Abstract
Myelodysplastic Neoplasms (MDS) are a group of clonal disorders characterized by ineffective hematopoiesis and morphologic dysplasia. Clinical manifestations of MDS vary widely and are dictated in large part by a range of genetic aberrations. The lack of robust in vitro models for MDS has limited the ability to conduct high throughput drug screens, which in turn has hampered the development of novel therapies for MDS. There are very few well-characterized MDS cell lines, and the available cell lines expand poorly in vitro. Conventional xenograft mouse models can provide an in vivo vessel to provide growth of cancer cells, but human MDS cells engraft poorly. Three-dimensional (3D) scaffold models that form human “ossicles” represent a promising new approach and can reproduce the intricate communication between hematopoietic stem and progenitor cells and their environment. Genetically engineered mice utilize specific mutations and may not represent the entire array of human MDS; however, genetically engineered mice provided in vivo proof of principle for novel agents such as luspatercept, demonstrating the clinical utility of this approach. This review offers an overview of available preclinical MDS models and potential approaches to accelerate accurate clinical translation.
Similar content being viewed by others
Introduction
Myelodysplastic neoplasms (MDS), also referred to as myelodysplastic syndromes and abbreviated MDS, are a group of malignant disorders of hematopoietic differentiation that are characterized by ineffective hematopoiesis and morphologic dysplasia, leading to abnormal peripheral blood counts, infectious complications, and increased risk of transformation to acute myeloid leukemia (AML) [1,2,3]. MDS represents a complex disease entity whose pathogenesis stems from an interplay between microenvironment, genetic and epigenetic elements. Compared to AML, advances in MDS therapies are lagging, in large part due to the paucity of successful preclinical models capable of reproducing the complex genetic landscape of this disease. MDS hematopoietic stem cells (HSCs) do not show long term proliferation in vitro [4] and available MDS cell lines are scarce [4]. In addition, xenotransplantation of human MDS cells in immunocompromised murine hosts has been limited by poor engraftment and maintenance of MDS in the host [5] not to mention the inability to reproduce a human bone marrow microenvironment (BME). Other efforts that generated genetically engineered mice (GEM) based on recurrent mutations identified in MDS patients failed to fully replicate all features of human MDS with numerous clinical, morphologic, and genetic aspects of the disease often lacking [6, 7] This made developing therapeutic strategies targeting recurrent genetic abnormalities very challenging. More recent efforts attempted to create “humanized” 3D structures that can mimic the human BME [8, 9]. These models can be quite sophisticated but are far from perfect tools as they require high level resources and expertise. In this synopsis, we will review the evolution of preclinical models from the simple single cell in-vitro systems to the complex in-vivo 3D models.
In vitro MDS cell lines
Numerous investigators have attempted to generate MDS cell lines, however, these efforts have been largely unsuccessful. An ideal MDS cell line would generate consistent and reproducible results, be easily transferable, and interactive with any artificial matrix that models the BME. Drexler et al. reviewed 31 putative MDS cell lines and used genomic DNA profiling and cytogenetic analysis, to assign each cell line into one of three categories: (1) false/non -malignant cell lines, (2) malignant cell lines in the leukemic phase and (3) valid MDS cell lines (3 of 31) [4]. Cell lines considered a valid representation of the MDS phenotype were M-TAT, TER-3, MDS92, and their derivatives (Table 1).
The M-TAT cell line was isolated from the peripheral blood of a 3-year-old patient with refractory anemia with excess blasts MDS (RAEB-MDS). These cells were responsive to various cytokines (such as erythropoietin (EPO) and granulocyte-macrophage colony-stimulating factor (GM-CSF)) for growth and maturation of their megakaryocytic and erythroid lineages [10].
The MDS92 cell line was derived from the bone marrow of a 52-year-old patient with refractory anemia and ringed sideroblasts (MDS-RARS) [11]. These cells were not in the leukemic stage and were able to be maintained in vivo for prolonged periods of time while retaining the ability to grow and mature with characteristics that are consistent with MDS. In addition, MDS92 cells proliferated in response to interleukin-3 (IL3) and were characterized by the deletion of 5q chromosome [del(5q)], monosomy 7 and a point mutation in codon 12 of the N-RAS oncogene [11, 12]. A blastic line, MDS-L, was later derived from MDS92 and was instrumental for the study of MDS with del(5q) as well as delineation of the therapeutic effects of lenalidomide on del(5q) MDS [13,14,15]. Additional blastic sublines were independently isolated from MDS92 and include the MDS-L-2007 and MDS-LGF. Kida et al. described these two major subclones of MDS-L and demonstrated that MDS-L-2007 proliferated in response to high dose (100 ng/mL) IL-3 while the MDS-LGF proliferated in response to much lower doses (1ng/mL). Authors also provided data on the relationship between histone H3 K27M mutation and IL-3 dependence. This data provided crucial evidence that the survival of MDS clones depends on an interplay between environmental factors and intrinsic genetic and epigenetic properties [12].
In 2002, Mishima et al. described TER-3, a new hematopoietic cell line derived from a patient that had recently progressed from refractory anemia (RA) to RAEB [16]. The TER-3 cell line had a complex karyotype and, similar to MDS92, displayed monosomy 7 and no chromosome 5 rearrangements. Cells were found to be constitutively dependent on cytokine signaling for growth and had the potential to differentiate into erythroid and megakaryocytic lineages [16]. Ultimately, Drexler and colleagues concluded that, given its availability and cytogenetic profile that includes a 5q deletion, the MDS92 cell line seemed to be the most promising in vitro MDS model [4].
Given the limited proliferative potential of MDS cell lines as well as the scarcity of dependable in vitro models, the study of genetically engineered murine models and xenografted mice emerged as alternate approaches. These in vivo models may provide a more realistic representation of human MDS as they allow the interaction between mammalian cells and their microenvironment.
Xenograft models
Patient-derived xenografts (PDX) represent an additional option to reproduce disease complexity in vivo and test therapeutic interventions. To generate a successful PDX model, an immunodeficient recipient is required to avoid immune rejection of the human cells. A landmark study demonstrated that a small subset of AML cells could be transplanted to Scid (severe combined immune deficient) mice; this study introduced the concept of cancer stem cells [17]. Combination of a non-obese diabetic (NOD) and severe combined immunodeficiency (SCID) strains resulted in mice that had defective T, B, and NK function [16, 17]. These models were further enhanced by removal of the common gamma chain of the interleukin 2 receptor (IL2rδnull) resulting in NOD SCID Gamma (NSG) mice that were more permissive recipients and allowed for significantly higher rates of AML engraftment [18,19,20].
