K = 15) [51,52,53,54,55,56,57,58,59,60,61,62,63,64,65], genetic (K = 2) [66, 67], ECG (K = 2) [68, 69], physical health (K = 1) [70], EEG and cognitive (K = 4) [71,72,73,74], sociodemographic and neuroimaging (K = 4) [75,76,77]294, clinical and cognitive (K = 4) [78,79,80,81], sociodemographic and cognitive (K = 2) [82, 83], cognitive and physical health (K = 2) [84, 85], genetic and neuroimaging (K = 2) [86, 87], sociodemographic and genetic (K = 1) [88], clinical and sociodemographic (K = 1) [89] EEG and EMG (K = 1) [90], sociodemographic and neuroimaging (K = 2) [91, 92], sociodemographic, clinical and cognitive (K = 3) [93,94,95], cognitive, sociodemographic and neuroimaging (K = 1) [96], clinical, sociodemographic and neuroimaging (k = 1) [97], clinical, cognitive and neuroimaging (k = 1) [98] and sociodemographic, clinical, cognitive and physical health (K = 2) [99, 100] predictors.
In the 5 prognostic prediction models, studies employed sociodemographic and clinical (K = 2) [101, 102], physical health and clinical (K = 1) [103], neuroimaging and genetic (K = 1) [104], and clinical and genetic (K = 1) [105] predictors.
In the 7 treatment-response prediction models, studies relied on neuroimaging (K = 1) [106], genetic (k = 1) [107], sociodemographic, clinical, cognitive, and physical health (K = 1) [108], genetic, cognitive and physical health (K = 1) [109], clinical, sociodemographic, service use and physical health (K = 1) [110], sociodemographic and clinical predictors (K = 1) [111], and sociodemographic, clinical and physical health (K = 1) predictors [112].
Performance of prediction models
The performance of ADHD prediction models was highly variable, with AUC ranging from 0.50 to 0.99. AUC ranged from 0.50 to 0.96 in diagnostic models, from 0.73 to 0.87 in prognostic models, and from 0.72 to 0.99 in models for predicting treatment-response. Accuracy ranged from 0.53 to 1.0 (0.53 to 1.0 for diagnostic models, 0.73 to 0.87 for prognostic models, and 0.72 to 0.88 for treatment-response models) (Tables S5–7, available online). Model calibration was assessed in 6.0% of the studies.
Meta-regression results
The performance of ADHD prediction models was increased in those models including (K = 26), as compared to those not including, clinical predictors (K = 36) (β = 6.540, p = 0.007). No significant findings emerged when considering type of validation (internal K = 58 vs external K = 4), age range (children and adolescents K = 33 vs adults K = 11 vs combined/not reported K = 18), type of prediction model (diagnostic K = 52 vs prognostic K = 3 vs treatment-response K = 7) (p > 0.05), number of predictors (K = 34), other types of predictors comparisons (clinical/service use/sociodemographic K = 13 vs biomarkers K = 33 vs combination K = 17) or quality of the studies (low risk K = 7 vs unclear risk K = 11 vs high risk K = 44) (all p > 0.05) (Table 2), according to our meta-regression analyses.
Quality of prediction models
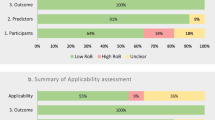
Sixty-seven (67.0%) of the included studies were deemed to be at high risk of bias according to the PROBAST tool. The results from the different domains were heterogeneous: 9 (9.0%) were at high risk of bias in the participants domain, 11 (11.0%) in the predictors domain, 19 (19.0%) in the outcomes domain, and 61 (61.0%) in the analysis domain. Only 8 (8.0%) of the included studies (seven diagnostic and one prognostic) were considered to be at overall low risk of bias; 86 (86.0%) of the studies were deemed at low risk of bias in the participants domain, 60 (60.0%) in the predictors domain, 57 (57.0%) in the outcomes domain, and 18 (18.0%) in the analysis domain (Table S8, available online; Fig. 3). In 13 studies (13.0%) the performance was evaluated in development dataset only, resulting in suboptimal validation (Table S9, available online).
Discussion
This is the first systematic review to quantitatively summarize the evidence regarding internally or externally validated diagnostic, prognostic, or treatment-response prediction models specifically in the field of ADHD, appraising the quality of the models and assessing possible factors affecting their performance in terms of AUC. Among the 100 prediction modelling studies included, 88% reported on diagnostic, 5% on prognostic, and 7% on treatment-response models. Furthermore, 35% of studies used clinical, 31.0% neuroimaging, and 27.0% cognitive predictors. Notably, only 7.0% of models were externally validated. The performance of ADHD prediction models was increased in those models including, compared to those models not including, clinical predictors. Meta-regressions did not detect any significant changes in the AUC according to other evaluated variables. Also, 67.0% of included studies were found to be at high risk of bias according to PROBAST quality assessment.
Our review shows that the number of prediction models in the field of ADHD is increasing exponentially over the years, with a wide range of predictors that might potentially support the diagnosis of ADHD, and, to a lesser extent, the understanding of the clinical progression of the disorder or the factor influencing the response to interventions.
However, the discrimination and accuracy of the models, although good, may not be enough for implementation into clinical practice. This emerging body of research is limited by not only a small number of externally validated models, but also, and crucially, by lack of implementation research in real-world clinical practice. Our findings align with previous evidence related to other mental health conditions suggesting that external validation of prediction models is still infrequent in psychiatry/mental health [113]. A similar review exploring prediction models across any mental health condition found that only 20.1% of all prediction models were externally validated [11]. Another review found that 30.3% of all models were externally validated following strict validation criteria (4.6% of the total models) [10]. This is in contrast with the status of prediction science in other areas of medicine. For instance, several models have been externally validated between five to seventeen times in the field of chronic obstructive pulmonary disease [114]. Similar approaches may move the field of ADHD forward, ensuring generalizability of the model to clinical populations not used to develop the model.
