Abstract
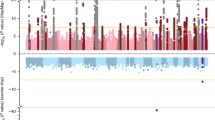
To newly identify loci for age at natural menopause, we carried out a meta-analysis of 22 genome-wide association studies (GWAS) in 38,968 women of European descent, with replication in up to 14,435 women. In addition to four known loci, we identified 13 loci newly associated with age at natural menopause (at P < 5 × 10−8). Candidate genes located at these newly associated loci include genes implicated in DNA repair (EXO1, HELQ, UIMC1, FAM175A, FANCI, TLK1, POLG and PRIM1) and immune function (IL11, NLRP11 and PRRC2A (also known as BAT2)). Gene-set enrichment pathway analyses using the full GWAS data set identified exoDNase, NF-κB signaling and mitochondrial dysfunction as biological processes related to timing of menopause.

Similar content being viewed by others
References
Burger, H.G. The menopausal transition. Baillieres Clin. Obstet. Gynaecol. 10, 347–359 (1996).
Broekmans, F.J., Soules, M.R. & Fauser, B.C. Ovarian aging: mechanisms and clinical consequences. Endocr. Rev. 30, 465–493 (2009).
te Velde, E.R. & Pearson, P.L. The variability of female reproductive ageing. Hum. Reprod. Update 8, 141–154 (2002).
te Velde, E.R., Dorland, M. & Broekmans, F.J. Age at menopause as a marker of reproductive ageing. Maturitas 30, 119–125 (1998).
Murabito, J.M., Yang, Q., Fox, C., Wilson, P.W. & Cupples, L.A. Heritability of age at natural menopause in the Framingham Heart Study. J. Clin. Endocrinol. Metab. 90, 3427–3430 (2005).
Snieder, H., MacGregor, A.J. & Spector, T.D. Genes control the cessation of a woman's reproductive life: a twin study of hysterectomy and age at menopause. J. Clin. Endocrinol. Metab. 83, 1875–1880 (1998).
van Asselt, K.M. et al. Heritability of menopausal age in mothers and daughters. Fertil. Steril. 82, 1348–1351 (2004).
Vink, J. & Boomsma, D.I. Modeling age at menopause. Fertil. Steril. 83, 1068 (2005).
He, C. et al. A large-scale candidate gene association study of age at menarche and age at natural menopause. Hum. Genet. 128, 515–527 (2010).
He, L.N. et al. Association study of the oestrogen signalling pathway genes in relation to age at natural menopause. J. Genet. 86, 269–276 (2007).
Weel, A.E. et al. Estrogen receptor polymorphism predicts the onset of natural and surgical menopause. J. Clin. Endocrinol. Metab. 84, 3146–3150 (1999).
Tempfer, C.B. et al. Polymorphisms associated with thrombophilia and vascular homeostasis and the timing of menarche and menopause in 728 white women. Menopause 12, 325–330 (2005).
van Asselt, K.M. et al. Factor V Leiden mutation accelerates the onset of natural menopause. Menopause 10, 477–481 (2003).
He, C. et al. Genome-wide association studies identify loci associated with age at menarche and age at natural menopause. Nat. Genet. 41, 724–728 (2009).
Stolk, L. et al. Loci at chromosomes 13, 19 and 20 influence age at natural menopause. Nat. Genet. 41, 645–647 (2009).
Kumar, P., Henikoff, S. & Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Pruim, R.J. et al. LocusZoom: regional visualization of genome-wide association scan results. Bioinformatics 26, 2336–2337 (2010).
Marini, F. & Wood, R.D. A human DNA helicase homologous to the DNA cross-link sensitivity protein Mus308. J. Biol. Chem. 277, 8716–8723 (2002).
Pullirsch, D. et al. The Trithorax group protein Ash2l and Saf-A are recruited to the inactive X chromosome at the onset of stable X X inactivation. Development 137, 935–943 (2010).
Copeland, W.C. & Longley, M.J. DNA polymerase gamma in mitochondrial DNA replication and repair. ScientificWorldJournal 3, 34–44 (2003).
