Abstract
Alu elements are highly abundant primate-specific short interspersed nuclear elements that account for ~10% of the human genome. Due to their preferential location in gene-rich regions, especially in introns and 3′ UTRs, Alu elements can exert regulatory effects on the expression of both host and neighboring genes. When two Alu elements with inverse orientations are positioned in close proximity, their transcription results in the generation of distinct double-stranded RNAs (dsRNAs), known as inverted Alu repeats (IRAlus). IRAlus are key immunogenic self-dsRNAs and post-transcriptional cis-regulatory elements that play a role in circular RNA biogenesis, as well as RNA transport and stability. Recently, IRAlus dsRNAs have emerged as regulators of transcription and activators of Z-DNA-binding proteins. The formation and activity of IRAlus can be modulated through RNA editing and interactions with RNA-binding proteins, and misregulation of IRAlus has been implicated in several immune-associated disorders. In this review, we summarize the emerging functions of IRAlus dsRNAs, the regulatory mechanisms governing IRAlus activity, and their relevance in the pathogenesis of human diseases.
Similar content being viewed by others
Introduction
Alu elements are ~300 base-pair (bp) long primate-specific short interspersed nuclear elements (SINEs) that constitute approximately 10% of the human genome1. They are originated from the fusion of two distinct short arms of 7SL RNA-derived sequences linked by an A-rich region2. They contain an internal RNA polymerase III (Pol III) promoter, which can initiate independent transcription3 and contribute to Alu amplification in the genome with the assistance of autonomous long interspersed nuclear elements, such as ORFp24. Throughout evolution, Alu retrotransposition has given rise to new copies with distinct mutations, leading to the formation of diverse subfamilies characterized by sequence variations and relative ages2.
Due to their high abundance and insertion preference in gene-rich regions, Alu elements can regulate gene expression at the transcriptional, post-transcriptional, and even translational level5. Alu insertions provide locations for DNA methylation and other epigenetic modifications, as Alu sequences are responsible for approximately 25% of CpG dinucleotides in the human genome6. Moreover, Alu elements can act as enhancers and activate the transcription of nearby genes by facilitating 3-dimensional long-range chromosomal interactions7. When transcribed, Alu RNAs impose additional gene regulatory effects independent of Alu amplification. During the heat shock response, Alu RNAs are transcribed by Pol III and can act as trans-acting transcriptional repressors by directly binding to Pol II and blocking transcription at the initiation step8. Interestingly, Alu elements embedded in introns and 3′ UTRs are transcribed as a part of the host mRNAs and serve as key post-transcriptional regulatory elements, with alternative splicing being a well-established biological process that is regulated by Alus. In this context, Alu sequences provide multiple potential splicing donor and acceptor sites for differential usage of splice sites of the host mRNAs9. Finally, Alu insertions in UTRs are known to modulate the translation of host genes by affecting the rate of initiation5,10,11,12.
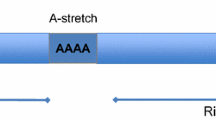
One special feature of Alu elements is provided by their sequence similarities and abundant copy numbers. A single gene typically contains multiple Alu elements with positive and negative orientations13,14. When transcribed, two neighboring Alus elements with opposite orientations can fold back and form an intramolecular hairpin structure with a long double-stranded stem (Fig. 1a). Known as inverted Alu repeats (IRAlus), IRAlus constitute an important class of endogenous double-stranded RNAs (dsRNAs) in human cells14. Originally, dsRNAs were considered hallmarks of viral infection because they are produced during viral transcription or replication15. In vertebrates, viral dsRNAs are recognized by pattern recognition receptors (PRRs), including protein kinase R (PKR) and melanoma differentiation-associated protein 5 (MDA5), and activate antiviral signaling pathways16. Recent studies have shown that endogenous dsRNAs, including IRAlus, can also activate PRRs and induce a viral mimicry state in uninfected cells17,18,19. Consequently, IRAlus are closely associated with immune-related diseases, and modulating IRAlus expression has been suggested as a potential therapeutic strategy for cancer19,20,21. In addition to the well-established immunogenic role, IRAlus participate in post-transcriptional gene regulation through distinct mechanisms of action. Intronic IRAlus, which account for almost half of all IRAlus, are strongly associated with the generation of thousands of circular RNAs (circRNAs) by facilitating the backsplicing of exons22. When IRAlus are located in the 3′ UTR, they suppress gene expression by inhibiting cytosolic export of host mRNAs, sequestering them in nuclear paraspeckles13,23. Therefore, properly regulating IRAlus activity is important, and to do so, cells employ various dsRNA-binding proteins (dsRBPs) that recognize the double-stranded secondary structure of IRAlus20,24,25,26,27. In this review, we aim to summarize recent findings on the multifaceted role of IRAlus and establish IRAlus as an essential gene regulatory element in human pathophysiological conditions.
a Two Alu elements with opposite directions can be present in a single gene. When transcribed, these Alu elements can form an intramolecular dsRNA structure, referred to as IRAlus. These IRAlus are predominantly found within introns and 3′ UTRs. b, c Recognition of IRAlus by dsRNA sensors. b The gain-of-function mutant MDA5 (G495R) undergoes aberrant oligomerization via robust sensing of IRAlus, which subsequently triggers an IFN response, whereas wild-type MDA5 does not normally recognize IRAlus. c PKR is activated by IRAlus and suppresses global translation. d Epigenetic dysregulation via a DNMT inhibitor (DNMTi) triggers cryptic transcription of intronic and intergenic IRAlus, inducing a viral mimicry state. e Disturbance of splicing by a PRMT inhibitor or spliceosome inhibitor induces the accumulation of mis-spliced mRNAs that retain intronic IRAlus and activate the I-IFN response. f RNA processing defects caused by TDP-43 or Dicer deficiency result in the accumulation of Pol III-transcribed Alu RNAs. These unprocessed Alu RNAs initiate the formation of both Alu duplex RNAs and Alu cDNAs, leading to the activation of RIG-I-dependent and cGAS-driven IFN responses, respectively.
Innate immune activation by IRAlus
Human dsRNAome captured using an anti-dsRNA J2 antibody by Kim et al. revealed that IRAlus are the most abundant class of cellular dsRNAs17. Furthermore, genome-wide sequencing analysis indicated that adenosine-to-inosine (A-to-I) editing, which occurs within dsRNAs longer than 50 bp, is observed primarily on IRAlus28. More importantly, an increasing number of reports suggests a close association of self-derived dsRNAs in the abnormal immune activation behind the pathogenesis of immune-related diseases, highlighting the importance of IRAlus regulation in cell pathophysiology14. In this section, we describe the immunogenic function of IRAlus as substrates for dsRNA sensors and share recently uncovered cellular contexts that can modulate IRAlus-mediated antiviral responses, including epigenetic and splicing dysregulation.
IRAlus: immunogenic self-dsRNAs
Our understanding of IRAlus that act as immunogenic self-dsRNAs has been significantly expanded by profiling substrates of dsRNA-recognizing immune sensors, including MDA5 and PKR. When bound to long dsRNAs, MDA5 undergoes filament assembly along the RNA and recruits adapter mitochondrial antiviral signaling (MAVS) proteins to induce a downstream type I interferon (I-IFN) response29. Ahmad et al. developed an MDA5 protection assay to identify endogenous RNA agonists of MDA5 in the cytosol19. The main idea was that a filament of MDA5 along the dsRNA would protect the RNA from RNases. Through such a strategy, the authors revealed that the IRAlus located in the 3′ UTR of mRNAs serve as primary ligands of MDA5. Intriguingly, wild-type MDA5 has limited ability to recognize IRAlus and inefficiently forms the filaments, compared to gain-of-function (GOF) mutant MDA5, which has enhanced RNA binding affinity and is often found in patients with inflammatory diseases, including Aicardi–Goutières syndrome (AGS)30. Indeed, robust recognition of IRAlus by the mutant MDA5 promotes aberrant oligomerization of MDA5 and subsequent activation of the I-IFN response (Fig. 1b).