Despite improved engraftment of AML cells in NSG mice, engraftment of MDS cells remained poor, therefore, additional alterations were made to improve the BME and support trilineage maturation of human progenitor cells [21, 22]. Introducing human cytokine genes into the NSG mouse genome led to the constitutive expression of stem cell factor (SCF), IL3 and granulocyte monocyte colony-stimulating-factor (GM-CSF) resulting in engraftment and expansion of myeloid cells (NSGS mice) [23, 24]. However, most MDS subtypes (except chronic myelomonocytic leukemia) did not engraft [25, 26]. A similar approach was used to generate MISTRG mice, in which human alleles of, macrophage-CSF (M-CSF), IL3, GM-CSF, signal regulatory protein alpha (SIRPα) and thrombopoietin (TPO) replaced their murine counterparts via homologous recombination; these alleles were then crossed onto an immunodeficient background (Rag−/−IL2rg−/−). This resulted in robust but transient engraftment of CD33+ myeloid cells and some cases of MDS [27,28,29,30]. The MISTRG model is promising in that engraftment was not limited to any specific MDS subtype, and MISTRG MDS PDX mice reliably reproduced patients’ dysplastic features and captured their mutational profiles and genetic complexities.
Potential pre-clinical utility of PDX models has been demonstrated by several investigators. In one study, MISTRG mice were engrafted with human MDS cells expressing an IDH2 R140Q mutant protein and treated with an oral IDH2 inhibitor. This resulted in differentiation of IDH2 mutant blasts and myeloid differentiation of the engrafted MDS cells [30]. Although there was no report of survival benefit or improved peripheral blood counts in the treated mice, these observations demonstrate the potential for this model in evaluating innovative and targeted therapies, alone or in combination. An additional study focused on treating mice engrafted with high risk MDS cells using omacetaxine mepesuccinate, a protein synthesis inhibitor [31]. These investigators showed a decrease in engraftment following omacetaxine treatment, which could be potentiated by adding 5-azacytidine or venetoclax. As an alternate to small molecules, antibodies to CD117 [the receptor for SCF] was shown to deplete human MDS stem cells using an in vivo xenograft model [32].
“Humanizing” the BM microenvironment
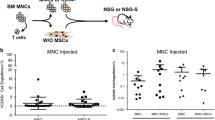
Generation of a reliable and readily transplantable MDS PDX remains a challenge due to inadequate engraftment and poor maintenance of donor MDS progenitor cells in the murine host. Some studies suggest a role for the BME in supporting MDS stem cell proliferation and differentiation [33]. This permissive “milieu” relies on the presence of human cytokines and mesenchymal stromal cells (MSCs), both of which have been shown to help engraft MDS cells under specific conditions [33,34,35,36]; a summary of humanized approaches is displayed in Fig. 1. Several research groups have attempted to co-engraft MSCs and MDS cells to promote stem cell myeloid differentiation and persistence. In one study, CD34+ bone marrow cells were intrafemorally injected alongside human stromal cells into NOG (NOD/Shi-scid/IL-2Rγnull) mice and the percentage of human CD45+ engraftment was as high as 89% with suppression of murine host hematopoiesis [35]. Similarly, Medyouf et al. demonstrated that the co-injection of patient-derived MDS CD34+ cells with MSCs led to efficient engraftments (70%) in NSG mice with the resulting xenograft model displaying dysplasia and characteristic molecular lesions [36]. This approach has been used as a pre-clinical platform to assess the efficacy of eltrombopag in producing platelet support for MDS in vivo [37], as well as to assess the ability of PXS-5505 (a pan-lysyl oxidase inhibitor) and 5-azacytidine to augment erythroid differentiation in vivo [38].
A common denominator of these models was sub-optimal, but improved, engraftment, leading to the concept that a BME that more closely mimicked human bone marrow might improve engraftment of primary MDS cells [39, 40]. The reasons behind sub-optimal engraftment are many and likely related to species-specific environmental requirements that promote stem cell survival, homing, and engraftment [23].
HSCs reside and flourish in the bone marrow and interact with an intricate network of cells that include MSCs, Schwann cells, vascular endothelial cells, and osteoblasts, all of which are necessary for proper function of HSCs [33, 41]. These “niches” are thought to support both hematopoietic and leukemic stem cells. Transfer of a scaffold structure that combines human hematopoietic BM cells with nonhematopoietic cells (referred to as an “ossicle”) into an immunodeficient mouse, can generate a “humanized” BM niche [8, 9, 42]. Reinisch et al. developed a humanized BMME by subcutaneous injection of extracellular matrix material with immature MSCs leading to ossicle formation in vivo. Following daily injection of human parathyroid hormone, these ossicles were able to support engraftment of human CD34+ hematopoietic stem and progenitor cells (HSPC). Compared to unmanipulated NSG mice, these “humanized” ossicles led to more robust engraftment of HSPCs as well as AML, acute promyelocytic leukemia (APL), and myelofibrosis samples [8].
Altrock et al. evaluated the feasibility of this approach [8, 9] in MDS and used a standardized approach to compare the 3D ossicle-based method to intrafemoral (IF) co-injection of MSCs and CD34+ HSPCs [41]. A combination of immunohistochemistry, flow cytometry and molecular analysis demonstrated that 3D humanized ossicle xenotransplantation had a significantly better engraftment rate of MDS samples compared to the historical IF injection of MSCs [36, 43]. A potential drawback of the 3D model was the poor recovery rates of MDS cells from the humanized matrices, which would impede downstream functional and molecular analyses. They suggested immortalizing matrix components and adding endothelial cells to help alleviate that concern [40, 41].
Additional studies have used the concept of 3D scaffold generation to produce ceramic matrices coated with MDS MSCs which were subsequently implanted into immunodeficient mice. The resulting human niche was associated with engraftment of AML samples across all major molecular and risk subgroups [44]. Abarrategi et al. used a gelatin-based scaffold to test distinct implantable niches (osteoblastic, endothelial and MSCs). The authors found that use of this scaffold was associated with improved engraftment of human AML samples compared with other approaches; it is important to note that these studies used human AML and not MDS samples [45]. Another study used a gelatin-based scaffold to create an in vivo vascularized disease model that allowed engraftment of 94% of MDS HSC patient samples irrespective of MDS subtype [46]. Of note, disease-associated “aging” was also demonstrated in this model with a trend toward myeloid differentiation, particularly, in high-risk samples [46].