Tschopp, J., Martinon, F. & Burns, K. NALPs: a novel protein family involved in inflammation. Nat. Rev. Mol. Cell Biol. 4, 95–104 (2003).
Segrè, A.V. et al. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genet. 6, pii, e1001058 (2010).
Raychaudhuri, S. et al. Identifying relationships among genomic disease regions: predicting genes at pathogenic SNP associations and rare deletions. PLoS Genet. 5, e1000534 (2009).
Bishop, D.K., Park, D., Xu, L. & Kleckner, N. DMC1: a meiosis-specific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell 69, 439–456 (1992).
Qin, Y. et al. NOBOX homeobox mutation causes premature ovarian failure. Am. J. Hum. Genet. 81, 576–581 (2007).
Rajkovic, A., Pangas, S.A., Ballow, D., Suzumori, N. & Matzuk, M.M. NOBOX deficiency disrupts early folliculogenesis and oocyte-specific gene expression. Science 305, 1157–1159 (2004).
Manolio, T.A. Genomewide association studies and assessment of the risk of disease. N. Engl. J. Med. 363, 166–176 (2010).
Johnson, F.B., Sinclair, D.A. & Guarente, L. Molecular biology of aging. Cell 96, 291–302 (1999).
Lee, C.K., Allison, D.B., Brand, J., Weindruch, R. & Prolla, T.A. Transcriptional profiles associated with aging and middle age-onset caloric restriction in mouse hearts. Proc. Natl. Acad. Sci. USA 99, 14988–14993 (2002).
Hamatani, T. et al. Age-associated alteration of gene expression patterns in mouse oocytes. Hum. Mol. Genet. 13, 2263–2278 (2004).
Promislow, D.E. DNA repair and the evolution of longevity: a critical analysis. J. Theor. Biol. 170, 291–300 (1994).
Du, X. et al. Telomere shortening exposes functions for the mouse Werner and Bloom syndrome genes. Mol. Cell. Biol. 24, 8437–8446 (2004).
Dorland, M., van Kooij, R.J. & te Velde, E.R. General ageing and ovarian ageing. Maturitas 30, 113–118 (1998).
Shiratori, A. et al. Assignment of the 49-kDa (PRIM1) and 58-kDa (PRIM2A and PRIM2B) subunit genes of the human DNA primase to chromosome bands 1q44 and 6p11.1-p12. Genomics 28, 350–353 (1995).
Pagnamenta, A.T. et al. Dominant inheritance of premature ovarian failure associated with mutant mitochondrial DNA polymerase gamma. Hum. Reprod. 21, 2467–2473 (2006).
Trifunovic, A. et al. Premature ageing in mice expressing defective mitochondrial DNA polymerase. Nature 429, 417–423 (2004).
Wong, J.C. & Buchwald, M. Disease model: Fanconi anemia. Trends Mol. Med. 8, 139–142 (2002).
Nebel, A. et al. A functional EXO1 promoter variant is associated with prolonged life expectancy in centenarians. Mech. Ageing Dev. 130, 691–699 (2009).
Wei, K. et al. Inactivation of exonuclease 1 in mice results in DNA mismatch repair defects, increased cancer susceptibility, and male and female sterility. Genes Dev. 17, 603–614 (2003).
Tye, B.K. Insights into DNA replication from the third domain of life. Proc. Natl. Acad. Sci. USA 97, 2399–2401 (2000).
Epstein, C.J., Martin, G.M., Schultz, A.L. & Motulsky, A.G. Werner's syndrome a review of its symptomatology, natural history, pathologic features, genetics and relationship to the natural aging process. Medicine (Baltimore) 45, 177–221 (1966).
Lee, S.J., Lee, S.H., Ha, N.C. & Park, B.J. Estrogen prevents senescence through induction of WRN, Werner syndrome protein. Horm. Res. Paediatr. 74, 33–40 (2010).
Gao, Y. et al. DNA repair gene polymorphisms and tobacco smoking in the risk for colorectal adenomas. Carcinogenesis 32, 882–887 (2011).
Welt, C.K. Primary ovarian insufficiency: a more accurate term for premature ovarian failure. Clin. Endocrinol. 68, 499–509 (2008).