Within the internally and externally validated models, there was no significant correlation seen between the internal and external performance measures. However, the number of models internally and externally validated (six studies) was low. Our findings may also reflect a suboptimal quality during the internal validation of the models, potentially leading to optimism in the reported performance measures and high risk of bias. In fact, 67.0% of the included studies were found to be at high risk of bias. The “analysis” domain, where 61.0% of the included studies were found to be at high risk of bias seems particularly problematic. Also, calibration was assessed in 6.0% of the studies only. Future prediction models need to make sure that: (1) the sample size is appropriate and there is an appropriate number of participants developing the outcome (which may vary depending on the population and outcome of interest); (2) the number of predictors is appropriate, (3) the missing data is handled appropriately (of note, >80% studies did not report how they handled missing data, 7.0% deleted missing data and only 4.0% carried out multiple imputation techniques, being low adherence to ADHD treatment frequent); (4) complexities in the data (e.g. competing risks, sampling of controls) are accounted for appropriately, and (5) model overfitting and optimism in model performance are accounted for, among other key criteria to develop and validate prediction models [9]. We note that risk of bias was heterogeneous across the different PROBAST domains: only 9.0% of models were at high risk of bias in the participants domain and only 11.0% were at high risk of bias in the predictors domain. Given the strictness of PROBAST scoring thresholds, this highlights some strengths among published ADHD prediction models in the selection of participants and predictors.
In terms of the aim of the models, most of them were intended to support the prediction of ADHD diagnosis (88/100), reflecting an increasing interest in developing diagnostic prediction models in the ADHD field, alongside other mental health conditions such as depression [115], first episode psychosis [116], or bipolar disorder [117], following a similar route, likely due to the perception of the suboptimal nature of a “subjective” diagnosis. Notably, unlike other mental health conditions, where performance in diagnostic models has been found to be superior to that in prognostic and treatment-response models [10], we did not find this to be the case in the field of ADHD. The limited number of available treatment-response models points to a critical need for carefully designed experimentally controlled trials (or high quality observational studies) to identify biomarkers that index inter-individual variability and predict treatment response [118]. While studies on treatment-response models are complex to perform, mostly due to the intervention-related components (particularly randomized clinical trials), as well as to ethical issues [10], observational studies relying on electronic health-care records on the long-term effectiveness and safety of the interventions could provide a meaningful alternative [119].
In terms of predictor types, a significant proportion of the reviewed prediction models included clinical predictors, followed closely by neuroimaging predictors and cognitive predictors. The performance of ADHD prediction models was higher in those models including clinical predictors, compared to those models not including clinical predictors. Thus, the use of clinical predictors, which may be combined with other type of predictors, may improve the performance of the models and their inclusion should be considered in prediction models [81]. However, it is important to note that further research is needed to validate these results across different populations, and including additional predictors [99]. While clinical predictors seem to be clearly predominant in other fields [10], in the ADHD field different biomarkers have commonly been used to aid the detection and correct characterization of ADHD. However, there is currently no biomarker in any neurodevelopmental condition, including ADHD, for which there is evidence from two or more studies from independent research groups, with results going into the same direction and of specificity and sensitivity of at least 80% [3]. This makes it difficult to recommend the use of any specific individual predictor in isolation, for future prediction models. Notably, we also found no evidence that multimodal prediction models achieved higher accuracy than unimodal models, arguing against the development of complex models with a wide variety of biomarkers and predictors (which would also be more difficult to apply and implement). In other words, we found no evidence that more complex prediction models encompassing biomarkers or a large number of predictors (which may be more prone to overfitting issues) outperformed less complex models. However, from a quality perspective, five of the six studies assessed at low risk of bias were multimodal, so caution is recommended in the interpretation of this finding.
Future studies should consider net benefit approaches for the evaluation of prediction models for ADHD, which were not used in any of the studies in this review. Net benefit approaches put the benefits and harms of using a prediction model on the same scale, to allow assessment of the relative value associated with using prediction models to guide clinical decision making, over other patient management strategies [120] an approach which is currently lacking in the ADHD prediction literature. 74% of the studies were case-control studies which tried to differentiate individuals with ADHD and healthy controls. Future studies should also try to differentiate ADHD from other relevant syndromes such as the cognitive disengagement syndrome (CDS) -or sluggish cognitive tempo-. CDS is an emerging condition -as opposed to a transdiagnostic phenomenon- in the field of child, adolescent and adult psychiatry [121, 122]. The presence of CDS is particularly important as misdiagnosis of this condition may result in a poor response to first-line treatment with methylphenidate and unwanted side effects [123]. Furthermore, among children crossing into adolescence with ADHD, CDS can result in poor physical activity and behaviour [122].
