Abstract
Multicolor imaging allows protein colocalizations and organelle interactions to be studied in biological research, which is especially important for single-molecule localization microscopy (SMLM). Here, we propose a multicolor method called excitation-resolved stochastic optical reconstruction microscopy (ExR-STORM). The method, which is based on the excitation spectrum of fluorescent dyes, successfully separated four spectrally very close far-red organic fluorophores utilizing three excitation lasers with cross-talk of less than 3%. Dyes that are only 5 nm apart in the emission spectrum were resolved, resulting in negligible chromatic aberrations. This method was extended to three-dimensional (3D) imaging by combining the astigmatic method, providing a powerful tool for resolving 3D morphologies at the nanoscale.
Similar content being viewed by others
Introduction
The development of single-molecule localization microscopy (SMLM)1,2,3 has provided a method for exploring nanostructures in cells that were previously concealed by diffraction limits, improving the spatial resolution of fluorescence microscopy to tens of nanometers4. In addition to the spatial resolution, multicolor imaging in SMLM is critical for investigating protein colocalizations and organelle interactions in biological research5,6,7.
The multicolor capability of SMLM is realized by single-molecule identification with intrinsic properties of different fluorescent dyes, such as fluorescence lifetime8, emission spectrum, and excitation spectrum.
Previous work has mainly focused on the emission spectra of fluorescent dyes. One approach used spectrally well-separated fluorophores9, requiring additional error-prone alignment procedures between channels10,11. Another approach used ratiometric detection to distinguish the fluorescence intensity in several channels12,13,14,15,16. A dispersive prism was also used to determine the spectral peaks of different dyes17,18, meanwhile, a dual-objective design was adopted because both locating single molecules and distinguishing spectra require a considerable number of photons.
Excitation spectra of fluorescent dyes can be also adopted in multicolor imaging. Exciting with different lasers has been proved in triple-color imaging of live cell19. Additionally, excitation spectral demixing has been applied in wide-field imaging20, and lasers with different excitation frequencies have been utilized for differentiating dyes in multicolor imaging21. However, these sequential imaging-based methods are not fully compatible with STORM imaging, due to the highly dynamic properties of single-molecule dyes.
Here we proposed a single molecule identification method that used the excitation spectrum of fluorescent dyes and was fully compatible with STORM imaging, called excitation-resolved stochastic optical reconstruction microscopy (ExR-STORM). This method is based on the fact that the fluorescence intensity is proportional to the excitation cross-section at each wavelength for a given laser intensity21. Using this method, we successfully resolved four spectrally very close far-red dyes simultaneously with negligible cross-talk and little chromatic aberrations by utilizing three excitation lasers with a single objective. A 4 kHz resonant mirror was adopted to enable fast switching among excitation channels; thus, three subimages could be obtained under different excitation wavelengths simultaneously within one exposure time22,23, enabling the identification of dyes in the realm of SMLM. ExR-STORM fully utilized photons in three light paths to locate and identify dyes through a co-fitting method, realizing high localization precision. Furthermore, by combining ExR-STORM with the astigmatic method24, we demonstrated the capability of 3D tetra-color imaging.
Results
Principle of ExR-STORM
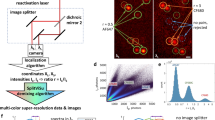
Most STORM dyes that exhibit superior performance are located in the far-red region25. In ExR-STORM, excitation lasers were selected for the candidate dyes, which could be differentiated based on the corresponding intensities under each laser. The optical setup scheme is shown in Fig. 1a. Four lasers (620 nm, 639 nm, 671 nm for excitation, and 405 nm for activation) were combined and modulated with an acousto-optic tunable filter (AOTF) before being focused on the back focal plane of the objective for illumination. In the imaging path, the fluorescence signals were focused on the surface of a 4 kHz resonant mirror, then sequentially reflected and refocused onto three subregions of the electron-multiplying charge-coupled device (EMCCD) with three pairs of lenses (Supplementary Fig. S1). The switching between subregions was synchronized with the AOTF to ensure that for each subregion, only the fluorescence signal excited by the corresponding laser was accumulated. The scanning cycle was 250 μs with a duty cycle of 84%, allowing brightness fluctuations caused by on-off state transitions and other effects to be minimized22,23,26 (Supplementary Fig. S2). With this method, three subimages of the same single molecule could be acquired under different excitation wavelengths within one exposure time (Supplementary Fig. S3). The molecule was then identified according to the intensities in the three subimages. For dyes in the multicolor sample, each kind of molecule can be recorded and identified simultaneously under the condition that the fluorophores are sparsely distributed in space to avoid overlapping.