In addition to vascularizing the niche, co-culturing of HSPCs with MSCs, and adding a 3D structural matrix, other elements of the human BM have been used to help mimic the dynamic interaction between hematopoietic cells and microenvironment. Khan et al. developed vascularized “living” organoids that incorporated human pluripotent stem cells that could generate myeloid and mesenchymal elements as well as vascular sinusoidal structures that can mimic the 3D architecture of the bone marrow [47]. Multimodal imaging and single cell RNA sequencing were used to demonstrate similarity between these organoids and human BMME. These organoids were also shown to support engraftment and proliferation of healthy and malignant human cells [47]. Proof of disease reproducibility was provided when organoid “remodeling and fibrosis” occurred after engraftment of myelofibrosis patients’ cells though application of fibrosis inhibitors such as TGFβ and BET inhibitors and ruxolitinib did not reverse hallmarks of fibrosis. Although MDS was not evaluated with this model, other diseases included were AML, acute lymphoblastic leukemia (ALL), chronic myelogenous leukemia (CML) and multiple myeloma [47]. Despite an impressive homology between this organoid’s vascular system and the human BM endothelial system, this model failed to reproduce adipocytes, lymphocytes and smooth muscle cells, limitations that likely could be enhanced by optimizing the growth factors, chemokines, and cytokines in this “hematopoietic milieu”.
Genetically engineered mice
In 1999, The Mouse Models of Human Cancers Consortium (MMHCC) emerged as an effort by the National Cancer Institute to develop mouse models of cancer that were corroborated by the scientific committee and used to better understand different pathways and processes involved in cancer development [48]. In 2002, the hematopathology subcommittee of the MMHCC, developed a consensus recommendation for the classification of hematopoietic neoplasms, including MDS, in murine models. Using clinical course, blood counts, histopathology and immunophenotypic features, they identified myeloid dysplasia which was defined by the presence of cytopenias in the peripheral blood of mice as well as dysplasia in at least 1 of the 3 myeloid lineages [48, 49]. Unlike human MDS, where the presence of dysplasia in 10% of a particular lineage, continues to hold as a diagnostic threshold, the minimal required dysplasia in mice is yet to be defined or validated. Nonetheless, this classification has provided scientists, pathologists, and investigators, with a standardized tool to diagnose MDS in GEM allowing these models to be reliably used to test novel diagnostic and treatment approaches to cancer management.
There are several approaches that have been used to generate MDS in mice; these approaches are distinct from xenografts, which engraft human MDS cells into mouse hosts. The first includes treating mice with known mutagens, such as benzene [50, 51], alkylating agents [52, 53] or viruses [54]. These strategies resulted in a wide spectrum of complex genetic phenotypes including MDS. However, despite providing insight into certain phenotypic aspects of MDS, these mutagenic approaches were largely unable to reliably define contributions of particular genes. In addition, since these approaches depended largely on random mutagenic events, they were not readily reproducible.
Two additional approaches are based on using mutations identified in patients with MDS to generate genetically engineered HSPC. Both approaches have been successful in replicating some, but not all, features of MDS, with certain clinical, morphological, or genetic aspects of the murine model disease lacking when compared to human MDS. In the first approach, murine HSPC are modified to express a mutant gene in vitro, typically using retroviral or lentiviral vectors. The modified HSPC are then transplanted into syngeneic hosts that have been lethally irradiated. Given the requirement for transplantation, this approach is primarily limited to the study of hematologic malignancy, and not easily adapted to the study of solid tumors, such as lung cancer.
A second general approach includes the creation of GEM models that have modified the germline mouse DNA. Transgenic, “knock-out” or “knock-in” mice, can be produced by a variety of techniques, including pronuclear injection of produced by injecting DNA into a fertilized egg or by homologous recombination of embryonic cells followed by blastocyst injection [55]. The general technique for generating GEM models has been refined and modified over the past forty years to allow use of endogenous promoters, as well as tissue specific and temporal expression of mutant genes; advantages and disadvantages for several of these modifications are listed in Table 2. These manipulations lead to modification of germline DNA, which can then be transmitted from generation to generation. A large cohort of mice with an identical genetic modification can then be observed to determine the incidence of MDS in the modified mice.
Although there are several advantages of using GEM models, such as knowledge of specific genes modified, transferability of GEM between laboratories, reproducibility, and ability to generate large numbers of genetically identical mice, there are several disadvantages as well. The mouse hematopoietic system, while similar to human, is not identical [56]. These mice are generally kept in specific pathogen free environments, which may not adequately mimic the diverse microbiome seen in humans. In addition, many investigators prefer to use inbred mice, which simplifies genetic analysis and facilitates HSC transplant experiments. This strategy may not adequately recapitulate human genetic diversity. Finally, single engineered mutations may be insufficient to adequately mimic the heterogeneity seen in human MDS, which may require combinations of mutant genes to produce a phenotype. In fact, several highly penetrant GEM models of leukemia are linked to spontaneous acquisition of somatic mutations, which collaborate with the mutation engineered in the mouse germline [57, 58].
TET2
Somatic mutations in the ten-eleven translocation 2 (TET2) gene, leading to loss of function, are found in up to 20% of MDS patients [59]. TET2 was shown to play a role in hematopoietic development, myeloid differentiation [60] and when mutated, malignant transformation [61,62,63]. Transplantation of Tet2−/− cells into lethally irradiated mice resulted in a CMML-like disease characterized by myeloid dysplasia in bone marrow as well as splenomegaly, neutrophilia, monocytosis and extramedullary hematopoiesis. Tet2 monoallelic loss had a similar but less pronounced phenotype [62].
EZH2
EZH2 is a histone methyltransferase located on chromosome 7 and found to be inactivated in clonal myeloid disorders such as MDS and MPN [64, 65]. Ezh2 knockout mice (Ezh2−/−) developed myelodysplastic disorders characterized by anemia, splenomegaly, and dysplasia of the bone marrow, and the concurrent loss of both Tet2 and Ezh2 resulted in either an MDS phenotype (pancytopenia, myelodysplasia) or MDS/MPN phenotype (monocytosis, splenomegaly) [63]. Interestingly, although latency of disease progression was markedly shortened when both genes were deleted, this genetic combination showed little to no leukemogenic potential. Consistent with observations that EZH2 mutated MDS have a very low risk of AML progression [66, 67], no Ezh2−/− or Tet2−/−Ezh2−/− mice developed AML in their experiments [63].
