Abstract
Although members of SOX family have been well documented for their essential roles in embryonic development, cell proliferation and disease, the functional role and molecular mechanism of SOX30 in cancer are largely unexplored. Here, we first identified SRY-box containing gene 30 (SOX30) as a novel preferentially methylated gene using genome-wide methylation screening. SOX30 hypermethylation was detected in 100% of lung cancer cell lines (9/9) and 70.83% (85/120) of primary lung tumor tissues compared with none (0/20) of normal and 8.0% (2/25) of peri-tumoral lung tissues (P<0.01). SOX30 was expressed in normal and peri-tumoral lung tissues in which SOX30 was unmethylated, but was silenced or downregulated in lung cancer cell lines and primary lung tumor tissues harboring a hypermethylated SOX30. De-methylation experiments further confirmed that silence of SOX30 was regulated by its hypermethylation. Ectopic expression of SOX30 induces cancer cell apoptosis with inhibiting proliferation in vitro and represses tumor formation in vivo, whereas knockdown of SOX30 demonstrates a reversed effect both in vitro and in vivo. At the molecular level, the antitumorigenic effect of SOX30 is mediated by directly binding to CACTTTG (+115 to +121) of p53 promoter region and activating p53 transcription, suggesting that SOX30 is a novel transcriptional activating factor of p53. Indeed, blockade of p53 attenuates the tumor inhibition of SOX30. Overall, these findings demonstrate that SOX30 is a novel epigenetic silenced tumor suppressor acting through direct regulation of p53 transcription and expression. This study provides novel insights on the mechanism of tumorigenesis in lung cancer.
Similar content being viewed by others
Introduction
Lung cancer is the most commonly diagnosed cancer, as well as the leading cause of cancer death in males and among females, it is the fourth most frequent cancer and the second leading cause of cancer death in 2008 globally.1, 2 It represents the most common malignancy and is rapidly increasing in China. Carcinogenesis is a complex multistep process presenting a variety of genetic and epigenetic abnormalities. Aberrant epigenetic changes are one of the most frequent events and are regarded as important mechanisms in carcinogenesis.3, 4 Moreover, methylation profiles have been used as potential biomarkers for early diagnosis, prognosis and screening in some cancers.5 Recently, accumulating evidence demonstrated that DNA hypermethylation of tumor-suppressor genes (TSGs) associated with gene silencing has an essential role in carcinogenesis.6, 7, 8, 9, 10 Increasing numbers of TSGs associated with epigenetic alterations have been identified in human cancers.9, 11, 12, 13 The identification of new useful biomarkers and new genes functionally involved in tumor development may provide alternative approaches for diagnostic and prognostic evaluation.
Through methylation-sensitive representational difference analysis, we have identified a novel preferentially methylated gene, SRY-box containing gene 30 (SOX30), in human lung cancer. SOX family members contain a highly conserved high mobility group (HMG) DNA-binding domain,14 and have critical roles in the regulation of cell fate and differentiation during embryonic and postnatal development.15 So far, SOX30 has been characterized in only a few species. It was first cloned from mouse and human.16 Recently, Sox30 was isolated from the Nile tilapia accidentally and was indicated to exist widely throughout the animal kingdom in our previous studies.17 In mouse and human, Sox30 is considered to be involved in mammalian spermatogonial differentiation and spermatogenesis.16, 18 In the Nile tilapia, Sox30 may be involved in female and male gonadal development.17 However, it remains unclear whether SOX30 has any role in cancer.
In this study, we observed a frequent loss of SOX30 expression because of DNA hypermethylation in human lung cancers. Gain- and loss-of-function studies demonstrated that SOX30 induced apoptosis with inhibiting proliferation of lung cancer cell lines in vitro, and repressed tumor growth in vivo. Further, SOX30 directly activated p53 transcription and expression, which mediated its function as a tumor suppressor.
Results
SOX30 is hypermethylated in lung cancer cell lines and lung cancers
To screen for differentially methylated DNA fragments and potential cancer-related genes with methylation, we used genome-wide methylation screening and identified a novel preferentially methylated gene SOX30 in lung cancer. Pairs of primers for methylation-specific polymerase chain reaction (MSP) and bisulfite genomic sequencing (BGS) were designed (Figure 1a). The MSP analysis showed that SOX30 was hypermethylated in lung cancer cell lines and a substantial proportion of cancer cases (Figures 1b and c). In contrast, SOX30 of non-tumor lung tissues exhibited an unmethylated status (Figures 1b and c). The MSP results were further validated by BGS analysis of SOX30 isolated from A549, H460, H358, T8 and N6 cell lines or tissue samples (Figures 1d and e).
Methylation status of SOX30 in lung cancer cell lines and tissues. (a) Schematic representation of the human SOX30. Open and closed boxes indicate the non-coding and coding regions, respectively, and an arrow denotes the transcriptional start site (+1). ATG is the start codon. Vertical bars show CpG sites. Arrows below the CpG sites indicate the regions subjected to MSP and BGS. (b) MSP analyses in lung cancer cell lines and normal lung tissues. M, methylation-specific primers; MK, marker; N, non-tumor; Nc, negative control (deionized water); U, unmethylated-specific primers. (c) Representative electrophoresis results of MSP analyses in non-tumor and tumor lung samples. T, tumor. (d) BGS of SOX30 in lung cancer cell lines was performed to confirm MSP results. Solid circles, methylated CpG sites; open circles, unmethylated CpG sites. (e) BGS of SOX30 in non-tumor and tumor samples was performed to confirm MSP results.
In total, we examined SOX30 methylation in 20 normal lung samples, 25 adjacent controls, 120 tumors and 9 lung cancer cell lines by MSP. The methylation incidence of SOX30 was 0% (0/20), 8% (2/25), 70.83% (85/120) and 100% (9/9) in these samples, respectively (Supplementary Table S2). The frequency of SOX30 methylation was lower in normal lung tissues from the control subjects than in lung cancer tissues from patients (0/20 (0%) vs 85/120 (70.83%); P<0.001), and it did not differ in adjacent tissues of cancer patients from normal lung tissues (2/25 (8%) vs 0/20 (0%); P=0.495) (Supplementary Table S2). When analyzing the relationship between SOX30 methylation status and clinical characteristics of these patients (after exclusing those with incomplete clinicopathological features, a total of 84 cases were analyzed), we did not find significant association of SOX30 methylation status with gender, age, grade, tumor size or tumor types (Supplementary Table S3).