a Scheme of the optical setup. Three excitation lasers (620, 639, and 671 nm) were combined and then modulated by an AOTF prior to the illumination of the sample. The laser switching was synchronized with the scanner, resulting in three subregions with different excitation wavelengths within one exposure time. b Absorption spectra of the eight fluorescent dyes. The three black lines correspond to the three excitation wavelengths. c Scatter plot of the photon number of Alexa Fluor 647 in the three excitation channels. d 2D projections of the photon number spatial occupation along the 671 path, 620 path, 639 path, and the isometric projection (which is an axonometric projection in which the three coordinate axes appear equally foreshortened and the angle between any two of them is 120°) for the investigated dyes, where the half-maximum contour of the distribution was selected as the boundary of the investigated dye; more than one million single molecules of each dye were selected for a statistical examination of the molecular behaviors
ExR-STORM identified dyes based on the fact that the fluorescence intensity is proportional to the absorption cross-section at each excitation wavelength21. In this work, candidate dyes were selected based on their absorption spectra (Fig. 1b) and evaluated by our system. For every single molecule, 2D Gaussian fitting was performed to extract the location and photon numbers in three subimages. A 3D scatter plot of the photon numbers demonstrated a narrow cluster distribution (Fig. 1c), indicating that the molecules possessed highly consistent fluorescence intensity properties under the three excitation wavelengths. Next, we calculated the photon number distribution for each dye, with the half-maximum contour of the distribution selected as the boundary of the investigated dye (Fig. 1d). We found that each dye exhibited a distinct cluster distribution of photon numbers under the three excitation wavelengths. Thus, multicolor STORM imaging could be conducted when the proper dyes were selected. Because this demixing method does not depend on the emission spectrum, dyes with identical emission spectra could be demixed as long as they possessed distinguishable excitation characteristics. Using this method, chromatic aberrations can be minimized, and the choice of dyes for multicolor imaging is enriched.
Demixing through ExR-STORM
The workflow of ExR-STORM is shown in Fig. 2. To illustrate our scheme, we selected DyLight 633, Alexa Fluor 647, Dyomics 654, and CF660C for tetra-color imaging. The recorded raw data (Fig. 2a and Supplementary Video 1) were first split and transformed to create 3 aligned subimage stacks (Fig. 2b). Following single-molecule detection and image extraction similar to the conventional STORM imaging process, the 3-channel images of every single molecule were co-fitted as modified 2D Gaussian models:
where Ai and Bi denote the amplitude intensity and the background of single molecules in subimage stack i (i = 1, 2, 3, which correspond to the 620 path, 639 path, and 671 path, respectively), respectively; σx and σy denote the standard deviation in the X and Y directions, respectively; and x0 and y0 denote the position of a single molecule. The above parameters were extracted by our fitting model (1). The photon numbers in each light path can be calculated as:
where Ni is the photon number in the subimage stack i, ADU is the analog-to-digital unit conversion factor of the EMCCD, and G is the gain of the EMCCD.
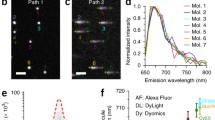
a One frame of the raw image stack. b Split channels of the raw image based on the ROI of the three light paths, and transformed by the affine matrices to the 639 nm light path. c Diagram of dye assignment along the projections of the 671 path, 620 path, 639 path, and the isometric projection (which is an axonometric projection in which the three coordinate axes appear equally foreshortened and the angle between any two of them is 120 degrees), with the yellow regions between the dashed lines, rejected. d Single-molecule of each dye in the three light paths from (b) based on the classification process shown in (c)
For molecule identification, we first plotted the photon numbers of every single molecule in the three light paths in a 3D logarithmic coordinate system, each with a point (N1, N2, N3). Then, molecule assignment was conducted in the projections of the 671, 620, and 639 paths, as well as the isometric projection (Fig. 2c). It should be noted that the classification result of each dye was the intersection of the selected regions in four projections; otherwise, the molecule was rejected. After molecule identification, multicolor superresolution images were reconstructed according to the classification results and positions obtained previously. Typical single-molecule images of each dye in the three channels are shown in Fig. 2d.
The co-fitting method was validated by microsphere images, as shown in Fig. 3. Three transformed subimages and the merged image are shown in Fig. 3a–d. The localization results obtained through independent fitting in each channel were compared with the co-fitting results, and the localization error and bias of each channel were analyzed (see Materials and methods). For the localization difference between independent fitting and co-fitting, the maximum mean bias and standard deviations were 2 and 3 nm, respectively (Fig. 3e), which were well below the localization precision of the ExR-STORM system (Supplementary Fig. S4).