U2AF1
Mutations in spliceosome genes are found in more than 80% of MDS patients [68]. Spliceosome gene aberrations are thought to often be an MDS initiating event that arises early in the disease process and disrupts healthy hematopoiesis [69, 70]. Mouse models that express mutant genes based on point mutations that have been identified in patients with MDS have been developed for some of these mutations including SF3B1, SRSF2 and U2AF1 [71,72,73,74]. Transgenic mice expressing a U2af1 S34F mutation were reported by Shirai et al. [72]. Cells containing a doxycycline-inducible human mutated (mut) U2AF1 or control U2AF1 cDNA were transplanted to lethally irradiated recipients. Following engraftment, both mut U2AF1 and control U2AF1 recipients received doxycycline [75]. Mice with mutated U2AF1 treated with doxycycline developed an isolated peripheral leukopenia (normal platelet counts and red blood cells) while their marrows had increased neutrophil count, apoptotic activity and reduced monocytes and B cells. Prolonged follow up after transplantation (500 days) did not reveal any dysplasia, MDS, or AML development [75]. More recently, U2af1 knock-in models with Cre-dependent knock-in alleles of U2af1S34F were generated and led to dysplasia, macrocytic anemia and leukopenia, but did not demonstrate decreased survival or increased leukemogenesis [76]. This model helped identify cooperating mutations via a study in which a combination of U2af1S34F and Runx1 deficiency led to abnormal hematopoiesis, but not MDS or AML. However, after mutagenesis with ethyl-nitrosourea, a minority of mice developed AML accompanied by mutations in Ikzf1, Idh1 and Gata2 [76].
SF3B1
SF3B1 mutations characterize a subset of MDS with RA, ring sideroblasts, and favorable outcome [72]. Several animal models have attempted to reproduce certain aspects of SF3B1 mutated MDS with varying degrees of success. Obeng et al. created an Sf3b1K700E model in which mice that expressed a SF3B1K700E “knock-in” allele developed erythroid dysplasia and macrocytic anemia but not clear MDS [77]. Independently, and using a “minigene” approach, Mupo et al. developed an Sf3b1K700E model that seemed to have an adequate heterozygous allele expression (50% by RNA-seq analysis) but failed to phenotypically produce disease: There was no difference in overall survival between Sf3b1K700E mice and wild-type, and no mouse developed MDS. Despite progressive normocytic anemia, platelet counts and white blood cell counts remained normal and morphological assessment of the bone marrow revealed no dysplasia or ring sideroblasts [78].
SRSF2
Mutations in SRSF2 are found in up to 15% of MDS patients and often associated with poorer outcomes and a shorter duration to leukemic transformation [71, 79]. Kon et al. developed a knock-in model in which Srsf2P95H/+ mice developed macrocytic anemia by week 15, along with leukopenia and a marked reduction in progenitor cell population. Dysplastic changes in erythroid and megakaryocytic lineages in the bone marrow were detected, although none of the mice developed MDS or AML by week 90 [80]. Another knock-in model, Srsf2P95H/+ mice, was developed by Smeets et al. who confirmed heterozygous expression and transcription by RNA-seq. Srsf2P95H/+ mice developed findings consistent with MDS by 12 months of age, with evidence of morphologic dysplasia in myeloid and erythroid lineages in bone marrow and peripheral blood, without progression into AML. Gene expression analysis found genetic signatures consistent with myeloid differentiation, loss of lymphoid maturation potential and MDS progression [81]. Unlike previous reports, in which Srsf2P95H/+ cells displayed poor engraftment, Smeets et al. were able to overcome this limitation by modifying the bone marrow competitor [81].
DICER
There is evidence that MDS pathogenesis can be driven, at least in part, by an interplay between somatic mutations and the BME [82]. This interaction is necessary for disease progression and maintenance of the malignant clone. Mimicking some of the dysfunction observed in the BME of MDS patients has been used to reproduce disease and create models to better understand it. Dicer1 is an RNase III endonuclease that acts to process pre-miRNA and synthesize microRNA necessary to regulate diverse cellular functions including hematopoiesis [83, 84]. Dicer1 deletion led to global downregulation of microRNAs and drove tumorigenesis in a mouse model of lung cancer mediated by expression of a mutant form of Kras [85]. In a similar effort, Raaijmakers et al. generated mice that deleted Dicer1 in a subset of mesenchymal osteolineage bone marrow cells (designated Oxs-GFP-Cre+Dicerfl/fl or OCDfl/fl) [86]. OCDfl/fl mice showed stunted growth with only 70% survival by week 8. Analysis at 4–6 weeks revealed impaired osteoblastic differentiation and decreased bone marrow mineral matrix deposition. Of interest, OCDfl/fl mice demonstrated ineffective hematopoiesis characterized by decreased peripheral blood counts (marked leukopenia and variable anemia and thrombocytopenia) despite having normal to increased bone marrow cellularity. Morphological examination of their bone marrows revealed dysplastic features, which, along with peripheral cytopenias, were consistent with a diagnosis of MDS per the Bethesda criteria [48, 86]. Other findings that are characteristic of human MDS included increased growth and apoptosis of primitive hematopoietic progenitor cells, decreased B-cell progenitor cells, marked vascularity of the bone marrow and a preferential maturation toward myeloid lineages. In addition, the authors demonstrated the role of BME in MDS pathogenesis by transplanting OCDfl/fl HSCs from mice with clinical MDS onto healthy wild-type recipients. Despite complete donor chimerism, indicating robust engrafted of the mutant OCDfl/fl cells, transplant recipients had normal peripheral blood counts and no dysplastic features. Alternatively, when HSCs from healthy wild-type mice were transplanted onto lethally irradiated OCDfl/fl mice, the wild-type cells developed features of MDS. This series of experiments demonstrated that the OCDfl/fl mouse model was successful at reproducing clinical MDS and delineating the interplay between disease pathogenesis and BME [86].
NUP98 translocations and MDS
The NUP98 gene is present on chromosome 11p15.5 and encodes a component of the nuclear pore complex, which can also function as a transcription scaffold [87]. Fusion proteins resulting from chromosomal translocations involving the NUP98 gene have been identified in lymphoid and myeloid malignancies and act as oncoproteins that can drive malignant transformation [88, 89]. NUP98 fused to several clustered homeobox (HOX) genes, including HOXA9, HOXA13, and HOXD13 have been identified in patients with MDS and AML [90]. Pineault et al. generated a NUP98-HOXD13 murine model in which mouse bone marrow stem cells that expressed a NUP98-HOXD13 fusion gene were generated via retroviral transduction. Colony forming unit spleen assay analysis of transduced bone marrow cells showed that myeloid cells (Gr+/Mac1+) were increased while erythroid precursors were reduced at day 12. In that study, some NUP98-HOXD13 mice developed myeloproliferative disease at the 4-week post-transplant mark but none progressed to AML (except those engineered to concurrently express the TALE homeobox gene Meis1, a strong mediator of NUP98-associated leukemia [91] during a 6-month study period [92].