Methylation of SOX30 is correlated with its transcriptional silencing
To determine the relationship between hypermethylation and expression of SOX30, a correlation analysis was performed on cell lines and pairs of primary tumor vs non-tumor tissues by reverse transcription–polymerase chain reaction (RT–PCR), quantitative RT–PCR (qRT–PCR), western blot (WB) and immunohistochemistry. The results showed that SOX30 was silenced or downregulated in cancer cell lines and tumor samples with hypermethylated SOX30 (Figures 2a and b, Supplementary Figure S1A). In contrast, SOX30 could be broadly detected in normal lung tissues in which SOX30 is unmethylated (Figure 2b, Supplementary Figure S1A). The expression of SOX30 was lower in lung tumors than that in adjacent non-tumor and normal lung tissues (Figure 2c, Supplementary Figures S1A–C), suggesting an aberrant gene silencing of SOX30 in lung cancer. These results indicate that silencing of SOX30 in tumors may be due to hypermethylation of SOX30. To test this hypothesis, we treated cell lines with 5-aza-2′-deoxycytidine, a pharmacological inhibitor of DNA methylation, which restored the expression of the silenced SOX30 in those cell lines (Figure 2d, Supplementary Figures S2A and B), suggesting that methylation is essential for silencing of SOX30. In addition, we also detected SOX30 expression levels during human lung development, and found that SOX30 was relatively highly expressed in human adult lung (Supplementary Figure S2C).
Expression patterns of SOX30 in lung cancer cell lines and lung tissues. (a) RT–PCR analysis of SOX30 mRNA levels in nine lung cancer cell lines. MK, Marker; NC, negative control (deionized water); PC, positive control (SOX30 plasmid DNA). ACTIN was used as an internal control. (b) QRT–PCR analysis of SOX30 expression levels in normal and tumor lung samples. (c) Immunohistochemistry (IHC) analysis of SOX30 expression levels in adjacent and tumor lung samples. SOX30 is highly expressed in adjacent lung tissues. Immunohistological staining assays were performed with an anti-SOX30 antibody (diaminobenzidine (DAB) staining, scale bars, 100 μm). (d) The de-methylation analysis of SOX30 expression. Lung cancer cell lines were treated with or without 5-aza-2’-deoxycytidine (5-aza-dc; 10 μm) for three days, and SOX30 expression was examined by qRT–PCR. *P<0.05; **P<0.01. Error bars indicate s.d. (n=3).
Genetic deletion and mutation of SOX30 is not detected in lung cancer
We next determined genetic deletion and mutation of SOX30 coding exons and promoter by DNA direct sequencing. We did not observe any homozygous deletion or mutation of SOX30 in three cancer cell lines (A549, H460 and H1975) and five primary cancers (chosen randomly), suggesting that genetic alteration does not contribute to the silencing of SOX30 in lung cancer (data not shown).
SOX30 overexpression inhibits proliferation and promotes apoptosis
To explore the potential role of SOX30 in tumorigenesis, we first generated gain-of-function cell models by transfecting a SOX30-expressing construct into the human adenocarcinoma A549 (A549) and large cell lung cancer NCI-H460 (H460) cell lines. The expression of exogenous SOX30 was confirmed by RT–PCR and WB (Figure 3a). We then examined the effect of SOX30 overexpression on cell proliferation and viability. Five-day growth curve analysis showed that overexpression of SOX30 inhibited proliferation of A549 and H460 cells (Figure 3b). The suppressive effect of SOX30 on cell proliferation was confirmed by colony formation and EdU assays. SOX30-expressing A549 and H460 (SOX30) formed about 20 and 40% colonies of their controls (Vector), respectively (P<0.01) (Figures 3c and d). SOX30 overexpression decreased the number of EdU+ cells in both A549 and H460 cell lines (Figure 3e).
Analyses of cell proliferation and apoptosis associated with SOX30 overexpression in A549 and H460 cells. (a) Transfectants of SOX30 and vector control in A549 and H460 cells were identified by RT–PCR and WB. Abundant SOX30 was detected after SOX30 transfection but not after the control vector transfection. (b) MTS assays were used to examine the effect of SOX30 on proliferation in A549 and H460 cells. Cell viability was evaluated in triplicate by CellTiter 96 AQueous One Solution Cell Proliferation Assay (Promega). *P<0.05; **P<0.01. (c, d) The effect of SOX30 on cell growth was further confirmed by colony formation assay. Surviving colonies were counted. Error bars indicate s.d. (n=3). (e) Photomicrographs of A549 and H460 cells at 46 h after transfection with SOX30 and vector control. EdU, 5-ethynyl-2’-deoxyuridine. (f) Flow cytometry assay with double staining in A549 and H460 cells. Cell apoptosis was detected by Annexin V-APC/7-amino-actinomycin D double staining. Error bars indicate s.d. (n=3).
To examine the effect of SOX30 overexpression on cell viability, we measured the percentage of sub-G1 phase cells, and conducted Annexin V-APC/7-amino-actinomycin D double staining followed by flow cytometry analysis and DNA ladder assay. Both A549 and H460 cells transfected with SOX30 had a significant higher percentage of sub-G1 apoptotic cells as compared with vector control transfectants (Supplementary Figure S3A). Annexin V-APC/7-amino-actinomycin D double staining analysis showed SOX30 overexpression in A549 and H460 cells resulted in an increase of early apoptotic cells and late apoptotic cells (Figure 3f, Supplementary Figure S3B). DNA ladder assay showed a typical DNA ladder formation in the SOX30-transfected cells, but not in the vector control-transfected cells (Supplementary Figure S3C). Taken together, these data demonstrated that SOX30 overexpression inhibited proliferation and promoted apoptosis in A549 and H460 cells.
Knockdown of SOX30 promotes proliferation and inhibits apoptosis
To further confirm the potential roles of SOX30 in lung cancer cells, the effect of knockdown SOX30 with micro RNA (miRNA) in a stable transfectants (A549-SOX30 stably) was investigated. RT–PCR and WB showed that SOX30 was significantly reduced in the transfectant (Figure 4a). The MTS assay, colony formation assay and flow cytometry analysis showed that knockdown of SOX30 promotes cell proliferation (Figures 4b and c) but inhibits cell apoptosis (Figure 4d), compared with cells transfected with the negative control. These data together with the aforementioned results from SOX30 overexpression study suggested that SOX30 might function as a potential tumor suppressor.