PKR is another innate immune protein that becomes activated upon binding to IRAlus17,18. To identify PKR-interacting endogenous dsRNAs, Kim et al. performed formaldehyde-mediated crosslinking and immunoprecipitation sequencing (fCLIP-seq) analysis17. The authors utilized formaldehyde to crosslink the dsRNA-RBP complex to overcome the low crosslinking efficiency of UV light on dsRNAs31. The authors found that more than 20% of the dsRNAs that interact with PKR are derived from Alu repeats, nearly all of which are IRAlus (Fig. 1c). In a recent study, intronic and intergenic inverted transposable elements (TEs), including IRAlus, were found to accumulate upon the inhibition of nuclear RNA decay by phosphorothioate DNAs. The increased expression of these inverted TEs resulted in the activation of dsRNA-sensing pathways, thereby providing additional evidence that IRAlus can activate PKR32. Overall, high-throughput sequencing methods tailored to study dsRNAs showed that IRAlus are the prominent source of immunogenic self-dsRNAs that activate multiple dsRNA sensors.
Dysregulation of Alu homeostasis and IRAlus-mediated aberrant innate immune responses
Due to their immunogenic activity, IRAlus generation is suppressed via multiple mechanisms. In most differentiated cells, TEs are subjected to transcriptional silencing through epigenetic mechanisms involving DNA methylation and histone modifications33,34. As a result, treating cells with 5-aza-2′-deoxycyctidine (5-Aza-CdR), a DNA methyltransferase inhibitor (DNMTi), derepressed the transcription of TEs and activated antiviral signaling pathways due to increased expression of self-dsRNAs35,36. In addition, in a recent study, the authors performed the MDA5 protection assay to identify drug-induced TEs; the results revealed enrichment of Alus20. Moreover, these drug treatment-induced Alus were mostly IRAlus originating from intronic and intergenic regions downstream of CpG islands. Intriguingly, some Alus adopted an intramolecular hairpin pairing with adjacent Alus within a single transcript rather than the previously suggested model of hybridization between sense and antisense transcripts. This observation indicated that inhibiting DNA methylation triggers cryptic transcription of IRAlus, which results in the formation of dsRNAs that stimulate MDA5 activation (Fig. 1d).
A substantial portion of Alu elements can be expressed from introns during the transcription of their host genes. Consequently, the dysregulation of mRNA splicing is closely associated with antiviral responses. Depletion of heterogeneous nuclear ribonucleoprotein C (hnRNPC) leads to increased dsRNA levels derived from Alu elements within pre-mRNA introns, thereby activating the dsRNA sensor retinoic acid-inducible gene I (RIG-I) and eventually inhibiting cell proliferation37. In addition, inhibiting type I protein arginine methyltransferase (PRMT) with a small molecule MS023 can upregulate the expression of intronic IRAlus38. Type I PRMTs are responsible for the asymmetric demethylation of proteins and can control RNA splicing and DNA repair39. In this context, inhibiting type I PRMTs alters mRNA splicing, which results in increased intron retention in triple-negative breast cancer cells. IRAlus embedded within retained introns of mis-spliced mRNAs activate RIG-I and Toll-like receptor 3 (TLR3) to facilitate cell death38. Together, these findings reveal intronic IRAlus-mediated innate immune activation within the context of splicing dysregulation (Fig. 1e).
Like those used for PRMT1 inhibition, small chemical inhibitors of the spliceosome have been employed in cancer therapy to induce viral mimicry through increased expression of intronic dsRNAs. Bowling et al. demonstrated that small spliceosome modulators, such as sudemycin D6 (SD6) and H3B-8800, led to widespread cytosolic accumulation of mis-spliced mRNAs, many of which adopt double-stranded secondary structures21 (Fig. 1e). These RNAs are recognized by dsRNA sensors, which trigger antiviral signaling and extrinsic apoptosis. Similarly, pharmacological modulation of pre-mRNA splicing factor 3b subunit 1 (SF3B1) using pladienolide B (Plad B) leads to aberrant production of intron-retained mRNAs, thereby activating the RIG-I-dependent I-IFN response and facilitating tumor cell death40. These studies highlight the fact that activating innate immune responses by IRAlus through DNMTis and spliceosome inhibitors can provide clinical benefit in cancer by inducing a viral mimicry state, suggesting that IRAlus are potential therapeutic targets for cancer immunotherapy.
Antiviral responses induced by IRAlus can be modulated through direct interaction with RBPs, such as TAR DNA-binding protein 43 (TDP-43) and Dicer. TDP-43 plays an essential role in mRNA metabolism, including transcription, splicing, stability, and transport41,42. Accumulating evidence has suggested that the loss of TDP-43 results in the accumulation of self-dsRNAs, including Pol III-transcribed Alu RNAs43,44. An increase in the cytosolic level of Alu RNAs is recognized by RIG-I and induces IFN-mediated cell death (Fig. 1f). Notably, the Alu consensus sequence is enriched in TDP-43 binding motifs, and the overexpression of wild-type TDP-43 diminishes the stability of Pol III transcripts and rescues IFN responses, whereas the RNA-binding mutant TDP-43 does not44. In addition, the RNase III Dicer can directly process Alu transcripts to downregulate their expression. Indeed, multiple studies revealed that Dicer depletion induces the accumulation of Alu RNAs, which promotes the development of geographic atrophy (GA), a form of age-related macular degeneration characterized by inflammation in the retinal pigment epithelium (RPE)45,46. Interestingly, upregulated Alu expression and the consequent cytotoxicity due to Dicer deficiency are mitigated by Pol III inhibition45. In addition, the authors found notable enrichment of Alu RNAs among J2-immunoprecipitated RNAs in GA patient samples that exhibited reduced Dicer levels, suggesting that Dicer might digest Pol III-transcribed IRAlus. In Dicer-deficient RPE cells, undegraded Alu RNAs activate the NLRP3 inflammasome and trigger TLR-independent MyD88 signaling, thereby promoting RPE cell death46. Kerur et al. further demonstrated that Dicer deficiency-induced inflammasome activation in RPE cells depends on cyclic GMP-AMP synthase (cGAS)-driven IFN signaling47. The authors suggested that Alu RNAs undergo reverse transcription to cDNA in the cytosol and that Alu cDNAs interact with cGAS, thereby facilitating the cytosolic release of mitochondrial DNA48. This event amplifies the activation of cGAS and induces IFNβ, consequently driving inflammasome activation and apoptosis. Overall, the immunogenic toxicity of the Alu element manifests in diverse manners, including Alu duplexes (mostly IRAlus) and Alu cDNAs (Fig. 1f).
Gene regulation by IRAlus
In addition to their immunogenic roles, IRAlus play a pivotal role in gene regulation. Alu elements are ubiquitously dispersed within tens of thousands of gene bodies, giving rise to the broad transcription of embedded Alu and IRAlus as parts of Pol II-transcribed RNAs, including mRNAs5,49. Over the past decade, an increasing number of studies have reported the unique regulatory roles of embedded IRAlus in gene expression, ranging from circRNA biogenesis to mRNA nuclear retention. Below, we highlight the gene regulation by IRAlus through their structural characteristics and genomic locations.
Intronic IRAlus: alternative splicing and circular RNA biogenesis
One of the most well-understood mechanisms of Alu-mediated alternative splicing is Alu exonization. In particular, the minus strand of the Alu consensus sequence contains splice donor and acceptor sites, suggesting that the antisense Alu serves as a primary reservoir for generating alternative exons49. RNA folding of intronic Alu elements can also facilitate Alu exonization50,51. In this context, slow transcription can strengthen the intronic Alu RNA structure and cause nearby cryptic splice sites to resemble canonical splice sites, ultimately promoting Alu retention in mRNAs50. The incorporation of Alu exons in the coding region can introduce premature termination codons or trigger a frameshift in the encoded protein52.
