Abstract
Mitochondria are of fundamental importance in programmed cell death, cellular metabolism, and intracellular calcium concentration modulation, and inheritable mitochondrial disorders via mitochondrial DNA (mtDNA) mutation cause several diseases in various organs and systems. Nevertheless, mtDNA editing, which plays an essential role in the treatment of mitochondrial disorders, still faces several challenges. Recently, programmable editing tools for mtDNA base editing, such as cytosine base editors derived from DddA (DdCBEs), transcription activator-like effector (TALE)-linked deaminase (TALED), and zinc finger deaminase (ZFD), have emerged with considerable potential for correcting pathogenic mtDNA variants. In this review, we depict recent advances in the field, including structural biology and repair mechanisms, and discuss the prospects of using base editing tools on mtDNA to broaden insight into their medical applicability for treating mitochondrial diseases.
Similar content being viewed by others
Introduction
Mitochondria, double membrane-bound organelles referred to as the “powerhouses of the cell”, play an indispensable role in eukaryotic cells, as they are associated with metabolism1,2 and orchestrate a variety of other cellular functions, such as apoptosis, cell pluripotency, autophagy, calcium homeostasis, and innate immunity3,4,5,6,7,8,9. The mitochondrial genome is independent and distinct from that of the nucleus10. Human mtDNA, with a few hundred to several hundred thousand copies contained in each cell11,12, encodes 37 genes (22 tRNAs, 2 rRNAs, and 13 oxidative phosphorylation protein subunits) in a small circular 16.5 kb double-stranded piece of DNA13,14. The exceeded heteroplasmic threshold of mtDNA mutations manifests in multiple disorders (Fig. 1a)14. Specifically, a vast array of more than 250 mitochondrial defects have been implicated in various pathogenic mtDNA variants that primarily affect muscle and nervous tissues15,16. Notably, among the 97 known pathogenic mtDNA variants, point mutations accounted for 92 variants (approximately 95%), which indicated transition point mutations, including A > G, T > C, G > A, and C > T, accounting for 87% (MITOMAP: A Human Mitochondrial Genome Database. www.mitomap.org) (Fig. 1b). For instance, the MT‑TL1 mutation m.3243 A > G can trigger multiple mitochondrial syndromes, such as chronic progressive external ophthalmoplegia (CPEO); maternally inherited diabetes and deafness (MIDD); and mitochondrial myopathy, encephalopathy, lactic acidosis, and stroke-like episodes (MELAS) syndrome. Meanwhile, three major mtDNA mutations, m.3460 G > A, m.11778 G > A, and m.14484 T > C, are present in more than 95% of patients suffering from Leber hereditary optic neuropathy (LHON), a maternally inherited disease that is associated with a loss of vision17,18. Myoclonus epilepsy and ragged-red fiber (MERRF) syndrome, a severe neurodegenerative defect, is predominantly caused by the point mutation m.8344 A > G in the MT-TK gene19. Furthermore, a spectrum of mutations in the SPG7 gene, including c.861dupT, c.2221 G > A, c.2224 G > A, c.2228 T > C, c.1672A > T, c.1192 C > T, c.1529 C > T, c.1454_1462del, c.1067 C > T, c.184-3 C > T, c.233 T > A, c.1046insC, c.1457 G > A, and c.1053dup, were revealed in patients suffering from many mitochondrial diseases, such as progressive external ophthalmoplegia (PEO), ptosis, ataxia, spastic paraparesis, cerebellar atrophy, and proximal myopathy20. Strikingly, the C5024T mutation in the mtDNA gene MT-TA was uncovered via mouse models of heteroplasmic mitochondrial defects in the smooth muscle surrounding the colon and cardiomyocytes21. Although single-base substitutions can theoretically be corrected using base editors, precise and efficient therapeutics for several mtDNA mutations and mitochondria-involved diseases, have not been reported thus far. The development of and advances in base editing technology for correcting mtDNA base variants promise a potential therapeutic strategy for mitochondrial diseases and the generation of disease models in living cells and animals.
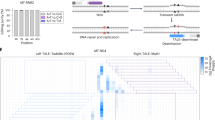
a The overall concept of heteroplasmic shifting. Although mutant and wild-type mtDNA can coexist, the exceeded threshold of mtDNA mutations compromises oxidative phosphorylation (OXPHOS). The mutations can be corrected by using gene editing platforms, which change the heteroplasmic level toward reducing the threshold. b Pie chart demonstrating the percentage of pathogenic mitochondrial DNA mutations in the MITOMAP database (accessed November 03, 2022). The white text reveals the distribution of transition point mutations, while the black text indicates transversion point mutations, deletions, insertions, or inversions. Parentheses indicate the number of pathogenic mitochondrial DNA mutations.
The development of and advances in mitochondrial gene editing tools
Research on mitochondrial DNA editing started with the use of restriction endonucleases, such as PstI, SmaI, and ApaLI, engineered to be expressed exclusively in mitochondria (called mitoRes)22,23,24. The specific elimination of pathogenic mtDNA is followed by repopulation with normal mitochondrial DNA via heteroplasmy. Since mitoRes cannot be easily redesigned, it is difficult to use them to correct various clinical mutations, as there is the limitation of potentially editing the off-target loci22,23,24.