Our study should be considered in the light of its limitations. Our study has several limitations that must be taken into consideration, mainly related to issues in the available studies rather than in our methods. The main limitation rests in the heterogeneity of the characteristics of prediction models developed in the included studies. The predictors used to develop the models varied considerably across studies. Therefore, in line with previous studies [10], we did not attempt to meta-analyse the categories of prediction models; rather, we presented only meta-regression analyses, stratifying the models for methodological features. We also could not conduct meta-regressions on the studies at low risk of bias and without suboptimal methodological strategies in regard to validation. The sample size and the quality of the studies was highly heterogeneous, with high risk of bias observed in 67.0% of included studies according to the PROBAST criteria, including 61.0% in the analysis domain. Final scores of the PROBAST should be taken with caution as the thresholds are stringent and an outcome is considered to be at high risk of bias when one or more of the questions is answered as not appropriate. We did not analyse the differences among validation measures, some of them being prone to data leakage and inflated accuracy or overfitting. We might have missed relevant studies, particularly if not published. Finally, we could not provide data about calibration as this was rarely reported.
In conclusion, several validated prediction models have been proposed to support the diagnosis of ADHD. However, efforts to predict prognostic outcomes or treatment response to ADHD have been limited. Advances in the field are limited by lack of implementation research in real-world clinical practice. A new generation of research should address these gaps by conducting high quality, replicable and externally validated models. Once an evidence-based model is available, efforts to disseminate it and implement it into clinical practice are recommended.
References
American Psychiatric Association. Washington DC. Diagnostic and statistical manual of mental disorders. 2013.
Cortese S. The neurobiology and genetics of Attention-Deficit/Hyperactivity Disorder (ADHD): What every clinician should know. Eur J Paediatr Neurol. 2012;16:422–33.
Cortese S, Solmi M, Michelini G, Bellato A, Blanner C, Canozzi A, et al. Candidate diagnostic biomarkers for neurodevelopmental disorders in children and adolescents: a systematic review. World Psychiatry. 2023;22:129–49.
Cortese S. Pharmacologic treatment of attention deficit-hyperactivity disorder. N. Engl J Med. 2020;383:1050–6.
Fernandes BS, Williams LM, Steiner J, Leboyer M, Carvalho AF, Berk M. The new field of ‘precision psychiatry. BMC Med. 2017;15:80.
Khanra S, Khess CRJ, Munda SK. Precision psychiatry”: a promising direction so far. Indian J Psychiatry. 2018;60:373–4.
Terry SF. Obama’s precision medicine initiative. Genet Test Mol Biomark. 2015;19:113–4.
Fusar-Poli P, Hijazi Z, Stahl D, Steyerberg EW. The science of prognosis in psychiatry: a review. JAMA Psychiatry. 2018;75:1289–97.
Wolff RF, Moons KGM, Riley RD, Whiting PF, Westwood M, Collins GS, et al. PROBAST: a tool to assess the risk of bias and applicability of prediction model studies. Ann Intern Med. 2019;170:51–58.
Salazar de Pablo G, Studerus E, Vaquerizo-Serrano J, Irving J, Catalan A, Oliver D, et al. Implementing precision psychiatry: a systematic review of individualized prediction models for clinical practice. Schizophr Bull. 2021;47:284–97.
Meehan AJ, Lewis SJ, Fazel S, Fusar-Poli P, Steyerberg EW, Stahl D, et al. Clinical prediction models in psychiatry: a systematic review of two decades of progress and challenges. Mol Psychiatry. 2022;27:2700–8.
Royston P, Altman DG. External validation of a Cox prognostic model: principles and methods. BMC Med Res Methodol. 2013;13:33.
Safari S, Baratloo A, Elfil M, Negida A. Evidence based emergency medicine; part 5 receiver operating curve and area under the curve. Emerg (Tehran). 2016;4:111–3.
Kassraian-Fard P, Matthis C, Balsters JH, Maathuis MH, Wenderoth N. Promises, pitfalls, and basic guidelines for applying machine learning classifiers to psychiatric imaging data, with autism as an example. Front Psychiatry. 2016;7:177.
Altinkaynak M, Dolu N, Guven A, Pektas F, Ozmen S, Demirci E, et al. Diagnosis of attention deficit hyperactivity disorder with combined time and frequency features. Biocybern Biomed Eng. 2020;40:927–37.
Finch HW, Davis A, Dean RS. Identification of individuals with ADHD using the Dean-Woodcock sensory motor battery and a boosted tree algorithm. Behav Res Methods. 2015;47:204–15.
Lee HJ, Cho S, Shin MS. Supporting diagnosis of attention-deficit hyperactive disorder with novelty detection. Artif Intell Med. 2008;42:199–212.
Predicting ADHD Risk from Touch Interaction Data. Proceedings of the 20th ACM International Conference on Multimodal Interaction (ICMI); 2018 Oct 16-20 2018; Boulder, CO2018.
Slobodin O, Yahav I, Berger I. A Machine-Based Prediction Model of ADHD Using CPT Data. Front Hum Neurosci. 2020;14:560021.
Altun S, Alkan A, Altun H. Application of deep learning and classical machine learning methods in the diagnosis of attention deficit hyperactivity disorder according to temperament features. Concurrency Comput Pract Experience. 2022; 34.
Bledsoe JC, Xiao C, Chaovalitwongse A, Mehta S, Grabowski TJ, Semrud-Clikeman M, et al. Diagnostic classification of ADHD versus control: support vector machine classification using brief neuropsychological assessment. J Atten Disord. 2020;24:1547–56.
Deserno MK, Bathelt J, Groenman AP, Geurts HM. Probing the overarching continuum theory: data-driven phenotypic clustering of children with ASD or ADHD. Eur Child Adolesc Psychiatry. 2023;32:1909–23
Duda M, Ma R, Haber N, Wall DP. Use of machine learning for behavioral distinction of autism and ADHD. Transl Psychiatry. 2016;6:e732.
Duda M, Haber N, Daniels J, Wall DP. Crowdsourced validation of a machine-learning classification system for autism and ADHD. Transl Psychiatry. 2017;7:e1133.