The splice sites within the antisense Alu can also regulate the alternative splicing of neighboring exons53. In the case of the RABL5 gene, two Alus (AluJo in the sense orientation and AluSx in the antisense orientation) are inserted within the intron upstream of exon 3. Here, AluSx suppresses exon 3 selection, resulting in exon skipping53. Interestingly, this exon skipping event is partly counteracted by AluJo located in the same intron. The interaction between these two Alus forms a dsRNA structure, and the deletion of AluJo or increase in the distance between the two Alus enhances AluSx-mediated exon skipping, suggesting that intronic IRAlus formation mediates the delicate balance between canonical and alternative splicing of downstream exons (Fig. 2a).
a An example of Alu-mediated alternative splicing in the RABL5 gene. The antisense-oriented AluSx that is located just upstream of exon 3 suppresses exon 3 selection, leading to exon skipping. This negative impact mediated by AluSx is partly reversed by the sense-oriented AluJo situated in the same intron. b circRNA formation is facilitated by backsplicing, which is driven by RNA pairs formed by IRAlus across flanking introns. c RNA pairs formed by IRAlus across neighboring introns compete with IRAlus within the same intron, leading to competition between backsplicing for circRNAs and canonical splicing for linear RNAs. d Competition between IRAlus across different pairs of flanking introns in the same gene locus promotes ABS, resulting in multiple circRNAs sharing the same backsplicing sites. e Trans-acting RBPs regulate circRNA biogenesis via direct interactions with intronic IRAlus flanking backspliced exons. SAM68 and NF90/NF110 positively regulate circRNA production by facilitating and stabilizing the formation of intronic IRAlus, while DHX9 represses circRNA production by destabilizing intronic IRAlus.
One of the key splicing processes through which intronic IRAlus exert a significant impact is the generation of circRNAs via the promotion of backsplicing. Backsplicing ligates a downstream splice donor site with an upstream splice acceptor site, leading to the formation of covalently closed circular transcripts or circRNAs54. Typically, backsplicing competes with canonical splicing for the spliceosomal machinery but occurs less efficiently, resulting in fewer circRNAs than their linear counterparts55. Nonetheless, due to their circular structure, circRNAs are highly stable, and human cells express tens of thousands of circRNAs56,57,58. Numerous studies have demonstrated that RNA pairings formed by IRAlus across flanking introns bridge distal splice sites in close proximity to facilitate backsplicing57,58,59,60 (Fig. 2b). Considering that RNA pairs can also be formed by IRAlus within the same intron, which favors canonical splicing of linear RNAs, competition among multiple Alu elements for binding to each other can influence the relative efficiency of backsplicing (Fig. 2c).
Binding of different intronic Alus within a single gene can lead to the generation of multiple circRNAs that share the same backsplicing site through alternative backsplicing (ABS)61. Comprehensive transcriptome analysis across 90 human tissue samples revealed that ABS events are prevalent during circRNA biogenesis and account for approximately 84% of circRNAs62. Predominant circRNAs exhibit longer flanking introns and accommodate more Alu elements than other circRNAs in the same ABS event. In addition, the greater pairing capacity of IRAlus for flanking introns enables the corresponding circRNAs to outcompete other circRNAs, thereby establishing them as the predominant circRNAs (Fig. 2d). Human-specific Alu species exhibit a more robust pairing capacity than their mouse counterpart, B1 SINEs, resulting in a greater prevalence of circRNAs in humans than in mice63. For instance, survival motor neuron (SMN) genes exhibit significant enrichment of Alu elements, which occupy approximately 40% of the transcribed region of the SMN64. While the coding sequences of the SMN genes are highly conserved in mammals, these frequent Alu insertions result in a greater diversity of exon-containing circRNAs in humans65. Therefore, intronic Alu elements diversify human RNA landscapes through alternative splicing regulation.
In addition to Alu-Alu interactions, trans-acting RBPs are involved in the regulation of circRNA biogenesis by directly interacting with intronic IRAlus flanking backspliced exons. For example, nuclear factor 90 (NF90) and its 110 kDa isoform NF110 promote circRNA production by stabilizing the intronic IRAlus structure through direct binding to facilitate backsplicing66. The depletion of NF90 or NF110 results in widespread downregulation of nascent circRNA expression, which can be restored by reintroducing wild-type NF90 but not NF90 that lacks the dsRNA-binding motif. Furthermore, ATP-dependent DExH-Box Helicase 9 (DHX9) suppresses circRNA formation by binding and destabilizing intronic IRAlus24. Loss of DHX9 promotes the generation of a subset of circRNAs from SMN genes with a high density of Alus65. In contrast to that of DHX9, the Src-associated in mitosis 68 kDa (SAM68) protein can promote SMN pre-mRNA circularization by binding to flanking sites in Alu-rich regions of SMN introns67. The interaction between Alu-rich introns and SAM68 results in SAM68 dimerization, which may bring distantly located intronic IRAlus closer to promote SMN pre-mRNA circularization. Together, the coordinated interplay between cis-acting intronic IRAlus and multiple trans-acting RBPs regulates the biogenesis of circRNAs (Fig. 2e).
IRAlus in 3′ UTRs: Nuclear sequestration and mRNA decay
In addition to introns, 3′ UTRs contain a large number of IRAlus elements, which provide an additional layer of post-transcriptional gene regulation. The most well-known gene regulatory mechanism of interest associated with the 3′ UTR IRAlus is the nuclear sequestration of host mRNAs13. In the nucleus, inosine-containing RNAs are recognized by the paraspeckle protein non-POU domain-containing octamer binding (NONO)68. Considering that most A-to-I editing occurs on the IRAlus element, NONO recognizes IRAlus RNAs and can transport them to nuclear paraspeckles. Moreover, Chen et al. reported that hundreds of mRNAs containing IRAlus in the 3′ UTR undergo A-to-I editing events, and these IRAlus-containing mRNAs are sequestered in the nucleus in a NONO-dependent manner13.
Paraspeckles are membrane-less nuclear organelles that primarily consist of two components: nuclear-enriched abundant transcript 1 (NEAT1), a long noncoding RNA (lncRNA), and several paraspeckle assembly proteins, including NONO13,23. The interaction between IRAlus and NONO results in the retention of mRNAs within the nucleus, subsequently leading to repression of protein translation in the cytosol (Fig. 3a). Hence, proper assembly of paraspeckles is essential for nuclear sequestration of mRNAs and subsequent gene silencing effects. A deficiency in the NEAT1 lncRNA in embryonic stem cells results in the loss of paraspeckle assembly and the cytosolic release of IRAlus-containing mRNAs23. Additionally, the transcriptional suppression of the NEAT1 lncRNA by coactivator-associated arginine methyltransferase 1 (CARM1) hinders the formation of paraspeckles and relieves IRAlus-mediated gene silencing69. CARM1 also methylates the coiled-coil domain of NONO to reduce its binding affinity to IRAlus to further free the 3′ UTR IRAlus-containing mRNAs from nuclear retention69. In pituitary cells, paraspeckle proteins and NEAT1 expression follow a rhythmic circadian pattern, resulting in the circadian assembly of paraspeckles70. This circadian pattern leads to rhythmic nuclear retention of IRAlus-containing reporter mRNAs, indicating that nuclear retention and gene silencing by 3′ UTR IRAlus may play a role in regulating the circadian rhythm.
a NONO recognizes 3′ UTR IRAlus in mRNAs and sequesters them in nuclear paraspeckles. b STAU1 facilitates the cytosolic export of IRAlus-containing mRNAs and enhances the translation of the corresponding mRNAs in the cytosol by preventing PKR binding. c STAU1 binds to the intermolecular Alu duplex and recruits the UPF1 RNA helicase to trigger SMD. d Long-distance enhancer-promoter interactions are facilitated by the formation of duplexes between the embedded Alu sequences in eRNAs and uaRNAs, which enables the transcription of multiple genes in the cluster. e R-loop formation through direct Alu RNA-DNA pairing facilitates the interaction between eRNAs and target promoters.
mRNAs with IRAlus in their 3′ UTR can escape nuclear retention by interacting with RBPs that compete with NONO for binding to IRAlus. The expression level of staufen1 (STAU1) is inversely correlated with NONO-IRAlus binding and is correlated with cytosolic export and translation of the corresponding mRNAs25. Notably, STAU1 also enhances the translation of IRAlus-containing mRNAs in the cytosol by preventing the binding of PKR25, which triggers global translational repression via eukaryotic translation initiation factor 2α (eIF2α) phosphorylation71 (Fig. 3b). Indeed, uncontrolled cytosolic release of IRAlus-containing mRNAs can lead to global translational suppression via PKR activation. While PKR and IRAlus-containing mRNAs are predominantly localized in the cytosol and nucleus, respectively, they can interact during mitosis when the nuclear envelope is disintegrated18. Subsequent PKR activation leads to global translational repression during mitosis. Similar nuclear sequestration of mRNAs was observed in the case of mouse CTN-RNA, which contains inverted repeats of B1 SINE elements with hyper A-to-I editing72. Interestingly, under stress, the inverted B1 repeats of CTN-RNA are removed from the 3′ UTR via alternative polyadenylation (APA), which leads to increased cytosolic export of RNA and increased translation of the encoded protein. This finding indicates that APA might be a key process that determines cell- or tissue-specific gene regulation by 3′ UTR IRAlus (see Future perspectives in the regulation and application of IRAlus for details)13,72,73.