Programmable DNA nucleases, such as clustered regularly interspaced short palindromic repeats (CRISPR)/CRISPR-associated protein 9 (Cas9), which recognize a short sequence known as the protospacer adjacent motif (PAM) and follow the DNA sequence targeted by a functional complex of Cas nucleases and a single guide RNA (sgRNA), has revolutionized gene modification and enabled the editing of mtDNA mutations25,26,27 (Fig. 2a). Of note, the CRISPR/Cas9 system, the most applied CRISPR construct, has recently been reported to efficiently cleave targeted mtDNA in HEK293T cells and zebrafish26,27. However, a key impediment to this approach is the import of the exogenous sgRNA into mitochondria28. Although several researchers have shown significant attempts at sgRNA delivery, this as-yet-unresolved limitation demonstrates the inefficiency of this approach26,29,30. Recently, one group demonstrated that not only Cas9 (type II) but also Cas12a (one of the type V CRISPR effectors) could access mitochondrial DNA editing31. However, although they tried to engineer the guide RNA using the RP loop related to importing mtRNA, clear evidence of efficient Cas12a-mediated mtDNA genome editing was not obtained31. Hence, to date, CRISPR-based systems have not been reliably applied to mtDNA manipulation.
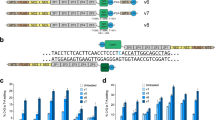
a A schematic of the architecture of CRISPR/Cas9 in the interactions with single-guide RNA and target DNA. The structures of b mtZFN, c ZFD, d mitochondrially targeted mitoTALEN, e DdCBE, and f TALED. ZFP zinc-finger protein, MTS mitochondrial targeting sequence, NES nuclear export signal, CTD (NG) C-terminal domain with NG repeat variable diresidues, NTD N-terminal domain.
Soon after the development of site-specific nucleases such as zinc-finger nuclease (ZFN) and transcription activator-like effector nuclease (TALEN)32,33,34, mitochondrially targeted ZFN (mtZFN) and mitochondrially targeted TALEN (mitoTALEN) versions emerged, optimized for delivery capacity into mitochondria, and they have been effectively used for heteroplasmic manipulation in several models of pathogenic mtDNA variants by inducing mtDNA double-strand breaks (DSBs)35,36. The sequence specificity of the mtZFN or mtTALEN monomers is demonstrated through interactions between the protein and DNA via tandemly organized repetitive elements derived from either zinc-finger protein (ZFP) or TALE DNA-binding protein (see Table 1 for a summary).
Development of the structure and use of ZFP technology for mtDNA editing
Zinc-finger nucleic acid-binding modules are some of the most profuse proteins in eukaryotes and comprise the first programmable nucleic acid-binding domain applied in epigenome engineering37,38,39. Each ZFN strategy for nuclear DNA modification includes two DNA‐binding domains recognizing and binding a specific DNA sequence and a nonspecific FokI cleavage domain, which is a mediator for DNA excision and fragment insertion or frameshift variants of the DNA target (Fig. 2b). Structurally, the DNA-binding domain, consisting of three ZFP groups (each a 30-amino-acid module) bound to a 3-nucleotide DNA sequence, is composed of two adjacent β-sheets and an α-helix that interact with a single zinc atom, whereas the nuclease activity of the FokI cleavage domain, which must contain two distinct yet complementary parts, requires its dimerization32. Based on the favorable qualities of the components, the combination of the flexible and specific DNA-binding characteristics of ZFPs and the vigorous ablation ability of the FokI restriction enzyme in the ZFN architecture makes it credible for genome editing. To date, the potential of ZFN technology has been harnessed and effectively used for mtDNA manipulation. Initial reports revealed a site-specific methylase status using a three-finger monomeric ZFP equipped with an N-terminal mitochondrial targeting sequence (MTS) and a nuclear export signal (NES) fused to DNA methyltransferase 3a (DNMT3A), specifically targeting the m.8993 T > G mutation40. Furthermore, a single ZFP monomer conjugated to two FokI endonuclease domains tethered together via a flexible linker of 35 amino acids was engineered to target m.8993 T > G in heteroplasmic cybrid cells. However, this approach raises a safety concern that the combination of a four-finger ZFP monomer binding only 12 bp and a constitutively active nuclease might excessively reduce the mtDNA copy number41. However, attempts to improve its characteristics are still far from being efficient42.
Alternatively, optimizing the heterodimeric mtZFN architecture consisting of NARPd(+), an mtZFN specifically binding to a mutant sequence, and COMPa(−), a nonmutant sequence-bound mtZFN monomer, caused an effective shift in the heteroplasmy of cybrid cells bearing the m.8993 T > G mutation and “common deletions”35. Changes in the heteroplasmy shifting efficiency have also been demonstrated in a fine-tuned m.8993 T > G model43,44. Specifically, approximately 80% of the cybrid cells bearing the m8993T > G mutation were successfully transfected with the specific mtZFN that induced a considerable shift toward the wild-type mtDNA. Strikingly, substantial depletion of the mtDNA copy number was revealed at 24 h, followed by a return to a level comparable to nontransfected cells 28 days post-transfection. Moreover, the model was further capitalized to investigate the metabolic defects in mtDNA mutation-bearing cells43. In a later study44, the authors exploited a model called mTUNE to produce several isogenic cell lines (mT7, mT45, and mT80) with various levels of m8993T > G mutation (7%, 45%, and 80%, respectively). These m8993T > G heteroplasmic cell types augmented cell migration, a finding supported by reports of MDH1 increasing ATP generation via glycolysis.