Effects of SOX30 knockdown by miRNA on cell proliferation and apoptosis in A549-SOX30 stably cells. (a) The knockdown expression of SOX30 was confirmed by RT–PCR and WB in A549-SOX30 stably cells. ACTIN was used as an internal control. (b) Cell proliferation assay was performed after SOX30 knockdown by MTS in A549-SOX30 stable cells. Cell viability was evaluated in triplicate by CellTiter 96 AQueous One Solution Cell Proliferation Assay (Promega). *P< 0.05; **P<0.01. (c) The effect of decreasing SOX30 expression on cell growth was further confirmed by colony formation assay in A549-SOX30 stable cells. (d) Cell apoptosis was detected by flow cytometry assay with Annexin V-APC/7-amino-actinomycin D double staining A549-SOX30 stably cells. Error bars indicate s.d. (n=3).
SOX30 inhibits tumor formation in nude mice
After a demonstration of the tumor suppression of SOX30 in vitro, we next wished to investigate whether SOX30 can suppress tumor growth in vivo. To this end, we used a xenograft tumor model to assess the growth of A549 or H460 stable transfectants in nude mice. The tumor volume was significantly smaller in mice receiving SOX30-transfected cells compared with those receiving cells transfected with vector control (Figure 5a). Overexpression of SOX30 resulted in a decrease of the mean weight of tumors collected 3 to 5 weeks after inoculation of the cells (P<0.05) (Figure 5b). In contrast, knockdown of SOX30 expression in SOX30 stable transfectants restored the malignant phenotype (Figures 5c and d). Collectively, these data indicated that SOX30 had a significant effect on impeding tumor formation, supporting SOX30 as a tumor suppressor in lung cancer in vivo.
Overexpression of SOX30 inhibits tumor formation in nude mice. (a) The tumor growth curve of SOX30-expressing cells was compared with vector control cells. Tumor growth in nude mice subcutaneously injected with A549 and H460 stable transfectants. A549 (3.5 × 106) and H460 (3.0 × 106) stable transfectants were injected into the left (SOX30) and right (vector control) flank of nude mice, respectively. Points represent the mean tumor volumes of three independent experiments (n=3). (b) Tumor weight from the SOX30 and vector control groups. The results were obtained from three independent experiments. *P<0.05; **P<0.01. (c) The tumor growth curve of miRNA1-expressing cells was compared with negative control-expressing cells. Tumor growth in nude mice subcutaneously injected with A549-SOX30 stable cells expressing miRNA1 or negative control. A549-SOX30 stable cells (3.5 × 106) transfected with miRNA1 or stable negative control were injected into the left (miRNA1) and right (negative control) flank of nude mice, respectively. (d) Tumor weight from the miRNA1-expressing and negative control-expressing groups. Error bars indicate s.d. (n=3).
Identification of potential target genes of SOX30
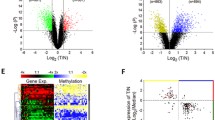
To gain insight into the molecular basis underlying the tumor-suppressive effect of SOX30, we performed a transcriptome analysis of SOX30 stable transfectants using gene expression microarray, and validated SOX30 targets by RT–PCR, qRT–PCR and WB. The antitumorigenesis effect of SOX30 was mediated by regulating genes mainly involved in controlling cell proliferation and apoptosis. Of particular interest, pro-apoptotic genes including p53 and its downstream target genes such as BAX, PMAIP1 and p21 were upregulated in SOX30 transfectants (Figures 6a and b, Supplementary Table S4, Supplementary Figure S4A). These results indicated that the antitumorigenicity of SOX30 might be mediated by activation of the p53 signaling pathway.
SOX30 accelerates expression of p53 signaling pathway-related genes. (a) Ectopic expression of SOX30 significantly (P<0.01) enhances endogenous expression of P53, P21, BAX and PMAIP1 in A549 and H460 cells. The mRNA expression was examined by qRT–PCR, and ACTIN was used as an internal control. *P<0.05; **P<0.01. (b) The protein levels of P53, P21, BAX and PMAIP1 were detected by WB after restoration of SOX30 in A549 and H460 cells. ACTIN was used as an internal control. (c, d) The mRNA and protein expression levels of SOX30 and P53 in xenograft tumors were examined by qRT–PCR and WB. ACTIN was used as an internal control. (e) SOX30 increases the number of terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling (TUNEL)-positive H460 cells in xenograft tumors. Two (1 and 2) representative images were randomly selected from each sample (H460-vector and H460-SOX30). The arrows indicate apoptotic cells.
To further determine whether p53 is the possible target gene of SOX30, the expression of SOX30 and p53 in xenograft tumors was examined by qRT–PCR and WB. The results indicated that p53 was increased markedly both at mRNA and protein levels in xenograft tumor tissues when SOX30 was expressed (Figures 6c and d, Supplementary Figure S4B). Furthermore, we detected cell proliferation and apoptosis in vivo using xenograft tumor tissues by Ki67 and terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling staining, respectively. The results showed that a decrease in proliferating cells and an increase in apoptotic cells were observed in SOX30-overexpressing tumors (Supplementary Figure S4C, Figure 6e). Taken together, these data suggested that p53 was the highly possible target gene of SOX30 in lung cancer.
P53 is a direct target of SOX30
To investigate whether p53 is a direct target of SOX30, we searched the putative SOX transcription factor-binding sites in p53 promoter using Jaspar (http://jaspar.genereg.net/). Several SOX transcription factor-binding sites were identified in p53 promoter region (Figure 7a, Supplementary Figure S4D). Regarding high similarity of the DNA-binding domains (HMG-boxes) of these SOX members and SOX30 (Supplementary Figure S4E), we speculated that SOX30 might regulate p53 transcription by directly binding to its promoter region. To examine this hypothesis, we first performed a luciferase reporter assay. The data showed that overexpression of SOX30 significantly enhanced the activity of p53 promoter (Figure 7b). Then, by using chromatin immunoprecipitation (ChIP)-PCR assays, we found direct binding of SOX30 to the promoter of p53, but not PMAIP1, p21 and BAX (Figure 7c, Supplementary Figures S5A–C). To further determined the SOX30-binding sites in the p53 promoter, different regions of the p53 promoter were analyzed by luciferase reporter assays, and the SOX30-binding site was very likely on the CACTTTG (+115 to +121) locus of the p53 promoter region using both Jaspar and ConSite (http://consite.genereg.net/cgi-bin/consite) searching (Figure 7d, Supplementary Figure S5D). These findings demonstrated that p53 was a direct target gene of SOX30 in lung cancer cell lines.