Another important role of 3′ UTR IRAlus is the regulation of host mRNA stability. In a process known as STAU1-mediated decay (SMD), STAU1 binds to the 3′ UTR and recruits the up-frameshift suppressor 1 (UPF1) RNA helicase to trigger mRNA decay upon translation termination upstream of the STAU1-binding site (SBS)74. SMD regulates the expression of genes harboring SBSs during myogenesis, cutaneous wound healing, and adipogenesis75,76,77. In particular, SBSs may be formed via intermolecular base pairing between two RNA molecules containing complementary Alu elements (Fig. 3c). For example, the Alu element within cytosolic polyadenylated lncRNA, known as 1/2-SBS RNA, can form a dsRNA structure with another complementary element in the 3′ UTR of an SMD target mRNA75. In addition, the Alu element of the sprouty RTK signaling antagonist 4 intronic transcript 1 (SPRY4-IT1) lncRNA, which is associated with aggressive behavior and poor prognosis in human cancers, can bind to the 3′ UTR of the transcription elongation factor B subunit 1 (TCEB1) mRNA, which inhibits TCEB1 expression to promote cell metastasis78. In summary, IRAlus in 3′ UTRs are key crosstalk hubs where several RBPs interact to post-transcriptionally regulate gene expression.
Transcriptional regulation by IRAlus
The gene regulatory function of IRAlus RNAs extends to transcription, where IRAlus act as a trans-acting factor. Traditionally, Alu DNA elements serve as enhancers that are regulated by H3K4me1 for tissue-specific regulation of gene expression7. A recent study revealed the novel role of Alu RNA duplexes in modulating long-range enhancer-promoter selectivity79. Enhancer-promoter interactions often occur over long distances. However, how enhancers find their cognate promoters has remained unclear. Interestingly, enhancers and promoters can be bidirectionally transcribed by Pol II, thereby generating enhancer RNAs (eRNAs) and promoter-antisense RNAs (uaRNAs), respectively80. Liang et al. employed RNA in situ confirmation sequencing, which maps RNA-RNA interactions, to explore enhancer-promoter connectivity using pairwise interactions between eRNAs and uaRNAs79. The authors constructed high-resolution enhancer-promoter RNA interaction maps in multiple cell lines and found significant enrichment of the Alu consensus sequence. Notably, enhancers containing Alu elements have a greater frequency of interaction with promoters than enhancers lacking Alu elements. The authors proposed a model in which long-distance enhancer-promoter looping is facilitated by the formation of duplexes between the embedded Alu sequences in eRNAs and uaRNAs. Ultimately, these Alu duplexes robustly direct an enhancer to its corresponding promoter, thereby establishing the specificity of enhancer-promoter interactions over extended genomic distances (Fig. 3d). Another study proposed a trans-acting R-loop model in which Alu elements potentially mediate the interaction between eRNAs and target promoters through direct RNA-DNA pairing81 (Fig. 3e). Collectively, these recent findings reveal an unprecedented role for Alu RNAs as regulators of enhancer-promoter selectivity.
Regulation of IRAlus via RNA editing
RNA modifications are important regulatory mechanisms in RNA metabolism, including RNA stability, transport, splicing, and structure82. Among over 100 distinct types of RNA modifications, RNA editing stands out as it diversifies the cellular functionality of RNA molecules by altering their sequence83. The most common type of RNA editing is A-to-I editing, which is catalyzed by enzymes encoded by the adenosine deaminase acting on RNA (ADAR) gene family. Mammals express three ADAR family members (ADAR1, ADAR2, and ADAR3), with ADAR1 accounting for the majority of editing events84. As discussed above, most A-to-I editing by ADAR1 occurs on IRAlus RNAs. Global analysis of ADAR1-RNA interactions in human cells using CLIP-seq and investigation of the ADAR1 editome revealed that ADAR1 primarily targets Alu RNAs27,85. In this section, we introduce the downstream effects of ADAR1-mediated A-to-I editing on the function of IRAlus.
Modulation of IRAlus immunogenicity by ADAR1
A key outcome of A-to-I editing is the alteration of dsRNA structural integrity by converting A-U pairs into less stable I-U wobble, resulting in decreased interactions with dsRNA sensors, such as MDA5 and PKR86 (Fig. 4a). Earlier studies revealed that editing of 3′ UTR IRAlus by ADAR1 resulted in reduced MDA5 activation by decreasing the efficiency of MDA5 filament assembly19,87. Indeed, under ADAR1 deficiency, 3′ UTR IRAlus become the primary ligands for MDA5 and can trigger MDA5 activation19 (Fig. 4b). Interestingly, one of the IFN-stimulated genes (ISGs), apolipoprotein L1 (APOL1), contains IRAlus in its 3′ UTR, which can activate MDA5 to upregulate its own expression through a positive feedback loop88. Moreover, MDA5 activation by unedited IRAlus can account for the frequent mutations in ADAR1 in patients with autoinflammatory disorders, including AGS89. A recent exploration of the cis-RNAs quantitative trait locus revealed that common genetic variants linked to RNA editing levels are substantially enriched in genome-wide association study (GWAS) signals of common autoimmune and immune-related disorders90. Remarkably, diminished editing levels of IRAlus are apparent in GWAS-associated genetic variants; these findings underscore the fact that RNA editing is a key mechanism underlying the genetic risk for immune-related diseases.
a A-to-I editing by ADAR1 reduces the structural integrity of dsRNAs by converting A-U pairs into the less stable I-U wobble. b Editing of IRAlus by ADAR1 diminishes the capacity of the RNA to activate MDA5 by reducing the efficiency of MDA5 filament assembly. Conversely, under ADAR1 deficiency, IRAlus become the primary ligands for MDA5, leading to MDA5 activation and the I-IFN response. c The two isoforms of ADAR1 include a constitutively expressed short nuclear isoform (p110) and a long IFN-inducible cytoplasmic isoform (p150). ADAR1 p150 contains the Zα domain in its extended N-terminus. d The interaction between the Zα domain-mediated ADAR1 with IRAlus in the Z-RNA conformation disrupts its secondary structure. In Zα mutant ADAR1 cells, unedited IRAlus are recognized by ZBP1 through its two Zα domains and trigger ZBP1-dependent cell death. e The dual roles of ADAR1 in modulating circRNA generation. A-to-I editing can destabilize or stabilize the dsRNA structure of IRAlus formed between flanking introns by creating I-U mismatches or correcting A-C mismatches to I-C pairs, respectively. f A-to-I editing affects the nuclear export of IRAlus-containing mRNAs by altering their binding to STAU1. In addition, ADAR1 prevents SMD of IRAlus-containing mRNAs through competitive binding with STAU1.
ADAR1 is expressed as two isoforms, including a constitutively expressed short nuclear isoform (p110) and a long IFN-inducible cytoplasmic isoform (p150). While both ADAR1s can bind to canonical A-form dsRNA, the ADAR1 p150 isoform can bind to an unusual left-handed double helix called Z-RNA through the Zα domain in its extended N-terminus91 (Fig. 4c). Numerous studies have shown that mutations in the Zα domain of ADAR1 are commonly observed in patients with AGS, and the loss of the ADAR1/Z-RNA interaction triggers a spontaneous MDA5-dependent type I IFN response in both human and mouse cells89,92,93,94. Comprehensive analysis of A-to-I editing sites between wild-type and Zα mutant ADAR1 in human embryonic kidney 293 (HEK293) cells revealed that most of the sites less edited by the mutant were mapped to Alu elements93. Indeed, putative Z-RNA-forming sequences are present in Alu repeats, suggesting that the Zα domain of ADAR1 might be needed to efficiently edit Alu RNAs in the Z-RNA form95. Consistent with this thought, de Reuver et al. revealed that IRAlus are recognized by Z-DNA binding protein 1 (ZBP1) through its two Zα domains and stimulate ZBP1-dependent cell death in cells harboring the Zα mutant ADAR196 (Fig. 4d). In summary, in addition to the recognition of dsRNA through its canonical dsRNA-binding domains, the Zα domain-mediated ADAR1 interaction with the IRAlus Z-RNA disrupts its secondary structure to maintain tolerance to endogenous dsRNAs.