Kim S, Lee HK, Lee K. Can the MMPI predict adult ADHD? an approach using machine learning methods. Diagnostics (Basel). 2021;11:976.
Shi Y, Schulte PJ, Hanson AC, Zaccariello MJ, Hu D, Crow S, et al. Utility of medical record diagnostic codes to ascertain attention-deficit/hyperactivity disorder and learning disabilities in populations of children. BMC Pediatr. 2020;20:510.
Silverstein M, Hironaka LK, Feinberg E, Sandler J, Pellicer M, Chen N, et al. Using clinical data to predict accurate ADHD diagnoses among urban children. Clin Pediatr (Philos). 2016;55:326–32.
Chen T, Tachmazidis I, Batsakis S, Adamou M, Papadakis E, Antoniou G. Diagnosing attention-deficit hyperactivity disorder (ADHD) using artificial intelligence: a clinical study in the UK. Front Psychiatry. 2023;14:1164433.
Chu KC, Huang HJ, Huang YS. Validity of diagnostic support model for attention deficit hyperactivity disorder: a machine learning approach. J Pers Med. 2023;13:1525.
Haque UM, Kabir E, Khanam R. Early detection of paediatric and adolescent obsessive-compulsive, separation anxiety and attention deficit hyperactivity disorder using machine learning algorithms. Health Inf Sci Syst. 2023;11:31.
Kurokami T, Kobayashi H, Nakajima M, Mikami M, Koeda T. Establishment of an objective index for the diagnosis of attention deficit/hyperactivity disorder by the continuous performance test “MOGRAZ. Brain Dev. 2022;44:664–71.
Bohland JW, Saperstein S, Pereira F, Rapin J, Grady L. Network, anatomical, and non-imaging measures for the prediction of ADHD diagnosis in individual subjects. Front Syst Neurosci. 2012.
Chaim-Avancini TM, Doshi J, Zanetti MV, Erus G, Silva MA, Duran FLS, et al. Neurobiological support to the diagnosis of ADHD in stimulant-naïve adults: pattern recognition analyses of MRI data. Acta Psychiatr Scand. 2017;136:623–36.
Cheng W, Ji X, Zhang J, Feng J. Individual classification of ADHD patients by integrating multiscale neuroimaging markers and advanced pattern recognition techniques. Front Syst Neurosci. 2012;6:58.
Dai D, Wang J, Hua J, He H. Classification of ADHD children through multimodal magnetic resonance imaging. Front Syst Neurosci. 2012;6:63.
Hart H, Chantiluke K, Cubillo AI, Smith AB, Simmons A, Brammer MJ, et al. Pattern classification of response inhibition in ADHD: toward the development of neurobiological markers for ADHD. Hum brain Mapp. 2014;35:3083–94.
Iannaccone R, Hauser TU, Ball J, Brandeis D, Walitza S, Brem S. Classifying adolescent attention-deficit/hyperactivity disorder (ADHD) based on functional and structural imaging. Eur child Adolesc psychiatry. 2015;24:1279–89.
Johnston BA, Mwangi B, Matthews K, Coghill D, Konrad K, Steele JD. Brainstem abnormalities in attention deficit hyperactivity disorder support high accuracy individual diagnostic classification. Hum brain Mapp. 2014;35:5179–89.
Lan Z, Sun Y, Zhao L, Xiao Y, Kuai C, Xue SW. Aberrant effective connectivity of the ventral putamen in boys with attention-deficit/hyperactivity disorder. Psychiatry Investig. 2021;18:763–9.
Differentiation between Resting-State fMRI data from ADHD and Normal Subjects : Based on Functional Connectivity and Machine Learning. Proceedings of the International Conference on Fuzzy Theory and Its Applications (iFUZZY); 2012 Nov 16-18 2012; Natl Chung Hsing Univ, Taichung, TAIWAN2012.
Luo Y, Alvarez TL, Halperin JM, Li X. Multimodal neuroimaging-based prediction of adult outcomes in childhood-onset ADHD using ensemble learning techniques. NeuroImage Clin. 2020;26:102238.
Mao Z, Su Y, Xu G, Wang X, Huang Y, Yue W, et al. Spatio-temporal deep learning method for ADHD fMRI classification. Inf Sci. 2019;499:1–11.
McNorgan C, Judson C, Handzlik D, Holden JG. Linking ADHD and behavioral assessment through identification of shared diagnostic task-based functional connections. Front Physiol. 2020;11:583005.
Sen B, Borle NC, Greiner R, Brown MRG. A general prediction model for the detection of ADHD and Autism using structural and functional MRI. PLoS One. 2018;13:e0194856.
Sun Y, Zhao L, Lan Z, Jia X-Z, Xue S-W. Differentiating boys with ADHD from those with typical development based on whole-brain functional connections using a machine learning approach. Neuropsychiatr Dis Treat. 2020;16:691–702.
Chen M, Li H, Fan H, Dillman JR, Wang H, Altaye M, et al. ConCeptCNN: a novel multi-filter convolutional neural network for the prediction of neurodevelopmental disorders using brain connectome. Med Phys. 2022;49:3171–84.
de Lacy N, Ramshaw MJ, McCauley E, Kerr KF, Kaufman J, Nathan Kutz J. Predicting individual cases of major adolescent psychiatric conditions with artificial intelligence. Transl Psychiatry. 2023;13:314.