Hoek, A., Schoemaker, J. & Drexhage, H.A. Premature ovarian failure and ovarian autoimmunity. Endocr. Rev. 18, 107–134 (1997).
Robb, L. et al. Infertility in female mice lacking the receptor for interleukin 11 is due to a defective uterine response to implantation. Nat. Med. 4, 303–308 (1998).
Northup, J., Griffis, K., Hawkins, J., Lockhart, L. & Velagaleti, G. Unusual pseudo dicentric, psu dic (1;19)(q10;q13.42), in a female with premature ovarian failure. Fertil. Steril. 87 697 e5–e8 (2007).
Tong, Z.B. & Nelson, L.M. A mouse gene encoding an oocyte antigen associated with autoimmune premature ovarian failure. Endocrinology 140, 3720–3726 (1999).
Anasti, J.N. et al. Karyotypically normal spontaneous premature ovarian failure: evaluation of association with the class II major histocompatibility complex. J. Clin. Endocrinol. Metab. 78, 722–723 (1994).
Jaroudi, K.A., Arora, M., Sheth, K.V., Sieck, U.V. & Willemsen, W.N. Human leukocyte antigen typing and associated abnormalities in premature ovarian failure. Hum. Reprod. 9, 2006–2009 (1994).
Gasser, S. & Raulet, D.H. The DNA damage response arouses the immune system. Cancer Res. 66, 3959–3962 (2006).
Kumar, T.R., Wang, Y., Lu, N. & Matzuk, M.M. Follicle stimulating hormone is required for ovarian follicle maturation but not male fertility. Nat. Genet. 15, 201–204 (1997).
McTavish, K.J. et al. Rising follicle-stimulating hormone levels with age accelerate female reproductive failure. Endocrinology 148, 4432–4439 (2007).
Lambalk, C.B., De Koning, C.H. & Braat, D.D. The endocrinology of dizygotic twinning in the human. Mol. Cell. Endocrinol. 145, 97–102 (1998).
Kottler, M.L. et al. A new FSHβ mutation in a 29-year-old woman with primary amenorrhea and isolated FSH deficiency: functional characterization and ovarian response to human recombinant FSH. Eur. J. Endocrinol. 162, 633–641 (2010).
Grigorova, M. et al. Increased prevalance of the -211 T allele of follicle stimulating hormone (FSH) β subunit promoter polymorphism and lower serum FSH in infertile men. J. Clin. Endocrinol. Metab. 95, 100–108 (2010).
Hoogendoorn, B. et al. Functional analysis of human promoter polymorphisms. Hum. Mol. Genet. 12, 2249–2254 (2003).
Kaku, U. et al. Ovarian histological findings in an adult patient with the steroidogenic acute regulatory protein (StAR) deficiency reveal the impairment of steroidogenesis by lipoid deposition. Endocr. J. 55, 1043–1049 (2008).
Pisarska, M.D., Bae, J., Klein, C. & Hsueh, A.J. Forkhead l2 is expressed in the ovary and represses the promoter activity of the steroidogenic acute regulatory gene. Endocrinology 145, 3424–3433 (2004).
Godinho, M., Meijer, D., Setyono-Han, B., Dorssers, L.C. & Agthoven, T.V. Characterization of BCAR4, a novel oncogene causing endocrine resistance in human breast cancer cells. J. Cell. Physiol. 226, 1741–1749 (2011).
Willer, C.J., Li, Y. & Abecasis, G.R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Göring, H.H. et al. Discovery of expression QTLs using large-scale transcriptional profiling in human lymphocytes. Nat. Genet. 39, 1208–1216 (2007).
Idaghdour, Y. et al. Geographical genomics of human leukocyte gene expression variation in southern Morocco. Nat. Genet. 42, 62–67 (2010).
Heap, G.A. et al. Complex nature of SNP genotype effects on gene expression in primary human leucocytes. BMC Med. Genomics 2, 1 (2009).
Dixon, A.L. et al. A genome-wide association study of global gene expression. Nat. Genet. 39, 1202–1207 (2007).
