Abstract
T cell antigen receptor (TCR) signaling in the thymus initiates positive selection, but the CD8+-lineage fate is thought to be induced by cytokines after TCR signaling has ceased, although this remains controversial and unproven. We have identified four cytokines (IL-6, IFN-γ, TSLP and TGF-β) that did not signal via the common γ-chain (γc) receptor but that, like IL-7 and IL-15, induced expression of the lineage-specifying transcription factor Runx3d and signaled the generation of CD8+ T cells. Elimination of in vivo signaling by all six of these 'lineage-specifying cytokines' during positive selection eliminated Runx3d expression and completely abolished the generation of CD8+ single-positive thymocytes. Thus, this study proves that signaling during positive selection by lineage-specifying cytokines is responsible for all CD8+-lineage-fate 'decisions' in the thymus.







Similar content being viewed by others
References
Starr, T.K., Jameson, S.C. & Hogquist, K.A. Positive and negative selection of T cells. Annu. Rev. Immunol. 21, 139–176 (2003).
Carpenter, A.C. & Bosselut, R. Decision checkpoints in the thymus. Nat. Immunol. 11, 666–673 (2010).
Teh, H.S. et al. Thymic major histocompatibility complex antigens and the αβ T-cell receptor determine the CD4/CD8 phenotype of T cells. Nature 335, 229–233 (1988).
Brugnera, E. et al. Coreceptor reversal in the thymus: signaled CD4+8+ thymocytes initially terminate CD8 transcription even when differentiating into CD8+ T cells. Immunity 13, 59–71 (2000).
Singer, A. New perspectives on a developmental dilemma: the kinetic signaling model and the importance of signal duration for the CD4/CD8 lineage decision. Curr. Opin. Immunol. 14, 207–215 (2002).
Singer, A., Adoro, S. & Park, J.H. Lineage fate and intense debate: myths, models and mechanisms of CD4- versus CD8-lineage choice. Nat. Rev. Immunol. 8, 788–801 (2008).
Littman, D.R. How Thymocytes Achieve Their Fate. J. Immunol. 196, 1983–1984 (2016).
He, X. et al. The zinc finger transcription factor Th-POK regulates CD4 versus CD8 T-cell lineage commitment. Nature 433, 826–833 (2005).
Kappes, D.J., He, X. & He, X. Role of the transcription factor Th-POK in CD4:CD8 lineage commitment. Immunol. Rev. 209, 237–252 (2006).
Xiong, Y. & Bosselut, R. The enigma of CD4-lineage specification. Eur. J. Immunol. 41, 568–574 (2011).
Collins, A., Littman, D.R. & Taniuchi, I. RUNX proteins in transcription factor networks that regulate T-cell lineage choice. Nat. Rev. Immunol. 9, 106–115 (2009).
Taniuchi, I. et al. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell 111, 621–633 (2002).
Park, J.H. et al. Signaling by intrathymic cytokines, not T cell antigen receptors, specifies CD8 lineage choice and promotes the differentiation of cytotoxic-lineage T cells. Nat. Immunol. 11, 257–264 (2010).
Yu, Q., Erman, B., Bhandoola, A., Sharrow, S.O. & Singer, A. In vitro evidence that cytokine receptor signals are required for differentiation of double positive thymocytes into functionally mature CD8+ T cells. J. Exp. Med. 197, 475–487 (2003).
Kimura, M.Y. et al. Timing and duration of MHC I positive selection signals are adjusted in the thymus to prevent lineage errors. Nat. Immunol. 17, 1415–1423 (2016).
Xiong, Y. & Bosselut, R. CD4-CD8 differentiation in the thymus: connecting circuits and building memories. Curr. Opin. Immunol. 24, 139–145 (2012).
Kurd, N. & Robey, E.A. T-cell selection in the thymus: a spatial and temporal perspective. Immunol. Rev. 271, 114–126 (2016).
McCaughtry, T.M. et al. Conditional deletion of cytokine receptor chains reveals that IL-7 and IL-15 specify CD8 cytotoxic lineage fate in the thymus. J. Exp. Med. 209, 2263–2276 (2012).