Of interest, mtZFNs exert their potential applications in vivo45. In particular, a construct containing an m.5024 C > T-specific monomer (MTM25) and a wild-type-specific monomer (WTM1) was produced by encoding the monomers in a distinct adeno-associated virus (AAV) genome, followed by their encapsidation into AAV9.45 vectors subsequently administered to mice21,45.
Recently, a ZFP-based base editor for mtDNA called ZFD has been developed for catalyzing the conversion of C to T in human cells46. ZFD constructs were created by connecting ZFPs to the split interbacterial toxin deaminase DddAtox and a uracil glycosylase inhibitor (UGI) via linkers (Fig. 2c). The optimized architecture of ZFDs showed successful base editing in HEK293T cells with frequencies of 2.6–30%46. Interestingly, the conversion of C to T mediated by ZFDs was performed in most Cs in both the TC and TCC sequence contexts. Although the rates of C to T conversion are currently low, further studies on engineering ZFDs to enhance both efficiency and precision may open a new door for correcting pathogenic mtDNA mutations in human embryos, fetuses, and specific adult tissues46.
Development of the structure and use of TALEs technology for mtDNA manipulation
MitoTALENs have been platform-engineered via the fusion of the DNA-binding domain of TALEs, comprising monomers from the plant pathogenic bacteria genus Xanthomonas, and the nonspecific cleavage domain of FokI endonucleases47,48. The DNA-binding domain that recognizes the specific nucleotide in the target DNA sequence consists of tandem conserved repeats of 33-35 amino acids (Fig. 2d). Among them, two positions (residues 12 and 13), known as the repeat variable diresidue (RVD), show high variation in the recognition ability for a specific nucleotide49. Notably, 4 out of 25 RVDs (HD, NI, NG, and NN), which were the most prevalent, could identify C, A, T, and G, respectively50. In contrast to ZFNs, the structural foundation of the TALE DNA-binding specificity is more amenable to engineering and requires a minimal number of RVD panels encoding all base specificities51.
The first study on the development of mitoTALENs demonstrated the elimination of common deletions and the m.14459 G > A point mutation in mtDNA by applying two engineered constructs: Δ5-mitoTALEN and 14459A-mitoTALEN, respectively36; these architectures cause heteroplasmic shifting in heteroplasmic cybrid cells. Interestingly, a design based on the requirement of a T at position 0 in the sequence responsible for DNA recognition, called the T0 strategy, was used for m.13513 G > A mutation elimination in heteroplasmic cybrid cells52. However, attaining the binding specificity for a single nucleotide at a mutation position that is neither N > A nor N > T is currently far from easy. Furthermore, the T0 strategy requires two monomers, thus limiting the packaging capacity in various vectors, especially AAVs. To improve these hurdles, the authors showed that a shortened mitoTALEN monomer could also be applied for the efficient elimination of m.8344 A > G mutations. In addition, later studies depicted changes in the heteroplasmic shifting efficiency in iPSCs harboring m.13513 G > A and m.3243 A > G53,54. Treatment with mitoTALENs targeting the CD5’ region of mtDNA was shown to induce DSBs, resulting in a common deletion, followed by an accumulation of mtDNA49,55,56. In addition, human LHON m.14459 G > A and NARP m.9176 T > C mutations were specifically eliminated by injecting mitoTALENs into MII stage mouse oocytes57.
Interestingly, a monomeric variant of mitoTALENs known as mitoTev-TALEs, which are engineered by fusing a TALE-binding domain with a T4 phage-derived I-TevI homing endonuclease catalytic domain through a linker, has been used to manipulate m.8344 A > G mutant mtDNA in heteroplasmic cybrids58. However, this architecture has apparent disadvantages, such as requiring a CNNNG cleavage site for I-TevI.
To further explore the potential of TALE, hybrid platforms have been constructed by the fusion of solely mtDNA-binding TALEs and base-editing deaminases for precise modification of mtDNA. Recently, DdCBEs that can precisely and specifically convert C to T in human mtDNA have been formed by adding TALE proteins and UGI to split-DddA (Fig. 2e)59. Combining two TALE proteins with the mtDNA target resulted in the activation of the inactively nontoxic split-DddA halves. Surprisingly, the addition of UGI, protecting U from base excision by glycosylase, improved the efficiency of base editing by approximately 8-fold. Most excitingly, DdCBEs have been successfully exploited for mtDNA base editing in various species, such as human embryos, zebrafish, mice (embryos and adults), rats, and plants60,61,62,63,64,65,66,67,68.