SOX30 stimulates p53 signaling pathway by directly binding to promoter of P53. (a) The putative SOX30 transcription factor-binding sites in p53 promoter region and different regions of the p53 promoter were constructed into luciferase reporter. The possible binding sites are underlined; TSS, transcription start site. (b) SOX30 directly targets P53 as detected by luciferase reporter assay. A luciferase reporter linked with the full-length native promoter of P53 was used for the luciferase reporter assay in A549 and H460 cells. Results were normalized with internal controls and presented as averages with s.d. from three experiments. (c) ChIP-PCR was performed to identify P53 as a direct binding target of SOX30. (d) Different partial regions of p53 promoter were analyzed by luciferase reporter assay. Luciferase reporters linked with partial native promoter regions of P53 were used for the luciferase reporter assay in HEK-293 cells. The section in blue and italic is the overlap region of p53-luciferase-1 and p53-luciferase-2. The SOX30-binding region in P53 promoter should be at +87 to +135 from the luciferase reporter assay results of p53-luciferase-1 and p53-luciferase-2. WB analysis of SOX30 expression was performed to exclude that the differences in transcriptional activity reflect changes in expression. According to the binding sites predicted by Jaspar and ConSite, the direct binding sites is likely located at CACTTTG (+115 to +121) of the P53 promoter region. (e) The effect of SOX30 on human wild-type and mutated/deleted P53 promoter activity in HEK-293 cells. Up, the effect of SOX30 on the wild-type and mutated/deleted P53 promoter function. Down, human P53 promoter (p53-luciferase-2) construct containing SOX30-binding sites and the constructs with mutation/deletion of the SOX30-binding sites. Vector, empty vector+wild-type p53-luciferase-2; SOX30, SOX30+wild-type p53-luciferase-2; p53-Mu-1, SOX30+Mutated p53-luciferase-2-1 (CACTTTG, bold indicates mutated); p53-Mu-2, SOX30+Mutated p53-luciferase-2-2 (CACTTTG, bold indicates mutated); p53-Dele, SOX30+Deleted p53-luciferase-2 (CACTTTG, bold indicates deleted). Error bars indicate s.d. (n=3). (f) The effect of wild-type and deleted SOX30 HMG-box on human p53 promoter activity in HEK-293 cells. Up, effect of wild-type and deleted SOX30 on the P53 promoter function. Down, human SOX30 constructs with two deletions of the SOX30. SOX30, wild-type SOX30+wild-type p53-luciferase-2; SOX30-Dele-1, SOX30-Dele-1 (without HMG-box)+wild-type p53-luciferase-2; SOX30-Dele-2, SOX30-Dele-2 (with HMG-box)+wild-type p53-luciferase-2; **P<0.01.
SOX30 fails to activate p53 promoter activity when binding sites are mutated or deleted
To further investigate whether SOX30 activates the p53 promoter through binding to the binding site (CACTTTG) by SOX30 HMG-box, constructs with site-directed mutagenesis of binding sites in the p53 promoter (mutations and deletions) and SOX30 HMG-box (deletions) were generated. When the SOX30-binding sites in the p53 promoter were mutated (CACTTTG, bold indicates mutated) or deleted (CACTTTG, bold indicates deleted), the stimulating effect was ablated (Figure 7e). Furthermore, the data of site-directed mutagenesis (deletion) of SOX30 HMG-box showed that SOX30 HMG-box was required for stimulating p53 promoter activity (Figure 7f).
Ablation of p53 attenuates the effects of SOX30
To define whether p53 was required for the antitumorigenesis function of SOX30, we blocked p53 activity or its signaling pathway by using the inhibitor pifithrin-α, or p53 small interfering RNA when overexpression of SOX30 (Figures 8a and b, Supplementary Figure S6A). Blockade of p53 or its signaling pathway by pifithrin-α or small interfering RNA significantly (P<0.01) diminished the effect of SOX30 overexpression on cell proliferation and apoptosis (Figures 8c–e, Supplementary Figure S6B). Besides, in order to avoid the off-target effects of the small interfering RNA for p53, we evaluated SOX30-positive regulatory role on p53's expression and functions in wild-type and p53-knockout HCT116 cells. The data indicated that it had a higher percentage of apoptosis in SOX30 overexpression wild-type HCT116 cells than p53-knockout HCT116 cells (Supplementary Figures S6C–E). These data indicated that p53 was required for the function of SOX30 as a tumor suppressor in lung cancer cells.
P53 is a functionally important target gene of SOX30 in A549 cells. (a, b) The expression of SOX30 and P53 was analyzed by qRT–PCR and WB at 48 h after treatment with pifithrin-α or transfected with P53 small interfering RNA (siRNA) in A549 cells. The ACTIN was used for normalization. The p53-specific inhibitor pifithrin-α was purchased from Calbiochem. The P53 siRNA was purchased from Santa Cruz Biotechnology. (c) Analysis of cell proliferation in A549 cells treated with pifithrin-α or transfected with P53 siRNA when overexpression SOX30. Cell numbers were counted every day after treatment for 5 days. *P<0.05; **P<0.01. (d, e) Analysis of the effects of the expression of SOX30 and P53, respectively, on cell apoptosis in A549 cells. Cell apoptosis was detected at 48 h after treatment with pifithrin-α or transfected with P53 siRNA by flow cytometry assay with Annexin V-APC/7-amino-actinomycin D double staining. (f) Schematic diagram for the mechanisms of SOX30 antitumorigenesis functions deriving from expression array analysis, and validated by RT–PCR, qRT–PCR, WB, luciferase reporter and ChIP-PCR.
Discussion
In this study, we identified SOX30 as a novel preferentially methylated gene in human lung cancer, and it is frequently silenced or downregulated in lung cancer cell lines and lung cancer samples, but is expressed in human normal and peri-tumoral lung tissues. The data suggest that the silencing or downregulation of SOX30 is closely related with its hypermethylation, as demonstrated by MSP and confirmed by BGS. De-methylation treatment with the de-methylating reagent 5-aza-2′-deoxycytidine restores the expression of SOX30 in silenced cancer cell lines, and mutation analysis showed that genetic deletion or mutation was not detected. These results indicate that hypermethylation of SOX30 mediates the transcriptional silence directly.
To investigate the clinical application of SOX30 in lung tumorigenesis in vivo, we examined the methylation of SOX30 by MSP and BGS in normal control, adjacent non-tumor and lung tumor samples. The data showed that frequent SOX30 hypermethylation occurs in lung tumor tissues, but not in normal lung or adjacent non-tumor tissues, indicating SOX30 methylation being tumor specific. These findings suggest that SOX30 methylation may be a putative epigenetic biomarker for lung cancer, which may have a high research value and application prospect.