A-to-I editing has received growing attention in cancer immunotherapies as increased expression of IRAlus by DNMTi treatment transforms the cells into the viral mimicry state. However, DNMTi treatment also promoted ADAR1 transcription, which counteracted the increase in IRAlus expression via increased A-to-I editing. Considering this negative feedback by ADAR1, depletion of ADAR1 in combination with DNMTi treatment substantially limited the growth of colorectal cancer, whereas ADAR1 depletion alone had minimal effects20. Moreover, a similar strategy was employed for splicing dysregulation, where a combination of hnRNPC and ADAR1 deficiency synergistically triggered the induction of ISGs, underscoring the protective function of ADAR1 in mitigating IRAlus-mediated innate immune responses resulting from splicing dysregulation26. Since dysregulation of splicing occurs frequently in cancers97, such a strategy could be a promising approach to improve the efficacy of cancer immunotherapy.
Modulation of IRAlus-mediated gene regulation by ADAR1
A-to-I editing can also influence intronic IRAlus-mediated gene regulation. Studies have reported the association of ADAR1-mediated editing with circRNA formation through disruption of the dsRNA structure of intronic IRAlus60,98. Under ADAR1-deficient conditions, the enhanced stability of IRAlus pairing promotes backsplicing, resulting in increased circRNA expression60. In contrast, ADAR1 can promote circRNA generation in specific cases. A recent study reported that some A-to-I edits can stabilize dsRNA structures formed between flanking introns by correcting A-C mismatches to I-C pairs98. This is possible because ADAR catalyzes A-to-I editing on multiple adenosine residues in the vicinity rather than specifically on A-U paired adenosines. Overall, these findings highlight the importance of orchestrating the base pairing capacity of IRAlus through RNA editing, which introduces an additional layer of complexity to the circRNA transcriptome (Fig. 4e).
ADAR1 can also edit 3′ UTR IRAlus and affect the nuclear sequestration of host mRNAs. However, the effect of A-to-I editing during nuclear retention remains controversial. While a correlation between editing levels and nuclear retention levels exists13, it has been proposed that long dsRNAs stretch rather than inosine residues is needed25. For certain IRAlus-containing antiapoptotic genes, such as X-linked inhibitor of apoptosis protein (XIAP) and mouse double minute 2 (MDM2), ADAR1 influences the cytosolic export of mRNAs by altering the dsRNA structures and subsequently affecting the interaction with STAU199 (Fig. 4f). Moreover, ADAR1 competes with STAU1 for binding to IRAlus and protects IRAlus-containing mRNAs from SBSs via SMD100 (Fig. 4f). Overall, the crosstalk between IRAlus and A-to-I editing by ADAR1 diversifies gene expression in cells.
Future perspectives on the regulation and application of IRAlus
Biomolecular condensates
Recent studies on cellular dsRNAs have proposed potential regulatory mechanisms for IRAlus. Membrane-less biomolecular condensates, such as stress granules (SGs) and paraspeckles, are aggregates that are abundant in RNAs and RBPs101. Notably, SGs can prevent the excessive activation of dsRNA-induced innate immunity by both viral dsRNAs and increased self-dsRNAs in ADAR1 deficiency102. Although the specific identity of self-dsRNAs regulated by SGs has not yet been elucidated, we speculate that SGs modulate the immunogenicity of IRAlus. This hypothesis further calls for the investigation of the potential role of SGs in immune disorders associated with IRAlus. In addition, one recent study reported novel cytosolic condensates, referred to as dsRNA-induced foci (dRIFs), that are formed in response to increased levels of self- and non-self-dsRNAs103. These dRIFs contain dsRNAs and multiple RBPs, including PKR, ADAR1, STAU1, and DHX9. The authors proposed that dRIFs serve as sites of PKR activation by facilitating efficient PKR-dsRNA interactions. Notably, overexpression of an EGFP reporter mRNA with IRAlus in its 3′ UTR also induced the formation of foci that were colocalized with PKR, suggesting that the immunostimulatory activity of IRAlus might be modulated through dRIF formation. Deciphering the connection between biomolecular condensates and IRAlus will provide novel perspectives on the regulatory potential of IRAlus.
m6A modification
In addition to A-to-I editing, another prevalent RNA modification, N6-methyladenosine (m6A), can affect local dsRNA structures and the recognition of IRAlus. Strikingly, depletion of m6A triggers a detrimental immune response, resulting in hematopoietic failure and perinatal lethality during murine fetal development104. The responsible dsRNAs are predominantly derived from protein-coding regions and marked by high levels of m6A modification in their native states. Although determining the precise regulatory mechanism of m6A modification on dsRNA structures will require further investigation, m6A may function as a structural switch in these RNAs by disrupting the thermostability of base pairing105. Several studies have shown that m6A modification can indirectly affect IRAlus by modulating A-to-I editing, suggesting the potential for interplay between these two types of RNA modifications106,107. While the adenosines targeted by A-to-I editing or m6A modification are unlikely to overlap, ADAR1 exhibits an unfavorable association with m6A transcripts for further A-to-I editing, and suppression of m6A catalyzing enzymes results in a global increase in A-to-I editing106. This crosstalk could determine the editing status of 3′ UTR IRAlus, which might modulate IRAlus activity. Since other RNA modifications, such as m1A and m5C, have been shown to affect RNA secondary structures82,108, the interplay between different RNA modifications and the regulation of IRAlus needs to be explored.
Alternative polyadenylation
Cell/tissue-specific alterations in 3′ UTR length present a novel perspective for regulating the functional landscape of IRAlus. A recent study by Ku et al. revealed that APA determines the inclusion or exclusion of IRAlus in 3′ UTR of hundreds of mRNAs, thereby globally affecting IRAlus-mediated gene regulation and essential cell signaling pathways73. For example, the authors revealed that global 3′ UTR shortening may contribute to tumorigenesis by excluding IRAlus in the 3′ UTR of MDM2 mRNA, which leads to translational activation of MDM2 and subsequent repression of the p53 signaling pathway (Fig. 5a). Conversely, they found that global 3′ UTR lengthening in neuronal progenitor cells facilitates 3′ UTR IRAlus inclusion, particularly in genes associated with the development of neurodegenerative diseases, including amyotrophic lateral sclerosis. Further investigations of cell/tissue-specific global APA changes and paraspeckle formation will advance our understanding of IRAlus-mediated gene regulation and its relevance to the pathogenesis of diverse human diseases.
In addition to IRAlus-mediated gene regulation, 3′ UTR length alteration may also modulate IRAlus-mediated innate immune activation. Recently, Dorrity et al. reported that extended 3′ UTRs in human neurons give rise to neuronal dsRNA structures, which results in high basal levels of self-dsRNAs and vulnerability to ADAR1 loss109 (Fig. 5b). The authors showed that the neuron-enriched genes ELAVL2, ELAVL3, and ELAVL4 (HuB, HuC, and HuD) cooperatively increase the length of their 3′ UTR to trigger inflammation due to elevated dsRNA levels. Although the specific identity of immunogenic self-dsRNAs derived from extended 3′ UTRs in neurons has not been determined, the incorporation of IRAlus in longer 3′ UTRs might increase the burden of dsRNA. Similarly, in a study by Ku et al., 3′ UTR lengthening caused by the downregulation of cleavage stimulation factor subunit 2 (CSTF2) and its paralog CSTF2T expression, which are cleavage factors responsible for polyadenylation, promoted 3′ UTR IRAlus incorporation and subsequently resulted in inflammatory responses, including PKR activation73. Both studies proposed the regulation of 3′ UTR length as a novel strategy to modulate dsRNA-mediated immune responses.