Decaluwe, H. et al. γc deficiency precludes CD8+ T cell memory despite formation of potent T cell effectors. Proc. Natl. Acad. Sci. USA 107, 9311–9316 (2010).
Hanada, T. et al. A mutant form of JAB/SOCS1 augments the cytokine-induced JAK/STAT pathway by accelerating degradation of wild-type JAB/CIS family proteins through the SOCS-box. J. Biol. Chem. 276, 40746–40754 (2001).
Voon, D.C., Hor, Y.T. & Ito, Y. The RUNX complex: reaching beyond haematopoiesis into immunity. Immunology 146, 523–536 (2015).
Vaillant, F. et al. A full-length Cbfa1 gene product perturbs T-cell development and promotes lymphomagenesis in synergy with myc. Oncogene 18, 7124–7134 (1999).
Yoshimura, A., Naka, T. & Kubo, M. SOCS proteins, cytokine signalling and immune regulation. Nat. Rev. Immunol. 7, 454–465 (2007).
Lee, K.S. et al. Runx2 is a common target of transforming growth factor β1 and bone morphogenetic protein 2, and cooperation between Runx2 and Smad5 induces osteoblast-specific gene expression in the pluripotent mesenchymal precursor cell line C2C12. Mol. Cell. Biol. 20, 8783–8792 (2000).
El-Asady, R. et al. TGF-beta-dependent CD103 expression by CD8+ T cells promotes selective destruction of the host intestinal epithelium during graft-versus-host disease. J. Exp. Med. 201, 1647–1657 (2005).
Konkel, J.E. et al. Control of the development of CD8αα+ intestinal intraepithelial lymphocytes by TGF-β. Nat. Immunol. 12, 312–319 (2011).
Ouyang, W. et al. TGF-β cytokine signaling promotes CD8+ T cell development and low-affinity CD4+ T cell homeostasis by regulation of interleukin-7 receptor α expression. Immunity 39, 335–346 (2013).
Shi, M.J. & Stavnezer, J. CBFα3 (AML2) is induced by TGF-β1 to bind and activate the mouse germline Ig alpha promoter. J. Immunol. 161, 6751–6760 (1998).
Kulkarni, A.B. & Karlsson, S. Transforming growth factor-β1 knockout mice. A mutation in one cytokine gene causes a dramatic inflammatory disease. Am. J. Pathol. 143, 3–9 (1993).
Shull, M.M. et al. Targeted disruption of the mouse transforming growth factor-β1 gene results in multifocal inflammatory disease. Nature 359, 693–699 (1992).
Egawa, T., Tillman, R.E., Naoe, Y., Taniuchi, I. & Littman, D.R. The role of the Runx transcription factors in thymocyte differentiation and in homeostasis of naive T cells. J. Exp. Med. 204, 1945–1957 (2007).
Kohu, K. et al. Overexpression of the Runx3 transcription factor increases the proportion of mature thymocytes of the CD8 single-positive lineage. J. Immunol. 174, 2627–2636 (2005).
Grueter, B. et al. Runx3 regulates integrin alpha E/CD103 and CD4 expression during development of CD4-/CD8+ T cells. J. Immunol. 175, 1694–1705 (2005).
Wang, L. et al. Distinct functions for the transcription factors GATA-3 and ThPOK during intrathymic differentiation of CD4+ T cells. Nat. Immunol. 9, 1122–1130 (2008).
Hunter, C.A. & Jones, S.A. IL-6 as a keystone cytokine in health and disease. Nat. Immunol. 16, 448–457 (2015).
Schroder, K., Hertzog, P.J., Ravasi, T. & Hume, D.A. Interferon-γ: an overview of signals, mechanisms and functions. J. Leukoc. Biol. 75, 163–189 (2004).
Ziegler, S.F. et al. The biology of thymic stromal lymphopoietin (TSLP). Adv. Pharmacol. 66, 129–155 (2013).
Travis, M.A. & Sheppard, D. TGF-β activation and function in immunity. Annu. Rev. Immunol. 32, 51–82 (2014).