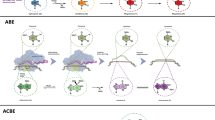
Although the discovery of DdCBEs is a huge step forward, this technology is limited by the dependence of DddA on editable sites where Cs in the 5’-TC-3’ contexts are preferably converted to Ts (conversion from 5′-TCC-3′ to 5′-TCT-3′ or 5′-TTT-3′ is also available depending on the target). As a consequence, methods based on the original DdCBEs could correct only 10% of pathogenic mtDNA point mutations69. Soon after, the authors developed DddA variants using rapid phage-assisted continuous evolution. DddA6-containing DdCBEs conduct C•G to T•A conversion at TC sites 3.3-fold more efficiently than canonical DdCBEs. Furthermore, base editing with the DddA11 variant revealed strong compatibility over a range of settings (TC, AC, and CC), in which the editable levels for CC and AC in both the nucleus and mtDNA were increased by 15–30%70. Despite effective base-editing improvements, engineered DdCBEs are still limited to only being able to edit C-to-T residues. However, this limitation has been circumvented by the advent of TALEDs, a novel tool that enables the conversion of A-to-G69. This platform consists of an mtDNA-targeting TALE, adenosine deaminase TadA8e, and inactive DddAtox as a cytosine deaminase (Fig. 2f). Moreover, TALEDs have been shown to successfully correct 47% of known pathogenic mtDNA point mutations in human cells69. More intriguingly, TadA8e, which supposedly operates on single-stranded DNA, was shown to possess an unexpected capability of adenine deamination in double-stranded DNA. This observation was explained by the presence of DddAtox, which enables the unwinding of the double-stranded DNA and provides TadA8e superfast access to the DNA for the necessary modifications69. However, this technology has the limitation of bystander editing, which converts nucleotides adjacent to the desired targets in the same editing window. Optimization in the engineering of DddAtox or TadA might eliminate this undesired editing69. In addition, it would be intriguing to enhance the specificity and efficiency of TALEDs, which might provide further possibilities for mtDNA editing. Novel methods for the TALED-mediated manipulation of ES cells or direct editing in mouse embryos are certain to be developed in the near future.
Off-target effects by mitochondrial gene editing tools
Solving unintended editing problems is of foremost importance for safe gene editing applications in clinical therapies. In the utilization of programmable editing platforms such as mtZFN and mitoTALEN, attenuating the temporary total mtDNA copy number is of paramount concern due to the depletion of mutant mtDNA before the repopulation of the wild-type mtDNA. Notably, the depletion might be significantly affected by the off-target editing of mtDNA as a consequence of inadequate reagent specificity or inappropriate concentrations of cleavage agents. Although off-target events of mtDNA editing using DdCBEs are relatively rare in HEK293T cells59, this platform can cause considerable mtDNA off-target editing in zebrafish and plants62,65,66,68. Most strikingly, two recent studies using DdCBEs have elucidated substantial off-target editing of the nuclear genome and mtDNA in mammalian cells and mice71,72. For instance, DdCBEs preferably caused undesired low-frequency mtDNA editing events at 5′-TC-3′ sequences, which were most prominent in the spacer regions to the left and right of the TALE binding sites, with some lying outside the regions71. Most unexpectedly, when delivered to fertilized 2-cell stage embryos by the genome-wide off-target analysis by two-cell embryo injection (GOTI) method, DdCBEs also induced remarkable sequence-independent off-target effects, resulting in single-nucleotide variants (1000–1500) in the nuclear DNA. The unexpected results are likely due to the unique characteristics of the DddAtox cytosine deaminase used in DdCBE, which favors dsDNAs as substrates, unlike the substrate preference of cytosine deaminase APOBEC1 in the BE3 protein for ssDNA71. Further studies will be necessary to clarify the effects of various MTSs on off-target editing events by DdCBEs as well as to develop novel approaches to reduce the adverse effects. Using an unbiased method (Detect-seq) to evaluate genome-wide specificity, Lei. et al.72 demonstrated that hundreds of nuclear off-target edited sites were induced by DdCBEs in both TALE array sequence-dependent and sequence-independent cases. The studies suggested that certain interactions between the DdCBE and the CTCF binding region seem to be associated with sequence-independent off-target effects. However, the exact mechanism underlying this observation remains to be determined. Interestingly, this off-target editing issue was significantly improved by applying several advanced DdCBE constructs: UGI-NES-DdCBE, to which NES sequences were added to hinder the localization of DdCBE in the nuclei, and DddIA-DdCBE, in which DddIA, a natural immunity protein of DddA, was used to preclude nuclear DNA editing by DdCBE by obturating the active center of DddAtox59,72. Nonetheless, further thorough analyses of off-target editing of both mtDNA and nuclear DNA by various mtDNA editing technologies will be necessary for both basic research and clinical applications in the future.
Conclusion
The potential of ZFPs and TALEs is promising, and their approaches for the base editing of mtDNA have been demonstrated. However, these platforms are limited by size compatibility, which makes their delivery to mitochondria less efficient using viral systems such as AAVs. In addition, there is a need for more efficient design and assembly methods for mtZFN and mitoTALEN monomers to recognize a wide spectrum of mtDNA sequences52,73. Consequently, the production of these proteins requires cost- and labor-intensive processes. These challenges could be overcome by using CRISPR/Cas9, a powerful editing system applicable to a wide number of organisms, including plants and mammals. CRISPR/Cas9 relies on only two components: (i) sgRNA, which recognizes a specific DNA sequence, and (ii) Cas9 nuclease, which cleaves the DNA sequence. However, as mentioned above, a major challenge in applying this system in mtDNA editing is to find effective methods for delivering exogenous sgRNA into the mitochondrion. If this could be achieved, the CRISPR/Cas9 system could potentially be broadly applied to manipulate mtDNA.