Synthetic Alu RNA for cancer immunotherapy
A novel approach for cancer immunotherapy has emerged with the development of a synthetic RNA virus platform that consists of in vitro transcribed viral RNAs (vRNAs) encapsulated in lipid nanoparticles110. The intravenous administration of synthetic vRNAs to tumors facilitates virus replication and assembly, thereby stimulating immune cell infiltration and augmenting the activity of immune checkpoint inhibitors. Interestingly, a recent study demonstrated that nanoparticle delivery of synthetic Alu RNA also promotes antitumor immunity by activating RNA-sensing pathways111. The authors synthesized AluJb RNA (Left Arm), which can form intramolecular local dsRNA structures, and verified its immunostimulatory activity in vivo. Given that IRAlus exhibited enhanced immunostimulatory effects than did a single Alu due to their elongated dsRNA structures18,19, synthetic IRAlus RNAs may serve as more potent innate immune agonists. Current cancer immunotherapies leveraging the antitumor immunity of endogenous IRAlus, such as DNMTis and spliceosome inhibitors, are associated with a risk of unforeseen side effects owing to their global impact on the genome or transcriptome and variable responsiveness depending on the cell-specific degree of dsRNA induction112,113. Therefore, administering synthetic IRAlus (or Alu) RNAs can be an alternative strategy for precisely modulating dsRNA-mediated innate activation while minimizing potential side effects.
Conclusion
High abundance and sequence similarity among Alu families allow multiple proximal Alu elements in an inverted orientation to form dsRNA structures called IRAlus. Numerous studies have attempted to unravel the physiological function of IRAlus and their roles in pathological conditions. These endeavors have significantly expanded our understanding of Alu elements. In this review, we highlight the multifaceted nature of IRAlus, with emphasis on their roles as self-immunogens and post-transcriptional gene regulators, and explain how IRAlus activity is regulated by RNA editing and interactions with RBPs. The recognition of IRAlus by dsRNA sensors and the downstream impacts are summarized in Table 1. In addition, the list of proteins modulating IRAlus activity and their corresponding functions are provided in Table 2.
By generating long dsRNAs, IRAlus are recognized as key endogenous activators of the innate immune response. Indeed, increased expression of IRAlus due to dysregulated epigenetics, as well as defects in RNA splicing and editing, is closely associated with the pathogenesis of immune-related disorders, including AGS20,21,92,93,94. Interestingly, recent studies have proposed the use of this immunogenicity of IRAlus in cancer therapy, in which small chemical inhibitors of DNA methylation or spliceosomes are used to transform the cancer cells into viral mimicry to induce apoptosis and sensitize cells to immunotherapy20,21. Inside the nucleus, IRAlus serve as a key post-transcriptional regulator of gene expression, depending on the genomic location. Intronic IRAlus contribute to the diversity of the human transcriptome by regulating circRNA biogenesis and alternative splicing53,58,59, while IRAlus in 3′ UTRs affect host gene expression by modulating the cellular localization and stability of mRNAs13,75. Recently, a study revealed the function of intermolecular IRAlus in transcription, where they mediate long-distance enhancer-promoter interactions79. Together, both intramolecular and intermolecular Alu duplexes exert their unique influence throughout the mRNA life cycle.
The activity of IRAlus is tightly modulated by ADAR1-mediated A-to-I editing, which melts base pairs and disrupts the secondary structure of IRAlus19,27,87. Emerging evidence suggests that IRAlus may exist in the Z-RNA conformation and require the Zα domain of ADAR1 p150 and ZBP1 to recognize and control these RNAs92,93,96. The involvement of the Z-RNA conformation in the interaction with specific RBPs and the potential cellular functions remain to be addressed. Additionally, IRAlus serve as a platform for the crosstalk of multiple types of RNA modifications. Moreover, recent studies have proposed that the functional landscape of IRAlus can be regulated by changes in 3′ UTR length through APA73,109. Given that RNA editing, modification, and APA exhibit cell line- and tissue-specific variability114,115,116,117, exploring tissue-specific IRAlus activity and their corresponding functions will augment our understanding and unravel the biological significance of highly abundant primate-specific Alu elements.
References
Lander, E. S. et al. Initial sequencing and analysis of the human genome. Nature 409, 860–921 (2001).
Batzer, M. A. & Deininger, P. L. Alu repeats and human genomic diversity. Nat. Rev. Genet. 3, 370–379 (2002).
Elder, J. T., Pan, J., Duncan, C. H. & Weissman, S. M. Transcriptional analysis of interspersed repetitive polymerase III transcription units in human DNA. Nucleic Acids Res. 9, 1171–1189 (1981).
Dewannieux, M., Esnault, C. & Heidmann, T. LINE-mediated retrotransposition of marked Alu sequences. Nat. Genet. 35, 41–48 (2003).
Chen, L. L. & Yang, L. ALUternative regulation for gene expression. Trends Cell Biol. 27, 480–490 (2017).
Xie, H. et al. High-throughput sequence-based epigenomic analysis of Alu repeats in human cerebellum. Nucleic Acids Res. 37, 4331–4340 (2009).
Su, M., Han, D., Boyd-Kirkup, J., Yu, X. & Han, J. D. J. Evolution of Alu elements toward enhancers. Cell Rep. 7, 376–385 (2014).
Mariner, P. D. et al. Human Alu RNA is a modular transacting repressor of mRNA transcription during heat shock. Mol. Cell 29, 499–509 (2008).
Payer, L. M. et al. Alu insertion variants alter mRNA splicing. Nucleic Acids Res. 47, 421–431 (2019).
Sobczak, K. & Krzyzosiak, W. J. Structural determinants of BRCA1 translational regulation. J. Biol. Chem. 277, 17349–17358 (2002).
Stuart, J. J., Egry, L. A., Wong, G. H. & Kaspar, R. L. The 3′ UTR of human MnSOD mRNA hybridizes to a small cytoplasmic RNA and inhibits gene expression. Biochem. Biophys. Res. Commun. 274, 641–648 (2000).
Landry, J. R., Medstrand, P. & Mager, D. L. Repetitive elements in the 5′ untranslated region of a human zinc-finger gene modulate transcription and translation efficiency. Genomics 76, 110–116 (2001).
Chen, L. L., DeCerbo, J. N. & Carmichael, G. G. Alu element-mediated gene silencing. EMBO J. 27, 1694–1705 (2008).
Kim, S., Ku, Y., Ku, J. & Kim, Y. Evidence of aberrant immune response by endogenous double-stranded RNAs: attack from within. BioEssays 41, 1900023 (2019).
Weber, F., Wagner, V., Rasmussen, S. B., Hartmann, R. & Paludan, S. R. Double-stranded RNA is produced by positive-strand RNA viruses and DNA viruses but not in detectable amounts by negative-strand RNA viruses. J. Virol. 80, 5059 (2006).
Hur, S. Double-stranded RNA sensors and modulators in innate immunity. Annu Rev. Immunol. 37, 349–375 (2019).
Kim, Y. et al. PKR senses nuclear and mitochondrial signals by interacting with endogenous double-stranded RNAs. Mol. Cell 71, 1051–1063.e6 (2018).
Kim, Y. et al. PKR is activated by cellular dsRNAs during mitosis and acts as a mitotic regulator. Genes Dev. 28, 1310–1322 (2014).
Ahmad, S. et al. Breaching self-tolerance to Alu duplex RNA underlies MDA5-mediated inflammation. Cell 172, 797–810.e13 (2018).
Mehdipour, P. et al. Epigenetic therapy induces transcription of inverted SINEs and ADAR1 dependency. Nature 588, 169–173 (2020).
Bowling, E. A. et al. Spliceosome-targeted therapies trigger an antiviral immune response in triple-negative breast cancer. Cell 184, 384–403.e21 (2021).
Liu, C. X. & Chen, L. L. Circular RNAs: characterization, cellular roles, and applications. Cell 185, 2016–2034 (2022).
Chen, L. L. & Carmichael, G. G. Altered nuclear retention of mRNAs containing inverted repeats in human embryonic stem cells: functional role of a nuclear noncoding RNA. Mol. Cell 35, 467–478 (2009).
Aktaş, T. et al. DHX9 suppresses RNA processing defects originating from the Alu invasion of the human genome. Nature 544, 115–119 (2017).
Elbarbary, R. A., Li, W., Tian, B. & Maquat, L. E. STAU1 binding 3′ UTR IRAlus complements nuclear retention to protect cells from PKR-mediated translational shutdown. Genes Dev. 27, 1495–1510 (2013).
