Abstract
Voltage-gated sodium channels (Nav) underlie the rapid upstroke of action potentials in excitable tissues. Binding of channel-interactive proteins is essential for controlling fast and long-term inactivation. In the structure of the complex of the carboxy-terminal portion of Nav1.5 (CTNav1.5) with calmodulin (CaM)–Mg2+ reported here, both CaM lobes interact with the CTNav1.5. On the basis of the differences between this structure and that of an inactivated complex, we propose that the structure reported here represents a non-inactivated state of the CTNav, that is, the state that is poised for activation. Electrophysiological characterization of mutants further supports the importance of the interactions identified in the structure. Isothermal titration calorimetry experiments show that CaM binds to CTNav1.5 with high affinity. The results of this study provide unique insights into the physiological activation and the pathophysiology of Nav channels.
Similar content being viewed by others
Introduction
Voltage-gated sodium channels (Nav) are transmembrane glycoproteins that underlie the rapid upstroke of action potentials in excitable tissues such as the heart, skeletal muscle and brain. Mendelian-inherited mutations in Nav channels result in diseases of excitability such as myotonias, paralyses, cardiac arrhythmias and ataxias and seizure disorders1. Na channels consist of an α subunit and one or more β subunits, but only the pore-forming α subunit is essential for function. Ten different isoforms of mammalian α subunits (Nav1.1–1.9, and Nax) have been described, with different properties and tissue distribution.
The transmembrane portion of the α subunit is formed by four homologous domains (DI–DIV), each containing six membrane-spanning helices (S1–S6) that form the ion-selective pore and contain the activation voltage sensors. Channel activation and opening is followed by prompt closure via a number of kinetically distinct inactivation states2. Fast inactivation (recovery τ<10 ms), the best characterized of these processes, involves occlusion of the cytoplasmic mouth of the channel by the interdomain DIII–DIV linker (DIII/IV). A triplet of hydrophobic residues of the linker, Ile-Phe-Met, is key for this inactivation3.
NaV channels are regulated by the interaction of their carboxyl terminal (CT) domain, located in the cytoplasm of responsive cells, with various channel-interactive proteins4,5. The importance of these interactions is highlighted by the effects of mutations in the Nav CT domain (CTNav) on channel function: gain-of-function mutations of the NaV1.5 CT domain (CTNav1.5) cause long QT syndrome and loss-of-function mutations result in Brugada syndrome.
The proximal portion of the CTNav (residues 1,776–1,929 in Nav1.5) is comprised of six α-helices (αI–αVI)6,7. The first four helices αI–αIV, form an EF-hand-like motif (EFL) that has a fold similar to that of a the Ca2+-binding EF hand8,9,10. The fifth helix (αV) and a flexible loop connect the EFL to a long sixth helix (αVI), which contains an IQ motif that binds calmodulin (CaM)7.
Several structural studies have explored the interaction of CaM with different regions of Nav channels. One structure, the complex of the C-lobe of CaM with the DIII/IV linker of NaV channels, suggests that CaM modulates fast inactivation by forming a bridge between the CTNav IQ motif and the DIII/IV linker of the channel11,12. In the structure of the ternary complex containing CTNaV1.5, apo-CaM and a fibroblast growth factor (FGF) homologous factor (FHF)—a long-term inactivator of Nav channels—FHF binds to the EFL of Nav1.5 and the C-lobe of CaM binds to the IQ motif7. Together, these structural data point to complex dynamic interactions among the participating components in regulating channel gating.
Despite the availability of biochemical, electrophysiological, biophysical and structural information, the participation of the CTNav in the regulation of NaV channels is still a matter of debate and may be isoform-specific8,13,14. Given the central importance of NaV1 channels, it is surprising that important details of the molecular mechanisms leading to their activation, inactivation and recovery from inactivation remain unknown. Missing, for example, are structures involving the cytoplasmic domain of Nav1.5 when the channel is poised for opening.
Here we present the structure of the complex of the C-terminal domain of the Nav1.5 channel with CaM–Mg2+, which we propose, represents the resting state of the cytoplasmic region of the channel after the recovery from inactivation, that is, the state in which the channel is poised for activation. We will refer to this state as ‘non-inactivated’ or resting. Site-specific mutations at the sites of interactions identified in the structure presented here alter the inactivation properties of Nav1.5, as determined by electrophysiological recordings. Calorimetric measurements show that CTNav1.5 binds tightly to full-length CaM, while the DIII–IV linker peptide cannot compete CTNav1.5 from the CTNav1.5–CaM complex. The results of this study provide unique insights into the physiological activation and the pathophysiology of Nav channels15.
Results
Overall structure of the CTNav1.5–CaM complex
The crystal of the CTNav1.5–CaM–Mg2+ complex (see below) determined to 2.8 Å contains five highly similar heterodimers in the asymmetric unit (ASU) (average r.m.s.d. 1.6 Å), each composed of a full-length CaM (residues 1–148) and one CTNav1.5 (residues 1,776–1,929) (Table 1; Fig. 1; Supplementary Table 1).
(a) CTNav1.5–CaM complex with CTNav1.5 in lime-green and CaM in yellow. The helices of CTNav1.5 are labelled αI–VI and CaM helices αA–G. (b) The five CTNav1.5–CaM complexes in the asymmetric unit. The CaM molecules are labelled A, B, C, D, and E; the CTNav1.5 are labelled F, G, H, I, J. For example, AF is the heterodimer with CaM molecule A and CTNav1.5 molecule F.
In each heterodimer the CTNav1.5 has a bipartite structure resembling a lollipop (Fig. 1a) in which the globular EFL domain (residues 1,776–1,866), the head, is connected via helix αV (residues 1,866–1,884) and a loop (residues 1,885–1,894) to the long stalk (helix αVI; residues 1,895–1,929) (Fig. 1a). The structure of the EFL domain (helices αI–αIV; Fig. 1) is similar to the nuclear magnetic resonance structures of the EFL domains of Nav1.5 (residues 1,773–1,865, protein data bank (PDB) id 2KBI, Supplementary Table 2)16 and Nav1.2 (PDB id 2KAV)17.