100 nm fluorescent microspheres were used for image registration and verification. a–d Transformed images of the three light paths and merged image. e Positions of independent fitting in each transformed channel relative to the co-fitting results in the X and Y directions. Error bars are presented as the mean ± standard deviation. Scale bars: 10 μm (a–d)
Tetra-color performance of ExR-STORM
The ExR-STORM identification process was conducted in three dimensions (N1, N2, and N3). Compared with the spectral demixing method12,13,14,16 that used a two-dimensional photon number distribution for dye classification, one additional dimension was used in our method. During our assignment process, four projections of the photon number distribution were selected, and the final classification was the intersection of the classifications in each projection, thus contributing to a more accurate classification result.
This innovation enabled tetra-color imaging in fixed COS-7 cells. DyLight 633, Alexa Fluor 647, Dyomics 654, and CF660C were selected to label microtubules, intermediate filaments, endoplasmic reticulum (ER), and the outer mitochondrial membrane, respectively. With ExR-STORM, the four dyes were well resolved, and tetra-color superresolution images were successfully reconstructed (Fig. 4). The images of each channel were consistent with images of the separately acquired structures, and no obvious misidentification or cross-talk was observed (Fig. 4a–e).
CF660C, Alexa Fluor 647, Dyomics 654, and DyLight 633 were selected to label the outer mitochondrial membrane (cyan), intermediate filaments (magenta), endoplasmic reticulum (green), and microtubules (yellow), respectively, in a fixed COS-7 cell. a Four-color superresolution reconstruction image in which each detected single molecule was classified based on the excitation spectrum of the fluorophores. b-e Split channels of the white box region in (a). f–i 2D projections of the photon number distributions along the 671 path, 620 path, 639 path, and the isometric projection (which is an axonometric projection in which the three coordinate axes appear equally foreshortened and the angle between any two of them is 120°). j Cross-talk and rejected fraction for the four dyes. Scale bars: 2 μm (a); 1 μm (b-e)
By imaging four dyes in the cell sample individually, the cross-talk can be evaluated. The photon number distribution of each dye is plotted in Fig. 4f–i, and each distribution contains more than 1 million points. The classification was performed for each dye as previously described, showing a cross-talk of <3% and a rejection fraction of <35% (Fig. 4j and Supplementary Table 1). Take CF660C as an example, 76.1%, 0.9%, 0.0%, and 0.0% of the total single molecule events were identified as CF660C, AF647, DY654, and DL633, respectively, and the rest 22.9% events were rejected because of ambiguity. (Fig. 4j and Supplementary Table 1).
3D tetra-color ExR-STORM imaging of COS-7 cells
Cell organelles and their interaction sites are distributed in 3D, so axial resolving capability is important for multicolor imaging. We achieved 3D tetra-color imaging by combining the astigmatic method with the ExR-STORM system. In 3D ExR-STORM, the resonant mirror introduced negligible influence on the point spread function (PSF), and the fitting process was compatible with the elliptical PSF (Supplementary Fig. S5). The localization precision was approximately 8 and 23 nm in the lateral and axial directions, respectively (Supplementary Fig. S4), which is consistent with previous single-color results24.
ER exhibits rich interactions with other organelles15,27. Therefore, ER, intermediate filaments, peroxisome, and the outer mitochondrial membrane were labeled in fixed COS-7 cells (Fig. 5a–e and Supplementary Video 2) for interaction analysis. We found that peroxisomes colocalized with ER in the XY, YZ, and XZ cross-sections (Fig. 5f–k), elucidating the relationships between peroxisomes and ER27,28. Additionally, a mitochondrion contacted peroxisome, and both of them colocalized with the ER (Fig. 5l–n), indicating the existence of tripartite contact sites between peroxisomes, ER, and mitochondria29,30. Colocalization between ER and intermediate filaments was also observed in Fig. 5a.
a-e CF660C, Alexa Fluor 647, Dyomics 654, and DyLight 633 were selected to label the outer mitochondrial membrane (cyan), intermediate filaments (magenta), endoplasmic reticulum (green), and peroxisomes (yellow), respectively, in a fixed COS-7 cell. a Reconstructed image of the four merged channels. b–e Color-coded four channels in (a). f–k Colocalization of peroxisomes and endoplasmic reticulum. f Zoomed-in view of boxed region i in (a). g XZ (120 nm in Y) cross-section of the boxed region in (f). h Intensity profile along the boxed region in (f). i Zoomed-in view of the boxed region ii in (a). j YZ (120 nm in X) cross-sections of the region between the two dashed lines in (i), where the distance between adjacent cross-sections in the X direction is 120 nm. k Intensity profiles along the Y direction in (i), with each profile corresponding to the cross-section image in (j). l The ER–mitochondria–peroxisome tripartite contact site in the boxed region iii in (a). m, n XZ (120 nm in Y) and YZ (120 nm in X) cross-sections of the boxed regions in (l). Scale bars: 2 μm (a–e); 1 μm (f–g); 500 nm (i, j, l–n)
Mitochondrial double-stranded DNA (dsDNA), intermediate filaments, and the outer mitochondrial membrane were also imaged with 3D ExR-STORM (Supplementary Fig. S6 and Supplementary Video 3). Cross-sections helped to reveal the relative 3D position of diffraction-limited structures–for example, the hollow structures of the outer mitochondrial membrane (Supplementary Fig. S6b–d), the dsDNA inside the mitochondria (Supplementary Fig. S6b–d) and intermediate filaments in contact with the mitochondria (Supplementary Fig. S6c, d).