Gaus R, Pölsterl S, Greimel E, Schulte-Körne G, Wachinger C. Can we diagnose mental disorders in children? A large-scale assessment of machine learning on structural neuroimaging of 6916 children in the adolescent brain cognitive development study. JCPP Adv. 2023;3:e12184.
Saha P, Sarkar D. Characterization and classification of ADHD subtypes: an approach based on the nodal distribution of eigenvector centrality and classification Tree Model. Child Psychiatry Hum Dev. 2024;55:622–34.
Yang CM, Shin J, Kim JI, Lim YB, Park SH, Kim BN. Classifying children with ADHD based on prefrontal functional near-infrared spectroscopy using machine learning. Clin Psychopharmacol Neurosci. 2023;21:693–700.
Chen M, Li H, Wang J, Dillman JR, Parikh NA, He L. A multichannel deep neural network model analyzing multiscale functional brain connectome data for attention deficit hyperactivity disorder detection. Radio Artif Intell. 2019;2:e190012.
Chen H, Song Y, Li X. A deep learning framework for identifying children with ADHD using an EEG-based brain network. Neurocomputing. 2019;356:83–96.
Dubreuil-Vall L, Ruffini G, Camprodon JA. Deep learning convolutional neural networks discriminate adult ADHD from healthy individuals on the basis of event-related spectral EEG. Front Neurosci. 2020;14:251.
An Auditory Brainstem Response-Based Expert System for ADHD Diagnosis Using Recurrence Qualification Analysis and Wavelet Support Vector Machine. Proceedings of the 23rd Iranian Conference on Electrical Engineering; 2015 May 10-14 2015; Sharif Univ of Technol, Tehran, IRAN2015.
Jahanshahloo HR, Shamsi M, Ghasemi E, Kouhi A. Automated and ERP-based diagnosis of attention-deficit hyperactivity disorder in children. J Med Signals Sens. 2017;7:26–32.
Kaur S, Singh S, Arun P, Kaur D, Bajaj M. Phase space reconstruction of EEG signals for classification of ADHD and control adults. Clin EEG Neurosci. 2020;51:102–13.
Kiiski H, Bennett M, Rueda-Delgado LM, Farina FR, Knight R, Boyle R, et al. EEG spectral power, but not theta/beta ratio, is a neuromarker for adult ADHD. Eur J Neurosci. 2020;51:2095–109.
Maniruzzaman M, Shin J, Hasan MAM, Yasumura A. Efficient feature selection and machine learning based ADHD detection using EEG signal. Cmc-Computers Mater Contin. 2022;72:5179–95.
Moghaddari M, Lighvan MZ, Danishvar S. Diagnose ADHD disorder in children using convolutional neural network based on continuous mental task EEG. Comput Methods Prog Biomed. 2020;197:105738.
Mueller A, Vetsch S, Pershin I, Candrian G, Baschera G-M, Kropotov JD, et al. EEG/ERP-based biomarker/neuroalgorithms in adults with ADHD: Development, reliability, and application in clinical practice. World J Biol Psychiatry. 2020;21:172–82.
Oztoprak H, Toycan M, Alp YK, Arikan O, Dogutepe E, Karakas S. Machine-based classification of ADHD and nonADHD participants using time/frequency features of event-related neuroelectric activity. Clin Neurophysiol. 2017;128:2400–10.
Pereda E, García-Torres M, Melián-Batista B, Mañas S, Méndez L, González JJ. The blessing of Dimensionality: Feature Selection outperforms functional connectivity-based feature transformation to classify ADHD subjects from EEG patterns of phase synchronisation. PLoS One. 2018;13:e0201660.
Tenev A, Markovska-Simoska S, Kocarev L, Pop-Jordanov J, Mueller A, Candrian G. Machine learning approach for classification of ADHD adults. Int J Psychophysiol. 2014;93:162–6.
Ahire N, Awale RN, Wagh A. Electroencephalogram (EEG) based prediction of attention deficit hyperactivity disorder (ADHD) using machine learning. Appl Neuropsychol Adult. 2023;30:1–2.
Chugh N, Aggarwal S, Balyan A. The hybrid deep learning model for identification of attention-deficit/hyperactivity disorder using EEG. Clin EEG Neurosci. 2024;55:22–33.
Wang X-H, Jiao Y, Li L. Identifying individuals with attention deficit hyperactivity disorder based on temporal variability of dynamic functional connectivity. Sci Rep. 2018;8:11789.
Zhu P, Pan J, Cai QQ, Zhang F, Peng M, Fan XL et al. MicroRNA profile as potential molecular signature for attention deficit hyperactivity disorder in children. Biomarkers. 2022;27:230–9.
Koh JEW, Ooi CP, Lim-Ashworth NS, Vicnesh J, Tor HT, Lih OS, et al. Automated classification of attention deficit hyperactivity disorder and conduct disorder using entropy features with ECG signals. Comput Biol Med. 2022;140:105120.
Loh HW, Ooi CP, Oh SL, Barua PD, Tan YR, Molinari F, et al. Deep neural network technique for automated detection of ADHD and CD using ECG signal. Comput Methods Prog Biomed. 2023;241:107775.
Lindhiem O, Goel M, Shaaban S, Mak KJ, Chikersal P, Feldman J, et al. Objective measurement of hyperactivity using mobile sensing and machine learning: pilot study. JMIR Form Res. 2022;6:e35803.
Abibullaev B, An J. Decision support algorithm for diagnosis of ADHD using electroencephalograms. J Med Syst. 2012;36:2675–88.
Biederman J, Hammerness P, Sadeh B, Peremen Z, Amit A, Or-Ly H, et al. Diagnostic utility of brain activity flow patterns analysis in attention deficit hyperactivity disorder. Psychol Med. 2017;47:1259–70.