CaM conformation in the Nav1.5–CaM–Mg2+ complex
The CaM N-lobe (residues 1–77; helices αA–αD; Supplementary Fig. 1) adopts a ‘closed’ conformation and the C-lobe (residues 82–148; helices αE–αH) a ‘semi-open’ conformation (Supplementary Fig. 1; Supplementary Fig. 2)18,19,20. A short linker (residues 78–81) connects the two lobes in such a way that the long axis of helix αD of the N-lobe (residues 65–77) is orthogonal to the αE helix of the C-lobe (residues 82–93) forming a ‘lock-washer’ type arrangement around helix αVI of the CTNav1.5 (Fig. 1a). This conformation of CaM is highly different from the one observed in the structure of the CTNav1.5–CaM-FGF13 (ref. 7) complex (PDB id 4DCK; FGF13=FHF2) in which helix αD, the linker and helix αE form a long nine-turn helix (residues 64–95) that places the CaM N-lobe at a significant distance from the CTNav1.5.
Ion binding in the Nav1.5–CaM–Mg2+complex
Both CaM lobes of all five CaMs in the ASU contain bound ions. The lack of anomalous scattering signal at Cu Kα, a wavelength at which Ca2+ and Mn2+ have significant anomalous signal, and the observed coordination suggest that the ions are Mg2+ from the crystallization solution19 (Supplementary Fig. 1). Significantly, the conformation of the N-lobe of CaM in the Nav1.5–CaM–Mg2+ complex reported here is highly similar to that of N-lobe of apo-CaM (PDB id 1DMO18; Supplementary Fig. 1) as well as to the N-lobe of the CaM–Mg2+ complex (PDB id 3UCW; Supplementary Table 3 (ref. 19)). In contrast, it is significantly different from that of the Ca2+-loaded CaM (PDB id 1CDM21). The C-lobe of CaM in the Nav1.5–CaM–Mg2+ complex has the semi-open conformation observed in CaM structures such as that of the Nav1.2-IQ–CaM complex, in which the cation-binding sites are not occupied by Ca2+ (PDB id 2KXW; PDB id 3WFN, Supplementary Fig. 2)22,23.
No ions are bound to the CTNav1.5 EFL domain in the CTNav1.5–CaM–Mg2+ complex. This domain does not show density at the expected ion positions (‘Ca+2-binding loops’; residues 1,802–1,812 and residues 1,838–1,850), and crucial acidic residues such as Glu1804 and Glu1846 are not in the conformations required for Ca+2 binding (Supplementary Fig. 2).
CTNav1.5–CaM interaction
In the complex reported here, CaM wraps around the long helix of Nav1.5 in such a way that the interface buries ~1,400 Å2 (Supplementary Table 4), ~950 Å2 between the C-lobe of CaM and the CTNav1.5 IQ and ~450 Å2 between of the N-lobe and the Nav1.5 EFL domain. The CaM–CTNaV1.5 interface is mostly hydrophobic (29 EFL hydrophobic interactions; Fig. 2a,b), but it is flanked at both ends by hydrogen bonds between the two proteins (7 H-bonds: Lys1899–Glu12, and Asn1831–Glu15, at one end and Arg1910–Asp81, Arg1913–Glu121, Arg1913–Glu115, Arg1919–Glu124 and Glu1901–Lys95 at the other; Fig. 2c).
(a) Overview of the heterodimer. (b) Surface representation of the residues involved at the interface of CTNav1.5 (orange) and the N- and C-lobe of CaM (magenta). The neighbouring CTNav1.5 is shown in green. (c) Hydrogen-bonding residues flanking the end of the interface (Glu12-Lys1899; Glu15-Asn1831; Glu115-Arg1913; Asp81-Arg1910).
Interaction of CTNav1.5 EFL domain and helix αVI
In the crystal structure reported here, the EFL domains (1,776–1,882) of each CTNav1.5 molecule contacts the C-terminal portion of helix αVI (1,910–1,926) of another CTNav1.5 molecule (Fig. 3). This CTNav1.5–CTNav1.5 interaction (buried area ~900 Å; complementarity index 0.55 (ref. 24)) is also mainly hydrophobic. It is flanked at both ends by salt bridges (Glu1804–Arg1910, Glu1799–Arg1914 and Asp1792–Lys1922) and hydrogen bonds (Tyr1795–Arg1914 and Asn1883–Ser1920) (Figs 3 and 4). A helix αVI–EFL interaction was postulated by Chazin et al.16, and measured by Glaaser25 by transition-metal ion fluorescence resonance energy transfer, but it was interpreted as an intramolecular interaction and not as an interaction between two molecules.
(a) Front view of the CTNav1.5–CTNav1.5 dimer. Residues of one CTNav1.5 molecule (green) interact with another CTNav1.5 molecule (orange). (b) Same as a but rotated 90° from a in the plane of the figure. (c) 90° rotation from b showing the concave cavity formed by the helices of the EFL domain; helix αI 1,788–1,801; helix αII 1,814–1,820; helix αIII 1,832–1,837; helix αIV 1,850–1,858; helix αV 1,866–1,882; helix αVI 1,897–1,926. (d) Close-up of the cavity formed by helices αI and αV with helix αVI. (e) Same as d, but with one CTNav1.5 coloured according to the electrostatic potential. It displays a hydrophobic surface with ionic patches at the beginning and end.
(a) Overview of the Nav1.5–Nav1.5 interaction. (b) Surface representation of the residues involved at the interface of CTNav1.5 helix αVI (orange and brown) with the neighbouring CTNav1.5 EFL (green and forest green) displaying the involved amino acids as sticks. (c) Ribbon representation of the interface showing the amino acid at the hydrogen-bonding distance. Hydrogen-bonding pattern of CTNav1.5 EFL (green and forest green) with the CTNav1.5 (orange and brown).
