Abstract
Melanoma patients treated with oncogenic BRAF inhibitors can develop cutaneous squamous cell carcinoma (cSCC) within weeks of treatment, driven by paradoxical RAS/RAF/MAPK pathway activation. Here we identify frequent TGFBR1 and TGFBR2 mutations in human vemurafenib-induced skin lesions and in sporadic cSCC. Functional analysis reveals these mutations ablate canonical TGFβ Smad signalling, which is localized to bulge stem cells in both normal human and murine skin. MAPK pathway hyperactivation (through BrafV600E or KrasG12D knockin) and TGFβ signalling ablation (through Tgfbr1 deletion) in LGR5+ve stem cells enables rapid cSCC development in the mouse. Mutation of Tp53 (which is commonly mutated in sporadic cSCC) coupled with Tgfbr1 deletion in LGR5+ve cells also results in cSCC development. These findings indicate that LGR5+ve stem cells may act as cells of origin for cSCC, and that RAS/RAF/MAPK pathway hyperactivation or Tp53 mutation, coupled with loss of TGFβ signalling, are driving events of skin tumorigenesis.
Similar content being viewed by others
Introduction
The development of epithelial tumours is generally accepted to take place over several years, involving the accumulation of mutations that drive tumour progression1. However, some tumours contain a relatively low mutation burden2 and develop rapidly, without progression from benign intermediary stages, suggesting a potential stem cell origin3. Data from murine model systems illustrate a tumour’s ability to form from both stem and differentiated cells. Within intestinal epithelium, loss of Apc in the LGR5+ve stem cell compartment leads to adenoma, whereas tumours rarely form from differentiated cells4. Conversely, we have shown that targeting Kras, in addition to Apc, can de-differentiate intestinal villi and permit tumour formation5. Thus, the tumour cell of origin remains unclear, as does the standard model of progression from benign tumour to malignant carcinoma.
Discord with the progression model is exemplified in the skin, which carries a high mutation burden6. Asymptomatic normal skin carries frequent mutations in TP53 (refs 7, 8) and NOTCH6,8. Classic chemical carcinogenesis 7,12-Dimethylbenz[a]anthracene (DMBA)/12-O-tetradecanoylphorbol-13-acetate (TPA) experiments demonstrate Hras mutations can lie dormant in the skin (without the addition of TPA), at no obvious consequence to the tissue9. Indeed even when Ras mutation is targeted to stem cell compartments (for example, LRIG1+ve cells or bulge stem cells10,11), this does not lead to cancer unless there is a disruption of tissue homeostasis through wounding. These findings support the hypothesis that homeostasis within stem cell compartments plays an important tumour suppressive role in highly organized structures such as the skin.
We reasoned that, in the absence of wounding, mutations in other oncogenic/tumour suppressor genes might facilitate rapid skin tumorigenesis. Using targeted sequence analysis and whole-exome sequencing (WES), we identify frequent mutation in both transforming growth factor-β (TGFβ) type 1 receptor (TGFBR1) and TGFβ type 2 receptor (TGFBR2) genes in human primary cutaneous squamous cell carcinoma (cSCC) samples. IntOgen mutation analysis reveals TGFβ signalling as a pathway significantly altered by mutation and functional analysis of several TGFβ receptor mutants indicates that many of these mutations result in loss of function. Pathway activation studies reveal highly localized TGFβ signalling in both normal human and mouse hair follicle bulge stem cells. In murine skin, targeted activation of the RAS/RAF/mitogen-activated protein kinase (MAPK) pathway, coupled with deletion of Tgfbr1 in LGR5+ve stem cells, promotes rapid development of cSCC, which, in the absence of wounding, may mimic the kinetics of tumour induction in vemurafenib-induced cSCC. Combined Tp53 mutation/inactivation coupled with Tgfbr1 loss in LGR5+ve stem cells also results in cSCC with longer latency, providing a model for cSCC development without RAS activation.
Results
TGFBR1 and TGFBR2 are frequently mutated in human cSCC
Cutaneous squamo-proliferative lesions (including keratoacanthomas and cSCC) arise in a significant proportion of patients treated with the type I RAF inhibitor vemurafenib. Such lesions develop within a few weeks of treatment12,13. Targeted sequencing has revealed that these lesions contain a high frequency of activating mutations in HRAS6,12,13. Cutaneous lesions isolated from patients treated with sorafenib (the ‘pan-RAF’ inhibitor) also harbour mutations in HRAS, TP53 and TGFBR1 (ref. 14). Employing targeted deep sequencing of 39 squamo-proliferative lesions from seven patients (including cSCC and actinic keratosis (Supplementary Table 1) treated with vemurafenib (using a percentage variance criterion of >10%), we identified frequent coding mutations in both TGFBR1 (8/39, 21% of samples) and TGFBR2 (5/39, 13% of samples), revealing mutation of TGFβ receptors in 28% of lesions (Fig. 1a and Supplementary Data 1). These mutational events were only surpassed in frequency by mutations in NOTCH1/NOTCH2 (56%) and activating mutations of HRAS (38%). TP53 mutations arose in 26% of lesions6 (Fig. 1a and Supplementary Data 2). In contrast to NOTCH (using our mutational call cutoff, see Methods), we did not detect any mutations in TGFβ receptors or HRAS in the normal or perilesional skin samples (n=6 from 4 patients, 3 of which had lesions containing TGFβ receptor mutations). These findings imply that a combination of potential mutational inactivation of TGFβ signalling and activation of HRAS may be important driving events in vemurafenib-induced skin lesions and skin tumorigenesis.
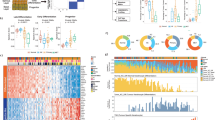
Mutation frequency, distribution and relationship with pathological features of skin lesions isolated from vemurafenib-treated patients (n=39) (a), sporadic cSCC (n=91) (b), 21 cSCC cell lines (RDEB, recessive dystrophic epidermolysis bullosa) (c); (a–c, adapted from ref. 6); 7 sporadic cSCC tumours (T) and normal distant/perilesional skin samples (N) collected in Dundee (d). Mutation status for nine genes is indicated and the overall percentage mutation is shown on the left. Each column represents a single case. Colours correspond to specific mutations as shown. Details of clinical parameters are included in Supplementary Tables 1–3 and mutations are included in Supplementary Data 1–5.
We next sought to investigate whether loss of TGFβ signalling is a frequent event in sporadic cSCC. We employed targeted 454 pyrosequencing of TGFBR1 and TGFBR2 in 91 human primary cSCC samples (Supplementary Table 2) and 21 human cell lines derived from primary cSCC15, all of which were recently sequenced for common genetic alterations6. Using a percentage variance criterion of >10%, we detected mutations of TGFBR1 in 22% and TGFBR2 in 30% of primary cSCC samples and 14% of cell lines (Fig. 1b,c and Supplementary Data 3). Overall, mutation of TGFβ receptors occurred in 43% of primary cSCC samples. These mutational events were only surpassed in frequency by mutations in NOTCH1/2 (86%) and this time TP53 (63%) (Fig. 1b, Supplementary Data 4 and ref. 6). In sporadic cSCC, oncogenic activation of RAS only occurred in 9% of samples (Fig. 1b, Supplementary Data 4 and ref. 6). We then sequenced normal blood samples from eight patients with sporadic cSCC, whose lesions harboured mutations in TGFβ receptors (Supplementary Data 3) and found no TGFβ receptor mutations. Next, we prospectively collected a further Dundee cohort of seven primary cSCC samples with complementary matched normal distant and perilesional skin (Supplementary Table 3). This cohort demonstrated a comparable spectrum of mutation in our selected gene panel and in both TGFβ receptors (Fig. 1d and Supplementary Data 5). TGFβ receptor mutations were again not identified in either distant or perilesional skin. To assess the potential lesion-specific, non-germline significance of TGFβ receptor mutations, we interrogated the pyrosequencing analysis in depth from all of the samples containing normal matched tissue (Supplementary Data 6). We observed only 8 variant reads out of 1,348 reads in total, in 4 out of 25 matched normal sample reads. Three of these samples were from perilesional skin and probably reflect rare contaminating tumour cells. In comparison, we observed 237 variant reads out of 1,340 reads in the tumour samples. Employing Fisher’s exact two-sided tests to compare variant allele frequencies (VAFs) in matched samples, we determined that 17/25 of the TGFβ receptor mutations reached tumour-specific VAF statistical significance, confirming the lesion-specific, non-germline nature of these mutations (Supplementary Data 6).