Mueller A, Candrian G, Kropotov JD, Ponomarev VA, Baschera G-M. Classification of ADHD patients on the basis of independent ERP components using a machine learning system. Nonlinear Biomed Phys. 2010;4:S1–S1.
Mueller A, Candrian G, Grane VA, Kropotov JD, Ponomarev VA, Baschera G-M. Discriminating between ADHD adults and controls using independent ERP components and a support vector machine: a validation study. Nonlinear Biomed Phys. 2011;5:5–5.
Eloyan A, Muschelli J, Nebel MB, Liu H, Han F, Zhao T, et al. Automated diagnoses of attention deficit hyperactive disorder using magnetic resonance imaging. Front Syst Neurosci. 2012;6:61.
Emser TS, Johnston BA, Steele JD, Kooij S, Thorell L, Christiansen H. Assessing ADHD symptoms in children and adults: evaluating the role of objective measures. Behav Brain Funct. 2018;14:11.
Itani S, Rossignol M, Lecron F, Fortemps P. Towards interpretable machine learning models for diagnosis aid: A case study on attention deficit/hyperactivity disorder. PLoS One. 2019;14:e0215720.
Han D, Fang Y, Luo H. A predictive model offor attention deficit hyperactivity disorder based on clinical assessment tools. Neuropsychiatr Dis Treat. 2020;16:1331–7.
Öztekin I, Finlayson MA, Graziano PA, Dick AS. Is there any incremental benefit to conducting neuroimaging and neurocognitive assessments in the diagnosis of ADHD in young children? A machine learning investigation. Dev Cogn Neurosci. 2021;49:100966.
Yeh S-C, Lin S-Y, Wu EH-K, Zhang K-F, Xiu X, Rizzo A, et al. A virtual-reality system integrated with neuro-behavior sensing for attention-deficit/hyperactivity disorder intelligent assessment. IEEE Trans Neural Syst Rehabilitation Eng. 2020;28:1899–907.
Mooney MA, Neighbor C, Karalunas S, Dieckmann NF, Nikolas M, Nousen E, et al. Prediction of attention-deficit/hyperactivity disorder diagnosis using brief, low-cost clinical measures: a competitive model evaluation. Clin Psychol Sci. 2023;11:458–75.
Acosta-Lopez JE, Suarez I, Pineda DA, Cervantes-Henriquez ML, Martinez-Banfi ML, Lozano-Gutierrez SG et al. Impulsive and omission errors: potential temporal processing endophenotypes in ADHD. Brain Sci. 2021;11:1218.
Cervantes-Henriquez ML, Acosta-Lopez JE, Martinez-Banfi ML, Velez JI, Mejia-Segura E, Lozano-Gutierrez SG, et al. ADHD endophenotypes in Caribbean families. J Atten Disord. 2020;24:2100–14.
Crippa A, Salvatore C, Molteni E, Mauri M, Salandi A, Trabattoni S, et al. The utility of a computerized algorithm based on a multi-domain profile of measures for the diagnosis of attention deficit/hyperactivity disorder. Front Psychiatry. 2017;8:189.
Mwamba HM, Fourie PR, den Heever DV. PANDAS: paediatric attention-deficit/hyperactivity disorder application software. Annu Int Conf IEEE Eng Med Biol Soc. 2019;2019:1444–7.
Kautzky A, Vanicek T, Philippe C, Kranz GS, Wadsak W, Mitterhauser M, et al. Machine learning classification of ADHD and HC by multimodal serotonergic data. Transl Psychiatry. 2020;10:104.
Yoo JH, Kim JI, Kim BN, Jeong B. Exploring characteristic features of attention-deficit/hyperactivity disorder: findings from multi-modal MRI and candidate genetic data. Brain Imaging Behav. 2020;14:2132–47.
Cervantes-Henríquez ML, Acosta-López JE, Martinez AF, Arcos-Burgos M, Puentes-Rozo PJ, Vélez JI. Machine Learning Prediction of ADHD Severity: Association and Linkage to ADGRL3, DRD4, and SNAP25. J Atten Disord. 2022;26:587–605.
Christiansen H, Chavanon M-L, Hirsch O, Schmidt MH, Meyer C, Mueller A et al. Use of machine learning to classify adult ADHD and other conditions based on the Conners’ Adult ADHD Rating Scales. Sci Rep. 2020;10:18871.
Muthuraman M, Moliadze V, Boecher L, Siemann J, Freitag CM, Groppa S, et al. Multimodal alterations of directed connectivity profiles in patients with attention-deficit/hyperactivity disorders. Sci Rep. 2019;9:20028.
Brown MRG, Sidhu GS, Greiner R, Asgarian N, Bastani M, Silverstone PH, et al. ADHD-200 global competition: diagnosing ADHD using personal characteristic data can outperform resting state fMRI measurements. Front Syst Neurosci. 2012;6:69–69.
Caye A, Agnew-Blais J, Arseneault L, Goncalves H, Kieling C, Langley K et al. A risk calculator to predict adult attention-deficit/hyperactivity disorder: generation and external validation in three birth cohorts and one clinical sample. Epidemiol Psychiatric Sci. 2020;29:e37.
Chen T, Antoniou G, Adamou M, Tachmazidis I, Su P. Automatic diagnosis of attention deficit hyperactivity disorder using machine learning. Appl Artif Intell. 2021;35:657–69.