Discussion
In this work, we combined multi-wavelength excitation with a fast switching scheme and successfully resolved four spectrally very close far-red dyes (DyLight 633, Alexa Fluor 647, Dyomics 654, and CF660C) simultaneously with negligible cross-talk and little chromatic aberrations.
Compared with other multicolor schemes in SMLM, our proposed method has several advantages. First, dyes with similar emission spectra could be resolved with ExR-STORM, resulting in negligible chromatic aberrations and providing more choices of dyes. In this study, we successfully resolved Dyomics 654 and Alexa Fluor 647, which have emission peaks that are only 5 nm apart. Second, ExR-STORM resolved four far-red dyes with only one objective, extending color channels with cross-talk of less than 3%. Third, differing from other sequential imaging with multi-wavelength excitation methods19,20,21, which would suffer from the blinking effects of STORM dyes, we used a 4 kHz scanner (250 μs per imaging cycle) to eliminate the brightness fluctuation, compatible with fast dynamics of single molecule dyes. Fourth, since the PSF of ExR-STORM occupied fewer pixels than the spectrum in spectrally resolved STORM did17,31, a higher molecule density could be realized, resulting in higher molecular throughput.
In our tetra-color experiment, the molecule density was 0.2–0.3 points/μm2. In this condition, most of the single molecules were not overlapped. We estimated the relationship between the molecule density and cross-talk ratio through simulations and found that the cross-talk ratio and the rejection ratio increased with increasing molecule density (Supplementary Fig. S7). Therefore, it is necessary to balance molecule density to a certain level where no evident cross-talk is introduced.
ExR-STORM successfully achieved tetra-color superresolution imaging, and more color channels could be extended by adopting the following two approaches. First, additional excitation lasers could be used to obtain more channels, which will also contribute to less cross-talk. Second, ExR-STORM could be combined with existing emission spectra-based multicolor schemes12,13,14,15,16, such as introducing a dichroic mirror before the resonant mirror. With these two modifications, more colors could be used in the palette of multicolor SMLM in the future.
Various cell organelles regularly interact with each other through direct membrane contact. Our ExR-STORM system provides a new approach to studying these interactions, and several organelle contacts and colocalization were observed in three-dimension in this work. We believe that ExR-STORM will allow further investigation of organelle organization and interactions in cell stress responses, migration, division, and differentiation.
Materials and methods
Optical setup
The optical setup was built based on an Olympus IX73 inverted microscope (Supplementary Fig. S1). Lasers with wavelengths of 620 nm (1 W, MPB Communications), 639 nm (1 W, CNI), and 671 nm (1 W, CNI) were coupled to an optical fiber after being modulated by the AOTF and then focused on the back focal plane of the objective (100×, 1.50 N.A., Olympus) to generate total internal reflection fluorescence illumination (TIRF). A dichroic mirror (FLD 681 DLP, Iridian) was used to reflect the excitation light and transmit the emission light. Emission filters (BLP01-664R, Semrock; ET740sp, Chroma) were installed to filter the excitation laser and infrared noise.
In the imaging path, a field diaphragm (GCM5711M, Daheng Optics) was placed at the first image plane to control the field of view (FOV) of the three subregions. A linear slide (GCM-TD50MX, Daheng Optics) was used for switching the cylindrical lenses (f = 500 mm, LJ1144L1-A, Thorlabs) into imaging paths, changing the image mode between 2D and 3D. The fluorescence images were then focused on a 4 kHz resonant scanning mirror (CRS 4 KHZ, CTI) after passing through 2 lenses (f = 180 mm, AC508-180-A-ML, Thorlabs; f = 150 mm, AC508-150-A-ML, Thorlabs). Then, the fluorescence images were reflected by the scanner to three light paths, each with a pair of lenses, generating three subimages on the EMCCD (iXon 897 EMCCD, Andor, Supplementary Fig. S3). Because the resonant mirror was placed at the image plane, the final image on the CCD was not blurred when the resonant mirror was scanning (Supplementary Fig. S8). During imaging, the excitation lasers were turned off when the fluorescence beam was located at the junction between the two lenses to avoid signal loss and cross-talk. During image acquisition, the resonant mirror generated a 4 kHz signal, then multiplied by a complex programmable logic device, and the output signal with a frequency of ~800 kHz was taken as a clock source for the data acquisition device (DAQ, NI USB-6343, National Instruments) to modulate the AOTF22,23. The details of the resonant scan sequence diagram and the image path-switching process are shown in Supplementary Fig. S2.