TGFBR1 and TGFBR2 mutations are driver events in human cSCC
Next, we examined a further cohort of 30 primary cSCC samples with matched normal tissue (Supplementary Table 4) employing next-generation WES (see Methods) and interrogated in detail NOTCH1, NOTCH2, TP53, CDKN2A, HRAS, KRAS, NRAS, TGFBR1 and TGFBR2 genes for mutational and copy number changes (Fig. 2a and Supplementary Data 7). We observed alterations in all of these genes with a similar frequency to that of our previous 454 pyrosequencing analysis. None of the mutational events were found in the matched normal samples and all except two of these were statistically significant (Fisher’s exact t-test) (Supplementary Data 8). Importantly, we observed changes in TGFBR1 in 30% of the samples and changes in TGFBR2 in 40% of the samples with a combined alteration in 53% of samples, confirming a frequent alteration of TGFβ receptor genes in cSCC. Copy number analysis also revealed that loss of heterozygosity occurred in both TGFBR1 and TGFBR2 genes including in tumours with missense mutations in TGFBR2 (Fig. 2a and Supplementary Data 8). Somatic single nucleotide variants (SNVs) of TGFβ receptors were detected in 30% of our samples consistent with our 454 pyrosequencing analysis and the recent sequencing analyses of two North American cSCC cohorts, which, when combined, detected TGFβ receptor proteins altering SNVs in 15.7% of samples16,17. Given the high mutational burden of cSCC, it is probable that many mutations identified will be passenger mutations with no functional consequence for tumorigenesis. We investigated the potential functional consequence of the mutations detected by WES employing MutsigCV18 and IntOgen analysis19. MutsigCV detected TP53, CDKN2A, NOTCH1 and NOTCH2 as significant drivers but no RAS genes and IntOgen detected TP53, CDKN2A NOTCH1 and HRAS as significant drivers but did not identify NOTCH2, KRAS or NRAS (Supplementary Data 9). Neither analysis detected TGFBR1 or TGFBR2 individually as significant drivers (Supplementary Data 9); however, IntOgen pathway analysis revealed TGFβ signalling as a significantly altered signalling pathway (Oncodrive-fm functional impact bias, functional mutation bias19, P=0.0019; Supplementary Data 10). We assessed the clonality of our candidate driver genes using the ABSOLUTE algorithm20. WES data were of sufficient quality for 24/30 exomes and ABSOLUTE analysis revealed purity and ploidy estimates ranging from 0.2 to 0.73 and from 1.78 to 5.79, respectively (Supplementary Data 11). ABSOLUTE clonality analysis indicated that all NOTCH1, CDKN2A and RAS mutations were clonal as were all bar one TP53, three NOTCH2 and one TGFBR1 mutation, which were subclonal (Fig. 2b,c and Supplementary Data 12). Mutations present in nearly all tumour cells (clonal) would suggest early events and therefore represent initiating ‘driver’ genes as appears to be the case here for NOTCH1, NOTCH2, CDKN2A, HRAS, KRAS, TP53 and, importantly, both TGFBR1 and TGBFR2.
(a) Mutation frequency, distribution and relationship with pathological features from 30 cSCC primary tumours. Mutation and copy number status (gain, loss and acquired uniparental disomy) for nine genes is indicated and the overall percentage alteration is shown on the left. Each column represents a single case. Colours correspond to specific mutations and copy number changes as shown. Split columns indicate where more than one mutation type is present in a single case. Details of clinical parameters and mutations are included in Supplementary Table 4 and Supplementary Data 7. (b) ABSOLUTE clonality analysis of potential driver genes of cSCC indicates that all nine genes are frequently clonal. (c) Cancer cell fraction clonality analysis indicates clonal and subclonal mutations in the indicated tumours.
Having established the probable driver event of mutation of TGFBR1 and TGFBR2 in our WES data set, we extended this analysis to include our samples assessed by targeted sequencing. We first calculated average percentage VAFs for our candidate drivers and these ranged from 48.7% for CDKN2A to 20% for TGFBR1 (Fig. 3a). TGFBR1 VAF was significantly lower than that of CDKN2A, TP53, HRAS, NOTCH1, TGFBR2 and NOTCH2 but not KRAS and NRAS (Fig. 3a and Supplementary Data 13). TGFBR2 VAF was only statistically significantly lower than CDKN2A and TP53 but equivalent to KRAS, NOTCH2 and NRAS (Fig. 3a and Supplementary Data 13). The VAFs of the TGFβ receptors are of a similar range to those observed in other cSCC driver genes. Ultraviolet light is the major oncogenic stimulus of cSCC and the % of mutations conforming to an ultraviolet signature (C-T or G-A transitions) of our candidate drivers ranged from 79.7% in CDKN2A to 30.4% in HRAS (Fig. 3b), with mutations in both TGFβ receptor genes lying within this range. VAFs were statistically significantly higher for ultraviolet signature mutations for NOTCH2, CDKN2A and TGFBR2 (Supplementary Fig. 1a and Supplementary Data 14). If these candidate genes represent potential driver genes then the mutational consequence should be predicted to change protein function. We classified these mutations as potentially damaging if they were predicted to be so by at least two of the four mutation function prediction programmes SIFT21, PolyPhen-2 (ref. 22), Provean23 and Mutation Assessor24 or were a splice site or PTC mutation (Fig. 3c). Damaging mutation rates ranged from 89% for TP53 to 53.5% for TGFBR1 (Fig. 3c and Supplementary Data 15–23) were statistically significantly higher for those with an ultraviolet signature for NOTCH2, TGFBR2 and TP53 (Fig. 3d and Supplementary Data 24), and damaging mutations had higher VAFs for NOTCH2, CDKN2A, TGFBR2 and NOTCH1 (Supplementary Fig. 1b and Supplementary Data 25). Together, our data suggest that ∼70% of TGFBR2 and 50% of TGFBR1 mutations will alter protein function with the potential to drive cSCC development. In its entirety, our analysis conservatively estimates functionally relevant TGFBR1 and TGFBR2 mutations in ∼10% and ∼16% of samples, respectively, and therefore 20% of cSCC samples could harbour damaging TGFβ receptor mutations.
Ranked driver gene mutations by (a) % VAF (P-values represent Student’s t-test (two-tailed) are shown above the figure, (b) % ultraviolet spectrum (c) predicted mutational consequence of Damaging/Non-damaging and (d) combined ultraviolet and damaging analysis (P-values are shown above the figure and represent χ2 Fisher’s exact-test). For all statistics, * defines statistical significance (***P<0.001, **P<0.01 and *P<0.05). Numbers of samples are contained in Supplementary Data 13. (e) Domain structures of TGFBR1 and TGFBR2 are shown. Exons are numbered and functional domains colour coded (see key). SNPs identified in TGFBR1 and TGFBR2 are labelled in AA sequence (using UniProtKB codes: P-36897-1, TGFBR1, HUMAN and P37173, P37173-2 and TGFBR2_HUMAN) for sporadic cSCC (above) and Vemurafenib-associated cSCC (below). Asterisked SNPs are those found in cSCC cell lines. Amino acid numbers for TGFBR2 refer to isoform 2.
TGFβ receptor mutation inactivates canonical Smad signalling
Identified missense and nonsense mutations were found throughout the coding exons of both TGFBR1 and TGFBR2, occurring in the extracellular and kinase domains of each protein (Fig. 3e). Structural analysis of the extracellular domains of TGFBR1 (Supplementary Fig. 2) and TGFBR2 (Supplementary Fig. 3) indicated mutations occur in, or in close proximity to, highly conserved disulphide bonds, ligand interaction motifs and/or receptor interaction motifs. These findings suggest significant potential for loss of function25,26.
TGFβ signals via activation of a heterotetrameric complex of TGFBR2:TGFBR1, resulting in TGFBR1-kinase driven carboxy-terminal phosphorylation of SMAD2 and SMAD3 (ref. 27). Once phosphorylated (PO4), SMAD2 and SMAD3 form hetero-oligomeric complexes with the co-Smad SMAD4 accumulate in the nucleus and regulate gene expression of hundreds of target genes28,29. Activity of SMAD-dependent reporter gene constructs and steady-state levels of SMAD2/3 C-terminal phosphorylation can be used as measures of canonical TGFβ signalling. To assess the functional consequence of these TGFβ receptor mutations, we generated a panel of four TGFBR1 and five TGFBR2 mutant expression plasmids from mutations identified in our original targeted sequencing series. We assayed each mutant receptor for functional activity in transient transfection reporter gene assays. TGFBR1 expression plasmids were co-transfected into TGFBR1-null mouse embryonic fibroblasts30 and TGFBR2 expression plasmids were co-transfected into TGFBR2-null T47D breast cancer cells, in addition to the TGFβ-responsive reporter construct SMAD7-Promoter Luciferase31 (Fig. 4a and b, respectively). Wild-type TGFβ receptor expression elevated reporter activity over empty vector controls, which was further elevated by TGFβ treatment (Fig. 4a,b). We confirmed this activity was dependent on intact SMAD binding elements in the SMAD7 promoter (Supplementary Fig. 4a,b). The TGFBR1 mutants H331R and W277C, and all of the TGFBR2 mutants (S474F, C486R, C96R, R2323W and A556T), failed to efficiently activate the reporter gene, despite similar levels of expression of the receptors, as assayed by western blotting (Fig. 4a,b). These findings indicate that mutation of TGFBR1 and TGFBR2 in cSCC frequently results in a loss of ability to activate canonical SMAD signalling. To demonstrate corollary of these findings in primary human tissue, we then established conditions to monitor C-terminal PO4-SMAD3 levels using a C-terminal Ser433/Ser435 PO4-SMAD3-specific antibody in cSCC by immunohistochemistry (IHC) (Supplementary Fig. 5). We measured PO4-SMAD3 activity in eight primary tumours harbouring wild-type receptors and eight primary tumours harbouring mutant TGFβ receptors with a combined VAF of >20% (Supplementary Data 26). Wild-type tumours exhibited readily detectable PO4-SMAD3 activity, whereas mutant tumours showed significantly reduced PO4-SMAD3 activity (Fig. 4c and Supplementary Fig. 6), consistent with our observation that mutation of TGFβ receptors results in loss of canonical SMAD signalling activity. Both wild-type and mutant tumours exhibited heterogeneity of staining, consistent with our previous observations that cSCC is heterogenous in nature6 and with the VAFs observed in mutant tumours.