In fact, several stem‒loop motifs mediating the shuttling of nuclear RNA molecules into mitochondria have been characterized at the 5′ or 3′ ends of a variety of endogenous RNAs, such as 5S rRNA, RNase MRP, and H1 RNA, which play an essential role in the functioning of mammalian mitochondria74,75,76. In particular, stem‒loop motifs from H1 RNA, which are appended to cytosolic RNA, facilitate the efficient importing of the RNA molecules into mitochondria56. Furthermore, it has been suggested that some 5S RNA motifs and two domains (yeast cytosolic tRNALys (CUU)-derived F-arm and D-hairpin) can also assist the uptake of synthetic RNA into mitochondria77,78,79. Moreover, it has been reported that the fusion of an MTS with the AAV2 capsid protein VP2 helped to target the NADH-encoding ND4 gene in mitochondria80. Strikingly, a recent study revealed that the use of an engineered Cas9 nuclease linked with N- or C-terminus MTSs for editing mtDNA mutations in HEK293T cells and zebrafish resulted in a significant reduction in mtDNA copy number27. These reports provide a critical proof of concept for the delivery of sgRNA appended to a stem‒loop element into mitochondria, followed by an interaction with the Cas9 nuclease and cleavage of specific mtDNA sequences. Moreover, not only Cas9 but also Cas12a can enter the mitochondria by using MTSs, and there have been attempts at mitochondrial delivery by engineering various stem‒loop motifs on guide RNAs31. Together, the studies suggest that efficient delivery of sgRNA via stem‒loop motifs show great promise for mtDNA-editing systems (Fig. 3).
ZFP- and TALE-based gene editors or engineered CRISPR/Cas systems modified with MTS and stem‒loop motifs can be imported into mitochondria via transfection with AAVs or eVLPs, which encapsulate plasmid DNA or mRNA encoding the gene editing platforms. mtZFN, mitochondrial zinc-finger nuclease; TALEN transcription activator-like effector nuclease, DdCBE DddA-derived cytosine base editor, ZFD zinc-finger deaminase, TALE-linked deaminase (TALED); CRISPR/Cas clustered regularly interspaced short palindromic repeats/Cas.
Recently, a transient ribonucleoprotein (RNP) delivery platform using engineered virus-like particles (eVLPs) has been reported81 (Fig. 3). Essentially, eVLPs are empty viral shells into which therapeutic RNPs can be packed and delivered to the target DNA with minimal off-target editing effects81. Specifically, a newly engineered variant of eVLPs (v4 eVLPs) showed 16- and 4.7-fold increases in the packaging capacities of base editor RNPs and Cas9, respectively, compared to earlier architectures82. Most excitingly, eVLPs have been successfully used for delivering therapeutic packages to various organs in mice (liver, brain, and retina). These studies underscore the possibility of utilizing eVLP to deliver gene-editing cargoes for mtDNA to cells or multiple targeted organs in both animals and humans with minimal off-target editing81. Despite the potential of transient and efficient delivery of eVLPs, further studies on their pharmacokinetics, such as half-lives, are imperative. Additionally, determining the adequate requirements for base-editing cargoes may be a genetic therapy milestone for mtDNA editing in the future. Recently, extraordinarily novel applications based on both TALEs and ZFPs have been developed for highly specific mtDNA editing platforms toward heteroplasmic shifting with minimal cytotoxicity.
Optimizing the engineering of these platforms will surely pave the way for genetic approaches toward clinical trials by reducing the size to improve the packaging capability of viral systems and promoting efficiency and specificity for mtDNA manipulation. In fact, novel types of CRISPR systems, mini CRISPR-AsCas12f1 and miniature CRISPR-SpaCas12f1, consisting of 422 and 497 amino acids, respectively, can be feasibly and readily delivered to bacteria, cells, and tissues employing various delivery methods, such as AAV, plasmid, and RNP, have been reported83,84. Due to the advantages of editing efficiency and delivery, miniature platforms should also be considered an effective approach for mtDNA editing. Although the successful delivery of exogenous sgRNA into mitochondria using CRISPR‒Cas systems could prompt a revolution in mtDNA editing, it is undeniable that there are still real-world obstacles that need to be resolved for exceptional competence in the application of TALEs and ZFPs.
References
Schell, J. C. et al. A role for the mitochondrial pyruvate carrier as a repressor of the Warburg effect and colon cancer cell growth. Mol. Cell 56, 400–413 (2014).
Herzig, S. et al. Identification and functional expression of the mitochondrial pyruvate carrier. Science 337, 93–96 (2012).
Filippi, M.-D. & Ghaffari, S. Mitochondria in the maintenance of hematopoietic stem cells: new perspectives and opportunities. Blood 133, 1943–1952 (2019).
Liu, X., Kim, C. N., Yang, J., Jemmerson, R. & Wang, X. Induction of apoptotic program in cell-free extracts: requirement for dATP and cytochrome c. Cell 86, 147–157 (1996).
Gomes, L. C., Benedetto, G. D. & Scorrano, L. During autophagy mitochondria elongate, are spared from degradation and sustain cell viability. Nat. Cell Biol. 13, 589–598 (2011).
Rambold Angelika, S., Kostelecky, B., Elia, N. & Lippincott-Schwartz, J. Tubular network formation protects mitochondria from autophagosomal degradation during nutrient starvation. Proc. Natl Acad. Sci. USA 108, 10190–10195 (2011).