Lin et al. developed NUP98-HOXD13 (NHD13) transgenic mice that utilized the HS21/45-Vav vector [93] to direct the expression of a human NHD13 fusion cDNA in hematopoietic tissues of mice. Like human MDS, NHD13 mice developed dysplasia and peripheral blood cytopenias in the setting of a hypercellular marrow, demonstrating ineffective hematopoiesis. In addition, this model reproduced the natural progression of MDS, as NHD13 transgenic mice show mild-moderate anemia and remain healthy for an extended period (typically 6–10 months) but develop progressively worsening cytopenias or transform to acute leukemia, most commonly between 10 and 14 months, with >90% of mice surviving less than 14 months [93]. Progression to acute leukemia was accompanied by spontaneous acquired mutations in genes commonly mutated in human MDS, such as Nras, Kras, Ptpn11, and Cbl [57].
The MDS seen in NHD13 mice was cell-autonomous, as WT mice transplanted with NHD13 bone marrow inevitably developed MDS, characterized by outcompeting WT bone marrow, macrocytic anemia, dysplasia, and transformation to AML. Additional transplant assays analyzed BME cellular elements from NHD13 mice, and found increased endothelial cells, decreased megakaryocytes and dysfunctional osteoblastic and mesenchymal cell populations [94]. Inflammatory cytokines shown to be increased in human MDS were also elevated in NHD13 mice suggesting that this model may capture aspects of the interplay between BMME and MDS pathogenesis [94]. In addition, transplantation of MHC matched WT bone marrow into NHD13 recipients led to increased survival, but not cure, of the transplant recipients, while transplant of MHC mismatched bone marrow led to an increased graft versus tumor effect [95]. The utility of this model for pre-clinical drug development is underscored by the fact that NHD13 mice were used to provide proof-of-concept for luspatercept, the only new drug to receive FDA approval for MDS in the past decade (100,101).
Conclusion
Successful MDS models should be reproducible, user-friendly, and dynamic tools that are able to accurately replicate the complex biological features of MDS. Ideal models would be “living” entities that reproduce the intricate communication between HSCs and their environment. In concept, these ideal models would allow one to individualize approach and evaluate treatment regimens while avoiding toxicities. However, although substantial progress has been made, such ideal models do not yet exist. But the MDS research community should not let “the perfect be the enemy of the good.” Despite the lack of ideal models, drugs (such as luspatercept) continue to be developed for patients with MDS using the available, less-than-ideal models described above. With the increased efforts over the past decade comes optimism that increasingly useful and dependable tools which accurately mimic MDS will continue to evolve.
Disclaimer
The views expressed in this work do not represent the official views of the National Institutes of Health or the United States Government.
References
Papaemmanuil E, Gerstung M, Malcovati L, Tauro S, Gundem G, Van Loo P, et al. Clinical and biological implications of driver mutations in myelodysplastic syndromes. Blood. 2013;122:3616–27.
Khoury JD, Solary E, Abla O, Akkari Y, Alaggio R, Apperley JF, et al. The 5th edition of the World Health Organization classification of haematolymphoid tumours: myeloid and histiocytic/dendritic neoplasms. Leukemia. 2022;36:1703–19.
Arber DA, Orazi A, Hasserjian RP, Borowitz MJ, Calvo KR, Kvasnicka HM, et al. International Consensus Classification of Myeloid Neoplasms and Acute Leukemias: integrating morphologic, clinical, and genomic data. Blood. 2022;140:1200–28.
Drexler HG, Dirks WG, Macleod RAF. Many are called MDS cell lines: one is chosen. Leuk Res. 2009;33:1011–6.
Martin MG, Welch JS, Uy GL, Fehniger TA, Kulkarni S, Duncavage EJ, et al. Limited engraftment of low-risk myelodysplastic syndrome cells in NOD/SCID gamma-C chain knockout mice. Leukemia. 2010;24:1662–4.
Rhyasen GW, Wunderlich M, Tohyama K, Garcia-Manero G, Mulloy JC, Starczynowski DT. An MDS xenograft model utilizing a patient-derived cell line. Leukemia. 2014;28:1142–5.
Gbyli R, Song Y, Halene S. Humanized mice as preclinical models for myeloid malignancies. Biochem Pharm. 2020;174:113794.
Reinisch A, Thomas D, Corces MR, Zhang X, Gratzinger D, Hong WJ, et al. A humanized bone marrow ossicle xenotransplantation model enables improved engraftment of healthy and leukemic human hematopoietic cells. Nat Med. 2016;22:812–21.
Reinisch A, Hernandez DC, Schallmoser K, Majeti R. Generation and use of a humanized bone-marrow-ossicle niche for hematopoietic xenotransplantation into mice. Nat Protoc. 2017;12:2169–88.
Minegishi N, Minegishi M, Tsuchiya S, Fujie H, Nagai T, Hayashi N, et al. Erythropoietin-dependent induction of hemoglobin synthesis in a cytokine-dependent cell line M-TAT. J Biol Chem. 1994;269:27700–4.
Tohyama K, Tsutani H, Ueda T, Nakamura T, Yoshida Y. Establishment and characterization of a novel myeloid cell line from the bone marrow of a patient with the myelodysplastic syndrome. Br J Haematol. 1994;87:235–42.
Kida JI, Tsujioka T, Suemori SI, Okamoto S, Sakakibara K, Takahata T, et al. An MDS-derived cell line and a series of its sublines serve as an in vitro model for the leukemic evolution of MDS. Leukemia. 2018;32:1846–50.
Matsuoka A, Tochigi A, Kishimoto M, Nakahara T, Kondo T, Tsujioka T, et al. Lenalidomide induces cell death in an MDS-derived cell line with deletion of chromosome 5q by inhibition of cytokinesis. Leukemia. 2010;24:748–55.
Krönke J, Fink EC, Hollenbach PW, MacBeth KJ, Hurst SN, Udeshi ND, et al. Lenalidomide induces ubiquitination and degradation of CK1α in del(5q) MDS. Nature. 2015;523:183–8.