Rochman, Y. et al. Thymic stromal lymphopoietin-mediated STAT5 phosphorylation via kinases JAK1 and JAK2 reveals a key difference from IL-7-induced signaling. Proc. Natl. Acad. Sci. USA 107, 19455–19460 (2010).
Adoro, S. et al. Coreceptor gene imprinting governs thymocyte lineage fate. EMBO J. 31, 366–377 (2012).
Chen, W. et al. Conversion of peripheral CD4+CD25− naive T cells to CD4+CD25+ regulatory T cells by TGF-beta induction of transcription factor Foxp3. J. Exp. Med. 198, 1875–1886 (2003).
Lee, Y.J., Holzapfel, K.L., Zhu, J., Jameson, S.C. & Hogquist, K.A. Steady-state production of IL-4 modulates immunity in mouse strains and is determined by lineage diversity of iNKT cells. Nat. Immunol. 14, 1146–1154 (2013).
Pobezinsky, L.A. et al. Let-7 microRNAs target the lineage-specific transcription factor PLZF to regulate terminal NKT cell differentiation and effector function. Nat. Immunol. 16, 517–524 (2015).
Dewas, C. et al. TSLP expression: analysis with a ZsGreen TSLP reporter mouse. J. Immunol. 194, 1372–1380 (2015).
Takahama, Y., Letterio, J.J., Suzuki, H., Farr, A.G. & Singer, A. Early progression of thymocytes along the CD4/CD8 developmental pathway is regulated by a subset of thymic epithelial cells expressing transforming growth factor β. J. Exp. Med. 179, 1495–1506 (1994).
Fadok, V.A. et al. Macrophages that have ingested apoptotic cells in vitro inhibit proinflammatory cytokine production through autocrine/paracrine mechanisms involving TGF-β, PGE2, and PAF. J. Clin. Invest. 101, 890–898 (1998).
Catlett, I.M. & Hedrick, S.M. Suppressor of cytokine signaling 1 is required for the differentiation of CD4+ T cells. Nat. Immunol. 6, 715–721 (2005).
Fry, T.J. & Mackall, C.L. Interleukin-7: master regulator of peripheral T-cell homeostasis? Trends Immunol. 22, 564–571 (2001).
Chou, C. et al. The transcription factor AP4 mediates resolution of chronic viral infection through amplification of germinal center B cell responses. Immunity 45, 570–582 (2016).
Egawa, T. & Littman, D.R. Transcription factor AP4 modulates reversible and epigenetic silencing of the Cd4 gene. Proc. Natl. Acad. Sci. USA 108, 14873–14878 (2011).
Al-Shami, A. et al. A role for thymic stromal lymphopoietin in CD4+ T cell development. J. Exp. Med. 200, 159–168 (2004).
Larsson, J. et al. TGF-β signaling-deficient hematopoietic stem cells have normal self-renewal and regenerative ability in vivo despite increased proliferative capacity in vitro. Blood 102, 3129–3135 (2003).
Egawa, T. & Littman, D.R. ThPOK acts late in specification of the helper T cell lineage and suppresses Runx-mediated commitment to the cytotoxic T cell lineage. Nat. Immunol. 9, 1131–1139 (2008).
Acknowledgements
We thank T. McCaughtry for initial contributions to the project; J.-H. Park, V. Lazarevic, D. Singer, X. Zhou and M. Kimura for critical reading of the manuscript; M. Kubo (Tokyo University of Science) for SOCSTg mice; W. Leonard (National Heart, Lung and Blood Institute) for TSLPRKO mice; W. Chen (National Institute of Dental and Craniofacial Research) for Tgfbr1fl mice; D. Littman (New York University) for Runx3dYFP mice; R. Bosselut (National Cancer Institute) for ThPOKKO mice; and S. Sharrow, A. Adams and L. Granger for flow cytometry. Supported by the Intramural Research Program of the US National Institutes of Health, the National Cancer Institute, the Center for Cancer Research and the US National Institutes of Health (R01AI097244-01A1 to T.E.).