Xiong, J., Camello, P. J., Verkhratsky, A. & Toescu, E. C. Mitochondrial polarisation status and [Ca2+]i signalling in rat cerebellar granule neurones aged in vitro. Neurobiol. Aging 25, 349–359 (2004).
Tang, S. et al. Mitochondrial Ca²+ uniporter is critical for store-operated Ca2+ entry-dependent breast cancer cell migration. Biochem. Biophys. Res. Commun. 458, 186–193 (2015).
Yasukawa, K. et al. Mitofusin 2 inhibits mitochondrial antiviral signaling. Sci. Signal. 2, ra47 (2009).
Kaniak-Golik, A. & Skoneczna, A. Mitochondria–nucleus network for genome stability. Free Radic. Biol. Med. 82, 73–104 (2015).
Chen, X. et al. Rearranged mitochondrial genomes are present in human oocytes. Am. J. Hum. Genet. 57, 239–247 (1995).
D’Erchia, A. M. et al. Tissue-specific mtDNA abundance from exome data and its correlation with mitochondrial transcription, mass and respiratory activity. Mitochondrion 20, 13–21 (2015).
Anderson, S. et al. Sequence and organization of the human mitochondrial genome. Nature 290, 457–465 (1981).
Schon, K. R., Ratnaike, T., van den Ameele, J., Horvath, R. & Chinnery, P. F. Mitochondrial diseases: a diagnostic revolution. Trends Genet. 36, 702–717 (2020).
Tuppen, H. A. L., Blakely, E. L., Turnbull, D. M. & Taylor, R. W. Mitochondrial DNA mutations and human disease. Biochim. Biophys. Acta Bioenerg. 1797, 113–128 (2010).
El-Hattab, A. W. & Scaglia, F. Mitochondrial cytopathies. Cell Calcium 60, 199–206 (2016).
Leber, T. Ueber hereditäre und congenital-angelegte Sehnervenleiden. Albrecht Graef. Arch. Ophthalmol. 17, 249–291 (1871).
Man, P. Y. W., Turnbull, D. M. & Chinnery, P. F. Leber hereditary optic neuropathy. J. Med. Genet. 39, 162 (2002).
Shoffner, J. M. et al. Myoclonic epilepsy and ragged-red fiber disease (MERRF) is associated with a mitochondrial DNA tRNALys mutation. Cell 61, 931–937 (1990).
Pfeffer, G. et al. Mutations in the SPG7 gene cause chronic progressive external ophthalmoplegia through disordered mitochondrial DNA maintenance. Brain 137, 1323–1336 (2014).
Kauppila, J. H. K. et al. A phenotype-driven approach to generate mouse models with pathogenic mtDNA mutations causing mitochondrial disease. Cell Rep. 16, 2980–2990 (2016).
Srivastava, S. & Moraes, C. T. Manipulating mitochondrial DNA heteroplasmy by a mitochondrially targeted restriction endonuclease. Hum. Mol. Genet. 10, 3093–3099 (2001).
Tanaka, M. et al. Gene therapy for mitochondrial disease by delivering restriction endonuclease SmaI into mitochondria. J. Biomed. Sci. 9, 534–541 (2002).
Bayona-Bafaluy Maria, P., Blits, B., Battersby Brendan, J., Shoubridge Eric, A. & Moraes Carlos, T. Rapid directional shift of mitochondrial DNA heteroplasmy in animal tissues by a mitochondrially targeted restriction endonuclease. Proc. Natl Acad. Sci. USA 102, 14392–14397 (2005).
Wang, H., La Russa, M. & Qi, L. S. CRISPR/Cas9 in genome editing and beyond. Annu. Rev. Biochem. 85, 227–264 (2016).
Jo, A. et al. Efficient mitochondrial genome editing by CRISPR/Cas9. Biomed. Res. Int. 2015, 305716–305716 (2015).
Bian, W.-P. et al. Knock-in strategy for editing human and zebrafish mitochondrial DNA using mito-CRISPR/Cas9 system. ACS Synth. Biol. 8, 621–632 (2019).
Gammage, P. A., Moraes, C. T. & Minczuk, M. Mitochondrial genome engineering: the revolution may not be CRISPR-Ized. Trends Genet. 34, 101–110 (2018).
Hussain, S. A., Yalvac, M. E., Khoo, B., Eckardt, S. & McLaughlin, K. J. Adapting CRISPR/Cas9 system for targeting mitochondrial genome. Front. Genet. 12, 627050 (2021).
Yin, T., Luo, J., Huang, D. & Li, H. Current Progress of Mitochondrial Genome Editing by CRISPR. Front. Physiol. https://doi.org/10.3389/fphys.2022.883459 (2022).
Antón, Z. et al. Mitochondrial import, health and mtDNA copy number variability seen when using type II and type V CRISPR effectors. J. Cell Sci. https://doi.org/10.1242/jcs.248468 (2020).
Kim, Y. G., Cha, J. & Chandrasegaran, S. Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc. Natl Acad. Sci. USA 93, 1156–1160 (1996).
Smith, J. et al. Requirements for double-strand cleavage by chimeric restriction enzymes with zinc finger DNA-recognition domains. Nucleic Acids Res. 28, 3361–3369 (2000).