Fang J, Liu X, Bolanos L, Barker B, Rigolino C, Cortelezzi A, et al. A calcium- and calpain-dependent pathway determines the response to lenalidomide in myelodysplastic syndromes. Nat Med. 2016;22:727–34.
Mishima Y, Terui Y, Mishima Y, Katsuyama M, Mori M, Tomizuka H, et al. New human myelodysplastic cell line, TER-3: G-CSF specific downregulation of Ca2+/calmodulin-dependent protein kinase IV. J Cell Physiol. 2002;191:183–90.
Lapidot T, Sirard C, Vormoor J, Murdoch B, Hoang T, Caceres-Cortes J, et al. A cell initiating human acute myeloid leukaemia after transplantation into SCID mice. Nature. 1994;367:645–8.
Bosma GC, Custer RP, Bosma MJ. A severe combined immunodeficiency mutation in the mouse. Nature 1983;301:527–30.
Prochazka M, Gaskins HR, Shultz LD, Leiter EH. The nonobese diabetic scid mouse: model for spontaneous thymomagenesis associated with immunodeficiency. Proc Natl Acad Sci USA. 1992;89:3290–4.
Shultz LD, Lyons BL, Burzenski LM, Gott B, Chen X, Chaleff S, et al. Human lymphoid and myeloid cell development in NOD/LtSz-scid IL2R gamma null mice engrafted with mobilized human hemopoietic stem cells. J Immunol. 2005;174:6477–89.
Brehm MA, Cuthbert A, Yang C, Miller DM, DiIorio P, Laning J, et al. Parameters for establishing humanized mouse models to study human immunity: analysis of human hematopoietic stem cell engraftment in three immunodeficient strains of mice bearing the IL2rgamma(null) mutation. Clin Immunol. 2010;135:84–98.
Ito R, Takahashi T, Katano I, Ito M. Current advances in humanized mouse models. Cell Mol Immunol. 2012;9:208–14.
Krevvata M, Shan X, Zhou C, Dos Santos C, Habineza Ndikuyeze G, Secreto A, et al. Cytokines increase engraftment of human acute myeloid leukemia cells in immunocompromised mice but not engraftment of human myelodysplastic syndrome cells. Haematologica. 2018;103:959–71.
Coughlan AM, Harmon C, Whelan S, O’Brien EC, O’Reilly VP, Crotty P, et al. Myeloid engraftment in humanized mice: impact of granulocyte-colony stimulating factor treatment and transgenic mouse strain. Stem Cells Dev. 2016;25:530–41.
Zhang Y, He L, Selimoglu-Buet D, Jego C, Morabito M, Willekens C, et al. Engraftment of chronic myelomonocytic leukemia cells in immunocompromised mice supports disease dependency on cytokines. Blood Adv. 2017;1:972–9.
Wunderlich M, Chou FS, Link KA, Mizukawa B, Perry RL, Carroll M, et al. AML xenograft efficiency is significantly improved in NOD/SCID-IL2RG mice constitutively expressing human SCF, GM-CSF and IL-3. Leukemia. 2010;24:1785–8.
Rongvaux A, Willinger T, Martinek J, Strowig T, Gearty SV, Teichmann LL. et al. Development and function of human innate immune cells in a humanized mouse model. Nat Biotechnol. 2014;32:364–72.
Saito Y, Ellegast JM, Rafiei A, Song Y, Kull D, Heikenwalder M, et al. Peripheral blood CD34+ cells efficiently engraft human cytokine knock-in mice. Blood. 2016;128:1829–33.
Sippel TR, Radtke S, Olsen TM, Kiem HP, Rongvaux A. Human hematopoietic stem cell maintenance and myeloid cell development in next-generation humanized mouse models. Blood Adv. 2019;3:268–74.
Song Y, Rongvaux A, Taylor A, Jiang T, Tebaldi T, Balasubramanian K, et al. A highly efficient and faithful MDS patient-derived xenotransplantation model for pre-clinical studies. Nat Commun. 2019;10:366.
Stevens BM, Khan N, D’Alessandro A, Nemkov T, Winters A, Jones CL, et al. Characterization and targeting of malignant stem cells in patients with advanced myelodysplastic syndromes. Nat Commun. 2018;9:3694.
Pang WW, Czechowicz A, Logan AC, Bhardwaj R, Poyser J, Park CY, et al. Anti-CD117 antibody depletes normal and myelodysplastic syndrome human hematopoietic stem cells in xenografted mice. Blood. 2019;133:2069–78.
Mendelson A, Frenette PS. Hematopoietic stem cell niche maintenance during homeostasis and regeneration. Nat Med. 2014;20:833–46.
Thanopoulou E, Cashman J, Kakagianne T, Eaves A, Zoumbos N, Eaves C. Engraftment of NOD/SCID-beta2 microglobulin null mice with multilineage neoplastic cells from patients with myelodysplastic syndrome. Blood. 2004;103:4285–93.
Muguruma Y, Matsushita H, Yahata T, Yumino S, Tanaka Y, Miyachi H, et al. Establishment of a xenograft model of human myelodysplastic syndromes. Haematologica. 2011;96:543–51.
Medyouf H, Mossner M, Jann JC, Nolte F, Raffel S, Herrmann C, et al. Myelodysplastic cells in patients reprogram mesenchymal stromal cells to establish a transplantable stem cell niche disease unit. Cell Stem Cell. 2014;14:824–37.
Schmitt N, Jann JC, Altrock E, Flach J, Danner J, Uhlig S, et al. Preclinical evaluation of eltrombopag in a PDX model of myelodysplastic syndromes. Leukemia. 2022;36:236–47.
Xu Q, Streuer A, Jann JC, Altrock E, Schmitt N, Flach J, et al. Inhibition of lysyl oxidases synergizes with 5-azacytidine to restore erythropoiesis in myelodysplastic and myeloid malignancies. Nat Commun. 2023;14:1497.
Altrock E, Sens-Albert C, Jann JC, Flach J, Riabov V, Schmitt N, et al. Humanized three-dimensional scaffold xenotransplantation models for myelodysplastic syndromes. Exp Hematol. 2022;107:38–50.
Abarrategi A, Foster K, Hamilton A, Mian SA, Passaro D, Gribben J, et al. Versatile humanized niche model enables study of normal and malignant human hematopoiesis. J Clin Investig. 2017;127:543–8.
Rouault-Pierre K, Mian SA, Goulard M, Abarrategi A, Di Tulio A, Smith AE, et al. Preclinical modeling of myelodysplastic syndromes. Leukemia. 2017;31:2702–8.