Mikolas P, Vahid A, Bernardoni F, Süß M, Martini J, Beste C, et al. Training a machine learning classifier to identify ADHD based on real-world clinical data from medical records. Sci Rep. 2022;12:12934.
Altun S, Alkan A, Altun H. Automatic diagnosis of attention deficit hyperactivity disorder with continuous wavelet transform and convolutional neural network. Clin Psychopharmacol Neurosci. 2022;20:715–24.
Yasumura A, Omori M, Fukuda A, Takahashi J, Yasumura Y, Nakagawa E, et al. Applied machine learning method to predict children with ADHD using prefrontal cortex activity: a multicenter study in Japan. J Atten Disord. 2020;24:2012–20.
Lohani DC, Rana B. ADHD diagnosis using structural brain MRI and personal characteristic data with machine learning framework. Psychiatry Res Neuroimaging. 2023;334:111689.
Öztekin I, Garic D, Bayat M, Hernandez ML, Finlayson MA, Graziano PA, et al. Structural and diffusion-weighted brain imaging predictors of attention-deficit/hyperactivity disorder and its symptomology in very young (4- to 7-year-old) children. Eur J Neurosci. 2022;56:6239–57.
Garcia-Argibay M, Zhang-James Y, Cortese S, Lichtenstein P, Larsson H, Faraone SV. Predicting childhood and adolescent attention-deficit/hyperactivity disorder onset: a nationwide deep learning approach. Mol Psychiatry. 2022;28:1232–9.
Ehrig L, Wagner AC, Wolter H, Correll CU, Geisel O, Konigorski S. FASDetect as a machine learning-based screening app for FASD in youth with ADHD. NPJ Digit Med. 2023;6:130.
Zhang Y, Sun Y, Yu Z, Sun Y, Chang X, Lu L, et al. Risk factors and an early prediction model for persistent methamphetamine-related psychiatric symptoms. Addict Biol. 2020;25:e12709.
Lavigne JV, Hopkins J, Ballard RJ, Gouze KR, Ariza AJ, Martin CP. A precision mental health model for predicting stability of 4-year-olds’ attention deficit/hyperactivity disorder symptoms to age 6 diagnostic status. Acad Pediatr. 2023;24:433–41.
Franz AP, Caye A, Lacerda BC, Wagner F, Silveira RC, Procianoy RS, et al. Development of a risk calculator to predict attention-deficit/hyperactivity disorder in very preterm/very low birth weight newborns. J Child Psychol Psychiatry. 2022;63:929–38.
Suresh P, Ray B, Duan K, Chen J, Schoenmacker G, Franke B, et al. Evaluating the neuroimaging-genetic prediction of symptom changes in individuals with ADHD. Annu Int Conf IEEE Eng Med Biol Soc. 2021;2021:1950–6.
Zhang-James Y, Chen Q, Kuja-Halkola R, Lichtenstein P, Larsson H, Faraone SV. MAchine-learning Prediction of comorbid substance use disorders in ADHD youth using Swedish registry data. J Child Psychol Psychiatry. 2020;61:1370–9.
Chang J-C, Lin H-Y, Lv J, Tseng W-YI, Gau SS-F. Regional brain volume predicts response to methylphenidate treatment in individuals with ADHD. BMC psychiatry. 2021;21:26.
Wang LJ, Kuo HC, Lee SY, Huang LH, Lin Y, Lin PH, et al. MicroRNAs serve as prediction and treatment-response biomarkers of attention-deficit/hyperactivity disorder and promote the differentiation of neuronal cells by repressing the apoptosis pathway. Transl Psychiatry. 2022;12:67.
Faraone SV, Gomeni R, Hull JT, Busse GD, Melyan Z, O’Neal W, et al. Early response to SPN-812 (viloxazine extended-release) can predict efficacy outcome in pediatric subjects with ADHD: a machine learning post-hoc analysis of four randomized clinical trials. Psychiatry Res. 2021;296:113664.
Kim J-W, Sharma V, Ryan ND. Predicting methylphenidate response in ADHD using machine learning approaches. Int J Neuropsychopharmacol. 2015;18:pyv052.
Morrow AS, Campos Vega AD, Zhao X, Liriano MM. Leveraging machine learning to identify predictors of receiving psychosocial treatment for attention deficit/hyperactivity disorder. Adm Policy Ment Health. 2020;47:680–92.
Setyawan J, Yang H, Cheng D, Cai X, Signorovitch J, Xie J, et al. Developing a risk score to guide individualized treatment selection in attention deficit/hyperactivity disorder. Value Health. 2015;18:824–31.
Wong HK, Tiffin PA, Chappell MJ, Nichols TE, Welsh PR, Doyle OM, et al. Personalized medication response prediction for attention-deficit hyperactivity disorder: learning in the model space vs. learning in the data space. Front Physiol. 2017;8:199.
Siontis GC, Tzoulaki I, Castaldi PJ, Ioannidis JP. External validation of new risk prediction models is infrequent and reveals worse prognostic discrimination. J Clin Epidemiol. 2015;68:25–34.
Bellou V, Belbasis L, Konstantinidis AK, Tzoulaki I, Evangelou E. Prognostic models for outcome prediction in patients with chronic obstructive pulmonary disease: systematic review and critical appraisal. BMJ. 2019;367:l5358.
Sajjadian M, Lam RW, Milev R, Rotzinger S, Frey BN, Soares CN, et al. Machine learning in the prediction of depression treatment outcomes: a systematic review and meta-analysis. Psychol Med. 2021;51:2742–51.