Christian, M. et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186, 757–761 (2010).
Gammage, P. A., Rorbach, J., Vincent, A. I., Rebar, E. J. & Minczuk, M. Mitochondrially targeted ZFNs for selective degradation of pathogenic mitochondrial genomes bearing large-scale deletions or point mutations. EMBO Mol. Med. 6, 458–466 (2014).
Bacman, S. R., Williams, S. L., Pinto, M., Peralta, S. & Moraes, C. T. Specific elimination of mutant mitochondrial genomes in patient-derived cells by mitoTALENs. Nat. Med. 19, 1111–1113 (2013).
Beerli, R. R., Segal, D. J., Dreier, B. & Barbas, C. F. 3rd Toward controlling gene expression at will: specific regulation of the erbB-2/HER-2 promoter by using polydactyl zinc finger proteins constructed from modular building blocks. Proc. Natl Acad. Sci. USA 95, 14628–14633 (1998).
Beerli, R. R., Dreier, B. & Barbas, C. F. 3rd Positive and negative regulation of endogenous genes by designed transcription factors. Proc. Natl Acad. Sci. USA 97, 1495–1500 (2000).
Gersbach, C. A., Gaj, T. & Barbas, C. F. Synthetic zinc finger proteins: the advent of targeted gene regulation and genome modification technologies. Acc. Chem. Res. 47, 2309–2318 (2014).
Minczuk, M., Papworth Monika, A., Kolasinska, P., Murphy Michael, P. & Klug, A. Sequence-specific modification of mitochondrial DNA using a chimeric zinc finger methylase. Proc. Natl Acad. Sci. USA 103, 19689–19694 (2006).
Minczuk, M., Papworth, M. A., Miller, J. C., Murphy, M. P. & Klug, A. Development of a single-chain, quasi-dimeric zinc-finger nuclease for the selective degradation of mutated human mitochondrial DNA. Nucleic Acids Res. 36, 3926–3938 (2008).
Minczuk, M., Kolasinska-Zwierz, P., Murphy, M. P. & Papworth, M. A. Construction and testing of engineered zinc-finger proteins for sequence-specific modification of mtDNA. Nat. Protoc. 5, 342–356 (2010).
Gammage, P. A. et al. Near-complete elimination of mutant mtDNA by iterative or dynamic dose-controlled treatment with mtZFNs. Nucleic Acids Res. 44, 7804–7816 (2016).
Gaude, E. et al. NADH shuttling couples cytosolic reductive carboxylation of glutamine with glycolysis in cells with mitochondrial dysfunction. Mol. Cell 69, 581–593.e587 (2018).
Gammage, P. A. et al. Genome editing in mitochondria corrects a pathogenic mtDNA mutation in vivo. Nat. Med. 24, 1691–1695 (2018).
Lim, K., Cho, S.-I. & Kim, J.-S. Nuclear and mitochondrial DNA editing in human cells with zinc finger deaminases. Nat. Commun. 13, 366 (2022).
Perez-Quintero, A. L. & Szurek, B. A decade decoded: spies and hackers in the history of TAL effectors research. Annu. Rev. Phytopathol. 57, 459–481 (2019).
Boch, J. et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326, 1509–1512 (2009).
Bogdanove, A. J. & Voytas, D. F. TAL effectors: customizable proteins for DNA targeting. Science 333, 1843–1846 (2011).
Richter, A., Streubel, J. & Boch, J. in TALENs: Methods and Protocols (eds R Kühn, W Wurst, & B Wefers) 9–25 (Springer New York, 2016).
Gaj, T., Gersbach, C. A. & Barbas, C. F. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol. 31, 397–405 (2013).
Hashimoto, M. et al. MitoTALEN: a general approach to reduce mutant mtDNA loads and restore oxidative phosphorylation function in mitochondrial diseases. Mol. Ther. 23, 1592–1599 (2015).
Yahata, N., Matsumoto, Y., Omi, M., Yamamoto, N. & Hata, R. TALEN-mediated shift of mitochondrial DNA heteroplasmy in MELAS-iPSCs with m.13513G>A mutation. Sci. Rep. 7, 15557 (2017).
Yang, Y. et al. Targeted elimination of mutant mitochondrial DNA in MELAS-iPSCs by mitoTALENs. Protein Cell 9, 283–297 (2018).
Phillips, A. F. et al. Single-molecule analysis of mtDNA replication uncovers the basis of the common deletion. Mol. Cell 65, 527–538.e526 (2017).
Tonin, Y. et al. Modeling of antigenomic therapy of mitochondrial diseases by mitochondrially addressed RNA targeting a pathogenic point mutation in mitochondrial DNA*. J. Biol. Chem. 289, 13323–13334 (2014).
Reddy, P. et al. Selective elimination of mitochondrial mutations in the germline by genome editing. Cell 161, 459–469 (2015).
Pereira, C. V. et al. mitoTev-TALE: a monomeric DNA editing enzyme to reduce mutant mitochondrial DNA levels. EMBO Mol. Med. 10, e8084 (2018).
Mok, B. Y. et al. A bacterial cytidine deaminase toxin enables CRISPR-free mitochondrial base editing. Nature 583, 631–637 (2020).