Calorimetric analysis
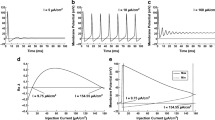
Isothermal titration calorimetry at 28 °C (Fig. 5a) shows that CTNav1.5 binds CaM in an exothermic reaction with Kds ranging from 46±33 to 105±15 nM depending on the buffer conditions. The lack of significant enthalpies of binding of the III–IV linker to the CTNav1.5–CaM complex argues against the competition between III–IV and the C-lobe of CaM (Fig. 5b).
(a) Thermodynamic analysis of CTNav1.5 and CaM binding. Isotherms of CTNav1.5 titrated with CaM. Top panel displays the heat evolved following each injection and the bottom panel shows the integrated heats of injection. All the curves are fitted to a one-binding site per monomer model. (b) Thermodynamic analysis of CTNav1.5–CaM and III–IV linker binding. Isotherm of CTNav1.5–CaM titrated with the III–IV linker (residues 1,489–1,502). (panels as in a). Note the change in scale on the y axis. (c) Attachment points of the N termini of the CTNav1.5 (coloured squares) to the transmembrane helices S6 of domain IV of the channel (not shown). (d) Overlap of CT helices αVI in the resting non-inactive conformation (orange) versus inactivated (grey). The rotation of the helix αVI is evidenced by the rotated positions of Lys1899, Arg1910, Arg1913 and Ser1920. This change highlights the disruption of the CTNav1.5–CTNav1.5 interaction caused by the rotation that results in the inactivated form of the channel. (e) Same as d, but seen from the C-terminal of helix αVI of the CTNav1.5.
Size-exclusion chromatography of CTNav1.5–CaM
Size-exclusion chromatography (SEC) analysis shows that, in solution, CTNav1.5–CaM behaves as a heterodimer with a retention time corresponding to a complex with one molecule of CTNav1.5 and one of CaM (Supplementary Fig. 3). The elution profile shows no indication of the presence of oligomers of this heterodimer.
Discussion
The most intriguing feature of the structure presented here is the presence of an intimate contact between the C-terminal portion of the Nav1.5 helix αVI and the EFL domain of another Nav1.5 molecule in an arrangement that can be described as an asymmetrical [CTNav1.5–CaM]–[CTNav1.5–CaM] homodimer (Figs 1, 3 and 4). This dimer may reflect the Nav1.5–Nav1.5 interaction reported in cells. Full-length Nav1.5 was shown by co-immunoprecipitation to form homodimers when expressed in cells26,27. Moreover, this dimerization of Nav1.5 explains the effect of some dominant-negative disease-causing mutations. Although there is no direct evidence that the interface observed in the structure described here is part of this dimerization, there are other strong indications that the contact described in this manuscript is part of an important physiological interaction. First, all five Nav1.5 monomers in the ASU participate in this exact same contact. The r.m.s.d. among the structures of the five heterodimers in the ASU range from 1.2 to 1.7 Å (for 270 Cα carbons aligned), indicating that this organization of two Nav1.5 and two CaM molecules is conserved regardless of crystal contacts. Had these contacts only been present between molecules related by crystal symmetry, an argument could have been made that they are a consequence of crystal packing. That all molecules in the ASU make this contact points to a fundamental interaction.
Second, this contact buries about 900 Å2 with a complementarity index of 0.55 (Supplementary Table 5). These values are more compatible with a true protein–protein interaction than with a crystal contact. Confirming this assessment, PISA28, the standard software for identifying protein–protein interactions, recognizes the interface between all complexes in the ASU, G and F, F and J, J and H, I and H and I and G24, as true dimers. Furthermore, residues that participate in this interface are well conserved among Nav isoforms (Supplementary Fig. 4).
Third, in the dimer, the N termini of the two CTNav1.5 monomers point in the same direction. This arrangement allows the N-terminal of both CTNav1.5 to simultaneously emerge as continuations of the helices S6 of their respective channels (Fig. 5c). In addition, in this arrangement, there are no clashes of any portion of the dimer of complexes (Nav1.5–CaM) with the membrane (Fig. 5c). The functional (physiological) effect of this dimerization is discussed in the next section.
The presence of a dimer of this construct is not detectable by SEC even at the highest concentrations achievable in solution (Supplementary Fig. 3). This is not surprising. While the CTNav1.5 is connected to the rest of the channel it is constrained to a very small volume close to the cytoplasmic side of the membrane (two-dimensional case) resulting in a very high local concentration. In contrast, in solution the molecules are free to occupy any position in the bulk (three-dimensional case). This effect was discussed extensively before29,30,31. The case for dimerization of the cytoplasmic portion of Nav1.5 is further strengthen by the observation that the full-length channel participates in functional dimers.
The physiological relevance of the structure reported here can be recognized by comparing it with that of CTNav1.5 in complex with CaM and FHF2. FHF2 has been shown to promote long-term inactivation of Nav channels5,32,33. This observation can be construed as suggesting that binding of the FHF stabilizes an inactivated conformation of the channel by locking the complex of CTNav1.5 and CaM in a configuration that is not compatible with channel conductance. This conformation of the CTNav1.5–CaM portion of the complex shows large differences with the structure reported here.