Validation of the co-fitting method
To validate the performance of the co-fitting method, one hundred groups of microsphere image stacks (each with ten frames) were obtained. The recorded raw data were first split and transformed to create three aligned subimage stacks. Then, three-channel co-fitting and independent fitting (fitting with one channel) were conducted. Points that persisted for ten frames were extracted, and the positions of the four fitting results were aligned by subtracting the center of the points in the co-fitting results. Then, statistical analyses of the registration errors of all groups were performed.
ExR-STORM imaging
The whole image is 512 × 512 pixels, and the three subimages were acquired in three different subregions of the EMCCD. The size of each subregion is approximately 220 × 220 pixels, with a pixel size of 160 nm, yielding a FOV of approximately 35 × 35 μm2.
For cellular imaging, the average excitation power densities (620, 639, and 671 nm) were 1100, 800, and 1100 W/cm2, respectively. A 405 nm laser was used for density control of single molecules during image acquisition. The frame rate of the EMCCD was 20 Hz, the EM gain factor was 100, and the ADU conversion factor was 4.96. A total of 20,000–30,000 frames were acquired for image reconstruction, and molecules with lifetimes exceeding 5 frames or photon numbers of less than 300 were rejected, as weak signals were more influenced by background noise, thus introducing more uncertainty into molecule identification.
In 3D imaging, a cylindrical lens was introduced, and the relative axial position was calculated according to the calibration results of 0.1 μm TetraSpeck microspheres before cellular imaging (Supplementary Fig. S5). The calibration curve was acquired by fitting the astigmatic PSF, enabling the 3D position to be determined in the axial range of ±400 nm. In the axial direction, the resonant mirror introduced negligible error to the PSF (Supplementary Fig. S5b). The difference of PSF widths in the x-direction and y-direction changed monotonically with the z-position during scanning so that the axial position could be determined according to a three-order polynomial fitting curve (Supplementary Fig. S5c).
The oxygen-depleted STORM imaging buffer consisted of 10% glucose, 1× phosphate-buffered saline (PBS) buffer, GLOX (catalase (0.06 mg/mL) and glucose oxidase (0.6 mg/mL), dissolved in Tris-HCl buffer) and 143 mM 2-mercaptoethanol.
Calibration and drift correction
The 0.1 μm TetraSpeck microspheres (T7279, Invitrogen) were used for image registration before cellular imaging. Image stacks of the bright microspheres were acquired and combined to increase the signal-to-noise ratio (SNR); and then, image splitting and registration were performed to obtain the transform matrix. All subimages were aligned to the 639 nm light path.
To ensure that the sample remained in the focal plane during image acquisition, a TIRF Lock system was used for real-time axial drift correction with PIEZO (P-725.4CD, PI), maintaining the focal plane according to the feedback from the center of the exiting TIRF beam collected by a CCD. To obtain the feedback coefficient for real-time focus locking, we scanned ±600 nm of the focal plane in the step of 10 nm and used least squares regression to fit the axial position and the center position of the exiting TIRF beam. The drift of the focal plane was kept within ±20 nm during image acquisition.
Redundant cross-correlation (RCC)32 was used for lateral drift correction during postprocessing. Five hundred frames were set for the correlation steps according to the structure distribution and point density.
Sample preparation
Cell culture
COS-7 cells (3111C0001CCC000033, National Infrastructure of Cell Line Resources, China) were grown in Dulbecco’s modified Eagle medium (DMEM) (C11995500BT, Gibco) supplemented with 10% fetal bovine serum (16000–044, Gibco) and 100 U/mL penicillin–streptomycin solution (SV30010, HyClone) at 37 °C with 5% CO2.
Nanobody and antibody conjugation
Nanobody against GFP (gt-250, Chromotek) was conjugated to Dyomics 654-NHS Ester (654-01, Dyomics). The conjugation reaction was carried out in 0.1 M sodium bicarbonate in the dark for 1 h. Then we used a Zeba Spin Desalting Column (89882, Thermo Scientific) with a molecular weight cutoff of 7 kDa to remove excess dye from the conjugation reaction. Nanobodies were incubated at 1:50 dilution at room temperature for 1 h.