Stranger, B.E. et al. Population genomics of human gene expression. Nat. Genet. 39, 1217–1224 (2007).
Kwan, T. et al. Genome-wide analysis of transcript isoform variation in humans. Nat. Genet. 40, 225–231 (2008).
Dimas, A.S. et al. Common regulatory variation impacts gene expression in a cell type-dependent manner. Science 325, 1246–1250 (2009).
Heinzen, E.L. et al. Tissue-specific genetic control of splicing: implications for the study of complex traits. PLoS Biol. 6, e1 (2008).
Zeller, T. et al. Genetics and beyond–the transcriptome of human monocytes and disease susceptibility. PLoS ONE 5, e10693 (2010).
Murphy, A. et al. Mapping of numerous disease-associated expression polymorphisms in primary peripheral blood CD4+ lymphocytes. Hum. Mol. Genet. 19, 4745–4757 (2010).
Emilsson, V. et al. Genetics of gene expression and its effect on disease. Nature 452, 423–428 (2008).
Webster, J.A. et al. Genetic control of human brain transcript expression in Alzheimer disease. Am. J. Hum. Genet. 84, 445–458 (2009).
Gibbs, J.R. et al. Abundant quantitative trait loci exist for DNA methylation and gene expression in human brain. PLoS Genet. 6, e1000952 (2010).
Liu, C. et al. Whole-genome association mapping of gene expression in the human prefrontal cortex. Mol. Psychiatry 15, 779–784 (2010).
Schadt, E.E. et al. Mapping the genetic architecture of gene expression in human liver. PLoS Biol. 6, e107 (2008).
Grundberg, E. et al. Population genomics in a disease targeted primary cell model. Genome Res. 19, 1942–1952 (2009).
Dabney, A., Storey, J.D. & Warnes, G.R. qvalue: Q-value estimation for false discovery rate control. R package version 1.22.0. (2010).
Li, J. & Ji, L. Adjusting multiple testing in multilocus analyses using the eigenvalues of a correlation matrix. Heredity 95, 221–227 (2005).
Acknowledgements
We are grateful to the study participants and staff from all cohorts involved in this study. Extended acknowledgments per cohort are in Supplementary Note.
Author information
Authors and Affiliations
Contributions
Individual study design and management were done by D.I.C., C. He, E.M.B., P.K., J.S.B., M. Boban, E.B., H.C., S.J.C., U.d.F., I.J.D., G.V.Z.D., S.E., J.G.E., L. Ferrucci, A.R.F., P.G., C.J.M.W.G., C.G., D.E.G., P.H., S.E.H., A.H., E.I., S.L.R.K., D.A.L., P.K.E.M., M. Marongiu, N.G.M., V.M., N.C.O.-M., G. Paré, A.N.P., N.L.P., P.H.M.P., O.P., B.M.P., K.R., I.R., J.M.S., R.P.S., V.J.M.P., V.G., G.E., T.B.H., L.J.L.,Y.V.S., A.T., Y.T.v.d.S., P.M.V., G. Waeber, H.E.W., J.F.W., B.H.R.W., L.M.L., D.I.B., J.E.B., L.C., F.B.H., D.J.H., G.W.M., B.A.O., P.M.R., D.S., T.D.S., K. Stefansson, E.A.S., M.U., C.M.v.D., H.V., S.B., A. Salumets and A. Metspalu. Data collection was done by