In the CTNav1.5–CaM-FGF13 (heterotrimer), CaM is in an extended conformation in which CaM helices αD–αE are fully extended, forming a single helix that includes the linker residues. The only interaction of CaM with the CTNav1.5 involves binding of the CaM C-lobe to the IQ motif. Because of the CaM extended conformation, the CaM N-lobe lies away from the CTNav1.5 in a position that would allow it to interact with the DIII–IV linker. In contrast, in the CTNav1.5–CaM structure, CaM is present in a conformation resembling a ‘lock-washer’ that allows its N-lobe to make significant contacts with the EFL domain of Nav1.5 (area buried 450 Å2, Fig. 6). Although the interaction of the CaM C-lobe with the IQ of helix αVI is the same in both complexes, in the structure of the complex reported here, the relation between helix αVI and the EFL domain is markedly different: helix αVI is rotated by ~90° around its axis (Fig. 5). Changes in the conformation of residues 1,892–1,895 of the Nav1.5, which link the EFL domain to helix αVI, accommodate this rotation without significant changes in either the long helix or the rest of the EFL domain. This configuration of the Nav1.5–CaM heterodimer is incompatible with the mode of binding of FGF13 in the tripartite inactivated complex. On the basis of these key differences, we propose that the structure reported here represents the non-inactivated (resting) state of the cytoplasmic portion of the Nav1.5, that is, the configuration that is poised for opening of the channel. In this configuration, the heterodimer can form the asymmetric CTNav1.5–CaM/CTNav1.5–CaM homodimer described above. This dimerization stabilizes the non-inactivated conformation of the channel.
(a) CTNav1.5–CaM is shown in lime-green/yellow (this work; PDB 4OVN). (b) The alignment highlights the rotation of helix αVI by 90° with respect to the EFL domain. The CaM rotates with the helix. CTNav1.5–CaM is shown in lime-green/yellow (this work) and Nav1.5–CaM-FGF13 (PDB 4DCK, red; CaM, sand, FHF in cyan). (c) Nav1.5–CaM-FGF13 (PDB 4DCK). (d) Close-up of the EFL area boxed in a; Nav1.5–CaM-FGF13 has an extra helix V’. (e) Overlap of CTNav1.5–CaM with Nav1.5–CaM-FGF13 showing that the extended conformation of CaM is not conducive to the formation of a CTNav1.5–CTNav1.5 dimer; two residues in helix VI show the rotation of the helix. The residues shown are part of the charge–charge interaction of Nav1.5 IQ–CaM.
In summary, the conformation of the CTNav1.5–CaM heterodimer appears to control the state of the channel. When in the conformation observed in the Nav1.5–CaM-FGF13 complex the channel is inactivated—it cannot conduct. This conformation is stabilized by binding FHFs. In the structure reported here, the CaM N-lobe interacts with the Nav1.5 EFL. As a result, the C-lobe changes its orientation with respect to the EFL by 90° and rotates with its helix αVI to the conformation that is ready for activation, that is, the non-inactivated state. This conformation is stabilized by the formation of the CaM–Nav1.5–Nav1.5–CaM homodimer.
We propose that under physiological conditions, when the two CTNav1.5 that form the homodimer are connected to the rest of their channels, only one of the two channels becomes non-inactivated and ready to open. Activation through the formation of an asymmetric homodimer has been proposed for another membrane protein, the EGF tyrosine kinase receptor, also based on an interaction observed in the crystal structure of its cytoplasmic C-terminal domain34.
All CaM molecules in the structure reported here contain Mg2+ in their EF hands and therefore adopt the conformation observed in the structures of other CaM–Mg2+ complexes and in the structures of apo-CaM. The arrangement of the CTNav1.5 and CaM in the homodimers is not compatible with the CaM conformation of the CaM–Ca2+ complexes, indicating that the non-inactivated state can only be achieved with apo or Mg2+-bound CaM. Since the cellular concentration of Mg2+ is maintained in the range 0.5–1.0 mM—that is, above the Kd of CaM for Mg2+—CaM most likely binds Mg2+ during the portion of the cycle when Ca2+ is at its resting concentration. Thus, when the channel changes from inactivated to non-inactivated, Ca2+ ions bound to the CaM EF hands are replaced by Mg2+, allowing the N-lobe to interact with the EFL domain and favouring formation of the Nav1.5–Nav1.5 homodimer.
Using the available structures, many mutations observed in cardiac arrhythmias map to the interfaces of CTNav1.5 with channel-interactive proteins or other regions of the channel including the DIII–IV linker. For example, Q1909R, associated with LQT3, produces a substantial depolarizing shift in steady-state inactivation (Fig. 7; Table 2). Another mutation Q1909A destabilizes CaM binding to the CTNav1.5 IQ domain in vitro (Table 213; Supplementary Fig. 5). In contrast, a mutation at the adjacent position, R1910A, has no significant effect on channel gating (Table 2; Supplementary Fig. 5).
(a,b) Electrophysiological features of Na+ channel variants altered at the CTNav1.5–CaM–Ca2+ interaction interface. Activation and steady-state inactivation of wild-type (circles) and mutant channels (squares). The data are fit to a Boltzman function as described in Methods. Mutant channels and Q1909R and K1922A exhibit a depolarizing shift in the V0.5 of inactivation (Supplementary Fig. 5). (c) Surface representation of one monomer (orange) interacting with a neighbouring molecule (lime green). CTNav1.5 with LQT3 mutations (1795, 1825, 1840, 1895, 1909 and 1924) shown in grey and brugada syndrome mutations in blue (1837, 1901, 1904 and 1919). Epilepsy mutations at the interface with the CaM-C-term lobe (1895 and 1852) are shown in purple; Brugada mutations (1901 and 1904) are at the interface of CTNav1.5 with the CaM-C-term lobe (blue) and 1705, 1825, 1840 and 1924 are at the CTNav1.5–CTNav1.5 interface. Residues 1788, 1790, 1792 and 1799, shown in dark red, line the EFL-binding site for helix αVI residues past the IQ motif14. Residues of the FGF13 that contact the EFL domain are coloured turquoise. (d) Same as c displaying the neighbouring CTNav1.5. (e) Same orientation as in c and d, including CaM as a magenta ribbon.