Unlabeled donkey anti-mouse IgG (ab6707, Abcam) was conjugated with Dyomics 678-NHS Ester (678-01, Dyomics), Dyomics 675-NHS Ester (675-01, Dyomics) or Alexa Fluor 660-NHS Ester (A20007, Invitrogen). The reaction was carried out in 0.1 M sodium bicarbonate in the dark for 1 h. Excess dye was removed from the antibody using a Pro-Spin Column (CS800, Princeton Separations) according to the manufacturer’s recommendations.
Sample labeling
Microtubule samples
To label the microtubules, COS-7 cells with a density of 50,000 were cultured in 35 mm glass-bottomed dishes (D35-14-1-N, Cellvis). Twenty-four hours later, the cells were rinsed with PBS (P4417, Sigma-Aldrich). Then, the cells were immobilized with 0.1% glutaraldehyde (16220, electron Microscopy Science) and 3% paraformaldehyde (157-8, electron Microscopy Science) in PBS at room temperature for 10 min. The reduction process was conducted in PBS with 0.1% sodium borohydride (71320, Sigma-Aldrich) for 7 min. Following washing with PBS, cell permeation was performed with 0.2% Triton X-100 (93443, Sigma-Aldrich) in PBS for 7 min. Samples were blocked with 10% NGS (C0265, Beyotime) and 0.05% Triton X-100 in PBS at room temperature for 90 min, then incubated with primary antibodies in antibody dilution buffer (0.05% Triton X-100, 5% NGS in PBS) for 60 min. The cells were then washed 5 times with washing buffer (0.05% Triton X-100, 1% NGS in PBS), each time for 15 min, and then stained for 60 min with secondary antibodies. Next, the cells were washed with washing buffer 5 times and fixed with 4% paraformaldehyde in PBS for 10 min. Then, the cells were washed with PBS 3 times, and ddH2O for 3 times, both 5 min per wash. At last, the samples in ddH2O were stored at 4 °C. The primary antibody used was mouse anti-α-tubulin (T9026, Sigma–Aldrich, used at 1:100 dilution). Secondary antibodies labeled with Alexa Fluor 647 (A31571, Invitrogen, used at 1:100 dilution), DyLight 633 (35512, Invitrogen, used at 1:300 dilution), and Dyomics 678, Dyomics 675, and Alexa Fluor 660 (conjugated in the laboratory; see Nanobody and antibody conjugation section; used at 1:50, 1:50, and 1:100 dilutions, respectively) were used to label primary antibodies.
ER samples
Lentiviral particles of GFP-Sec61β were packaged by OBiO Technology (Shanghai) according to the authors’ designs and requirements. The titer of the lentivirus used in the experiments was 5.4 × 108 infectious units/mL. COS-7 cells were infected with lentivirus according to the manufacturer’s protocols provided by OBiO Technology. After 48 h, the infected cells with a density of 50,000 were cultured in 35 mm glass-bottomed dishes. The following immunofluorescence steps were performed as described in the Microtubule samples section. Nanobody GFP-binding protein conjugated with Dyomics 654 (see Nanobody and antibody conjugation section; used at 1:50 dilution) was used to label GFP-Sec61β.
Mitochondria samples
To label mitochondria, COS-7 cells with a density of 50,000 were cultured in 35 mm glass-bottomed dishes. After 24 h, immunofluorescence was performed as described in the Microtubule samples section. The primary antibody used was rabbit anti-TOMM20 (ab186735, Abcam, used at 1:250 dilution). Secondary antibodies labeled with CF660C (20815, Biotium, used at 1:100 dilution) and CF680 (20820, Biotium, used at 1:500 dilution) were used to label primary antibodies.
Intermediate filament samples
To label intermediate filament samples, COS-7 cells with a density of 50,000 were cultured in 35 mm glass-bottomed dishes. After 24 h, immunofluorescence was performed as described in the Microtubule samples section. The primary antibody used was chicken anti-vimentin (ab24525, Abcam, used at 1:200 dilution). Secondary antibodies labeled with Alexa Fluor 647 (A21449, Invitrogen, used at 1:150 dilution) were used to label primary antibodies.
For the mitochondria/dsDNA/intermediate filament three-color sample, COS-7 cells with a density of 50,000 were cultured in 35 mm glass-bottomed dishes. After 24 h, immunofluorescence was performed as described in the Microtubule samples section. The rabbit anti-TOMM20, mouse anti-dsDNA (ab27156, Abcam), and chicken anti-vimentin dilution ratios were 1:250, 1:2000, and 1:200, respectively. Secondary antibodies of the three-color sample were labeled with CF660C (1:100)/DyLight 633 (1:2 000)/Alexa Fluor 647 (1:150).