Herzner, A. M. et al. ADAR and hnRNPC deficiency synergize in activating endogenous dsRNA-induced type I IFN responses. J. Exp. Med. 218, e20201833 (2021).
Chung, H. et al. Human ADAR1 prevents endogenous RNA from triggering translational shutdown. Cell 172, 811–824.e14 (2018).
Ramaswami, G. et al. Accurate identification of human Alu and non-Alu RNA editing sites. Nat. Methods 9, 579–581 (2012). 2012 9:6.
Del Toro Duany, Y., Wu, B. & Hur, S. MDA5-filament, dynamics and disease. Curr. Opin. Virol. 12, 20–25 (2015).
Oda, H. et al. Aicardi-Goutières syndrome is caused by IFIH1 mutations. Am. J. Hum. Genet. 95, 121 (2014).
Kim, B., Jeong, K. & Kim, V. N. Genome-wide mapping of DROSHA cleavage sites on primary MicroRNAs and noncanonical substrates. Mol. Cell 66, 258–269.e5 (2017).
Chitrakar, A. et al. Introns encode dsRNAs undetected by RIG-I/MDA5/ interferons and sensed via RNase L. Proc. Natl. Acad. Sci. USA 118, e2102134118 (2021).
Conti, A. et al. Identification of RNA polymerase III-transcribed Alu loci by computational screening of RNA-Seq data. Nucleic Acids Res. 43, 817–835 (2015).
Moqtaderi, Z. et al. Genomic binding profiles of functionally distinct RNA polymerase III transcription complexes in human cells. Nat. Struct. Mol. Biol. 17, 635–640 (2010).
Chiappinelli, K. B. et al. Inhibiting DNA methylation causes an interferon response in cancer via dsRNA including endogenous retroviruses. Cell 162, 974–986 (2015).
Roulois, D. et al. DNA-demethylating agents target colorectal cancer cells by inducing viral mimicry by endogenous transcripts. Cell 162, 961 (2015).
Wu, Y. et al. Function of HNRNPC in breast cancer cells by controlling the dsRNA-induced interferon response. EMBO J. 37, e99017 (2018).
Wu, Q. et al. PRMT inhibition induces a viral mimicry response in triple-negative breast cancer. Nat. Chem. Biol. 18, 821–830 (2022).
Guccione, E. & Richard, S. The regulation, functions and clinical relevance of arginine methylation. Nat. Rev. Mol. Cell Biol. 20, 642–657 (2019).
Chang, A. Y. et al. Modulation of SF3B1 in the pre-mRNA spliceosome induces a RIG-I-dependent type I IFN response. J. Biol. Chem. 297, 101277 (2021).
Jo, M. et al. The role of TDP-43 propagation in neurodegenerative diseases: integrating insights from clinical and experimental studies. Exp. Mol. Med. 52, 1652–1662 (2020).
Prasad, A., Bharathi, V., Sivalingam, V., Girdhar, A. & Patel, B. K. Molecular mechanisms of TDP-43 misfolding and pathology in amyotrophic lateral sclerosis. Front. Mol. Neurosci. 12, 436464 (2019).
Saldi, T. K. et al. TDP-1, the caenorhabditis elegans ortholog of TDP-43, limits the accumulation of double-stranded RNA. EMBO J. 33, 2947–2966 (2014).
Dunker, W. et al. TDP-43 prevents endogenous RNAs from triggering a lethal RIG-I-dependent interferon response. Cell Rep. 35, 108976 (2021).
Kaneko, H. et al. DICER1 deficit induces Alu RNA toxicity in age-related macular degeneration. Nature 471, 325–330 (2011).
Tarallo, V. et al. DICER1 loss and Alu RNA induce age-related macular degeneration via the NLRP3 inflammasome and MyD88. Cell 149, 847–859 (2012).
Kerur, N. et al. cGAS drives noncanonical-inflammasome activation in age-related macular degeneration. Nat. Med. 24, 50–61 (2017).
Fukuda, S. et al. Alu complementary DNA is enriched in atrophic macular degeneration and triggers retinal pigmented epithelium toxicity via cytosolic innate immunity. Sci. Adv. 7, 3658–3687 (2021).
Häsler, J. & Strub, K. Alu elements as regulators of gene expression. Nucleic Acids Res 34, 5491–5497 (2006).
Saldi, T., Riemondy, K., Erickson, B. & Bentley, D. L. Alternative RNA structures formed during transcription depend on elongation rate and modify RNA processing. Mol. Cell 81, 1789–1801.e5 (2021).
Borovská, I., Vořechovský, I. & Královičová, J. Alu RNA fold links splicing with signal recognition particle proteins. Nucleic Acids Res. 51, 8199–8216 (2023).
Nekrutenko, A. & Li, W. H. Transposable elements are found in a large number of human protein-coding genes. Trends Genet. 17, 619–621 (2001).
Lev-Maor, G. et al. Intronic Alus influence alternative splicing. PLoS Genet. 4, e1000204 (2008).
Chen, L. L. The biogenesis and emerging roles of circular RNAs. Nat. Rev. Mol. Cell Biol. 17, 205–211 (2016).
Zhang, Y. et al. The biogenesis of nascent circular RNAs. Cell Rep. 15, 611–624 (2016).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013).
Zhang, X. O. et al. Complementary sequence-mediated exon circularization. Cell 159, 134–147 (2014).
Liang, D. & Wilusz, J. E. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 28, 2233–2247 (2014).
Ivanov, A. et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 10, 170–177 (2015).
Zhang, X. O. et al. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 26, 1277–1287 (2016).
Zhang, P. et al. Comprehensive identification of alternative back-splicing in human tissue transcriptomes. Nucleic Acids Res. 48, 1779 (2020).
Dong, R., Ma, X. K., Chen, L. L. & Yang, L. Increased complexity of circRNA expression during species evolution. RNA Biol. 14, 1064 (2017).
Ottesen, E. W., Seo, J., Singh, N. N. & Singh, R. N. A multilayered control of the human survival motor neuron gene expression by Alu elements. Front. Microbiol. 8, 2252 (2017).
Ottesen, E. W., Luo, D., Seo, J., Singh, N. N. & Singh, R. N. Human survival motor neuron genes generate a vast repertoire of circular RNAs. Nucleic Acids Res. 47, 2884–2905 (2019).
Li, X. et al. Coordinated circRNA Biogenesis and Function with NF90/NF110 in Viral Infection. Mol. Cell 67, 214–227.e7 (2017).
Pagliarini, V. et al. Sam68 binds Alu-rich introns in SMN and promotes pre-mRNA circularization. Nucleic Acids Res. 48, 633–645 (2020).
Zhang, Z. & Carmichael, G. G. The fate of dsRNA in the nucleus: a p54nrb-containing complex mediates the nuclear retention of promiscuously A-to-I edited RNAs. Cell 106, 465–476 (2001).
Hu, S. B. et al. Protein arginine methyltransferase CARM1 attenuates the paraspeckle-mediated nuclear retention of mRNAs containing IRAlus. Genes Dev. 29, 630–645 (2015).
Torres, M. et al. Circadian RNA expression elicited by 3′-UTR IRAlu-paraspeckle associated elements. Elife 5, e14837 (2016).
Meurs, E. F. et al. Constitutive expression of human double-stranded RNA-activated p68 kinase in murine cells mediates phosphorylation of eukaryotic initiation factor 2 and partial resistance to encephalomyocarditis virus growth. J. Virol. 66, 5805–5814 (1992).
Prasanth, K. V. et al. Regulating gene expression through RNA nuclear retention. Cell 123, 249–263 (2005).
Ku, J. et al. Alternative polyadenylation determines the functional landscape of inverted Alu repeats. Mol Cell (2024) https://doi.org/10.1016/J.MOLCEL.2024.01.008.
Kim, Y. K., Furic, L., DesGroseillers, L. & Maquat, L. E. Mammalian Staufen1 recruits Upf1 to specific mRNA 3′UTRs so as to elicit mRNA decay. Cell 120, 195–208 (2005).
Gong, C. & Maquat, L. E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature 470, 284–290 (2011).
Cho, H. et al. Staufen1-mediated mRNA decay functions in adipogenesis. Mol. Cell 46, 495–506 (2012).