The physiological effects of many mutations in the CT were previously difficult to understand. Some of these can now be explained using the Nav1.5–Nav1.5 homodimer interface identified here (Fig. 7). For example, the mutation of K1922A produces a depolarizing shift in steady-state inactivation. Interestingly, K1922 is involved in a salt bridge that links helix VI to EFL residue E1788 (Figs 4 and 7). Rotation of helix αVI in the inactivated conformation disrupts this interaction (Fig. 5d). In contrast, mutations of R1914, which is part of the same interface, produce a smaller shift in gating (Table 2; Supplementary Fig. 5). Residues 1,924 and 1,914, mutated in Brugada syndrome, are at the Nav1.5–Nav1.5 interaction interface. A1924T was predicted to be at the CaM–IQ motif interface35; however, Chazin et al.36 showed that this residue was not in contact with CaM but with the EFL domain and suggested that the alanine residue is in a dynamic region as a possible explanation. The structure of the CTNav1.5–CaM provides a direct explanation for the dysfunction: the intermolecular interaction of the Nav1.5 helix αVI with the Nav1.5 EFL domain of another channel (Fig. 7), brings residue 1,924 in close proximity to residues Met1851 and Met1875 of the interacting CTNav1.5 (Fig. 7).
Other mutations that result in significant alterations in inactivation of gating include mutations in the NaV1.5–NaV1.5 interface that can only be explained by the structure reported here. Notably, mutations in the αI helix of the EFL (for example, D1790G and E1788A/D1790A/D1792A/E1799A) markedly shift the steady-state inactivation (V0.5) in the hyperpolarizing direction and result in slow recovery from inactivation (Table 2).
Direct binding measurements provided additional information about binding among cytoplasmic components. On the basis of isothermal titration calorimetry (ITC) experiments, Sarhan et al.12 suggested that the DIII–IV linker of Nav1.5 (residues 1,489–1,522) binds the C-lobe of CaM (Kd≈19 μM) and that this is the sole anchor point of binding of the DIII–IV peptide. In the structure reported here, however, the IQ portion of the helix αVI of Nav1.5 binds to the same region of CaM as the DIII–IV linker, but in the opposite orientation (Supplementary Fig. 6). In solution, CTNav1.5 binds CaM with high affinity (Kd≈50 nM). The ITC titration of the CTNav1.5–CaM complex with the DIII–IV linker peptide (see Methods) does not show significant enthalpy of binding, suggesting that the DIII–IV linker may not be able to displace the IQ from CaM binding (Fig. 5b). This result suggests that after a dimerized channel opens a change takes place in the Nav1.5–Nav1.5 interaction that releases one of the two lobes of CaM for fast inactivation by DIII–IV binding (Fig. 8). We propose that as part of this change, it is the N-lobe of CaM that is released from its interaction with the EFL and binds the DIII/IV of the same channel. In addition, we also propose that the C-lobe of CaM remains bound to the IQ and that the linker between the two CaM lobes adopts the extended helical configuration observed in the Nav1.5–CaM–FHF structure. This arrangement would set up Nav1.5 for binding FHF for long-term inactivation. A schematic representation of the relevant states is shown in Fig. 8. An important question can be asked: How does the increase in the intracellular Ca2+ concentration affect these states? One possible explanation is that Ca2+ ions will bind to CaM and change its conformation, including the conformation of both lobes. These changes would have a significant impact on the way that CaM interacts with the CTNav1.5 and as a result, on the regulation of channel activation.
Nav1.5 is coloured in green with a serrated marker in helix αVI to highlight the 90° rotation. Calmodulin is shown in purple. (a) Resting state showing the intermolecular interaction of the Nav1.5 poised for activation. (b) Active. N-lobe of CaM interacts with the EFL domain of Nav1.5; C-lobe of CaM interacts with IQ of helix αVI. (c) Long inactivated. FGF13 bound to the EFHL. (d) Fast inactivated. N-lobe of CaM releases the EFL-like and possible interacts with the III–IV linker. Helix αVI rotates 90°.
The intricate set of interactions observed in the structure of the complex presented here, that includes binding of both CaM lobes to the CTNav1.5, as well as an apparent asymmetric dimerization of two Nav1.5s, suggests a basis for an exquisite mechanism for switching from the inactivated to the non-inactivated form of the Nav1.5 channel (Fig. 8). This model also provides a rationale for disease-causing mutations that until now have escaped explanation.
Methods
Cloning
Complementary DNA (cDNA) corresponding to amino acids 1,773–1,929 of the Homo sapiens Nav1.5 sodium channel α-subunit (CTNav1.5) was cloned into the pGEX-6-P1 vector. The entire H. sapiens CaM gene was cloned into the pET24b vector using the BamHI and NdeI sites.
Expression and purification of the CTNav1.5–CaM complex
B834 (DE3) methionine auxotroph cells were transformed with both the CTNav1.5-containing and CaM-containing plasmids simultaneously. The cells were grown overnight at 37 °C in 50 ml of LB medium supplemented with 50 μg ml−1 kanamycin and 100 μg ml ampicillin. The overnight culture was centrifuged, the LB media was discarded and the cells were resuspended in 50 ml of 1 × M9 media containing 50 μg ml kanamycin and 100 μg ml−1 ampicillin. Ten ml of the resuspension was used to inoculate 1 l of M9 media containing 0.05 mg ml−1 selenomethionine, 50 μg ml−1 kanamycin, 100 μg ml−1 ampicillin and additives as described by Leahy et al.37 The cells were grown at 37 °C to an absorbance of 0.6 and protein expression was induced with 0.1 mM isopropyl β-D-1-thiogalactopyranoside. The cells were grown for an additional 5 h, centrifuged and the pellet was frozen at −80 °C.