B.Z.A., S.B., L.B.L., J.S.B., M. Boban, A.B., H.C., P.dA., U.dF., I.J.D., G.V.Z.D., S.E., J.G.E., L. Ferreli, L. Ferrucci, A.R.F., D.E.G., P.H., S.E.H., A.C.H., A.H., A.C.J.W.J., I.K., S.L., D.A.L., P.K.E.M., N.G.M., I.M.K., A.B.N., N.L.P., P.H.M.P., O.P., B.M.P., K.R., I.R., A. Scuteri, S.N.S., J.M.S., M.G.S., U.S., B.T., L.T., S.U., V.J.M.P., V.G., G.E., Y.T.v.d.S., C.H.v.G., J.M.V., G. Waeber, H.E.W., A.F.W., T.Z., L.Z., M.C.Z., M.Z., L.M.L., A.M.A., J.E.B., E.A.S., A.G.U., J.M.M., P.v.N. and A. Metspalu. Genotyping was done by J.K., L.S., T.E., E.B., F.J.B., S.J.C., P.dA., G.D., P.D., C. Hayward, A.C.H., J. Liu, B.C.J.M.F., S.E.M., V.M., P.N., D.R.N., N.C.O.-M., A.S.P., A.N.P., B.M.P., J.I.R., A. Singleton, K. Stirrups, A.T., K.T., Y.T.v.d.S., M.V., E.W., T.Z., L.M.L., F.B.H., G.W.M., B.A.O. and U.T. Genotype preparation was done by J.R.B.P., M. Barbalic, N.F., E.P., S.-Y.S., W.V.Z., L.B.L., P.dA., G.D., C.J.M.W.G., C.G., C. Hayward, E.I., S.L.R.K., P.K.E.M., I.M.K., C.M., S.E.M., P.N., D.R.N., N.C.O.-M., A.P., G. Pistis, E.R., C.S., J.A.S., H.S., N.S., Y.V.S., A.T., M.V., E.W., T.Z., L.M.L., F.B.H., D.J.H., S.S., K.L.L. and M.H. Phenotype preparation was done by J.R.B.P., L.S., C. He, M. Mangino, M. Barbalic, L.B., E.M.B., F.E., N.F., D.F.G., J.-J.H., P.K., G.Z., W.V.Z., L.B.L., M. Boban, E.J.C.d.G., I.J.D., G.V.Z.D., M.G.S., S.E., J.G.E., C.H.v.G., L. Ferreli, K.F., M.H., C. Hayward, E.I., A.C.J.W.J., S.L.R.K., I.K., J. Lahti, S.L., T.L., D.A.L., L.M.L., P.K.E.M., I.M.K., C.M., P.H.M.P., G. Pistis, O.P., E.R., C.S., A. Scuteri, J.A.S., B.T., S.U., R.M.v.D., V.G., G.E., T.A., Y.T.v.d.S., H.W., G. Willemsen, B.H.R.W., A.F.W., M.C.Z., L.M.L., A.M.A., E.W.D., A. Metspalu, K.L.L. and J.M.M. Analysis plan development was done by L.S., D.I.C., C. He, E.M.B., P.K., T.C., P.G., D.K., D.P.K., N.G.M., D.T., D.J.H., G.W.M., K.L.L., J.M.M. and A. Murray. Analysis plan review was done by L.S., D.I.C., C. He, A.V.S., A.D.C., N.G., D.K., D.P.K., B.M., A.B.N., B.M.P., A.M.A., D.J.H., K.L.L., J.M.M. and A. Murray. Study data analysis was done by L.S., D.I.C., C. He, M. Mangino, M. Barbalic, L.B., E.M.B., F.E., T.E., N.F., D.F.G., J.-J.H., P.K., P.F.M., E.P., S.-Y.S., A.V.S., S.v.W., G.Z., W.V.Z., E.A., C.C., M.C.C., T.C., N.G., T.H., C. Hayward, Z.K., J. Lahti, D.A.L., L.M.L., D.M., N.C.O.M., G. Paré, G. Pistis, A.S.P., V.E., E.R., J.A.S., M.G.S., S.U., Y.T.vdS., M.V., L.M.Y.-A., L.Z., S.S. and U.T. Review and interpretation of analyses were done by J.R.B.P., L.S., D.I.C., C. He, P.S., M. Barbalic, E.M.B., N.F., P.K., P.F.M., A.V.S., B.Z.A., S.J.C., A.D.C., I.J.D., S.E.H., E.I., D.K., S.L.R.K., D.P.K., L.M.L., P.K.E.M., N.G.M., G. Paré, A.S.P., J.A.S., H.S., L.M.Y.-A., J.E.B., E.W.D., F.B.H., D.J.H., G.W.M., P.M.R., C.M.v.D., H.V., K.L.L., J.M.M., P.v.N. and A. Murray. Meta-analyses were done by L.S., P.S. and K.L.L. Pathway and other analyses were done by J.R.B.P., D.I.C., C. He, A.D.J., M. Mangino, G. Paré and J.A.V. Menopause study was designed by J.R.B.P., L.S., D.I.C., C. He, D.E.G., P.H.M.P., Y.T.v.d.S., C.H.v.G., A. Metspalu, K.L.L., J.M.M. and A. Murray. Manuscript was prepared by