For the ER/microtubule/mitochondria/intermediate filament four-color sample, COS-7 cells were infected with lentiviral particles of GFP-Sec61β. The infected cells with a density of 50,000 were cultured in 35 mm glass-bottomed dishes, and immunofluorescence was performed as described in the Microtubule samples section. The mouse anti-α-tubulin, rabbit anti-TOMM20, and chicken anti-vimentin dilution ratios were 1:400, 1:250, and 1:200, respectively. Secondary antibodies of the four-color sample were labeled with Dyomics 654 (1:50)/DyLight 633 (1:400)/CF660C (1:100)/Alexa Fluor 647 (1:150).
For the ER/peroxisome/mitochondria/intermediate filament four-color sample, COS-7 cells were infected with lentiviral particles of GFP-Sec61β. The infected cells with a density of 50,000 were cultured in 35 mm glass-bottomed dishes, and immunofluorescence was performed as described in the Microtubule samples section. The mouse anti-PMP70 (SAB4200181, Sigma-Aldrich), rabbit anti-TOMM20, and chicken anti-vimentin dilution ratios were 1:100, 1:250 and 1:200, respectively. Secondary antibodies of the four-color sample were labeled with Dyomics 654 (1:50)/DyLight 633 (1:50)/CF660C (1:100)/Alexa Fluor 647 (1:150).
Data availability
The data that support the findings of this study are available from the corresponding author upon request.
References
Betzig, E. et al. Imaging intracellular fluorescent proteins at nanometer resolution. Science 313, 1642–1645 (2006).
Hess, S. T., Girirajan, T. P. & Mason, M. D. Ultra-high resolution imaging by fluorescence photoactivation localization microscopy. Biophys. J. 91, 4258–4272 (2006).
Rust, M. J., Bates, M. & Zhuang, X. Sub-diffraction-limit imaging by stochastic optical reconstruction microscopy (STORM). Nat. Methods 3, 793–796 (2006).
Lelek, M. et al. Single-molecule localization microscopy. Nat. Rev. Methods Prim. 1, 1–27 (2021).
Huang, B., Jones, S. A., Brandenburg, B. & Zhuang, X. Whole-cell 3D STORM reveals interactions between cellular structures with nanometer-scale resolution. Nat. Methods 5, 1047–1052 (2008).
Valm, A. M. et al. Applying systems-level spectral imaging and analysis to reveal the organelle interactome. Nature 546, 162–167 (2017).
Guo, Y. et al. Visualizing intracellular organelle and cytoskeletal interactions at nanoscale resolution on millisecond timescales. Cell 175, 1430–1442.e1417 (2018).
Oleksiievets, N. et al. Fluorescence lifetime DNA-PAINT for multiplexed super-resolution imaging of cells. Commun. Biol. 5, 1–8 (2022).
Shroff, H. et al. Dual-color superresolution imaging of genetically expressed probes within individual adhesion complexes. Proc. Natl Acad. Sci. USA 104, 20308–20313 (2007).
Pertsinidis, A., Zhang, Y. & Chu, S. Subnanometre single-molecule localization, registration and distance measurements. Nature 466, 647–651 (2010).
Erdelyi, M. et al. Correcting chromatic offset in multicolor super-resolution localization microscopy. Opt. Express 21, 10978–10988 (2013).
Bossi, M. et al. Multicolor far-field fluorescence nanoscopy through isolated detection of distinct molecular species. Nano Lett. 8, 2463–2468 (2008).
Lampe, A., Haucke, V., Sigrist, S. J., Heilemann, M. & Schmoranzer, J. Multi‐colour direct STORM with red emitting carbocyanines. Biol. Cell 104, 229–237 (2012).
Lampe, A., Tadeus, G. & Schmoranzer, J. Spectral demixing avoids registration errors and reduces noise in multicolor localization-based super-resolution microscopy. Methods Appl. Fluorescence 3, 034006 (2015).
Zhang, Y. et al. Nanoscale subcellular architecture revealed by multicolor three-dimensional salvaged fluorescence imaging. Nat. Methods 17, 225–231 (2020).
Gimber, N., Strauss, S., Jungmann, R. & Schmoranzer, J. Simultaneous multicolor DNA-PAINT without sequential fluid exchange using spectral demixing. Nano Lett. 22, 2682–2690 (2022).
Zhang, Z., Kenny, S. J., Hauser, M., Li, W. & Xu, K. Ultrahigh-throughput single-molecule spectroscopy and spectrally resolved super-resolution microscopy. Nat. Methods 12, 935–938 (2015).
Xiang, L., Chen, K. & Xu, K. Single molecules are your quanta: a bottom-up approach toward multidimensional super-resolution microscopy. ACS Nano 15, 12483–12496 (2021).
Wilmes, S. et al. Triple‐color super‐resolution imaging of live cells: resolving submicroscopic receptor organization in the plasma membrane. Angew. Chem. 124, 4952–4955 (2012).