After thawing, pellets were resuspended in 40 ml 1 × phosphate-buffered saline (PBS) with a mixture of protease inhibitors: 2 μg ml−1 pepstatin A, 50 μg ml−1 leupeptin and 35 μg ml−1 phenylmethanesulfonyl fluoride (PMSF) per liter of culture. An additional 40 ml of 1 × PBS containing 2 mg ml−1 lysozyme were added, along with Triton X-100 to bring the final detergent concentration to 1%. The solution was incubated at 4 °C for 1.5 h with shaking, sonicated on ice in 15 s bursts for a total of 1 min and spun at 15,000 r.p.m. for 30 min. The supernatant was filtered through a 0.22 μm polyethersulfone (PES) filter, and loaded on a 25 ml Glutathione Sepharose 4 Fast Flow resin using a hydrostatic pump at 0.5 ml min−1. The column was washed with 1 × PBS at 1 ml min−1 for a total of six resin volumes and protein was eluted in aliquots of 15 ml with an elution buffer containing 154 mg of reduced L-glutathione in 50 mM Tris-HCl at pH 8. Protein concentration in the samples was measured using a Nanodrop, and purity was assessed on a 4–12% Bis-Tris gel stained using Coomasie blue. Samples were stored on ice overnight at 4 °C. Prescission protease (0.3 mg) was added to the GST-CTNav1.5–CaM complex per 10 mg of protein. The protein was dialysed overnight against 50 mM Tris pH 7.0, 1 mM dithiothreitol. The next day, the protein was loaded onto a Source Q column. The eluate from the Source Q column was concentrated to 55 mg ml−1 and stored at −80 °C. Purity was assessed at each step by SDS–polyacrylamide gel electrophoresis. Visual inspection revealed a purity of at least 95%.
Structure determination
The CTNav1.5–CaM wild-type and selenomethionine-derivatized protein complex were crystallized by hanging-drop vapour diffusion at 18 °C by 1.2 :0.8 ratio of protein at 35–55 mg ml−1 and a well solution containing 15% polyethylene glycol 4K, 0.2 M MgSO4 and 10% glycerol. Prior to data collection, crystals were transferred to the reservoir solution supplemented with additional glycerol and flash-frozen in liquid nitrogen.
Data for native protein crystals were collected at the home source; multiwavelength anomalous diffraction (MAD) data for the selenomethionine-derivatized crystals were collected at APS on the LRL-CAT beamline 31A and at the National Synchrotron Light Source beamline X6A at the Se peak wavelength of 0.97989 Å. Indexing and data reduction were carried out with HKL2000 (ref. 38). The crystal contained five CTNav1.5–CaM complexes in the ASU.
Phases were determined with SHARP39 using data from the SeMet crystal. The positions of 47 out of the 90 seleniums atoms (8 methionine residues in CTNav1.5, 10 methionine residues in CaM with 5 heterodimers in the ASU) were determined by the program followed by phase determination, map calculation, density modification and initial model building to 3.3 Å. Non-crystallographic symmetry averaging with the Solomon (density modification) program of the AutoSharp suite40 was used to extend the phases to 3.0 Å using a solvent content of 55%. The map generated using these improved phases allowed a combination of automatic and manual tracing of the protein backbone followed by iterative cycles of manual model building with Coot41 and refinement with Refmac5 in the CCP4i suite.40
The model of the SeMet-derivatized protein was used as a molecular replacement search model42 for the native protein, which diffracted to a higher resolution. Model building was completed with iterative cycles of manual model building with Coot and refinement with Refmac5 (ref. 40). The correctness of the alignment of the sequence to the electron density was ascertained using the selenomethionine sites determined in SHARP. The final model was refined to 2.8 Å with Rwork and Rfree values of 21.6 and 28.3, respectively (Supplementary Table 1). The quality of the structure was validated with Procheck43 and Molprobity44. Residues (82.6%) are in the favoured region of the Ramachandran plot, 14.9% in additional allowed region and no residues in the disallowed region. The final model contains a total of 1352 amino acids. Buried surface areas were calculated using Protein Interfaces, Surfaces, and Assemblies (PISA) software24. Native data collected at Cu Kα was used to calculate anomalous maps to verify the absence of Ca2+ bound to CTNav1.5–CaM–Mg2+.
Size-exclusion chromatography
Gel filtration of 200 μl CTNav1.5–CaM 40 mg ml−1 was performed (n=5) on a Superdex 75 10/300 GL column on an AKTA FPLC (GE Lifesciences) in 50 mM Tris Cl pH 7.0, 150 mM NaCl, 1 mM dithiothreitol with and without 20 mM MgSO4. The protein standards albumin (66 kDa) and carbonic anhydrase (29 kDa) were used to calibrate the column.
Isothermal calorimetry
ITC experiments were performed with CTNav1.5 75 μM and 1 mM CaM. The protein in the sample cell was titrated with 25 10 μl injections of CaM. ITC experiments were performed with CTNav1.5–CaM and III–IV peptide (residues 1,489–1,502). The protein complex was diluted to a concentration of 40 μM (in monomers) in a buffer containing 10 mM Na2HPO4, 2 mM KH2PO4, 2.7 mM KCl, 137 mM NaCl, pH 7.4. The III–IV peptide was prepared in the same buffer at a concentration of 0.5 mM. Protein (1.8 ml) in the sample cell was titrated with twenty-five 10-μl injections. The data were analysed with Origin-5.0 software and fitted to a single binding site per monomer.
Cellular electrophysiology
The NaV1.5 mutations were made by site-directed mutagenesis of NaV1.5-GFP cDNAs using the Strategene Quickchange XL kit14. Approximately 0.75 × 106 human embryonic kidney cells (HEK293; American Type Culture Collection, Manassas, VA) were cultured in six-well tissue culture dishes in Dulbecco's modified Eagle's medium supplemented with foetal bovine serum 10%, L-glutamine (2 mmol l−1). The cells were co-transfected with plasmids encoding the appropriate NaV1.5-GFP and NaVβ1 subunit cDNAs as previously described14. Cells were transfected using Lipofectamine 2000 (Invitrogen) according to the manufacturer’s instructions and were studied at 48 h post transfection. The total amount of DNA for all transfections was kept constant.
