Abstract
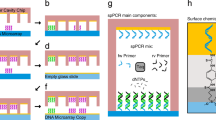
We describe a method, DNA array to protein array (DAPA), which allows the 'printing' of replicate protein arrays directly from a DNA array template using cell-free protein synthesis. At least 20 copies of a protein array can be obtained from a single DNA array. DAPA eliminates the need for separate protein expression, purification and spotting, and also overcomes the problem of long-term functional storage of surface-bound proteins.


Similar content being viewed by others
References
Bertone, P. & Snyder, M. FEBS J. 272, 5400–5411 (2005).
Wingren, C. & Borrebaeck, C.A. Expert Rev. Proteomics 1, 355–364 (2004).
Horn, S. et al. Proteomics 6, 605–613 (2006).
Wulfkuhle, J.D., Edmiston, K.H., Liotta, L.A. & Petricoin, E.F. III. Nat. Clin. Pract. Oncol. 3, 256–268 (2006).
Stoll, D., Templin, M.F., Bachmann, J. & Joos, T.O. Curr. Opin. Drug Discov. Dev. 8, 239–252 (2005).
He, M. & Taussig, M.J. Nucleic Acids Res. 29, e73 (2001).
Angenendt, P., Kreutzberger, J., Glokler, J. & Hoheisel, J.D. Mol. Cell. Proteomics 5, 1658–1666 (2006).
Nord, O., Uhlen, M. & Nygren, P.A. J. Biotechnol. 106, 1–13 (2003).
Ramachandran, N. et al. Science 305, 86–90 (2004).
Tao, S.C. & Zhu, H. Nat. Biotechnol. 24, 1253–1254 (2006).
He, M., Stoevesandt, O. & Taussig, M.J. Curr. Opin. Biotechnol. (in the press).
Khan, F., He, M. & Taussig, M.J. Anal. Chem. 78, 3072–3079 (2006).
MacBeath, G. & Schreiber, S.L. Science 289, 1760–1763 (2000).
Langlais, C. et al. BMC Biotechnol. 7, 64 (2007).
Murakami, H., Ohta, A., Ashigai, H. & Suga, H. Nat. Methods 3, 357–359 (2006).
Devirgiliis, C., Gaetani, S., Apreda, M. & Bellovino, D. Biochem. Biophys. Res. Commun. 332, 504–511 (2005).
Acknowledgements
We thank H. Liu for expert technical assistance, B. Korn (DKFZ, Heidelberg) and J. Taipele (University of Helsinki) for providing clones. Research at the Babraham Institute is supported by the Biotechnology and Biological Sciences Research Council. Work on this project has been supported through the European Commission 6th Framework Programme Integrated Project MolTools.
Author information
Authors and Affiliations
Contributions
M.H. and M.J.T. devised the principle of DAPA; M.H. developed and exemplified the method; O.S. optimized the method, devised the DAPA apparatus and designed and performed the experiments shown in Figures 1 and 2; E.A.P. performed initial DAPA experiments, contributing examples to the data shown in Supplementary Figures 3 and 4; F.K. characterized the double-6His tag for protein immobilization and protein detection methods; O.E. spotted the DNA template slides for experiments shown in Figure 1; M.H., O.S. and M.J.T. prepared the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4, Supplementary Methods (PDF 379 kb)
Rights and permissions
About this article
Cite this article
He, M., Stoevesandt, O., Palmer, E. et al. Printing protein arrays from DNA arrays. Nat Methods 5, 175–177 (2008). https://doi.org/10.1038/nmeth.1178
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1178
- Springer Nature America, Inc.
This article is cited by
-
Cell-free gene expression: an expanded repertoire of applications
Nature Reviews Genetics (2020)
-
How to copy and paste DNA microarrays
Scientific Reports (2019)
-
Immunoprofiling of Chlamydia trachomatis using whole-proteome microarrays generated by on-chip in situ expression
Scientific Reports (2018)
-
Using airbrushes to pattern reagents for microarrays and paper-fluidic devices
Microsystems & Nanoengineering (2017)
-
Personalised proteome analysis by means of protein microarrays made from individual patient samples
Scientific Reports (2017)