Chen, K., Yan, R., Xiang, L. & Xu, K. Excitation spectral microscopy for highly multiplexed fluorescence imaging and quantitative biosensing. Light 10, 1–12 (2021).
Gómez-García, P. A., Garbacik, E. T., Otterstrom, J. J., Garcia-Parajo, M. F. & Lakadamyali, M. Excitation-multiplexed multicolor superresolution imaging with fm-STORM and fm-DNA-PAINT. Proc. Natl Acad. Sci. USA 115, 12991–12996 (2018).
Gu, L. et al. Molecular resolution imaging by repetitive optical selective exposure. Nat. Methods 16, 1114–1118 (2019).
Gu, L. et al. Molecular-scale axial localization by repetitive optical selective exposure. Nat. Methods 18, 369–373 (2021).
Huang, B., Wang, W., Bates, M. & Zhuang, X. Three-dimensional super-resolution imaging by stochastic optical reconstruction microscopy. Science 319, 810–813 (2008).
Dempsey, G. T., Vaughan, J. C., Chen, K. H., Bates, M. & Zhuang, X. Evaluation of fluorophores for optimal performance in localization-based super-resolution imaging. Nat. Methods 8, 1027–1036 (2011).
Balzarotti, F. et al. Nanometer resolution imaging and tracking of fluorescent molecules with minimal photon fluxes. Science 355, 606–612 (2017).
Wu, H., Carvalho, P. & Voeltz, G. K. Here, there, and everywhere: the importance of ER membrane contact sites. Science 361, eaan5835 (2018).
Smith, J. J. & Aitchison, J. D. Peroxisomes take shape. Nat. Rev. Mol. Cell Biol. 14, 803–817 (2013).
Shai, N., Schuldiner, M. & Zalckvar, E. No peroxisome is an island—peroxisome contact sites. Biochim. Biophys. Acta 1863, 1061–1069 (2016).
Murley, A. & Nunnari, J. The emerging network of mitochondria-organelle contacts. Mol. Cell 61, 648–653 (2016).
Song, K.-H., Zhang, Y., Brenner, B., Sun, C. & Zhang, H. F. Symmetrically dispersed spectroscopic single-molecule localization microscopy. Light 9, 1–10 (2020).
Wang, Y. et al. Localization events-based sample drift correction for localization microscopy with redundant cross-correlation algorithm. Opt. Express 22, 15982–15991 (2014).
Acknowledgements
We thank Dr. Junjie Hu for the help in interpreting the 3D multicolor imaging results. This work was supported by the National Key Research and Development Program of China (Grant no. 2021YFA1301500 to W.J.), the National Natural Science Foundation of China (Grant no. 32027901 to T.X., Grant no. 91954202 to W.J., Grant no. 31700743, 32170704 and 91954201 to L.G., and Grant no. 31900419 to C.F.), the Strategic Priority Research Program of the Chinese Academy of Sciences (Grant no. XDB37040104 to W.J.), the Instrument Development Project of the CAS (Grant no. GJJSTD20210001 to T.X.), National Science and Technology Innovation 2030 Major Program (Grant no. 2022ZD0211900 to L.G.), Beijing Nova Program from Beijing Municipal Science & Technology Commission (Grant no. Z202003 to L.G.), and the Youth Innovation Promotion Association of the CAS (Grant no. 2013066 to W.J., Grant no. 2017135 to L.G.).
Author information
Authors and Affiliations
Contributions
W.J., T.X., and L.G. designed the experiments. W.W. and S.L. developed the ExR-STORM system and performed data analyses. S.L. and C.F. prepared the samples. W.W. and S.L. performed imaging. W.J., T.X., L.G., W.W., S.L., and C.F. interpreted the results. W.J., T.X., L.G., W.W., and S.L. wrote the paper, which was modified by all the other authors.
Corresponding authors
Ethics declarations
Competing interests
A patent describing the ExR-STORM system and image analysis in this work has been filed. W.J., L.G., W.W., S.L., C.F., and T.X. are the co-inventors. The remaining authors declare no competing interests.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wu, W., Luo, S., Fan, C. et al. Tetra-color superresolution microscopy based on excitation spectral demixing. Light Sci Appl 12, 9 (2023). https://doi.org/10.1038/s41377-022-01054-6
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41377-022-01054-6
- Springer Nature Limited
This article is cited by
-
Multicolor single-molecule localization microscopy: review and prospect
PhotoniX (2024)
-
RegiSTORM: channel registration for multi-color stochastic optical reconstruction microscopy
BMC Bioinformatics (2023)
-
Fluorescence-based super-resolution-microscopy strategies for chromatin studies
Chromosoma (2023)