Previous studies have considered that Sox30 may be involved in gonadal development.16, 17, 18 In this study, SOX30 is highly methylated and frequently silenced or downregulated in human lung cancers, suggesting it may be involved in regulation lung cancer development. Ectopic expression of SOX30 in silenced lung cancer cell lines significantly induces cell apoptosis and inhibits proliferation. Conversely, knockdown of SOX30 inhibits cell apoptosis and increases proliferation. Furthermore, SOX30 significantly suppresses tumor growth in nude mice. These in vitro and in vivo studies indicate for the first time that SOX30 functions as a tumor suppressor in lung carcinogenesis.
This study improved our understanding of the molecular basis of SOX30 as a tumor suppressor. Transcriptome analysis showed that the expression of p53 and its targets, including BAX, PMAIP1 and p21, are upregulated by SOX30. Luciferase reporter and ChIP assays indicated that SOX30 directly binds to p53 promoter region, and induces its transcription and expression. Moreover, the effect of SOX30 on cancer cell proliferation and apoptosis is dependent on p53. P53 is well-known as a TSG. Induction of p53 has been reported to downregulate BCL-2 in breast cancer,19 and upregulate BAX in lung cancer.20 BAX is identified by its interaction with BCL-2, and it induces apoptotic cell death in response to cytokine deprivation.21, 22 BAX is repressed by BCL-2 either directly or indirectly through titrating BH3-only proteins, and the ratio of BCL-2/BAX constitutes a rheostat that sets the threshold of susceptibility to apoptosis for the intrinsic pathway.23 PMAIP1, a ‘BH3-only’ member of the Bcl-2 family, was shown to be a target of p53 and/or p73-mediated trans-activation; it first translocates to mitochondria and then functions through Bax and/or Bak to induce apoptosis.24, 25 Recent studies demonstrated that PMAIP1 could induce apoptosis of some cancer cells.25, 26, 27 In this study, we show that increased levels of p53 and its downstream targets, BAX and PMAIP1, are associated with overexpression of SOX30. Therefore, upregulation of p53 and its downstream targets by SOX30 could explain the effect of inducing cancer cell apoptosis with inhibiting proliferation. Blockade of p53 could reverse the effects of SOX30 overexpression, suggesting that p53 is a functionally important direct target gene of SOX30.
It has been reported that p53 can directly enhance the transcription of p21,28, 29 which is critical for the G1 checkpoint response because p21 loss compromises p53-mediated G1 arrest in response to DNA damage.30, 31 From our data, restoration of SOX30 increases p21 expression. However, it does not cause cell cycle arrest in G0/G1 phase (Supplementary Figure S2A). Further studies are required to solve this discrepancy.
To define whether p53 is required for the antitumor effects of SOX30, we blocked p53 activity or its signaling pathway using the inhibitor pifithrin-α when overexpressing SOX30. Blockade of p53 signaling pathway by pifithrin-α diminished the effect of SOX30 overexpression on cell proliferation and apoptosis. In general, pifithrin-α is considered to act endogenous p53-responsive genes. However, in this study, pifithrin-α could downregulate p53 mRNA. There may be a specific mechanism whereby pifithrin-α inhibits p53 transcriptional activity.32, 33, 34 The previous observations indicate that pifithrin-α effects are not limited to p53 and downstream components of p53 pathway but involve other cellular factors and signal transduction pathways.35, 36, 37 Thus, whether pifithrin-α binds directly to p53 or other factors effected by pifithrin-α regulate p53 transcriptional activity is still unknown, and further study is required to clarify this.
In conclusion, we have identified a novel functional TSG SOX30 inactivated by methylation. SOX30 has important roles in inducing cell apoptosis, acting through aberrant activation of p53 by directly binding to its promoter, as well as its downstream targets (Figure 8f). Importantly, p53 is a functionally key target gene of SOX30 as a TSG.
Materials and methods
Cell lines and patient samples
The lung cancer cell lines (A549, H460, H1395, SPC-A-1, H1975, 95D, H358, H1650 and LTEP) were obtained from the Cell Bank of the Chinese Academy of Science (Shanghai, China) and the American Type Culture Collection (ATCC, Manassas, VA, USA), cultured in RPMI-1640 (HyClone, Logan, UT, USA)/F12K (Sigma, St Louis, MO, USA) supplemented with 10% fetal bovine serum (HyClone), and incubated in 5% CO2 at 37 °C. A total of 120 lung cancer patients and 45 controls (including 25 matched tumor and adjacent non-tumor samples) were recruited from the Southwest Hospital in Chongqing, China. This study was approved by the ethics committee of the Southwest Hospital. Informed consent was signed by all of the recruited patients.
DNA and RNA extraction
DNA was extracted with a DNA Purification Wizard kit (Promega, Madison, WI, USA). Total RNAs were extracted using TRIzol Reagent (Invitrogen, Carlsbad, CA, USA), and were treated with DNase I to eliminate the genomic DNA contamination. Then, complementary DNAs were synthesized using M-MLV First Strand Kit (Invitrogen), and stored at −20 °C.
Screening methylated fragments by MS-AP-PCR
DNA from the samples (mixture from 10 normal lung tissues or 20 lung cancer tissues) was separately digested by restriction enzyme. PCR was used to amplify the restriction digest DNA using a single primer MLG2 or a combination of two primers MGE2 and MGF2 that arbitrarily binds within GC-rich regions of DNA. Amplified products were purified, subcloned into vectors and sequenced. Sequence data were used to determine genomic information, using the NCBI’s BLAST program (http://blast.ncbi.nlm.nih.gov).
Methylation analysis of SOX30 CpG islands by MSP and BGS
DNA samples were modified using the EZ DNA Methylation-Gold Kit (Zymo Research, Orange, CA, USA). The MSP and BGS were performed as previously reported.38 Primers are listed in Supplementary Table S1.
Mutation analysis
Five exons and promoter of SOX30 were amplified using genomic DNA with primers (Supplementary Table S1). The target fragments were purified, subcloned into vector and sequenced. Sequence homologies were analyzed using NCBI’s BLAST.