Schaeffer A, Czlonka E, Tirado-González I, Böse T, Beauvarlet J, Kur I, et al. Development and exploitation of a fully human and modular organotypic bone marrow niche model to study the role of stroma-produced factors in human MDS. Blood. 2020;136:23.
Antonelli A, Noort WA, Jaques J, de Boer B, de Jong-Korlaar R, Brouwers-Vos AZ, et al. Establishing human leukemia xenograft mouse models by implanting human bone marrow-like scaffold-based niches. Blood. 2016;128:2949–59.
Mian SA, Abarrategi A, Kong KL, Rouault-Pierre K, Wood H, Oedekoven CA, et al. Ectopic humanized mesenchymal niche in mice enables robust engraftment of myelodysplastic stem cells. Blood Cancer Discov. 2021;2:135–45.
Khan AO, Rodriguez-Romera A, Reyat JS, Olijnik AA, Colombo M, Wang G, et al. Human bone marrow organoids for disease modeling, discovery, and validation of therapeutic targets in hematologic malignancies. Cancer Discov. 2023;13:364–85.
Kogan SC, Ward JM, Anver MR, Berman JJ, Brayton C, Cardiff RD, et al. Bethesda proposals for classification of nonlymphoid hematopoietic neoplasms in mice. Blood. 2002;100:238–45.
Zhou T, Kinney MC, Scott LM, Zinkel SS, Rebel VI. Revisiting the case for genetically engineered mouse models in human myelodysplastic syndrome research. Blood. 2015;126:1057–68.
Das M, Chaudhuri S, Law S. Benzene exposure-an experimental machinery for induction of myelodysplastic syndrome: stem cell and stem cell niche analysis in the bone marrow. J Stem Cells. 2012;7:43–59.
Li W, Schnatter AR. Benzene risk assessment: does new evidence on myelodysplastic syndrome justify a new approach? Crit Rev Toxicol. 2018;48:417–32.
Fenske TS, McMahon C, Edwin D, Jarvis JC, Cheverud JM, Minn M, et al. Identification of candidate alkylator-induced cancer susceptibility genes by whole genome scanning in mice. Cancer Res. 2006;66:5029–38.
Mahgoub N, Taylor BR, le Beau MM, Gratiot M, Carlson KM, Atwater SK, et al. Myeloid malignancies induced by alkylating agents in Nf1 mice. Blood. 1999;93:3617–23.
Suzuki T, Shen H, Akagi K, Morse HC, Malley JD, Naiman DQ, et al. New genes involved in cancer identified by retroviral tagging. Nat Genet. 2002;32:166–74.
Beachy SH, Aplan PD. Mouse models of myelodysplastic syndromes. Hematol Oncol Clin N Am. 2010;24:361–75.
Parekh C, Crooks GM. Critical differences in hematopoiesis and lymphoid development between humans and mice. J Clin Immunol. 2013;33:711–5.
Slape C, Liu LY, Beachy S, Aplan PD. Leukemic transformation in mice expressing a NUP98-HOXD13 transgene is accompanied by spontaneous mutations in Nras, Kras, and Cbl. Blood 2008;112:2017–9.
Gough SM, Goldberg L, Pineda M, Walker RL, Zhu YJ, Bilke S, et al. Progenitor B-1 B-cell acute lymphoblastic leukemia is associated with collaborative mutations in 3 critical pathways. Blood Adv. 2017;1:1749–59.
Delhommeau F, Dupont S, della Valle V, James C, Trannoy S, Massé A, et al. Mutation in TET2 in myeloid cancers. N Engl J Med. 2009;360:2289–301.
Figueroa ME, Abdel-Wahab O, Lu C, Ward PS, Patel J, Shih A, et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell. 2010;18:553–67.
Ko M, Huang Y, Jankowska AM, Pape UJ, Tahiliani M, Bandukwala HS, et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature. 2010;468:839–43.
Moran-Crusio K, Reavie L, Shih A, Abdel-Wahab O, Ndiaye-Lobry D, Lobry C, et al. Tet2 loss leads to increased hematopoietic stem cell self-renewal and myeloid transformation. Cancer Cell. 2011;20:11–24.
Muto T, Sashida G, Oshima M, Wendt GR, Mochizuki-Kashio M, Nagata Y, et al. Concurrent loss of Ezh2 and Tet2 cooperates in the pathogenesis of myelodysplastic disorders. J Exp Med. 2013;210:2627–39.
Ernst T, Chase AJ, Score J, Hidalgo-Curtis CE, Bryant C, Jones AV, et al. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat Genet. 2010;42:722–6.
Nikoloski G, Langemeijer SMC, Kuiper RP, Knops R, Massop M. Tönnissen ERLTM, et al. Somatic mutations of the histone methyltransferase gene EZH2 in myelodysplastic syndromes. Nat Genet. 2010;42:665–7.
Raza A, Galili N. The genetic basis of phenotypic heterogeneity in myelodysplastic syndromes. Nat Rev Cancer. 2012;12:849–59.
Shih AH, Abdel-Wahab O, Patel JP, Levine RL. The role of mutations in epigenetic regulators in myeloid malignancies. Nat Rev Cancer. 2012;12:599–612.
Yoshida K, Sanada M, Shiraishi Y, Nowak D, Nagata Y, Yamamoto R, et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature. 2011;478:64–9.
Sperling AS, Gibson CJ, Ebert BL. The genetics of myelodysplastic syndrome: from clonal haematopoiesis to secondary leukaemia. Nat Rev Cancer. 2017;17:5–19.
Haferlach T, Nagata Y, Grossmann V, Okuno Y, Bacher U, Nagae G, et al. Landscape of genetic lesions in 944 patients with myelodysplastic syndromes. Leukemia. 2014;28:241–7.
Kim E, Ilagan JO, Liang Y, Daubner GM, Lee SCW, Ramakrishnan A, et al. SRSF2 mutations contribute to myelodysplasia by mutant-specific effects on exon recognition. Cancer Cell. 2015;27:617–30.
Shirai CL, Ley JN, White BS, Kim S, Tibbitts J, Shao J, et al. Mutant U2AF1 expression alters hematopoiesis and pre-mRNA splicing in vivo. Cancer Cell. 2015;27:631–43.
Smeets MF, Tan SY, Xu JJ, Anande G, Unnikrishnan A, Chalk AM, et al. Srsf2P95H initiates myeloid bias and myelodysplastic/myeloproliferative syndrome from hemopoietic stem cells. Blood. 2018;132:608–21.