Gong, C., Kim, Y. K., Woeller, C. F., Tang, Y. & Maquat, L. E. SMD and NMD are competitive pathways that contribute to myogenesis: effects on PAX3 and myogenin mRNAs. Genes Dev. 23, 54–66 (2009).
Zhao, L. et al. NF-κB-activated SPRY4-IT1 promotes cancer cell metastasis by downregulating TCEB1 mRNA via Staufen1-mediated mRNA decay. Oncogene 40, 4919–4929 (2021).
Liang, L. et al. Complementary Alu sequences mediate enhancer–promoter selectivity. Nature 2023 1–8 https://doi.org/10.1038/s41586-023-06323-x (2023).
Kim, T. K. & Shiekhattar, R. Architectural and functional commonalities between enhancers and promoters. Cell 162, 948–959 (2015).
Bai, X., Li, F. & Zhang, Z. A hypothetical model of trans-acting R-loops-mediated promoter-enhancer interactions by Alu elements. J. Genet. Genom. 48, 1007–1019 (2021).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA modifications in gene expression regulation. Cell 169, 1187–1200 (2017).
Boccaletto, P. et al. MODOMICS: a database of RNA modification pathways. 2021 update. Nucleic Acids Res. 50, D231–D235 (2022).
Mannion, N., Arieti, F., Gallo, A., Keegan, L. P. & O’Connell, M. A. New insights into the biological role of mammalian ADARs; the RNA editing proteins. Biomolecules 5, 2338–2362 (2015).
Bahn, J. H. et al. Genomic analysis of ADAR1 binding and its involvement in multiple RNA processing pathways. Nat. Commun. 6, 1–13 (2015).
Bass, B. L. & Weintraub, H. An unwinding activity that covalently modifies its double-stranded RNA substrate. Cell 55, 1089–1098 (1988).
Liddicoat, B. J. et al. RNA editing by ADAR1 prevents MDA5 sensing of endogenous dsRNA as nonself. Science 349, 1115–1120 (2015).
Riella, C. V. et al. ADAR regulates APOL1 via A-to-I RNA editing by inhibition of MDA5 activation in a paradoxical biological circuit. Proc. Natl. Acad. Sci. USA 119, e2210150119 (2022).
Rice, G. I. et al. Mutations in ADAR1 cause Aicardi-Goutières syndrome associated with a type I interferon signature. Nat. Genet. 44, 1243–1248 (2012).
Li, Q. et al. RNA editing underlies genetic risk of common inflammatory diseases. Nature 608, 569–577 (2022).
Placido, D., Brown, B. A., Lowenhaupt, K., Rich, A. & Athanasiadis, A. A left handed RNA double helix bound by the Zα domain of the RNA editing enzyme ADAR1. Structure 15, 395 (2007).
Tang, Q. et al. Adenosine-to-inosine editing of endogenous Z-form RNA by the deaminase ADAR1 prevents spontaneous MAVS-dependent type I interferon responses. Immunity 54, 1961–1975.e5 (2021).
de Reuver, R. et al. ADAR1 interaction with Z-RNA promotes editing of endogenous double-stranded RNA and prevents MDA5-dependent immune activation. Cell Rep. 36, 109500 (2021).
Nakahama, T. et al. Mutations in the adenosine deaminase ADAR1 that prevent endogenous Z-RNA binding induce Aicardi-Goutières-syndrome-like encephalopathy. Immunity 54, 1976–1988.e7 (2021).
Herbert, A. Z-DNA and Z-RNA in human disease. Commun. Biol. 2, 1–10 (2019).
de Reuver, R. et al. ADAR1 prevents autoinflammation by suppressing spontaneous ZBP1 activation. Nature 607, 784–789 (2022).
Obeng, E. A., Stewart, C. & Abdel-Wahab, O. Altered RNA processing in cancer pathogenesis and therapy. Cancer Discov. 9, 1493–1510 (2019).
Shen, H. et al. ADARs act as potent regulators of circular transcriptome in cancer. Nat. Commun. 13, 1508 (2022).
Yang, C. C. et al. ADAR1-mediated 3′ UTR editing and expression control of antiapoptosis genes fine-tunes cellular apoptosis response. Cell Death Dis. 8, e2833–e2833 (2017).
Sakurai, M. et al. ADAR1 controls apoptosis of stressed cells by inhibiting Staufen1-mediated mRNA decay. Nat. Struct. Mol. Biol. 24, 534–543 (2017).
Roden, C. & Gladfelter, A. S. RNA contributions to the form and function of biomolecular condensates. Nat. Rev. Mol. Cell Biol. 22, 183–195 (2020).
Paget, M. et al. Stress granules are shock absorbers that prevent excessive innate immune responses to dsRNA. Mol. Cell 83, 1180–1196.e8 (2023).
Corbet, G. A., Burke, J. M., Bublitz, G. R., Tay, J. W. & Parker, R. dsRNA-induced condensation of antiviral proteins modulates PKR activity. Proc. Natl. Acad. Sci. USA 119, e2204235119 (2022).
Gao, Y. et al. m6A modification prevents formation of endogenous double-stranded RNAs and deleterious innate immune responses during hematopoietic development. Immunity 52, 1007–1021.e8 (2020).
Liu, N. et al. N6-methyladenosine-dependent RNA structural switches regulate RNA–protein interactions. Nature 518, 560–564 (2015).
Xiang, J. F. et al. N6-Methyladenosines modulate A-to-I RNA Editing. Mol. Cell 69, 126–135.e6 (2018).
Terajima, H. et al. N6-methyladenosine promotes induction of ADAR1-mediated A-to-I RNA editing to suppress aberrant antiviral innate immune responses. PLoS Biol. 19, e3001292 (2021).
Safra, M. et al. The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution. Nature 551, 251–255 (2017).
Dorrity, T. J. et al. Long 3′UTRs predispose neurons to inflammation by promoting immunostimulatory double-stranded RNA formation. Sci. Immunol. 8, eadg2979 (2023).
Kennedy, E. M. et al. Development of intravenously administered synthetic RNA virus immunotherapy for the treatment of cancer. Nat. Commun. 13, 1–13 (2022).
Garland, K. M. et al. Nanoparticle delivery of immunostimulatory Alu RNA for cancer immunotherapy. Cancer Res. Commun. 3, 1800 (2023).
Kagan, A. B. et al. DNA methyltransferase inhibitor exposure–response: challenges and opportunities. Clin. Transl. Sci. 16, 1309–1322 (2023).
Kang, M. et al. Double-stranded RNA induction asa potential dynamic biomarkerfor DNA-demethylating agents. Mol. Ther. Nucleic Acids 29, 370–383 (2022).
Schaffer, A. A. et al. The cell line A-to-I RNA editing catalogue. Nucleic Acids Res. 48, 5849–5858 (2020).
Cuddleston, W. H. et al. Cellular and genetic drivers of RNA editing variation in the human brain. Nat. Commun. 13, 1–15 (2022).
Shulman, E. D. & Elkon, R. Cell-type-specific analysis of alternative polyadenylation using single-cell transcriptomics data. Nucleic Acids Res. 47, 10027–10039 (2019).
Xia, Z. et al. Dynamic analyses of alternative polyadenylation from RNA-seq reveal a 3′-UTR landscape across seven tumour types. Nat. Commun. 5, 1–13 (2014).
Acknowledgements
We thank the members of the Kim laboratory for helpful discussions. This study was supported by the Basic Research Laboratory Program through the National Research Foundation of Korea (grant number NRF-2021R1A4A3032789), funded by the Korean government’s Ministry of Science and ICT.
Author information
Authors and Affiliations
Contributions
K.L., J.K. and Y.K. organized and wrote the manuscript. K.L. prepared the figures. K.L., J.K., D.K. and Y.K. revised the manuscript. Y.K. supervised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lee, K., Ku, J., Ku, D. et al. Inverted Alu repeats: friends or foes in the human transcriptome. Exp Mol Med 56, 1250–1262 (2024). https://doi.org/10.1038/s12276-024-01177-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-024-01177-3
- Springer Nature Limited
This article is cited by
-
A sequence of SVA retrotransposon insertions in ASIP shaped human pigmentation
Nature Genetics (2024)
-
Regulatory RNA: from molecular insights to therapeutic frontiers
Experimental & Molecular Medicine (2024)