Analysis of SOX30 expression by RT–PCR and qRT–PCR
The analysis of SOX30 mRNA expression levels was performed by RT–PCR and qRT–PCR. A series of PCRs with different cycle numbers were performed to determine the linear phase of amplification for RT–PCR. Based on these pilot experiments, the appropriate cycles were chosen. The qRT–PCR was performed using an iQ5 real-time detection system (Bio-Rad Laboratories, Hercules, CA, USA) and SYBR Premix Ex Taq II (Takara, Dalian, China). The relative expression levels were calculated using 2−ΔΔct method. Primers are listed in Supplementary Table S1.
De-methylation treatment with 5-aza-2′-deoxycytidine
De-methylation experiments using 5-aza-2′-deoxycytidine were performed as previously described.38
Tissue microarray
All samples from lung cancer patients were reviewed histologically by hematoxylin and eosin staining, and two cores were taken from each representative tumor tissue and from lung tissue adjacent to the tumor within a distance of 10 μm to construct tissue microarray slides (in collaboration with the Shanghai Biochip Company Ltd, Shanghai, China). Duplicate cores from two different areas, intratumoral and peri-tumoral, were obtained. Then, tissue microarray sections with tumors and matched peri-tumoral samples were constructed.39
Immunohistochemistry and WB analysis
Immunohistochemical staining was performed using the antibody against SOX30 and Ki67 (1:100 or 1:50; Santa Cruz Biotechnology, Heidelberg, Germany) as described previously.40
Fifty micrograms of protein was run on 10–15% sodium dodecyl sulfate–polyacrylamide gel electrophoresis and transferred to polyvinylidene difluoride membrane (Millipore Corporation, Bedford, MA, USA). After blocking with 5% milk for 2 h at room temperature, membranes was incubated overnight at 4 °C with primary antibodies. After incubation with the secondary antibody, the proteins were detected by chemiluminescence (Pierce, Rockford, IL, USA). The same membrane was stripped and incubated with ACTIN monoclonal antibody (1:2000; Sigma), serving as an internal control. Primary antibodies were SOX30 rabbit polyclonal antibody (1:1000; Santa Cruz Biotechnology), p53 mouse polyclonal antibody (1:1000; Santa Cruz Biotechnology), BAX rabbit polyclonal antibody (1:1000; Santa Cruz Biotechnology), P21 rabbit polyclonal antibody (1:1000; Cell Signaling Technology, Boston, MA, USA) and PMAIP1 (NOXA) rabbit polyclonal antibody (1:1000; Santa Cruz Biotechnology). Secondary antibodies were horseradish peroxidase-conjugated donkey anti-rabbit and horseradish peroxidase-conjugated rabbit anti-mouse (1:3000, Jackson ImmunoResearch Laboratories, Inc., West Grove, PA, USA).
Construction of SOX30, miRNA expression vectors and cell transfection
The expression vector encoding full-length open reading frame of human SOX30 was constructed by synthesis and PCR amplification. Briefly, single-stranded oligonucleotides were designed and synthesized. The synthetic oligonucleotides were spliced into complete sequence by PCR, and validated by sequencing. Then, it was subcloned into the pIRES2-EGFP expression vector (Invitrogen Preservation, Carlsbad, CA, USA) and validated by sequencing. For knockdown, four pairs of oligomeric single-stranded oligonucleotides and a pair of negative oligomeric single-stranded oligonucleotides were synthesized, and then inserted into miRNA expression vector pcDNA6.2-GW/EmGFP-miR using the vector construction kit BLOCK-iT Pol II miR RNAi Expression Vector Kit with EmGFP (Invitrogen). The plasmids were transfected using ViaFect Transfection Reagent (Promega) or Lipofectamine2000 Reagent (Invitrogen). The stably transfected cells were screened under G418 (Calbiochem, La Jolla, CA, USA) or Blasticidin (Sigma). Cell clones were obtained by the cylinder method.
Colony formation assay
A549 and H460 cells were plated in 12-well plates at 2 × 105 cells per well. For knockdown, A549-SOX30 stable cells were plated in 12-well plates at 3 × 105 cells per well. After culturing for 24 h, cells were transfected with SOX30 or vector control and miRNA or negative control, respectively. After 48 h of transfection, cells were collected, diluted 1:5, plated in 12-well plates and selected with 0.4 mg/ml of G418 or 4 μg/ml of Blasticidin for 14 days to establish stable clones in which the plasmids had stably integrated into genomic DNA. Surviving colonies (⩾50 cells per colony) were stained by using Giemsa’s azur eosin methylene blue solution (Merck, Darmstadt, Germany) and counted. The experiment was carried out in triplicate wells for three times.
MTS and EdU assay
MTS and EdU assays were used to assess cell proliferation. A549 and H460 cells were plated at 3 × 103 cells per well on 96-well plates, and transfected with SOX30 or vector control. For knockdown, A549-SOX30 stable cells were plated at 3 × 103 cells per well on 96-well plates, and transfected with SOX30 miRNA or negative control. Cell proliferation was evaluated on days 1, 2, 3, 4 and 5 by determining the number of cells with Cell Proliferation Reagent MTS (Promega). The assay was carried out in triplicate for three independent experiments.
The effect of SOX30 suppression on cell proliferation was also tested by the EdU assay. Briefly, cells were cultured in 96-well plates and transfected with SOX30 or vector control for 46 h. Then, cells were incubated with 50 μm of EdU for additional 2 h at 37 °C. Cells were fixed with 4% formaldehyde for 30 min, incubated with glycine (2 mg/ml) for 5 min and treated with 0.5% Triton X-100 for 10 min to permeabilize cells. After being washed with phosphate-buffered saline, cells were incubated with Apollo reaction cocktail for 30 min and treated twice with 0.5% Triton X-100. DNA was stained with Hoechst 33342 stain for 30 min and visualized with fluorescence microscopy. Five groups of confluent cells were randomly selected from each sample image.
Flow cytometry assay
SOX30 or vector control-transfected cells (3 × 105 cells per well) were harvested at 48-h post-transfection, and fixed in 70% ethanol overnight at 4 °C. The cells were stained with propidium iodide (BD Pharmingen, San Jose, CA, USA). A total of 30 000 cells were sorted by FACSCalibur System (BD Biosciences, Franklin Lakes, NJ, USA) and cell cycle profiles were analyzed using the ModFit software (Verity Software House, Topsham, ME, USA). Apoptosis was also determined by dual staining with Annexin V-APC/7-amino-actinomycin D (KeyGEN, Nanjing, China). Briefly, cells were harvested and washed with phosphate-buffered saline twice, and were resuspended in 500 μl binding buffer, 5 μl Annexin V-APC were added and mixed gently. Then, 5 μl 7-ADD were added, mixed gently and incubated for 5–15 min in the dark and 50 000 cells were analyzed using the FACSCalibur System. The relative proportion of Annexin V-positive cells was determined using the ModFit software and counted as apoptotic cells. The assays were carried out in triplicate for three times.