Mupo A, Seiler M, Sathiaseelan V, Pance A, Yang Y, Agrawal AA, et al. Hemopoietic-specific Sf3b1-K700E knock-in mice display the splicing defect seen in human MDS but develop anemia without ring sideroblasts. Leukemia. 2017;31:720–7.
Shirai CL, White BS, Tripathi M, Tapia R, Ley JN, Ndonwi M, et al. Mutant U2AF1-expressing cells are sensitive to pharmacological modulation of the spliceosome. Nat Commun. 2017;8:14060.
Fei DL, Zhen T, Durham B, Ferrarone J, Zhang T, Garrett L, et al. Impaired hematopoiesis and leukemia development in mice with a conditional knock-in allele of a mutant splicing factor gene U2af1. Proc Natl Acad Sci USA. 2018;115:E10437–46.
Patnaik MM, Tefferi A. Refractory anemia with ring sideroblasts (RARS) and RARS with thrombocytosis (RARS-T): 2017 update on diagnosis, risk-stratification, and management. Am J Hematol. 2017;92:297–310.
Obeng EA, Chappell RJ, Seiler M, Chen MC, Campagna DR, Schmidt PJ, et al. Physiologic expression of Sf3b1 K700E causes impaired erythropoiesis, aberrant splicing, and sensitivity to therapeutic spliceosome modulation. Cancer Cell. 2016;30:404–17.
Wu SJ, Kuo YY, Hou HA, Li LY, Tseng MH, Huang CF, et al. The clinical implication of SRSF2 mutation in patients with myelodysplastic syndrome and its stability during disease evolution. Blood 2012;120:3106–11.
Gangat N, Patnaik MM, Tefferi A. Myelodysplastic syndromes: contemporary review and how we treat. Am J Hematol. 2016;91:76–89.
Kon A, Yamazaki S, Nannya Y, Kataoka K, Ota Y, Nakagawa MM, et al. Physiological Srsf2 P95H expression causes impaired hematopoietic stem cell functions and aberrant RNA splicing in mice. Blood. 2018;131:621–35.
Mian SA, Bonnet D. Nature or nurture? Role of the bone marrow microenvironment in the genesis and maintenance of myelodysplastic syndromes. Cancers. 2021;13:4116.
Lu J, Guo S, Ebert BL, Zhang H, Peng X, Bosco J, et al. MicroRNA-mediated control of cell fate in megakaryocyte-erythrocyte progenitors. Dev Cell. 2008;14:843–53.
Krol J, Fiszer A, Mykowska A, Sobczak K, de Mezer M, Krzyzosiak WJ. Ribonuclease dicer cleaves triplet repeat hairpins into shorter repeats that silence specific targets. Mol Cell. 2007;25:575–86.
Kumar MS, Lu J, Mercer KL, Golub TR, Jacks T. Impaired microRNA processing enhances cellular transformation and tumorigenesis. Nat Genet. 2007;39:673–7.
Raaijmakers MHGP MukherjeeS, Guo S, Kobayashi ZhangS. T, Schoonmaker JA, et al. Bone progenitor dysfunction induces myelodysplasia and secondary leukaemia. Nature 2010;464:852–7.
Radu A, Moore MS, Blobel G. The peptide repeat domain of nucleoporin Nup98 functions as a docking site in transport across the nuclear pore complex. Cell 1995;81:215–22.
Nakamura T, Largaespada DA, Shaughnessy JD, Jenkins NA, Copeland NG. Cooperative activation of Hoxa and Pbx1-related genes in murine myeloid leukaemias. Nat Genet. 1996;12:149–53.
Borrow J, Shearman AM, Stanton VP, Becher R, Collins T, Williams AJ, et al. The t(7;11)(p15;p15) translocation in acute myeloid leukaemia fuses the genes for nucleoporin NUP98 and class I homeoprotein HOXA9. Nat Genet. 1996;12:159–67.
Gough SM, Slape CI, Aplan PD. NUP98 gene fusions and hematopoietic malignancies: common themes and new biologic insights. Blood. 2011;118:6247–57.
Kawagoe H, Humphries R, Blair A, Sutherland H, Hogge D. Expression of HOX genes, HOX cofactors, and MLL in phenotypically and functionally defined subpopulations of leukemic and normal human hematopoietic cells. Leukemia. 1999;13:687–98.
Pineault N, Buske C, Feuring-Buske M, Abramovich C, Rosten P, Hogge DE, et al. Induction of acute myeloid leukemia in mice by the human leukemia-specific fusion gene NUP98-HOXD13 in concert with Meis1. Blood. 2003;101:4529–38.
Lin YW, Slape C, Zhang Z, Aplan PD. NUP98-HOXD13 transgenic mice develop a highly penetrant, severe myelodysplastic syndrome that progresses to acute leukemia. Blood. 2005;106:287–95.
Balderman SR, Li AJ, Hoffman CM, Frisch BJ, Goodman AN, LaMere MW, et al. Targeting of the bone marrow microenvironment improves outcome in a murine model of myelodysplastic syndrome. Blood. 2016;127:616–25.
Chung YJ, Fry TJ, Aplan PD. Myeloablative hematopoietic stem cell transplantation improves survival but is not curative in a pre-clinical model of myelodysplastic syndrome. PLoS ONE. 2017;12:e0185219.
Kubasch AS, Fenaux P, Platzbecker U. Development of luspatercept to treat ineffective erythropoiesis. Blood Adv. 2021;5:1565–75.
Suragani RNVS, Cadena SM, Cawley SM, Sako D, Mitchell D, Li R, et al. Transforming growth factor-β superfamily ligand trap ACE-536 corrects anemia by promoting late-stage erythropoiesis. Nat Med. 2014;20:408–14.
Acknowledgements
This work was supported by the Intramural Research Program of the National Cancer Institute, National Institutes of Health (grant numbers ZIA SC 010378 and BC 010983).
Funding
Open access funding provided by the National Institutes of Health.
Author information
Authors and Affiliations
Contributions
All three authors, Peter D. Aplan, Steven Pavletic and Alain Mina, contributed to idea generation, provided expertise and were involved in both writing and editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
PDA receives royalties from the NIH Office of Technology Transfer for the invention of NUP98-HOXD13 (NHD13) mice. No other conflict of interest to report.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mina, A., Pavletic, S. & Aplan, P.D. The evolution of preclinical models for myelodysplastic neoplasms. Leukemia 38, 683–691 (2024). https://doi.org/10.1038/s41375-024-02181-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-024-02181-2
- Springer Nature Limited