(a) Indicated TGFBR1 plasmids were co-transfected into TGFBR1-null mouse embryonic fibroblasts and assayed for SMAD7-Promoter Luciferase (SMAD7 Promoter-Luc) reporter gene activity and receptor expression levels by western blotting (lower panels) with and without TGFβ stimulation for 4 h. β-Actin is used as a loading control. EV, empty vector control; WT, wild type. Data are mean ±s.d., n=3. (b) Indicated TGFBR2 plasmids were co-transfected into TGFBR2-null T47D cells and assayed for SMAD7-Promoter Luciferase (SMAD7 Promoter-Luc) reporter gene activity and receptor expression levels by western blotting (lower panels) with and without TGFβ stimulation for 4 h. β-Actin is used as a loading control. EV, empty vector control; WT, wild type. Data are mean ±s.d., n=3. (c) PO4-SMAD3 activity was assessed by IHC in wild-type and mutant tumours (n=8, ***P=0.001, Mann–Whitney U-test). Representative images are shown. Scale bar, 100 μM. (d) Effects of TGFβ stimulation on growth of NHK and cSCC cell lines. Data represent Cell Titre Glo measurement of cell proliferation over the indicated time course of cells treated with the indicated dose of TGFβ1. NHKs and cell lines harbouring mutant TGFBR2 (SCCIC8 and SCCIC12) are shown. Data represent the mean ±s.d. n=6. (e) Restoration of wild-type TGFBR2 restores growth inhibition. SCC1C8 and SCCIC12 cells were co-transfected with empty vector control (EV) or wild-type TGFBR2 expression plasmids (TGFBR2) and a GFP expression plasmid. Proliferation of GFP+ve cells was assessed using real-time Incucyte Zoom imaging over 6 days. Data represent the mean ±s.d. n=6. *P<0.05, **P<0.01 and ***P<0.001 (Student’s t-test).
Finally, we used primary human cSCC cell lines to assess whether TGFβ receptor mutation results in a loss of TGFβ signalling. Exogenous treatment of normal human keratinocytes (NHKs) with TGFβ1 resulted in a dose-dependent decrease in cell proliferation (Fig. 4d). The TGFBR2 mutant harbouring cell lines SCCIC8 and SCCIC12 (Supplementary Data 3) failed to respond to exogenous TGFβ stimulation by either PO4-SMAD activation (Supplementary Fig. 7) or by any effect on cell proliferation (Fig. 4d). Co-transfecting these TGFBR2 mutant cells with either empty vector, or wild-type TGFBR2 expression plasmids in addition to a green fluorescent protein (GFP) expression plasmid, we measured cell proliferation in real time using Incucyte-Zoom imager over 6 days. Cell proliferation of the GFP+ve cells indicated that cells expressing wild-type TGFBR2 proliferated at a slower rate in the presence of exogenous TGFβ (Fig. 4e). The degree of inhibition was commensurate to the degree of restoration of SMAD activity as measured using the multimerized SMAD binding element reporter gene CAGA12-Luciferase32 (Supplementary Fig. 7c,d). These findings indicate that re-expression of wild-type TGFBR2 restores canonical TGFβ signalling and proliferative inhibition, confirming mutational loss of TGFβ tumour suppressive activity.
Matrix cells exhibit active TGFβ signalling
Given this potential aetiological loss of TGFβ signalling, we sought to identify sites of active TGFβ signalling in normal skin, to gain insight into the cellular origin of cSCC RAF inhibitor-induced lesions. PO4-SMAD3 activity was barely detectable by IHC analysis in normal human epidermis (Supplementary Fig. 8) but showed strong immunoreactivity in the hair matrix zone of anagen hair follicles (Fig. 5a and Supplementary Fig. 8). PO4-SMAD3 positivity was also detected in the hair matrix of anagen hair follicles in mouse back skin (Fig. 5a). In anagen, the hair follicle transit-amplifying (TA) cells are localized in the matrix and are positive for Sonic hedgehog (SHH)11. Elegant studies by Blanpain and colleagues11 have demonstrated these cells are unable to act as a cell of origin for papilloma formation, even when both oncogenic Kras and Tp53 were targeted. This suggests that these PO4-SMAD3+ve hair matrix cells are unlikely to be the cell of origin for the rapid cSCC observed in humans following RAF inhibitor treatment. To investigate this in the mouse, we tested whether Tgfbr1 deletion could permit the transformation of TA cells. RAF inhibitors stimulate paradoxical activation of the MAPK pathway in cells with wild-type BRAF harbouring upstream pathway activation, via mechanisms such as the following: upregulated receptor tyrosine kinases, oncogenic RAS via RAF dimer formation33,34,35 or relief of inhibitory auto-phosphorylation36. Circumventing pharmacological enhancement of MAPK signalling in the presence of mutated RAS, we modelled hyperactivation of the MAPK pathway in the SHH+ve compartment by targeting downstream oncogenic BrafV600E and oncogenic activation of KrasG12D. We crossed our previously described LSL-BrafV600E mice37, which allow inducible expression of BrafV600E from the endogenous Braf gene, with the ShhCREER strain38. This permits tamoxifen-inducible activation of the Cre recombinase in SHH+ve cells. To assess the role of TGFβ signalling in the SHH+ve cells, we then crossed these animals with Tgfbr1fl mice30 (Supplementary Fig. 9a,b). No tumours formed in the skin of ShhCREER BrafV600E and ShhCREER BrafV600E Tgfbr1fl/+ mice (Fig. 5b and Supplementary Fig. 10a,b). A small percentage of ShhCREER BrafV600E Tgfbr1fl/fl mice developed minimally proliferative papillomatous lesions (as evidenced by low level 5-bromodeoxyuridine (BrdU) staining) mainly in the lips, but only at long latency (Fig. 5b and Supplementary Fig. 10c). No mice developed cSCC. Mice failed to develop any skin lesions following oncogenic activation of KrasG12D with or without deletion of Tgfbr1 in this cell compartment (Fig. 5c). Together, these studies indicate that the SHH+ve cells are unlikely to be the cell of origin for either rapid onset vemurafenib-induced cSCC or sporadic cSCC.
(a) PO4-SMAD3 IHC in anagen human (left panel) and mouse (right panel) hair follicles reveals immunoreactivity in the hair matrix. CTL, connective tissue layer; DP, dermal papilla; IRS, inner root sheath; ORS, outer root sheath. Scale bar, 100 μm (b) Kaplan–Meier survival curve of ShhCreER BrafV600E (n=13), ShhCreER BrafV600E Tgfbr1fl/+ (n=28) and ShhCreER BrafV600E Tgfbr1fl/fl (n=39) mice. (c) Kaplan–Meier survival curve of ShhCreER KrasG12D (n=17) and ShhCreER KrasG12D Tgfbr1fl/fl (n=21) mice.
TGFβ signalling is active in telogen bulge stem cells
Approximately 90% of human hair follicles are present in the anagen phase of the hair cycle with the remaining 10% existing in catagen or the resting telogen phase. Analysis of human telogen hair follicles revealed highly localized PO4-SMAD3 staining in the bulge stem cells, characterized in part by Keratin 15 staining (Fig. 6a). This pattern was recapitulated in mouse telogen hair follicles (Fig. 6b), characterized by the expression of the stem cell marker LGR5 (ref. 39). To investigate further, we used the Lgr5-EGFP-Ires-CREERT2 knockin mouse (hereafter termed Lgr5CREER), where the endogenous Lgr5 promoter controls expression of enhanced GFP (EGFP) and the CREERT2 fusion protein40. IHC analysis for GFP revealed a staining pattern similar of that observed for PO4-SMAD3 (Fig. 6c). Furthermore, co-immunofluorescence revealed LGR5+ve cells (stained for EGFP) are highly enriched for both PO4-SMAD3 and TGFBR1 (Fig. 6d and Supplementary Fig. 11).
(a) IHC analysis of PO4-SMAD3 (left panels) and Keratin 15 (right panels) in human normal skin. Insert shows strong PO4-SMAD3 staining in the telogen hair follicle Keratin 15+ve bulge stem cells. Scale bar, 100 μm. (b) IHC analysis of PO4-SMAD3 in murine normal skin. BG, bulge; DP, dermal papilla; IFE, inter-follicular epidermis; SG, sebaceous gland. (c) IHC analysis of LGR5-GFP in murine skin in the telogen phase of the hair cycle. Scale bar, 100 μm. (d) Immunofluorescence (IF) analysis of LGR5-GFP and PO4-SMAD3 in murine telogen skin. Nuclei are counterstained with DAPI. Scale bar, 50 μm. (e) qRT–PCR analysis of Tgfbr1, Tgfbr2, Tgfb1, Tgfb2, Tgfb3 and Smad7 in LGR5+ve (n=3 biological replicates) and LGR5−ve cells (n=3 biological replicates) freshly isolated from back skin in the telogen phase. Data are shown as ratios to the internal Gapdh control with error bars representing s.e.m. Statistical significance *P=0.04 (Mann–Whitney U-test, one-tailed test).
Recent studies indicate that the dermal papilla may provide a source of TGFβ2, activating SMAD signalling in overlying hair germ stem cells41. We sorted epithelial EGFP-positive LGR5+ve stem cells from murine back skin. Quantitative reverse-transcriptase PCR (qRT–PCR) analysis revealed LGR5+ve cells express enhanced levels of Tgfbr1, Tgfb1 and Tgfb3 messenger RNA when compared with LGR5−ve cells, with negligible amounts of Tgfb2 (Fig. 6e). Expression of Tgfbr2 was readily detected in GFP+ve and GFP−ve compartments (Fig. 6e). This indicates enriched autocrine TGFβ signalling in the LGR5+ve compartment. We observed high levels of the TGFβ target gene Smad7 (ref. 42) in LGR5+ve cells (Fig. 6e). Together, these findings indicate that autocrine TGFβ signalling is highly localized to the LGR5+ve hair follicle bulge stem cells in the mouse and the Keratin 15+ve hair follicle bulge stem cells in humans, and that this cell compartment may give rise to both vemurafenib-induced and sporadic cSCC.