Lee R, Leighton SP, Thomas L, Gkoutos GV, Wood SJ, Fenton SH, et al. Prediction models in first-episode psychosis: systematic review and critical appraisal. Br J Psychiatry. 2022;220:1–13. Spec Iss 4 Themed Iss Precision Medicine and Personalised Healthcare in Psychiatry.
Colombo F, Calesella F, Mazza MG, Melloni EMT, Morelli MJ, Scotti GM, et al. Machine learning approaches for prediction of bipolar disorder based on biological, clinical and neuropsychological markers: A systematic review and meta-analysis. Neurosci Biobehav Rev. 2022;135:104552.
Buitelaar J, Bölte S, Brandeis D, Caye A, Christmann N, Cortese S, et al. Toward precision medicine in ADHD. Front Behav Neurosci. 2022;16:900981.
Wong ICK, Banaschewski T, Buitelaar J, Cortese S, Döpfner M, Simonoff E, et al. Emerging challenges in pharmacotherapy research on attention-deficit hyperactivity disorder-outcome measures beyond symptom control and clinical trials. Lancet Psychiatry. 2019;6:528–37.
Vickers AJ, Van Calster B, Steyerberg EW. Net benefit approaches to the evaluation of prediction models, molecular markers, and diagnostic tests. BMJ. 2016;352:i6.
Becker SP, Willcutt EG, Leopold DR, Fredrick JW, Smith ZR, Jacobson LA, et al. Report of a work group on sluggish cognitive tempo: key research directions and a consensus change in terminology to cognitive disengagement syndrome. J Am Acad Child Adolesc Psychiatry. 2023;62:629–45.
Fredrick JW, Becker SP. Cognitive disengagement syndrome (sluggish cognitive tempo) and social withdrawal: advancing a conceptual model to guide future research. J Atten Disord. 2023;27:38–45.
Froehlich TE, Becker SP, Nick TG, Brinkman WB, Stein MA, Peugh J et al. Sluggish cognitive tempo as a possible predictor of methylphenidate response in children with ADHD: a randomized controlled trial. J Clin Psychiatry. 2018; 79.
Funding
Prof. Fusar-Poli is supported by #NEXTGENERATIONEU (NGEU), funded by the Ministry of University and Research (MUR), National Recovery and Resilience Plan (NRRP), project MNESYS (PE0000006) – A Multiscale integrated approach to the study of the nervous system in health and disease (DN. 1553 11.10.2022). Samuele Cortese, NIHR Research Professor (NIHR303122) is funded by the NIHR for this research project. Samuele Cortese is also supported by NIHR grants NIHR203684, NIHR203035, NIHR130077, NIHR128472, RP-PG-0618-20003 and by grant 101095568-HORIZONHLTH- 2022-DISEASE-07-03 from the European Research Executive Agency. This paper represents independent research part-funded by the National Institute for Health Research (NIHR) Maudsley Biomedical Research Centre at South London and Maudsley NHS Foundation Trust and King’s College London. The views expressed in this publication are those of the author(s) and not necessarily those of the NIHR, NHS or the UK Department of Health and Social Care.
Author information
Authors and Affiliations
Contributions
GSdP and SC conceived the study. GSdP conducted the analyses and drafted the first version of the manuscript. RI provided statistical support. SC supervised the project and study. RI, AB, AC, MD, VP, MG-A, LL, AC, MHA and HS obtained the data. LA, AJM, MS, PF-P, ZC, SVF and HC revised the manuscript and provided a substantial conceptual contribution. All authors proofread and approved the final draft of the manuscript.
Corresponding author
Ethics declarations
Competing interests
Dr. Salazar de Pablo has received honoraria from Janssen Cilag, Lundbeck and Angelini. Dr Archer is supported by funding from the NIHR Birmingham Biomedical Research Centre at the University Hospitals Birmingham NHS Foundation Trust and the University of Birmingham. Dr. Solmi has received honoraria/has been a consultant for Abbvie, Angelini, Lundbeck, Otsuka. Prof. Fusar-Poli has received grant support from Lundbeck and honoraria fees from Angelini, Menarini, and Lundbeck. In the past year, Dr. Faraone received income, potential income, travel expenses continuing education support and/or research support from Aardvark, Aardwolf, AIMH, Tris, Otsuka, Ironshore, KemPharm/Corium, Akili, Supernus, Atentiv, Noven, Sky Therapeutics, Axsome and Genomind. With his institution, he has US patent US20130217707 A1 for the use of sodium-hydrogen exchange inhibitors in the treatment of ADHD. He also receives royalties from books published by Guilford Press: Straight Talk about Your Child’s Mental Health, Oxford University Press: Schizophrenia: The Facts and Elsevier: ADHD: Non-Pharmacologic Interventions. He is Program Director of www.ADHDEvidence.org and www.ADHDinAdults.com. Prof Larsson reports receiving grants from Shire Pharmaceuticals; personal fees from and serving as a speaker for Medice, Shire/Takeda Pharmaceuticals and Evolan Pharma AB; all outside the submitted work. Henrik Larsson is editor-in-chief of JCPP Advances. Prof. Cortese has declared reimbursement for travel and accommodation expenses from the Association for Child and Adolescent Central Health (ACAMH) in relation to lectures delivered for ACAMH, the Canadian AADHD Alliance Resource, the British Association of Psychopharmacology, and from Healthcare Convention for educational activity on ADHD.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Salazar de Pablo, G., Iniesta, R., Bellato, A. et al. Individualized prediction models in ADHD: a systematic review and meta-regression. Mol Psychiatry (2024). https://doi.org/10.1038/s41380-024-02606-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41380-024-02606-5
- Springer Nature Limited