Author information
Authors and Affiliations
Contributions
R.E. designed the study, performed experiments, analyzed data and contributed to the writing of the manuscript; T.K., X.T., T.I.G. and T.E. performed experiments, analyzed data and provided helpful discussions; A.A. and B.E. generated experimental mice; and A.S. designed and supervised the study, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cytokine receptors on developing thymocytes that can potentially signal Runx3d expression.
(a) Characterization of γc-independent SP8 cells. Stainings for maturation markers (top) and CD8 lineage markers (bottom) are shown for B6 (black) and γccKO (red) SP8 (CD8SP TCRβhi) cells, compared to DP (CD4+ CD8+) cells (shaded curves). (b) List of cytokines, cytokine receptor chains, and signaling molecules examined in this study. (c) Identification of intermediate stage (CD4+CD8loCD69+) thymocytes that have been signaled by MHC class I-dependent ligands to undergo positive selection and to differentiate into SP8 thymocytes. MHC class I-signaled thymocytes were obtained from MHCIIKOCD1dKO mice because MHCI-signaled thymocytes in these mice do not differentiate into CD4+ NKT cells. (d) Surface expression of non-γc cytokine receptor chains on intermediate stage thymocytes from MHCIIKOCD1dKO mice. Cytokine receptor stainings are shown in red, control stainings are in gray. One of two experiments is shown. (e) Schematic of the DP stimulation assay. Pre-selection DP thymocytes (CD69-CD4+CD8+) were electronically sorted and stimulated with PMA+Ionomycin for 16h; washed and rested in medium for 10h; and then cultured with cytokines for 16-20h. Cells were then harvested and their gene expression analyzed by quantitative PCR. (f) List of cytokines that we tested in the B6 DP stimulation assay and their upregulation of mRNAs encoding Runx3d, Runx1, Runx2, Cbfb and Bcl2 (−, no gene up-regulation; +, minimal gene up-regulation; ++, intermediate gene up-regulation; +++, high gene up-regulation).
Supplementary Figure 2 Effect of cytokines on the generation of SP8 cells in fetal thymic organ culture.
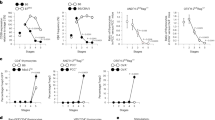
(a) Thymocyte profiles of E16.5 B6 thymic lobes before and after 5 days FTOC in medium (top and middle rows). Profile of TCRβhi thymocytes on d5 of FTOC (bottom row) and numbers in boxes within the profiles indicate frequency of cells within that box. (b) SP8 cell generation in γccKO FTOCs by non-γc cytokines. Bar graph of SP8 cell frequencies from γccKO FTOCs cultured with the indicated cytokines, with SP8 cell frequencies in each cytokine group compared to medium alone (horizontal red dashed line). As an additional comparison, SP8 cell frequencies from B6 FTOCs are also shown. Data are from 4-17 individual lobes combined from 2-6 experiments. (c) IL-13 fails to induce SP8 cell generation in FTOCs. Comparison of SP8 cell frequencies (left) and numbers (right) in γccKO FTOCs cultured for 5 days with or without IL-13. SP8 cells from B6 FTOC are shown for comparison (black bar). (d) Exogenous addition of any lineage-specifying cytokine increases SP8 cell generation in B6 FTOCs. Bar graph of SP8 cell frequencies (left) and numbers (right) in B6 FTOCs cultured for 5 days with the indicated cytokines compared to medium alone cultures. Mean and s.e.m. are shown. *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Supplementary Figure 3 Lymph node SP8 cells in CytoQuad mice.
Numbers of CD8 TCRβ+ LN T cells in the indicated mice. Mean and s.e.m. are shown.
Supplementary Figure 4 γc cytokines and non-γ c cytokines signal overlapping CD8+ T cell populations.