Rapid cSCC formation from Lgr5+ve stem cells
To investigate the consequence of hyperactivation of the MAPK pathway coupled with ablation of TGFβ signalling in LGR5+ve stem cells, we crossed the Lgr5CREER mice with the LSL-BrafV600E mice (Supplementary Fig. 9c), or LSL-KrasG12D mice and Tgfbr1fl mice (Supplementary Fig. 9d). Loss of TGFβ signalling alone was not sufficient to initiate tumorigenesis (Fig. 7a). Targeted activation of BRAF to LGR5+ve cells resulted in decreased survival, with all mice killed 300 days post induction of the transgene by injection of tamoxifen (median survival 276 days). Although 6 out of 14 mice succumbed to adrenal tumours, 50% of these mice presented with papillomas consistent with LGR5 expression in murine skin (Fig. 7a,c). However, the long latency period suggests Braf mutation requires additional events to facilitate papilloma development. The combined targeted inactivation of one allele of Tgfbr1 reduced survival (median survival 231 days) and enhanced both the number of mice with skin lesions and the number of lesions per mouse (Fig. 7a,c and Supplementary Fig. 12a). Inactivation of both Tgfbr1 alleles significantly increased the number of tumours per mouse and dramatically shortened both skin tumour-free survival (all mice developing skin lesions within 63 days of induction) and overall survival (median survival 51 days) (Fig. 7a,c and Supplementary Fig. 12a). Phenotypically, these lesions appeared as differentiated papillomas in Tgfbr1 wild-type and heterozygous mice (Fig. 7c–e and Supplementary Fig. 12b). Remarkably, in the homozygous Tgfbr1fl/fl mice, tumours appeared as ulcerative cSCC (Fig. 7c and Supplementary Fig. 12b). Elegant work by the laboratories of Blanpain and colleagues11, and Jensen and colleagues10 have shown when Kras is targeted to skin stem cells, there is long latency to papilloma formation (similar to the BrafV600E allele described here) and most of these lesions form around areas associated with wounding. Targeted activation of Kras alone mainly failed to produce skin lesions; however, when we targeted inactivation of both alleles of Tgfbr1 and the KrasG12D mutation to the LGR5+ve compartment, mice developed rapid cSCC with kinetics comparable to BrafV600E mice (Fig. 7b,c). In addition, Keratin 1 (Fig. 7d) and Keratin 5 staining (Fig. 7e) revealed that cSCC lesions in both the Braf and Kras mice are poorly differentiated cSCC. Importantly, these lesions were highly proliferative (Supplementary Fig. 12c) and never progressed via a papillomatous stage, recapitulating the rapid cSCC onset observed in humans12,13. PO4-SMAD3 activity exhibited a dose-dependent reduction in tumours isolated from these mice, indicating loss of TGFβ signalling (Supplementary Fig. 12d,e). qRT–PCR analysis of these tumours revealed loss of Tgfbr1 expression (Supplementary Fig. 12f) without any significant change in ligand mRNA expression (Supplementary Fig. 13).
(a) Kaplan–Meier survival curve (left panel) of Lgr5CreER Tgfbr1fl/fl (n=12), Lgr5CreER BrafV600E(n=14), Lgr5CreER BrafV600E Tgfbr1fl/+ (n=23) and Lgr5CreER BrafV600E Tgfbr1fl/fl (n=26) mice (P≤0.0001 by log-rank, Mantel–Cox). Skin tumour-free survival curve (right panel) of Lgr5CreER BrafV600E (n=5), Lgr5CreER BrafV600E Tgfbr1fl/+ (n=10) and Lgr5CreER BrafV600E Tgfbr1fl/fl (n=19) mice. (b) Kaplan–Meier survival curve of Lgr5CreER KrasG12D (n=9) and Lgr5CreER KrasG12D Tgfbr1fl/fl (n=14) mice (P<0.0001 by log-rank, Mantel–Cox). (c) Macroscopic pictures of skin tumours from Lgr5CreER BrafV600E, Lgr5CreER BrafV600E Tgfbr1fl/+, Lgr5CreER BrafV600E Tgfbr1fl/fl and Lgr5CreER KrasG12D Tgfbr1fl/fl mice. (d) Representative staining of Keratin 1 (K1) and (e) Keratin 5 (K5) on Lgr5CreER BrafV600E, Lgr5CreER BrafV600E Tgfbr1fl/+, Lgr5CreER BrafV600E Tgfbr1fl/fl and Lgr5CreER KrasG12D Tgfbr1fl/fl mice. Scale bar, 100 μm.
Skin tissue compartmentalization has been recently proposed as a mechanism involved in tissue maintenance10. To test whether Tgfbr1 deletion perturbed such compartmentalization, we lineage traced LGR5+ve cells by intercrossing Lgr5CreER with the RosaLSL-RFP reporter mice (Supplementary Fig. 9e). We observed that red fluorescent protein (RFP)-positive cells were confined to the hair follicle39 and were never detected in the sebaceous gland, or interfollicular epidermis regions of Lgr5CreER BrafV600E or Lgr5CreER BrafV600ETgfbr1fl/+ mice, at early time points post induction (Supplementary Fig. 14). The cSCC arising within Lgr5CreER BrafV600ETgfbr1fl/fl mice were fully recombined and RFP positive. In the normal skin comparator for these tumours, but also at earlier time points, the LGR5+ve cells and their progeny were localized in their normal compartment (Supplementary Fig. 14). These results indicate that perturbation of TGFβ signalling is insufficient to disrupt compartmentalization, but acts as a tumour suppressor in LGR5+ve stem cells.
Given the infrequent coincident activation of RAS genes and mutational inactivation of TGFβ receptors in sporadic cSCC, we finally sought to model this disease by inactivating Tp53 function coupled with deletion of Tgfbr1 in LGR5+ve cells (Supplementary Fig. 9f). Knockin of mutant Tp53 (R172H) coupled with deletion of the wild-type allele had no discernible phenotype (Fig. 8a). Heterozygous knockin or deletion of Tp53 coupled with homozygous deletion of Tgfbr1 resulted in the emergence of skin tumours in a few mice (30% and 20%, respectively) with long latency. Combined knockin of mutant Tp53 with deletion of the wild-type allele of Tp53 coupled with deletion of Tgfbr1 resulted in skin tumour development in 81% of mice with increased tumour number at a shorter latency (Fig. 8a,b). These tumours exhibited loss of differentiation expressing low levels of Keratin 1 and higher levels of Keratin 5 (Fig. 8c).
(a) Kaplan–Meier survival curve of Lgr5CreER Tp53R172H/fl (n=31), Lgr5CreER Tp53R172/+ Tgfbr1fl/fl (n=10), Lgr5CreER Tp53fl/+ Tgfbr1fl/fl (n=5) and Lgr5CreER Tp53R172H/fl Tgfbr1fl/fl (n=11) mice (P≤0.0001 by log-rank, Mantel–Cox). (b) Tumour number of indicated genotypes. (c) Macroscopic picture of a Lgr5CreER Tp53R172H/fl Tgfbr1fl/fl mouse with a skin tumour. Representative staining of haematoxylin and eosin, Keratin 1 (K1) and Keratin 5 (K5) is shown. Scale bar, 100 μm.
Discussion
Recent studies have revealed an exceptionally high mutation burden (50 mutations per megabase of DNA6) in cSCC6,16,17. This rate is second only to that of the commonest skin malignancy basal cell carcinoma43. This translates to potentially thousands of mutations per tumour, providing a particular challenge in identifying driver mutations. This challenge is further compounded by varying efficiencies in deep-sequencing technologies and profound tumour heterogeneity2,6,44,45. Our studies here reveal that targeted deep sequencing using fluidigm PCR amplification and Roche 454 pyrosequencing can provide a robust platform to identify mutations in NOTCH1, NOTCH2 (ref. 6), TGFBR1 and TGFBR2 genes. This approach has also implicated alterations of NOTCH, TP53 and RAS in cSCC tumour development6,46. We further these studies by revealing mutation of TGFβ receptors in 43% of sporadic human cSCC and 28% of vemurafenib-induced skin lesions (Fig. 1). The prevalent tumour-initiating event in cSCC is ultraviolet-induced damage, which manifests as C-T and G-A transitions39. Approximately 68% of all nucleotide changes observed in our cSCC samples present with this signature6. Analysis of mutational signatures in TGFβ receptors reveals that 42% conform to an ultraviolet signature (Fig. 3b and Supplementary Data 13). This figure increases to 56.1% when scored as possibly damaging events via protein function prediction programmes (Fig. 3d and Supplementary Data 24). This indicates that ultraviolet damage may also be responsible for inactivation of TGFβ receptors. Mutation prediction programmes scored 53.5% of TGFBR1 and 71.1% of TGFBR2 receptor mutants as damaging, indicating that ∼20% of cSCC harbour TGFβ receptor inactivation (Supplementary Data 15 and 16, and Fig. 3c). Subsequent functional analysis of four TGFBR1 mutants and five TGFBR2 mutants indicated that half of the TGFBR1 mutants and all five TGFBR2 mutants were loss of function for canonical Smad signalling, and that tumours harbouring TGFβ receptor mutations had reduced PO4-SMAD3 activity (Fig. 4). Restoration of TGFBR2 expression to TGFBR2-null cell lines restricted cell proliferation (Fig. 4). Taken together, these findings indicate that loss of TGFβ tumour suppressor function is a common event in cSCC.
The assessment of VAFs provides an indication of the clonality of tumours and aids the potential identification of early driver mutations during tumour development2. We ranked mutational events by potential order of occurrence in the seven genes we have previsouly implicated in cSCC development and TGFβ receptors by measurement of VAF (Supplementary Data 15 and 16). These analyses indicate that potentially damaging mutations in TGFBR1 occur early in 25% of tumours harbouring these mutations and in 42% of tumours harbouring potentially damaging TGFBR2 mutations. Although this analysis is limited to the 9 genes studied in-depth here (but importantly including NOTCH genes previously identified as gatekeeper mutations in cSCC6), 11 samples exhibited TGFβ receptor mutations with the highest VAF, indicating that this could be an initiating event in the development of cSCC. Strong support for this hypothesis comes from ABSOLUTE clonality analysis of our WES samples, which revealed that 7/8 TGFβ receptor mutations were clonal and represent probable driver events in these lesions.