DNA ladder assay
Cells at 48-h post-transfection were harvested and resuspended in lysis buffer for 30 min on ice. After centrifugation, the supernatant was treated with RNase (100 μg/ml) for 30 min at 55 °C and then with proteinase K (400 μg/ml) for another 1 h at 55 °C. The cell lysates were extracted with phenol–chloroform. DNA was precipitated with ethanol, and electrophoresed on 1.5% agarose gels.
Terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling assay
The apoptotic cells were detected in situ using the in situ cell death detection kit, POD (Roche, Penzberg, Germany). The terminal deoxynucleotidyl transferase-mediated dUTP nick end labeling method was performed according to the manufacturer's protocol. The sections after treatment were followed by incubation in diaminobenzidine substrate (Roche), and the reaction was observed under a microscope. Then, the nuclei were counterstained with hematoxylin buffer. The assay was carried out for three independent experiments.
In vivo tumorigenicity
To assess the effect of SOX30 expression on tumor formation in vivo, A549 and H460 cells with SOX30 or empty vector stably expression, for knockdown, A549-SOX30 stably cells with miRNA or negative control stably expression, were injected subcutaneously into the left and right flanks of 6-week-old male Balb/c nude mice, respectively. The tumor volume was calculated as 0.5236L1(L2)2, where L1 is the long axis and L2 is the short axis of the tumor.41 The developing tumors were observed over the next 4 to 8 weeks, and the mice were then killed. All experiments on mice were approved by the Institutional Animal Care and Use Committee of Third Military Medical University, China.
Expression array analysis
Expression profiles were generated using Agilent 4 × 44 K expression arrays (Agilent, Mississauga, ON, USA). Total RNA was extracted, checked and purified by RNeasy mini kit (QIAGEN, GmBH, Hilden, Germany). After RNA amplification and labeling, each slide was hybridized with 1.65 μg labeled sample. Slides were washed in staining dishes after 17-h hybridization, and scanned using an Agilent Microarray Scanner (Agilent Technologies, Santa Clara, CA, USA). Data were extracted with the Feature Extraction software 10.7 (Agilent Technologies) and normalized by Quantile algorithm, Gene Spring Software 11.0 (Agilent Technologies). Genes with fold changes>or <2.0 were considered to be of biological significance.
Site-directed mutagenesis of binding sites in p53 promoter and SOX30 HMG-box
The binding sites in the p53 promoter and SOX30 HMG-box constructs were mutated or deleted using a QuikChange Lightning Multi Site-Directed Mutagenesis Kit or a QuikChange Lightning Site-Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA), and the mutations or deletions were confirmed by DNA sequencing.
Luciferase reporter assay
The cells were plated in 24-well plates (3–5 × 104 cells per well) in triplicate for each condition. After overnight incubation, cells were transfected with a DNA mix contained pGL3-p53 promoter-luciferase, pIRES2-EGFP-SOX30 or empty vector, and pRL-TK plasmids. Luciferase activities were measured 36-h post-transfection using a Dual-luciferase reporter kit (Promega). Each experiment was performed in triplicate and repeated three times.
ChIP assay
ChIP analysis was performed using a ChIP Assay Kit (Cell Signaling Technology). The immunoprecipitated and input DNA was used as a template for RT–PCR analysis using the primers listed in Supplementary Table S1.
Statistical analysis
Statistical analyses were performed with the SPSS 16.0 software (SPSS, Inc., Chicago, IL, USA). The results are expressed as the mean±s.d. Differences between two groups were analyzed using the X2 test and the Student’s t-test. The Mann–Whitney U-test was performed to compare the pathological variables of two sample groups in functional assay. Clinical and pathologic characteristics of the patients were compared by Pearson X2 test or Fisher exact test. P-values <0.05 were taken as statistically significant.
References
Jemal A, Bray F, Center MM, Ferlay J, Ward E, Forman D . Global cancer statistics. CA Cancer J Clin 2011; 61: 69–90.
Ferlay J, Shin HR, Bray F, Forman D, Mathers C, Parkin DM . Estimates of worldwide burden of cancer in 2008: GLOBOCAN 2008. Int J Cancer 2010; 127: 2893–2917.
Jones PA, Baylin SB . The fundamental role of epigenetic events in cancer. Nat Rev Genet 2002; 3: 415–428.
Yang IV, Schwartz DA . Epigenetic control of gene expression in the lung. Am J Respir Crit Care Med 2011; 183: 1295–1301.
Minguez B, Lachenmayer A . Diagnostic and prognostic molecular markers in hepatocellular carcinoma. Dis Markers 2011; 31: 181–190.
Herman JG, Baylin SB . Gene silencing in cancer in association with promoter hypermethylation. New Engl J Med 2003; 349: 2042–2054.
Jones PA, Baylin SB . The epigenomics of cancer. Cell 2007; 128: 683–692.
Tsao CM, Yan MD, Shih YL, Yu PN, Kuo CC, Lin WC et al. SOX1 functions as a tumor suppressor by antagonizing the WNT/beta-catenin signaling pathway in hepatocellular carcinoma. Hepatology 2012; 56: 2277–2287.
Grady WM, Carethers JM . Genomic and epigenetic instability in colorectal cancer pathogenesis. Gastroenterology 2008; 135: 1079–1099.
Khare S, Verma M . Epigenetics of colon cancer. Methods Mol Biol 2012; 863: 177–185.
Zitt M, Zitt M, Muller HM . DNA methylation in colorectal cancer—impact on screening and therapy monitoring modalities? Dis Markers 2007; 23: 51–71.
Baylin SB, Jones PA . A decade of exploring the cancer epigenome- biological and translational implications. Nat Rev Cancer 2011; 11: 726–734.
Patai AV, Molnar B, Kalmar A, Scholler A, Toth K, Tulassay Z . Role of DNA methylation in colorectal carcinogenesis. Dig Dis 2012; 30: 310–315.
Gubbay J, Collignon J, Koopman P, Capel B, Economou A, Münsterberg A et al. A gene mapping to the sex-determining region of the mouse Y chromosome is a member of a novel family of embryonically expressed genes. Nature 1990; 346: 245–250.