Whole-cell INa was measured under voltage-clamp with an Axopatch 200B patch-clamp amplifier (Molecular Devices Corp.) at room temperature (22 °C). Voltage command protocols were generated by custom-written software and PCLAMP 10. Capacitance compensation was optimized and series resistance was compensated by 40–80%. Membrane currents were filtered at 5 kHz and digitized with 12-bit resolution through a DigiData-1200 interface (Molecular Devices Corp.). The patch pipettes had 1–2 MΩ tip resistances when filled with a pipette solution containing: 10 mM NaF, 20 mM CsCl, 100 mM CsF, 10 mM HEPES, 5 mM BAPTA, 4 mM CaCl2, pH 7.3 with CsOH. The bath solution contained 145 mM NaCl, 4 mM KCl, 1.8 mM CaCl2, 1 mM MgCl2, 10 mM glucose, 10 mM HEPES (pH 7.4 with NaOH).
Standard voltage-clamp protocols were used to characterize the voltage dependence of activation and inactivation and recovery from inactivation. To determine the V1/2 and slope factor k, steady-state inactivation data were fit with a Boltzmann function of the form: I/Imax={1+exp[(V−V1/2)/k]}−1. Recovery from inactivation data were fit with a bi-exponential function of the form:

using a nonlinear least squares minimization.
Additional information
Accession codes: Coordinates and structure factors have been deposited in the Protein Data Bank with accession code 4OVN.
How to cite this article: Gabelli, S. B. et al. Regulation of the NaV1.5 cytoplasmic domain by calmodulin. Nat. Commun. 5:5126 doi: 10.1038/ncomms6126 (2014).
Accession codes
References
Zimmer, T. & Surber, R. SCN5A channelopathies--an update on mutations and mechanisms. Prog. Biophys. Mol. Biol. 98, 120–136 (2008).
Stuhmer, W., Methfessel, C., Sakmann, B., Noda, M. & Numa, S. Patch clamp characterization of sodium channels expressed from rat brain cDNA. Eur. Biophys. J. 14, 131–138 (1987).
West, J. W. et al. A cluster of hydrophobic amino acid residues required for fast Na(+)-channel inactivation. Proc. Natl Acad. Sci. USA 89, 10910–10914 (1992).
Abriel, H. Roles and regulation of the cardiac sodium channel Na v 1.5: recent insights from experimental studies. Cardiovasc. Res. 76, 381–389 (2007).
Savio-Galimberti, E., Gollob, M. H. & Darbar, D. Voltage-gated sodium channels: biophysics, pharmacology, and related channelopathies. Front. Pharmacol. 3, 124 (2012).
Cormier, J. W., Rivolta, I., Tateyama, M., Yang, A. S. & Kass, R. S. Secondary structure of the human cardiac Na+ channel C terminus: evidence for a role of helical structures in modulation of channel inactivation. J. Biol. Chem. 277, 9233–9241 (2002).
Wang, C., Chung, B. C., Yan, H., Lee, S. Y. & Pitt, G. S. Crystal structure of the ternary complex of a NaV C-terminal domain, a fibroblast growth factor homologous factor, and calmodulin. Structure 20, 1167–1176 (2012).
Biswas, S. et al. Calmodulin regulation of Nav1.4 current: role of binding to the carboxyl terminus. J. Gen. Physiol. 131, 197–209 (2008).
Wingo, T. L. et al. An EF-hand in the sodium channel couples intracellular calcium to cardiac excitability. Nat. Struct. Mol. Biol. 11, 219–225 (2004).
Kim, J. et al. Calmodulin mediates Ca2+ sensitivity of sodium channels. J. Biol. Chem. 279, 45004–45012 (2004).
Sarhan, M. F., Tung, C. C., Van Petegem, F. & Ahern, C. A. Crystallographic basis for calcium regulation of sodium channels. Proc. Natl Acad. Sci. USA 109, 3558–3563 (2012).
Sarhan, M. F., Van Petegem, F. & Ahern, C. A. A double tyrosine motif in the cardiac sodium channel domain III-IV linker couples calcium-dependent calmodulin binding to inactivation gating. J. Biol. Chem. 284, 33265–33274 (2009).
Deschenes, I. et al. Isoform-specific modulation of voltage-gated Na(+) channels by calmodulin. Circ. Res. 90, E49–E57 (2002).
Biswas, S., DiSilvestre, D., Tian, Y., Halperin, V. L. & Tomaselli, G. F. Calcium-mediated dual-mode regulation of cardiac sodium channel gating. Circ. Res. 104, 870–878 (2009).
Haitin, Y., Carlson, A. E. & Zagotta, W. N. The structural mechanism of KCNH-channel regulation by the eag domain. Nature 501, 444–448 (2013).
Chagot, B., Potet, F., Balser, J. R. & Chazin, W. J. Solution NMR structure of the C-terminal EF-hand domain of human cardiac sodium channel NaV1.5. J. Biol. Chem. 284, 6436–6445 (2009).
Miloushev, V. Z. et al. Solution structure of the NaV1.2 C-terminal EF-hand domain. J. Biol. Chem. 284, 6446–6454 (2009).
Zhang, M., Tanaka, T. & Ikura, M. Calcium-induced conformational transition revealed by the solution structure of apo calmodulin. Nat. Struct. Biol. 2, 758–767 (1995).
Senguen, F. T. & Grabarek, Z. X-ray structures of magnesium and manganese complexes with the N-terminal domain of calmodulin: insights into the mechanism and specificity of metal ion binding to an EF-hand. Biochemistry 51, 6182–6194 (2012).
Feldkamp, M. D., Yu, L. & Shea, M. A. Structural and energetic determinants of apo calmodulin binding to the IQ motif of the Na(V)1.2 voltage-dependent sodium channel. Structure 19, 733–747 (2011).
Meador, W. E., Means, A. R. & Quiocho, F. A. Modulation of calmodulin plasticity in molecular recognition on the basis of X-ray structures. Science 262, 1718–1721 (1993).
Swindells, M. B. & Ikura, M. Pre-formation of the semi-open conformation by the apo-calmodulin C-terminal domain and implications binding IQ-motifs. Nat. Struct. Biol. 3, 501–504 (1996).