Several of our samples display VAFs of 10–20% for the TGFβ receptors, which is not too dissimilar to the VAFs of the other known cSCC tumour suppressors studied here, and probably reflects the heterogeneous nature of cSCC. It is however an intriguing possibility that in some cases low TGFβ receptor VAF may reflect spontaneous regression of TGFβ receptor mutant clones as observed in multiple self-healing squamous epithelioma patients who harbour germline mutations in TGFBR1 (ref. 47). TGFβ signalling has been demonstrated to play both positive and negative roles in cSCC development in various mouse models48 acting to limit tumour cell proliferation, and also to promote tumour-initiating capacity and drug resistance49. Effects are dependent on the timing of aberrant TGFβ signalling and the cooperating oncogenic driving events (reviewed in refs 48, 50, 51, 52). We provide evidence that a dose-dependent loss of TGFβ signalling drives tumour progression, emphasizing its role as a major tumour suppressor in the skin. Although the cell of origin in cSCC in humans remains poorly defined3, our observations indicate that tumours can initiate efficiently and rapidly from LGR5+ve stem cells compared with TA cells, and mutational modulation of two signalling pathways within this cellular compartment is sufficient to drive rapid progression directly to carcinoma, without the need for protracted tumour evolution. The kinetics of this event mimic exactly that observed during development of cutaneous lesions in RAF inhibitor-treated patients12,13. Intriguingly this rapid process requires MAPK pathway activation as targeted intereference with Tp53 function coupled with Tgfbr1 loss results in the development of skin tumours with long latency. Importantly, our studies revealed highly localized TGFβ signalling in Keratin 15+ve bulge stem cells in human telogen hair follicles, mirrored exquisitly the location of specific autocrine TGFβ signalling activity identified in LGR5+ve bulge stem cells of murine telogen hair follicles. It has been proposed that stem cell quiescence acts as a tumour suppresive mechanism in murine skin, and that LGR5+ve stem cells are refractory to oncogenic transformation53. Our data clearly indicate that oncogenic activation of the RAS/RAF/MAPK pathway, or p53 modulation, coupled with loss of TGFβ signalling, is capable of leading to tumour development from this compartment. As we demonstrate that mutational inactivation of TGFβ receptors is a frequent event in human cSCC, and that TGFβ signalling is highly localized to stem cells in normal skin, we propose that these cells represent a cell of origin for human cSCC. It remains possible that loss of TGFβ signalling may also contribute to cSCC development from other cell compartments in the skin and this warrants further investigation.
Our data, both in human and mice, indicate that TGFβ signalling inactivation can be an initiating event in sporadic cSCC. This is clearly the case in multiple self-healing squamous epithelioma where germline loss-of-function mutations in TGFBR1 have been identified as the underlying genetic lesion47. We speculate that activation of the RAS/RAF/MAPK pathway, or p53 modulation, may be a cooperating event in the development of this disease, and that these tumours may originate from the bulge stem cell compartment. TGFβ signalling inactivation may also occur following the acquisition of other driving mutational events and act as a limiting factor for tumour development. Intriguingly, initial clinical trials targeting systemic TGFβ inhibition with GC1008 (a pan-TGFβ neutralizing antibody) have also reported the occurrence of spontaneous cSCC as a side effect54. This provides further compelling support for the tumour suppressive role of TGFβ in skin carcinogenesis.
Methods
Samples
Ethical approval for this investigation was obtained from the East London and City Health Authority and the Tayside Tissue bank local ethics committee, and the study was conducted according to the Declaration of Helsinki Principles. All patients participating in this study were from dermatology and plastic surgery units in the United Kingdom and all provided written, informed consent. Punch biopsies of cSCC tissue were collected and processed as previously reported6. NHKs were isolated from normal skin samples according to previously published protocols55. Human tumour cell lines SCCIC1, SCCIC4, SCCIC8, SCCIC15, SCCIC12, SCCIC18, SCCIC19, SCCIC21, SCCT1, SCCT2, SCCT6, SCCT8, PM1, MET1, MET4, SCCT9, SCCT10, SCCT11, RDEBSCC2, RDEBSCC3, RDEBSCC4 and NTERT cells were established by our laboratories and were cultured as described15. TGFBR1-null MEFs30 and T47D cells (ATCC) were maintained in DMEM and RPMI medium supplemented with 10% FCS, respectively. Mycoplasma contamination checks were carried out on all cultures as routine and all lines were confirmed mycoplasma negative.
454 Sequencing
TGFBR1 and TGFBR2 primers were designed and validated by Fluidigm (Fluidigm Corporation, San Francisco, CA) as per recommended guidelines for Roche Titanium sequencing (Roche, Mannheim, Germany). Primers for NOTCH1, NOTCH2, TP53, CDKN2A, HRAS, KRAS and NRAS were previously described6 and all primer sequences are listed in Supplementary Data 27. Each primer included sample-specific Fluidigm 454 barcode primer and adapter sequences. Sequencing was performed in the same manner as our previous study56. Briefly, for thermal cycling a Fluidigm FC1 Cycler was used. The libraries were normalized and pooled before purification using Agencourt AMPure XP system (Beckman, UK). Library components were clonally amplified utilising the GSJunior emPCR Lib-A Kit (Roche) by inputting one molecule of library DNA per capture bead. Pyrosequencing was done using the GS Junior system (Roche/454 Life Sciences).
454 Variant analyses
Variant analysis was performed as previously described6. Briefly, reads were mapped to the hg19 build of the human genome using LASTZ via the public GALAXY instance and filtered to exclude those mapping to <100 loci using tools available through GALAXY.
Coding variants and splice site detection
Pileup files were generated and filtered using SAMTools57. Variants present in a single read or <10% of the total reads were excluded using a custom java programme available from https://github.com/mattsouth/laszt-variant-filter (last accessed 6 May 2013). Coding variants were called against the RefSeq gene list using the amino acid tool via the public GALAXY instance. Variants present in <3 reads were excluded. Variants present in >1 independent sample and adjacent to a homopolymer >3 bases were excluded, unless present in COSMIC58. Variants present in >30 samples were excluded, unless present in COSMIC. All variants present in the exome variant server database (http://evs.gs.washington.edu/EVS/) were excluded, unless present in COSMIC. Splice sites were called from the pileup variant list if present in >4 reads and within 2 bases of Refseq coding sequence using Excel (Microsoft Inc., CA).
WES data analysis
Twenty previously published cSCC whole exomes6 were re-analysed with the addition of ten new cSCC whole exomes with the overall mean coverage of 63 × (Supplementary Table 4), using a previous pipeline59. SNVs and short indels were identified using the Strelka pipeline60 with a minimum coverage of ten reads at the targeted sites. Annotation of somatic variants was performed using the Oncotator tool61. Mutations in our targeted genes were further identified across the 30 cSCC WES samples.
Copy number analysis using WES data
Two independent approaches were applied. First, to generate SNP and indel variant genotyping information, the tumour-normal pair was processed together against the reference genome using the VarScan2 germline variants calling method mpileup2cns62. The minimum coverage for identified sites was ten reads for both tumour and normal. Next, the logR and BAF (B-allele frequency) files were created based on the tumour-normal pair genotyping information, with the depth information normalized by dividing the depth of each variant by the median depth across all variants. The ASCAT R packages63 were then used to perform allele-specific copy number analysis, to identify copy number aberrations and loss-of-heterozygosity regions. The second approach was based on number of reads aligned to each exon between the tumour and normal pair. VarScan2 copy number calling method was first applied. Raw copy number calls were adjusted as previously reported64. Finally, results from the two approaches were cross-compared, to produce the final copy number aberrations and acquired uniparental disomy calls for targeted genes.
Identification of potential cancer drivers and significantly mutated pathways
Based on all mutations identified from the 30 cSCC WES data set, we used the IntOGen platform19 to identify significantly mutated genes and pathways, based on the significance (P-value) of the FM bias (that is, the bias towards the accumulation of mutations with high functional impact). The significantly mutated signalling pathways (based on the IntOGen Oncodrive-fm functional impact bias, FM bias P<0.05) were further selected (Supplementary Data 10). MutsigCV18 was also used to detect significant genes with point mutations above the background mutation rate.
Estimating the clonality of mutations
For the somatic mutations of TGFBR1/2, TP53, CDKN2A, NOTCH1/2 and RAS genes identified by WES (Supplementary Data 7), we further classified them as clonal or subclonal on the basis of the posterior probability that the cancer cell fraction exceeded 0.95 using ABSOLUTE20. Numbers of reads supporting the reference and alternative alleles were extracted and the copy number segmentation files were generated based on the DNAcopy CBS segments using WES data. Mutations with the somatic clonal probability >0.5 were classified as clonal with high confidence. Those mutations with clonal probability >0.25 but with very small subclonal probability scores were also called clonal (Supplementary Data 12). Tumour purity and ploidy were also estimated (Supplementary Data 11). For samples with TGFBR1/2 mutations, cancer cell fractions for TGFBR1/2 were further compared with those for other genes, to determine the clonality orders.
Functional prediction of mutations
A combination of four approaches were used to predict the functional impact of identified mutations by targeted sequencing, (i) SIFT21, which uses sequence homology and protein conservation to predict the effects of all possible substitutions at each position in the protein sequence; (ii) PolyPhen-2 (ref. 22), which predicts possible functional impact of an amino acid substitution on the structure and function level using physical and comparative considerations; (iii) Provean23, which predicts the damaging effects of SNVs and indels using a versatile alignment-based score; and (iv) Mutation Assessor24, which measures the functional impact scores for amino acid residue changes using evolutionary conservation patterns derived from aligned families and sub-families of sequence homologues within and between species. Mutations predicted as functional damaging by at least two of the four approaches were classified being potentially damaging/deleterious.