Chew LJ, Gallo V . The Yin and Yang of Sox proteins: Activation and repression in development and disease. J Neurosci Res 2009; 87: 3277–3287.
Osaki E, Nishina Y, Inazawa J, Copeland NG, Gilbert DJ, Jenkins NA et al. Identification of a novel Sry-related gene and its germ cell-specific expression. Nucleic Acids Res 1999; 27: 2503–2510.
Han F, Wang Z, Wu F, Liu Z, Huang B, Wang D . Characterization, phylogeny, alternative splicing and expression of Sox30 gene. BMC Mol Biol 2010; 11: 98.
Ballow D, Meistrich ML, Matzuk M, Rajkovic A . Sohlh1 is essential for spermatogonial differentiation. Dev Biol 2006; 294: 161–167.
Haldar S, Negrini M, Monne M, Sabbioni S, Croce CM . Down-regulation of bcl-2 by p53 in breast cancer cells. Cancer Res 1994; 54: 2095–2097.
Miyashita T, Reed JC . Tumor suppressor p53 is a direct transcriptional activator of the human bax gene. Cell 1995; 80: 293–299.
Hockenbery DM, Zutter M, Hickey W, Nahm M, Korsmeyer SJ . BCL2 protein is topographically restricted in tissues characterized by apoptotic cell death. Proc Natl Acad Sci USA 1991; 88: 6961–6965.
Oltvai ZN, Milliman CL, Korsmeyer SJ . Bcl-2 heterodimerizes in vivo with a conserved homolog, Bax, that accelerates programmed cell death. Cell 1993; 74: 609–619.
Danial NN, Korsmeyer SJ . Cell death: critical control points. Cell 2004; 116: 205–219.
Oda E, Ohki R, Murasawa H, Nemoto J, Shibue T, Yamashita T et al. Noxa, a BH3-only member of the Bcl-2 family and candidate mediator of p53-induced apoptosis. Science 2000; 288: 1053–1058.
Hassan M, Alaoui A, Feyen O, Mirmohammadsadegh A, Essmann F, Tannapfel A et al. The BH3-only member Noxa causes apoptosis in melanoma cells by multiple pathways. Oncogene 2008; 27: 4557–4568.
Seo YW, Shin JN, Ko KH, Cha JH, Park JY, Lee BR et al. The molecular mechanism of Noxa-induced mitochondrial dysfunction in p53-mediated cell death. J Biol Chem 2003; 278: 48292–48299.
Suzuki S, Nakasato M, Shibue T, Koshima I, Taniguchi T . Therapeutic potential of proapoptotic molecule Noxa in the selective elimination of tumor cells. Cancer Sci 2009; 100: 759–769.
Harper JW, Adami GR, Wei N, Keyomarsi K, Elledge SJ . The p21 Cdk-interacting protein Cip1 is a potent inhibitor of G1 cyclin-dependent kinases. Cell 1993; 75: 805–816.
el-Deiry WS, Tokino T, Velculescu VE, Levy DB, Parsons R, Trent JM et al. WAF1, a potential mediator of p53 tumor suppression. Cell 1993; 75: 817–825.
Deng C, Zhang P, Harper JW, Elledge SJ, Leder P . Mice lacking p21CIP1/WAF1 undergo normal development, but are defective in G1 checkpoint control. Cell 1995; 82: 675–684.
Brugarolas J, Chandrasekaran C, Gordon JI, Beach D, Jacks T, Hannon GJ . Radiation-induced cell cycle arrest compromised by p21 deficiency. Nature 1995; 377: 552–557.
Culmsee C, Zhu X, Yu QS, Chan SL, Camandola S, Guo Z et al. A synthetic inhibitor of p53 protects neurons against death induced by ischemic and excitotoxic insults, and amyloid beta-peptide. J Neurochem 2001; 77: 220–228.
Plesnila N, von Baumgarten L, Retiounskaia M, Engel D, Ardeshiri A, Zimmermann R et al. Delayed neuronal death after brain trauma involves p53-dependent inhibition of NF-kappaB transcriptional activity. Cell Death Differ 2007; 14: 1529–1541.
Kim HR, Jung MH, Lee SY, Oh SM, Chung KH . Marijuana smoke condensate induces p53-mediated apoptosis in human lung epithelial cells. J Toxicol Sci 2013; 38: 337–347.
Gorgoulis VG, Zacharatos P, Kotsinas A, Kletsas D, Mariatos G, Zoumpourlis V et al. p53 activates ICAM-1 (CD54) expression in an NF-kappaB-independent manner. EMBO J 2003; 22: 1567–1578.
Cronauer MV, Schulz WA, Burchardt T, Ackermann R, Burchardt M . Inhibition of p53 function diminishes androgen receptor-mediated signaling in prostate cancer cell lines. Oncogene 2004; 23: 3541–3549.
Gudkov AV, Komarova EA . Prospective therapeutic applications of p53 inhibitors. Biochem Biophys Res Commun 2005; 331: 726–736.
Liu WB, Han F, Jiang X, Yang LJ, Li YH, Liu Y et al. ANKRD18A as a novel epigenetic regulation gene in lung cancer. Biochem Biophys Res Commun 2012; 429: 180–185.
Gao Q, Qiu SJ, Fan J, Zhou J, Wang XY, Xiao YS et al. Intratumoral balance of regulatory and cytotoxic T cells is associated with prognosis of hepatocellular carcinoma after resection. J Clin Oncol 2007; 25: 2586–2593.
Liu WB, Liu JY, Ao L, Zhou ZY, Zhou YH, Cui ZH et al. Epigenetic silencing of cell cycle regulatory genes during 3-methylcholanthrene and diethylnitrosamine-induced multistep rat lung cancer. Mol Carcinog 2010; 49: 556–565.
Zi X, Guo Y, Simoneau AR, Hope C, Xie J, Holcombe RF et al. Expression of Frzb/secreted Frizzled-related protein 3, a secreted Wnt antagonist, in human androgen-independent prostate cancer PC-3 cells suppresses tumor growth and cellular invasiveness. Cancer Res 2005; 65: 9762–9770.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (nos. 81172714 and 81071695).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Oncogene website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Han, F., Liu, W., Jiang, X. et al. SOX30, a novel epigenetic silenced tumor suppressor, promotes tumor cell apoptosis by transcriptional activating p53 in lung cancer. Oncogene 34, 4391–4402 (2015). https://doi.org/10.1038/onc.2014.370
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/onc.2014.370
- Springer Nature Limited