Onion diagram for schematic representation of cytokine redundancy. We think that, in normal mice, all SP8 cell generation is signaled by the γc cytokine IL-7. In the absence of IL-7 signaling, fewer SP8 cells are generated and these are signaled by the γc cytokine IL-15. In the absence of γc signaling, still fewer SP8 cells are generated and these are signaled by four non-γc cytokines (IL-6, IFN-γ, TSLP, and TGF-β). It is only in the absence of in vivo signaling by the other three non-γc cytokines (IL-6, IFN-γ, and TSLP) that the contribution of TGF-β signaling to SP8 cell generation is appreciated. Most importantly, elimination of in vivo signaling by all six of these cytokines eliminates SP8 cell generation.
Supplementary Figure 5 Analysis of STAT- and SMAD-binding sites in the Runx3d locus.
(a) Vista conservation analysis of murine and human Runx3d genomic regions. Peaks represent degree of conservation. Colored circles indicate potential binding sites for STATs (red) and SMADs (blue). (b) Comparison of surface CD103 expression on SP8 thymocytes from the indicated mice.
Supplementary Figure 6 Lineage-specifying transcription factors and the generation of SP8 cells.
(a) The Runx3dYFP/YFP DP stimulation assay (schematized in supplementary Fig. 1e) was performed with pre-selection DP thymocytes from Runx3d-deficient (Runx3dYFP/YFP) mice. After PMA+ionomycin stimulation, the Runx3dYFP/YFP DP thymocytes were stimulated with various cytokines and assessed for expression of Runx1 mRNA. (b) Generation of MHC-II specific SP8 cells in ThPOK-deficient mice requires cytokine signaling. Thymocyte profiles from the indicated mice. Numbers in boxes in the profiles indicate CD8SP cell frequencies. Mean and s.e.m. are shown.
Supplementary Figure 7 Signaling events in the thymus that generate SP8 thymocytes: all generation of SP8 cells requires signaling by lineage-specifying cytokines.
TCR signaling of DP thymocytes selectively terminates Cd8 gene expression, causing surface CD8 protein expression to decline. (a) In wildtype mice, MHC-I-specific TCR signaling eventually ceases because of the loss of surface CD8 protein expression, which allows cells to then be signaled by lineage-specifying cytokines. Signaling of positively selected thymocytes by lineage-specifying cytokines induces high Runx3d expression which both inhibits Runx1 and mediates co-receptor reversal (i.e. Cd4 gene silencing and Cd8 gene re-expression), resulting in differentiation into SP8 cells. (b) In the absence of intra-thymic signaling by lineage-specifying cytokines, Runx3d expression is not induced and Runx1 is unable to generate SP8 cells. (c) In Runx3d-deficient mice, intra-thymic signaling by lineage-specifying cytokines augments the ability of Runx1 to mediate coreceptor reversal and promote SP8 cell generation. (d) In ThPOK-deficient mice, CD4-lineage specification cannot occur. Because MHC-II-specific TCR signaling in the thymus must eventually cease, perhaps because of limiting ligand expression, cessation of TCR signaling allows the cells to then be signaled by lineage-specifying cytokines that induce high Runx3d expression which then inhibits Runx1 and mediates co-receptor reversal with the result that MHC-II-specific thymocytes differentiate into SP8 cells.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7. (PDF 1348 kb)
Rights and permissions
About this article
Cite this article
Etzensperger, R., Kadakia, T., Tai, X. et al. Identification of lineage-specifying cytokines that signal all CD8+-cytotoxic-lineage-fate 'decisions' in the thymus. Nat Immunol 18, 1218–1227 (2017). https://doi.org/10.1038/ni.3847
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3847
- Springer Nature America, Inc.
This article is cited by
-
How autoreactive thymocytes differentiate into regulatory versus effector CD4+ T cells after avoiding clonal deletion
Nature Immunology (2023)
-
Reversal of the T cell immune system reveals the molecular basis for T cell lineage fate determination in the thymus
Nature Immunology (2022)
-
The molecular basis and cellular effects of distinct CD103 expression on CD4 and CD8 T cells
Cellular and Molecular Life Sciences (2021)
-
Leaving no one behind: tracing every human thymocyte by single-cell RNA-sequencing
Seminars in Immunopathology (2021)
-
The order and logic of CD4 versus CD8 lineage choice and differentiation in mouse thymus
Nature Communications (2021)