In vivo analyses
All experiments were performed under the UK Home Office guidelines. Mice were segregating for C57BL6J and S129 background. Alleles used throughout this study were: Lgr5-cre-ERT2(ref. 40), ShhCreER (ref. 38), BrafV600E (ref. 37), KrasG12D (ref. 65), Tgfbr1fl30, RosaLSL-RFP (ref. 66), Tp53fl (ref. 6767) and Tp53R172H (ref. 6868). A mix of males and females were used. Recombination in the Lgr5-cre-ERT2 mouse model was induced with one intraperitoneal injection of 3 mg Tamoxifen (Sigma) followed by one injection of 2 mg Tamoxifen for 3 days. Mice were induced post 7 weeks of age. Recombination in the ShhCreER mouse model was induced with one intraperitoneal injection of 2.5 mg Tamoxifen. Mice were induced post 28 days of age. For proliferation analysis, mice were injected with 250 μl of BrdU (Amersham Biosciences) 2 h before being sampled.
FACS analysis
Epidermis was prepared as previously described69. Briefly, fat was scraped from the mouse back and left at 37 °C in a dish (dermis down) in 0.25% of Trypsin/EDTA (Invitrogen, Carlsbad, CA) for 90 min. Epidermis was removed using a scalpel and dissociated by pipetting. Cells were filtered through a 40 μm strainer, centrifuged at 250 g for 5 min and washed with PE (PBS/EDTA). Cells were washed with 0.1% BSA/PE, centrifuged at 250 g for 5 min and used for Lgr5-GFP sorting.
Immunohistochemistry
IHC was performed on formalin-fixed skin sections. Standard IHC techniques were used throughout this study. Primary antibodies were as follows: TGFBR1 (Santa Cruz, V22, 1:100), PO4-SMAD3 (Abcam, EP823Y, (52903), 1:50), GFP (Abgent, 168AT1211, 1:100), Keratin 1 (Covance, AF109, 1:1,000), Keratin 5 (Covance, AF138, 1:4,000), Keratin 15 (Abcam, 80522 (LHK15), 1:1,000), KI67 (Thermo, RM-9106-S) and BrdU (BD Biosciences, 347580, 1:200). Mouse PO4-SMAD3 score was performed in a blinded manner. For each antibody, staining was performed on at least three mice of each genotype and at least six sections of normal human skin. Representative images are shown for each staining. PO4-SMAD3 antibody was optimized for IHC use using formalin-fixed paraffin-embedded SCCIC4 cells treated with and without recombinant TGFβ1 or the TGFBR1 kinase inhibitor SB-431542 (ref. 70) (Supplementary Fig. 5). PO4-SMAD3 IHC scoring was performed in a blinded manner using the histoscore method.
RNA isolation and quantitative PCR
RNA was isolated using a Qiagen RNeasy Mini Kit (Qiagen, Crawly, West Sussex, UK) according to the manufacturer’s instructions. DNA-free kit (Ambion/Applied Biosystems, Warrington, UK) was used to remove genomic DNA contamination according to the manufacturer’s instructions. One microgram of RNA was reverse transcribed to complementary DNA using a DyNAmo SYBR Green 2-step quantitative PCR kit (Finnzymes, Espoo, Finland) in a reaction volume of 20 μl. Glyceraldehyde 3-phosphate dehydrogenase (GAPDH) was used to normalize for differences in RNA input.
qRT–PCR primers
qRT–PCR primers were as follows. mTgfbr1 F-5′-TGCCATAACCGCACTGTCA-3′, mTgfbr1 R-5′-AATGAAAGGGCGATCTAGTGATG-3′; mTgfbr2 F-5′-CCGGAAGTTCTAGAATCCAG-3′, mTgfbr2 R-5′-TAATCCTTCACTTCTCCCAC-3′; mTgfb1 F-5′-AGCCCGAAGCGGACTACTAT-3′, mTgfb1 R-5′-TTCCACATGTTGCTCCACAC-3′; mTgfb2 F-5′-TTTAAGAGGGATCTTGGATGGA-3′, mTgfb2 R-5′-AGAATGGTCAGTGGTTCCAGAT-3′; mTgfb3 F-5′-CGCACAGAGCAGAGAATTGA-3′, mTgfb3 R-5′-GTGACATGGACAGTGGATGC-3′; mSmad7 F-5′-TCAAGAGGCTGTGTTGCTGT-3′, mSmad7 R-5′-TGGGTATCTGGAGTAAGGAGGA-3′; and mGapdh F-5′-GAAGGCCGGGGCCCACTTGA-3′, mGapdh R-5′-CTGGGTGGCAGTGATGGCATGG-3′.
Western blotting
Cells were lysed directly in 4 × SDS sample buffer at 60–80% confluence. Lysates were subjected to standard SDS–PAGE. Bands were detected using enhanced chemiluminescence solution (Amersham). Secondary antibodies used throughout were horseradish peroxidase-conjugated polyclonal goat anti-mouse Ig (Dako, P0448, 1:2,000) and horseradish peroxidase-conjugated polyclonal goat anti-rabbit Ig (Dako, P0260, 1:2,000). Primary antibodies were PO4-SMAD3 (Abcam, 52903, 1:1,000), SMAD3 (Cell Signaling, 9523, 1:1,000), TGFBR1 (Santa Cruz, 398 (V22), 1:500) and TGFBR2 (Santa Cruz, 17792, (E6), 1:500). For TGFBR2 western blottings, lysates were prepared directly from transfected cells using the Dual-luciferase cell lysis buffer (Promega, UK). For TGFBR1 western blottings, parallel transfections to the luciferase assays were performed and samples were lysed directly in 4 × SDS sample buffer. Original uncropped western blot scans are also provided (Supplementary Fig. 15).
Plasmids
The full-length wild-type human TGFBR1 and pathogenic mutants, amplified with BglII/NotI restriction sites, were shuttled into pCMV5 mammalian cell expression vectors onto the BamHI/NotI sites. The full-length wild-type human TGFBR2 and pathogenic mutants were sub-cloned into pCMV5 using the BamHI/NotI restriction sites. Site-directed mutagenesis was carried out using the QuickChange method (Stratagene) but substituting the Taq with KOD Hot Start DNA polymerase (Novagen). All DNA constructs were verified by DNA sequencing (by the DNA Sequencing Service at University of Dundee; www.dnaseq.co.uk). GFP expression plasmid was from Amaxa.
Transient transfection analysis
All transfections were performed in 24-well format in biological triplicate using LipofectAMINE 2000 (Invitrogen) according to the manufacturer’s instructions. Cells were transfected overnight with 100 ng of reporter gene (SMAD7-Promoter Luciferase or CAGA12-Luciferase) and 10 ng of internal Renilla luciferase control (pRL-TK, E2241, Promega) with empty vector (pCMV5, 211175, Stratagene), wild-type or mutant TGFβ receptor plasmids (range 150–300 ng). Recombinant human TGFβ1 (Peprotech) was dissolved in 4 mM HCL/1 mg ml−1 BSA and used at final concentration of 5 ng ml−1, and cells were treated for 4 h before harvest. Luciferase activities were measured using the Dual Luciferase assay (Promega) and firefly luciferase activity was normalized to Renilla luciferase activity.
Cell proliferation assays
Cells were seeded at a density of 500–1,000 cells per well of 96-well plates in keratinocyte media (RM+) without growth factors and incubated overnight. Cells were fed 50 μl of medium supplemented with treatment and controls every 2 days until harvest. All cultures were performed in sextuplet (n=6). Cells were assayed for proliferation using the CellTitreGlo Luminescent Cell Viability assay (NHKs) as per the manufacturer’s instructions (Promega; luminesence was measured on a Berthold Orion II microplate luminometer) or IncucyteZoom Live cell imager.
Data availability
The WES data for the 30 samples have been deposited in the European Genome-phenome Archive under accession code EGAS00001001892. The authors declare that all other relevant data supporting the findings of this study are available within the article and its Supplementary Information files. Additional information can be obtained from the corresponding authors (G.J.I. and O.J.S.).
Additional information
How to cite this article: Cammareri, P. et al. Inactivation of TGFβ receptors in stem cells drives cutaneous squamous cell carcinoma. Nat. Commun. 7:12493 doi: 10.1038/ncomms12493 (2016).
References
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Blanpain, C. Tracing the cellular origin of cancer. Nat. Cell Biol. 15, 126–134 (2013).
Barker, N. et al. Crypt stem cells as the cells-of-origin of intestinal cancer. Nature 457, 608–611 (2009).
Schwitalla, S. et al. Intestinal tumorigenesis initiated by dedifferentiation and acquisition of stem-cell-like properties. Cell 152, 25–38 (2013).
South, A. P. et al. NOTCH1 mutations occur early during cutaneous squamous cell carcinogenesis. J. Invest. Dermatol. 134, 2630–2638 (2014).
Stahl, P. L. et al. Sun-induced nonsynonymous p53 mutations are extensively accumulated and tolerated in normal appearing human skin. J. Invest. Dermatol. 131, 504–508 (2011).
Martincorena, I. et al. Tumor evolution. High burden and pervasive positive selection of somatic mutations in normal human skin. Science 348, 880–886 (2015).
Nelson, M. A., Futscher, B. W., Kinsella, T., Wymer, J. & Bowden, G. T. Detection of mutant Ha-ras genes in chemically initiated mouse skin epidermis before the development of benign tumors. Proc. Natl Acad. Sci. USA 89, 6398–6402 (1992).