Chichili, V. P., Xiao, Y., Seetharaman, J., Cummins, T. R. & Sivaraman, J. Structural basis for the modulation of the neuronal voltage-gated sodium channel NaV1.6 by calmodulin. Sci. Rep. 3, 2435 (2013).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Glaaser, I. W. et al. Perturbation of sodium channel structure by an inherited long QT Syndrome mutation. Nat. Commun. 3, 706 (2012).
Hoshi, M. et al. Brugada syndrome disease phenotype explained in apparently benign sodium channel mutations. Circ. Cardiovasc. Genet. 7, 123–131 (2014).
Clatot, J. et al. Dominant-negative effect of SCN5A N-terminal mutations through the interaction of Na(v)1.5 alpha-subunits. Cardiovasc. Res. 96, 53–63 (2012).
Krissinel, E. Crystal contacts as nature’s docking solution. J. Comput. Chem. 31, 133–143 (2009).
Bell, G. I. Models for the specific adhesion of cells to cells. Science 200, 618–627 (1978).
Bell, G. I., Dembo, M. & Bongrand, P. Cell adhesion. Competition between nonspecific repulsion and specific bonding. Biophys. J. 45, 1051–1064 (1984).
Wu, Y., Vendome, J., Shapiro, L., Ben-Shaul, A. & Honig, B. Transforming binding affinities from three dimensions to two with application to cadherin clustering. Nature 475, 510–513 (2011).
Dover, K., Solinas, S., D’Angelo, E. & Goldfarb, M. Long-term inactivation particle for voltage-gated sodium channels. J. Physiol. 588, 3695–3711 (2010).
Liu, C. J., Dib-Hajj, S. D., Renganathan, M., Cummins, T. R. & Waxman, S. G. Modulation of the cardiac sodium channel Nav1.5 by fibroblast growth factor homologous factor 1B. J. Biol. Chem. 278, 1029–1036 (2003).
Jura, N., Shan, Y., Cao, X., Shaw, D. E. & Kuriyan, J. Structural analysis of the catalytically inactive kinase domain of the human EGF receptor 3. Proc. Natl Acad. Sci. USA 106, 21608–21613 (2009).
Tan, H. L. et al. A calcium sensor in the sodium channel modulates cardiac excitability. Nature 415, 442–447 (2002).
Chagot, B. & Chazin, W. J. Solution NMR structure of Apo-calmodulin in complex with the IQ motif of human cardiac sodium channel NaV1.5. J. Mol. Biol. 406, 106–119 (2011).
Leahy, D. J., Erickson, H. P., Aukhil, I., Joshi, P. & Hendrickson, W. A. Crystallization of a fragment of human fibronectin: introduction of methionine by site-directed mutagenesis to allow phasing via selenomethionine. Proteins 19, 48–54 (1994).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in OS. Methods Enzymol. 276, 307–326 (1997).
Bricogne, G., Vonrhein, C., Flensburg, C., Schiltz, M. & Paciorek, W. Generation, representation and flow of phase information in structure determination: recent developments in and around SHARP 2.0. Acta Crystallogr. D Biol. Crystallogr. 59, 2023–2030 (2003).
Winn, M. D. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. D67, 235–242 (2011).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Navaza, J. Implementation of molecular replacement in AMoRe. Acta Crystallogr. D Biol. Crystallogr. 57, 1367–1372 (2001).
Laskowski, R., MacArthur, M., Moss, D. & Thornton, J. M. PROCHECK- a program to check the stereochemical quality of protein structures. J. Appl. Crystallogr. 26, 283–291 (1993).
Lovell, S. et al. Structure validation by Ca geometry. Proteins 50, 437–450 (2003).
Acknowledgements
This work was funded in part by NIH HL050411. Use of the Advanced Photon Source at Argonne National Laboratory was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under Contract No. DE-AC02-06CH11357. Use of the Lilly Research Laboratories Collaborative Access Team beamline at Sector 31 of the Advanced Photon Source was provided by Eli Lilly Company, which operates the facility. Data collection was carried out at beamline X6A, funded by the National Institute of General Medical Sciences, National Institute of Health under agreement GM-0080. The NSLS, Brookhaven National Laboratory is supported by the US Department of energy under contract No. DE AC02-98CH10886. We acknowledge the use and services of the JHU SOM Mass Spectrometry and Proteomics Core, supported by the NHI NIDDK center grant P30 DK089502. The rendition of Fig. 8 was contributed by T. Phelps, M.S.
Author information
Authors and Affiliations
Contributions
S.B.G., A.B., V.H.K., M.A.B., F.F., S.A., J.Y. and J.J. performed the experiments. S.B.G., M.A.B., F.F. analysed data. S.B.G., G.F.T. and L.M.A. designed and supervised research. S.B.G., M.A.B., G.F.T. and L.M.A. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-6 and Supplementary Tables 1-5 (PDF 977 kb)
Rights and permissions
About this article
Cite this article
Gabelli, S., Boto, A., Kuhns, V. et al. Regulation of the NaV1.5 cytoplasmic domain by calmodulin. Nat Commun 5, 5126 (2014). https://doi.org/10.1038/ncomms6126
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms6126
- Springer Nature Limited
This article is cited by
-
Fibroblast growth factor homologous factors serve as a molecular rheostat in tuning arrhythmogenic cardiac late sodium current
Nature Cardiovascular Research (2022)
-
Ca2+-dependent regulation of sodium channels NaV1.4 and NaV1.5 is controlled by the post-IQ motif
Nature Communications (2019)
-
Chemical shift assignments of a calmodulin intermediate with two Ca2+ bound in complex with the IQ-motif of voltage-gated Ca2+ channels (CaV1.2)
Biomolecular NMR Assignments (2019)
-
The voltage-gated sodium channel EF-hands form an interaction with the III-IV linker that is disturbed by disease-causing mutations
Scientific Reports (2018)
-
Backbone resonance assignments of complexes of apo human calmodulin bound to IQ motif peptides of voltage-dependent sodium channels NaV1.1, NaV1.4 and NaV1.7
Biomolecular NMR Assignments (2018)