Wei, Y. et al. Human cleaving embryos enable efficient mitochondrial base-editing with DdCBE. Cell Discov. 8, 7 (2022).
Sabharwal, A. et al. The FusX TALE base editor (FusXTBE) for rapid mitochondrial DNA programming of human cells in vitro and zebrafish disease models in vivo. CRISPR J. 4, 799–821 (2021).
Guo, J. et al. Precision modeling of mitochondrial diseases in zebrafish via DdCBE-mediated mtDNA base editing. Cell Discov. 7, 78 (2021).
Guo, J. et al. DdCBE mediates efficient and inheritable modifications in mouse mitochondrial genome. Mol. Ther. 27, 73–80 (2022).
Lee, H. et al. Mitochondrial DNA editing in mice with DddA-TALE fusion deaminases. Nat. Commun. 12, 1190 (2021).
Qi, X. et al. Precision modeling of mitochondrial disease in rats via DdCBE-mediated mtDNA editing. Cell Discov. 7, 95 (2021).
Kang, B.-C. et al. Chloroplast and mitochondrial DNA editing in plants. Nat. Plants 7, 899–905 (2021).
Silva-Pinheiro, P. et al. In vivo mitochondrial base editing via adeno-associated viral delivery to mouse post-mitotic tissue. Nat. Commun. 13, 750 (2022).
Nakazato, I. et al. Targeted base editing in the plastid genome of Arabidopsis thaliana. Nat. Plants 7, 906–913 (2021).
Cho, S.-I. et al. Targeted A-to-G base editing in human mitochondrial DNA with programmable deaminases. Cell 185, 1764–1776.e1712 (2022).
Mok, B. Y. et al. CRISPR-free base editors with enhanced activity and expanded targeting scope in mitochondrial and nuclear DNA. Nat. Biotechnol. https://doi.org/10.1038/s41587-022-01256-8 (2022).
Wei, Y. et al. Mitochondrial base editor DdCBE causes substantial DNA off-target editing in nuclear genome of embryos. Cell Discov. 8, 27 (2022).
Lei, Z. et al. Mitochondrial base editor induces substantial nuclear off-target mutations. Nature https://doi.org/10.1038/s41586-022-04836-5 (2022).
Moraes, C. T. A magic bullet to specifically eliminate mutated mitochondrial genomes from patients’ cells. EMBO Mol. Med. 6, 434–435 (2014).
Smirnov, A., Entelis, N., Martin, R. P. & Tarassov, I. Biological significance of 5S rRNA import into human mitochondria: role of ribosomal protein MRP-L18. Genes Dev. 25, 1289–1305 (2011).
Puranam Ram, S. & Attardi, G. The RNase P associated with HeLa cell mitochondria contains an essential RNA component identical in sequence to that of the nuclear RNase P. Mol. Cell. Biol. 21, 548–561 (2001).
Mercer, TimR. et al. The human mitochondrial transcriptome. Cell 146, 645–658 (2011).
Zelenka, J. & Ježek, P. in Mitochondrial DNA: Methods and Protocols (ed Matthew McKenzie) 175–181 (Springer New York, 2016).
Towheed, A., Markantone, D. M., Crain, A. T., Celotto, A. M. & Palladino, M. J. Small mitochondrial-targeted RNAs modulate endogenous mitochondrial protein expression in vivo. Neurobiol. Dis. 69, 15–22 (2014).
Smirnov, A. et al. Two distinct structural elements of 5S rRNA are needed for its import into human mitochondria. RNA 14, 749–759 (2008).
Yu, H. et al. Gene delivery to mitochondria by targeting modified adenoassociated virus suppresses Leber’s hereditary optic neuropathy in a mouse model. Proc. Natl Acad. Sci. USA 109, E1238–E1247 (2012).
Banskota, S. et al. Engineered virus-like particles for efficient in vivo delivery of therapeutic proteins. Cell 185, 250–265.e216 (2022).
Mangeot, P. E. et al. Genome editing in primary cells and in vivo using viral-derived Nanoblades loaded with Cas9-sgRNA ribonucleoproteins. Nat. Commun. 10, 45 (2019).
Wu, Z. et al. Programmed genome editing by a miniature CRISPR-Cas12f nuclease. Nat. Chem. Biol. 17, 1132–1138 (2021).
Wang, Y. et al. Guide RNA engineering enables efficient CRISPR editing with a miniature Syntrophomonas palmitatica Cas12f1 nuclease. Cell Rep. 40, 111418 (2022).
Acknowledgements
This study was supported by the Chung Yang, Cha Young Sun, and Jang Hi Joo Memorial Fund. This work was also supported by the Bio & Medical Technology Development Program of the National Research Foundation (NRF) of Korea (NRF-2020M3A9D5A01082439 and NRF-2019R1A2C2087198) and the Korea Research Institute of Bioscience and Biotechnology (KRIBB) Research Initiative Program. The illustrations were generated using BioRender.com.
Author information
Authors and Affiliations
Contributions
H.T.L.P., H.L., and K.K. discussed and commented on the paper. H.L. and K.K. supervised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Phan, H.T.L., Lee, H. & Kim, K. Trends and prospects in mitochondrial genome editing. Exp Mol Med 55, 871–878 (2023). https://doi.org/10.1038/s12276-023-00973-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s12276-023-00973-7
- Springer Nature Limited