J.R.B.P., L.S., D.I.C., C. He, T.E., N.F., A.D.C., D.K., D.P.K., E.W.D., A. Metspalu, K.L.L., J.M.M., A. Murray and J.A.V. Manuscript was reviewed by J.R.B.P., L.S., D.I.C., C. He, N.F., P.K., P.F.M., A.V.S., B.Z.A., T.A., L.B.L., J.S.B., M. Boban, F.J.B., H.C., S.J.C., C.C., M.C.C., A.D.C., G.D., U.d.F., I.J.D., G.E., B.C.J.M.F., M.E.G., N.G., D.E.G., P.H., S.E.H., C. Hayward, E.I., D.K., D.P.K., S.L.R.K., J.A.S., I.K., Z.K., T.L., T.E., A. Salumets, A. Metspalu, J.S.E.L., J. Liu, L.M.L., Y.V.L., P.K.E.M., I.M.K., B.M., V.M., P.N., A.B.N., N.C.O.-M., G. Paré, A.N.P., N.L.P., P.H.M.P., O.P., B.M.P., J.I.R., I.R., H.S., J.M.S., R.P.S., A.T., R.M.v.D., Y.T.v.d.S., C.H.v.G., P.M.V., M.V., G. Waeber, J.F.W., B.H.R.W., A.F.W., L.M.Y.-A., T.Z., L.Z., M.C.Z., V.J.M.P., L.M.L., A.M.A., D.I.B., J.E.B., E.W.D., V.G., T.B.H., F.B.H., D.J.H., L.J.L., P.M.R., T.D.S., E.A.S., H.V., K.L.L., J.M.M., A. Murray, P.v.N. and J.A.V. Consortium was overseen by K.L.L., J.M.M., A. Murray and J.A.V.
Corresponding authors
Ethics declarations
Competing interests
V.M. was a full-time employee of GlaxoSmithKline when this work was done. B.C.J.M.F. has received fees and grant support from the following companies (in alphabetical order): Andromed, Ardana, Ferring, Genovum, Merck Serono, MSD, Organon, Pantharei Bioscience, PregLem, Schering, Schering Plough, Serono and Wyeth. F.J.B. is a member of the external advisory board for Merck Serono, does consultancy work for MSD and carries out educational activities for Ferring. B.M.P. discloses (i) service on a data and safety monitoring board for a clinical trial of a device funded by the manufacturer and (ii) service on the steering committee for the Yale Open-Data Project, which is funded by Medtronic for the review of clinical trial data on recombinant morphogenic protein 2.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–7 and Supplementary Note (PDF 640 kb)
Rights and permissions
About this article
Cite this article
Stolk, L., Perry, J., Chasman, D. et al. Meta-analyses identify 13 loci associated with age at menopause and highlight DNA repair and immune pathways. Nat Genet 44, 260–268 (2012). https://doi.org/10.1038/ng.1051
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.1051
- Springer Nature America, Inc.
This article is cited by
-
Micronutrients intake and genetic variants associated with premature ovarian insufficiency; MASHAD cohort study
BMC Women's Health (2024)
-
Mechanisms of ovarian aging in women: a review
Journal of Ovarian Research (2023)
-
Premature ovarian insufficiency is associated with global alterations in the regulatory landscape and gene expression in balanced X-autosome translocations
Epigenetics & Chromatin (2023)
-
Landscape of pathogenic mutations in premature ovarian insufficiency
Nature Medicine (2023)
-
mTOR pathway candidate genes and energy intake interaction on breast cancer risk in Black women from the Women’s Circle of Health Study
European Journal of Nutrition (2023)