Page, M. E., Lombard, P., Ng, F., Gottgens, B. & Jensen, K. B. The epidermis comprises autonomous compartments maintained by distinct stem cell populations. Cell Stem Cell 13, 471–482 (2013).
Lapouge, G. et al. Identifying the cellular origin of squamous skin tumors. Proc. Natl Acad. Sci. USA 108, 7431–7436 (2011).
Oberholzer, P. A. et al. RAS mutations are associated with the development of cutaneous squamous cell tumors in patients treated with RAF inhibitors. J. Clin. Oncol. 30, 316–321 (2012).
Su, F. et al. RAS mutations in cutaneous squamous-cell carcinomas in patients treated with BRAF inhibitors. N. Engl. J. Med. 366, 207–215 (2012).
Arnault, J. P. et al. Skin tumors induced by sorafenib; paradoxic RAS-RAF pathway activation and oncogenic mutations of HRAS, TP53, and TGFBR1. Clin. Cancer Res. 18, 263–272 (2012).
Purdie, K. J., Pourreyron, C. & South, A. P. Isolation and culture of squamous cell carcinoma lines. Methods Mol. Biol. 731, 151–159 (2011).
Pickering, C. R. et al. Mutational landscape of aggressive cutaneous squamous cell carcinoma. Clin. Cancer Res. 20, 6582–6592 (2014).
Lee, C. S. et al. Recurrent point mutations in the kinetochore gene KNSTRN in cutaneous squamous cell carcinoma. Nat. Genet. 46, 1060–1062 (2014).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Gonzalez-Perez, A. et al. IntOGen-mutations identifies cancer drivers across tumor types. Nat. Methods 10, 1081–1082 (2013).
Carter, S. L. et al. Absolute quantification of somatic DNA alterations in human cancer. Nat. Biotechnol. 30, 413–421 (2012).
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Choi, Y., Sims, G. E., Murphy, S., Miller, J. R. & Chan, A. P. Predicting the functional effect of amino acid substitutions and indels. PLoS ONE 7, e46688 (2012).
Reva, B., Antipin, Y. & Sander, C. Predicting the functional impact of protein mutations: application to cancer genomics. Nucleic Acids Res. 39, e118 (2011).
Groppe, J. et al. Cooperative assembly of TGF-beta superfamily signaling complexes is mediated by two disparate mechanisms and distinct modes of receptor binding. Mol. Cell 29, 157–168 (2008).
De Crescenzo, G. et al. Three key residues underlie the differential affinity of the TGFbeta isoforms for the TGFbeta type II receptor. J. Mol. Biol. 355, 47–62 (2006).
Massague, J. TGFbeta signalling in context. Nat. Rev. Mol. Cell. Biol. 13, 616–630 (2012).
Ross, S. & Hill, C. S. How the Smads regulate transcription. Int. J. Biochem. Cell Biol. 40, 383–408 (2008).
Massague, J., Seoane, J. & Wotton, D. Smad transcription factors. Genes Dev. 19, 2783–2810 (2005).
Larsson, J. et al. Abnormal angiogenesis but intact hematopoietic potential in TGF-beta type I receptor-deficient mice. EMBO J. 20, 1663–1673 (2001).
Denissova, N. G., Pouponnot, C., Long, J., He, D. & Liu, F. Transforming growth factor beta -inducible independent binding of SMAD to the Smad7 promoter. Proc. Natl Acad. Sci. USA 97, 6397–6402 (2000).
Dennler, S. et al. Direct binding of Smad3 and Smad4 to critical TGF beta-inducible elements in the promoter of human plasminogen activator inhibitor-type 1 gene. EMBO J. 17, 3091–3100 (1998).
Heidorn, S. J. et al. Kinase-dead BRAF and oncogenic RAS cooperate to drive tumor progression through CRAF. Cell 140, 209–221 (2010).
Poulikakos, P. I., Zhang, C., Bollag, G., Shokat, K. M. & Rosen, N. RAF inhibitors transactivate RAF dimers and ERK signalling in cells with wild-type BRAF. Nature 464, 427–430 (2010).
Hatzivassiliou, G. et al. RAF inhibitors prime wild-type RAF to activate the MAPK pathway and enhance growth. Nature 464, 431–435 (2010).
Holderfield, M. et al. RAF inhibitors activate the MAPK pathway by relieving inhibitory autophosphorylation. Cancer Cell 23, 594–602 (2013).
Mercer, K. et al. Expression of endogenous oncogenic V600EB-raf induces proliferation and developmental defects in mice and transformation of primary fibroblasts. Cancer Res. 65, 11493–11500 (2005).
Harfe, B. D. et al. Evidence for an expansion-based temporal Shh gradient in specifying vertebrate digit identities. Cell 118, 517–528 (2004).
Jaks, V. et al. Lgr5 marks cycling, yet long-lived, hair follicle stem cells. Nat. Genet. 40, 1291–1299 (2008).
Barker, N. et al. Identification of stem cells in small intestine and colon by marker gene Lgr5. Nature 449, 1003–1007 (2007).
Oshimori, N. & Fuchs, E. Paracrine TGF-beta signaling counterbalances BMP-mediated repression in hair follicle stem cell activation. Cell Stem Cell 10, 63–75 (2012).
Nakao, A. et al. Identification of Smad7, a TGFbeta-inducible antagonist of TGF-beta signalling. Nature 389, 631–635 (1997).
Jayaraman, S. S., Rayhan, D. J., Hazany, S. & Kolodney, M. S. Mutational landscape of basal cell carcinomas by whole-exome sequencing. J. Invest. Dermatol. 134, 213–220 (2014).
Hudson, A. M. et al. Discrepancies in cancer genomic sequencing highlight opportunities for driver mutation discovery. Cancer Res. 74, 6390–6396 (2014).
Helleday, T., Eshtad, S. & Nik-Zainal, S. Mechanisms underlying mutational signatures in human cancers. Nat. Rev. Genet. 15, 585–598 (2014).
Wang, N. J. et al. Loss-of-function mutations in Notch receptors in cutaneous and lung squamous cell carcinoma. Proc. Natl Acad. Sci. USA 108, 17761–17766 (2011).
Goudie, D. R. et al. Multiple self-healing squamous epithelioma is caused by a disease-specific spectrum of mutations in TGFBR1. Nat. Genet. 43, 365–369 (2011).
Glick, A. B. The role of TGFbeta signaling in squamous cell cancer: lessons from mouse models. J. Skin Cancer 2012, 249063 (2012).
Oshimori, N., Oristian, D. & Fuchs, E. TGF-beta promotes heterogeneity and drug resistance in squamous cell carcinoma. Cell 160, 963–976 (2015).
White, R. A. et al. Epithelial stem cell mutations that promote squamous cell carcinoma metastasis. J. Clin. Invest. 123, 4390–4404 (2013).
Connolly, E. C. & Akhurst, R. J. The complexities of TGF-beta action during mammary and squamous cell carcinogenesis. Curr. Pharm. Biotechnol. 12, 2138–2149 (2011).
Oshimori, N. & Fuchs, E. The harmonies played by TGF-beta in stem cell biology. Cell Stem Cell 11, 751–764 (2012).
White, A. C. et al. Stem cell quiescence acts as a tumour suppressor in squamous tumours. Nat. Cell Biol. 16, 99–107 (2014).
Morris, J. C. et al. Phase I study of GC1008 (fresolimumab): a human anti-transforming growth factor-beta (TGFbeta) monoclonal antibody in patients with advanced malignant melanoma or renal cell carcinoma. PLoS ONE 9, e90353 (2014).
Rheinwald, J. G. & Green, H. Serial cultivation of strains of human epidermal keratinocytes: the formation of keratinizing colonies from single cells. Cell 6, 331–343 (1975).
Kluk, M. J. et al. Gauging NOTCH1 activation in cancer using immunohistochemistry. PLoS ONE 8, e67306 (2013).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Forbes, S. A. et al. the catalogue of somatic mutations in cancer (COSMIC). Curr. Protoc. Hum. Genet. Chapter 10, Unit 10–Unit 11 (2008).
Okosun, J. et al. Integrated genomic analysis identifies recurrent mutations and evolution patterns driving the initiation and progression of follicular lymphoma. Nat. Genet. 46, 176–181 (2014).
Saunders, C. T. et al. Strelka: accurate somatic small-variant calling from sequenced tumor-normal sample pairs. Bioinformatics 28, 1811–1817 (2012).
Lo Nigro, C. et al. Methylated tissue factor pathway inhibitor 2 (TFPI2) DNA in serum is a biomarker of metastatic melanoma. J. Invest. Dermatol. 133, 1278–1285 (2013).
Koboldt, D. C. et al. VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res. 22, 568–576 (2012).
Van Loo, P. et al. Allele-specific copy number analysis of tumors. Proc. Natl Acad. Sci. USA 107, 16910–16915 (2010).
Okosun, J. et al. Recurrent mTORC1-activating RRAGC mutations in follicular lymphoma. Nat. Genet. 48, 183–188 (2016).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
Luche, H., Weber, O., Nageswara Rao, T., Blum, C. & Fehling, H. J. Faithful activation of an extra-bright red fluorescent protein in “knock-in” Cre-reporter mice ideally suited for lineage tracing studies. Eur. J. Immunol. 37, 43–53 (2007).
Jonkers, J. et al. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nature Genetics. 29, 418–425 (2001).
Olive, K. P. et al. Mutant p53 gain of function in two mouse models of Li-Fraumeni syndrome. Cell. 119, 847–860 (2004).
Nowak, J. A. & Fuchs, E. Isolation and culture of epithelial stem cells. Methods Mol. Biol. 482, 215–232 (2009).
Inman, G. J. et al. SB-431542 is a potent and specific inhibitor of transforming growth factor-beta superfamily type I activin receptor-like kinase (ALK) receptors ALK4, ALK5, and ALK7. Mol. Pharmacol. 62, 65–74 (2002).
Acknowledgements
O.S. is supported by a Cancer Research UK core grant (A21139) and an ERC starting grant (311301). P.C. is supported by FP7 Health CP-IP - Large-scale integrating project grant (278568). D.F.V. is supported by ERC Starting grant (311301). A.M.R. is supported by Cancer Research UK centre grant (A12976). L.C.S. was supported by WWCR grant 11-0788. G.J.I. was supported by WWCR fellowship (03-0900). G.J.I., I.M.L., C.M.P., C.A.H., K.J.P., A.M., C.P. and A.P.S. were supported by a Cancer Research UK programme grant (A13044) and an ERC grant (250170). J.W., A.N. and C.C. were supported by a Cancer Research UK centre award to Barts Cancer Institute. We thank the research and scientific services at the CRUK Beatson Institute in general.
Author information
Authors and Affiliations
Contributions
O.J.S., G.J.I., P.C., A.M.R., D.F.V., J.W. and A.P.S. contributed to study design. P.C., A.M.R., D.F.V., S.L. and D.A. contributed to the data acquisition. P.C., A.M.R., D.F.V., J.W., A.N., S.L., R.A.R., D.A., P.J.C.V., A.M., C.P., J.H.S.D., J.L., S.W., L.C.S., G.P.S., K.J.P., C.M.P., C.A.H., I.R.L., H.C., N.B., S.K., C.P., R.M., C.C., A.P.S., O.J.S. and G.J.I. contributed to the data analysis and interpretation of the data. O.J.S., G.J.I., P.C., A.M.R., D.F.V., A.P.S. and J.W. contributed to drafting the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-15 and Supplementary Tables 1-4. (PDF 1508 kb)
Supplementary Data 1
TGFBR1 and TGFBR2 mutations in Vemurafenib-induced lesions by 454 pyrosequencing. (XLSX 11 kb)
Supplementary Data 2
454 pyrosequencing in Vemurafenib-induced lesions. 454 pyrosequencing data of NOTCH1, NOTCH2, TP53, CDKN2A, HRAS, NRAS and KRAS in vemurafenib-induced lesions (adapted from South et al, 2014). (XLSX 20 kb)
Supplementary Data 3
TGFBR1 and TGFBR2 mutations in sporadic cSCC primary tumors and cell lines by 454 pyrosequencing. TGW = Well Differentiated SCC, TGM = Moderately Differentiated SCC, TGP = Poorly Differentiated SCC. (XLSX 18 kb)
Supplementary Data 4
454 pyrosequencing in sporadic cSCC. 454 pyrosequencing data of NOTCH1, NOTCH2, TP53, CDKN2A, HRAS, NRAS and KRAS in sporadic cSCC primary tumors and cell lines (adapted from South et al, 2014). TGW = Well Differentiated SCC, TGM = Moderately Differentiated SCC, TGP = Poorly Differentiated SCC. (XLSX 62 kb)
Supplementary Data 5
TGFBR1 and TGFBR2 mutations in sporadic cSCC by 454 sequencing-Dundee Cohort. 454 pyrosequencing data of NOTCH1, NOTCH2, TP53, CDKN2A, HRAS, NRAS, KRAS, TGFBR1 and TGFBR2 in sporadic cSCC primary tumors and matched normal distant and perilesional skin isolated in Dundee. (XLSX 15 kb)
Supplementary Data 6
Statistical analysis of TGFBR mutations. Tumour Vs Normal VAF in 454 sequencing for a) TGFBR1, b) TGFBR2 and c) Combined. (XLSX 13 kb)
Supplementary Data 7
Somatic mutations of TGFBR1/2, TP53, CDKN2A, NOTCH1/2 and RAS genes across 30 cSCC exomes. The copy number / LOH status for each mutation is also shown. (XLSX 17 kb)
Supplementary Data 8
Statistical analysis of WES mutations. Tumour Vs Normal VAF in whole exome sequencing for TGFBR1/2, TP53, CDKN2A, NOTCH1/2 and RAS genes across 30 cSCC exomes, and associated p-values from two-sided fisher's exact test based on numbers of reads for mut and wt alleles between tumor and normal samples. (XLSX 17 kb)
Supplementary Data 9
MutsigCV and IntOgen analysis of selected candidate driver genes. (XLSX 10 kb)
Supplementary Data 10
ntogen Mutated Signalling pathways. (XLSX 9 kb)
Supplementary Data 11
ABSOLUTE estimates of purity and ploidy from exome-seq data. (XLSX 8 kb)
Supplementary Data 12
ABSOLUTE analysis for targeted mutations based on exome-seq. (XLSX 17 kb)
Supplementary Data 13
VAF analysis. (XLSX 114 kb)
Supplementary Data 14
VAF and UV analysis. (XLSX 8 kb)
Supplementary Data 15
PolyPhen-2, SIFT, Provean and Mutation assessor mutational impact (upper table) and ranking analysis of TGFBR1 mutations isolated from all samples (lower left table). The nature of the base change and its characteristic of a UV signature (C-T or G-A transitions) or a T-C or A-G signature are shown. Variant allele frequencies (% read) are shown, as is the ranking of the mutation according to variant allele frequency of NOTCH1, NOTCH2, TP53, CDKN2A, HRAS, NRAS, KRAS, TGFBR1 and TGFBR2 mutations identified in each tumour with rank 1 indicating the highest variant allele frequency. The total number of mutations in the 9 genes is also shown. Right hand lower table shows average (AV) read %, average rank and average number of mutations for total samples and samples separated by damaging or non-damaging prediction. The % of mutations exhibiting a UV signature or a T-C + A-G signature and ranked 1 or 2, ranked > median are also shown. The total number of TGFb receptor mutations ranked 1 is also indicated. (XLSX 22 kb)
Supplementary Data 16
Potential functional consequence of TGFBR2 mutations. PolyPhen-2, SIFT, Provean and Mutation assessor mutational impact (upper table) and ranking analysis of TGFBR2 mutations isolated from all samples (lower left table). The nature of the base change and its characteristic of a UV signature (C-T or G-A transitions) or a T-C or A-G signature are shown. Variant allele frequencies (% read) are shown, as is the ranking of the mutation according to variant allele frequency of NOTCH1, NOTCH2, TP53, CDKN2A, HRAS, NRAS, KRAS, TGFBR1 and TGFBR2 mutations identified in each tumour with rank 1 indicating the highest variant allele frequency. The total number of mutations in the 9 genes is also shown. Right hand lower table shows average (AV) read %, average rank and average number of mutations for total samples and samples separated by damaging or non-damaging prediction. The % of mutations exhibiting a UV signature or a T-C + A-G signature and ranked 1 or 2, ranked > median are also shown. The total number of TGFb receptor mutations ranked 1 is also indicated. (XLSX 23 kb)
Supplementary Data 17
Potential functional consequence of NOTCH1 mutations. PolyPhen-2, SIFT, Provean and Mutation assessor mutational impact analysis of NOTCH1 mutations. (XLSX 39 kb)
Supplementary Data 18
Potential functional consequence of NOTCH2 mutations. PolyPhen-2, SIFT, Provean and Mutation assessor mutational impact analysis of NOTCH2 mutations. (XLSX 27 kb)
Supplementary 19
Potential functional consequence of CDKN2A mutations. PolyPhen-2, SIFT and Provean mutational impact analysis of CDKN2A mutations. (XLSX 14 kb)
Supplementary Data 20
Potential functional consequence of TP53 mutations. PolyPhen-2, SIFT, Provean and Mutation assessor mutational impact analysis of TP53 mutations. (XLSX 21 kb)
Supplementary Data 21
Potential functional consequence of HRAS mutations. PolyPhen-2, SIFT, Provean and Mutation assessor mutational impact analysis of HRAS mutations. (XLSX 14 kb)
Supplementary Data 22
Potential functional consequence of NRAS mutations. PolyPhen-2, SIFT, Provean and Mutation assessor mutational impact analysis of NRAS mutations.. (XLSX 11 kb)
Supplementary Data 23
Potential functional consequence of KRAS mutations. PolyPhen-2, SIFT, Provean and Mutation assessor mutational impact analysis of HRAS KRAS mutations. (XLSX 12 kb)
Supplementary Data 24
UV versus damaging analysis. (XLSX 110 kb)
Supplementary Data 25
Damaging and Non Damaging. (XLSX 8 kb)
Supplementary Data 26
PO4-SMAD3 Histoscores of human SCC tumour panel: Wild-type (WT) Vs Mutant (MT) (XLSX 9 kb)
Supplementary Data 27
454 pyrosequencing primers. (XLSX 53 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Cammareri, P., Rose, A., Vincent, D. et al. Inactivation of TGFβ receptors in stem cells drives cutaneous squamous cell carcinoma. Nat Commun 7, 12493 (2016). https://doi.org/10.1038/ncomms12493
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms12493
- Springer Nature Limited
This article is cited by
-
Driver gene combinations dictate cutaneous squamous cell carcinoma disease continuum progression
Nature Communications (2023)
-
Bifunctional anti-PD-L1/TGF-βRII agent SHR-1701 in advanced solid tumors: a dose-escalation, dose-expansion, and clinical-expansion phase 1 trial
BMC Medicine (2022)
-
Emerging precision diagnostics in advanced cutaneous squamous cell carcinoma
npj Precision Oncology (2022)
-
miR-22 promotes stem cell traits via activating Wnt/β-catenin signaling in cutaneous squamous cell carcinoma
Oncogene (2021)
-
Loss of retinoic acid receptor-related receptor alpha (Rorα) promotes the progression of UV-induced cSCC
Cell Death & Disease (2021)