Abstract
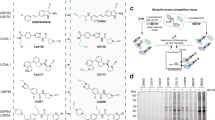
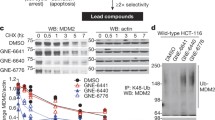
Given the importance of ubiquitin-specific protease 7 (USP7) in oncogenic pathways, identification of USP7 inhibitors has attracted considerable interest. Despite substantial efforts, however, the development of validated deubiquitinase (DUB) inhibitors that exhibit drug-like properties and a well-defined mechanism of action has proven particularly challenging. In this article, we describe the identification, optimization and detailed characterization of highly potent (IC50 < 10 nM), selective USP7 inhibitors together with their less active, enantiomeric counterparts. We also disclose, for the first time, co-crystal structures of a human DUB enzyme complexed with small-molecule inhibitors, which reveal a previously undisclosed allosteric binding site. Finally, we report the identification of cancer cell lines hypersensitive to USP7 inhibition (EC50 < 30 nM) and demonstrate equal or superior activity in these cell models compared to clinically relevant MDM2 antagonists. Overall, these findings demonstrate the tractability and druggability of DUBs, and provide important tools for additional target validation studies.



Similar content being viewed by others
References
Yau, R. & Rape, M. The increasing complexity of the ubiquitin code. Nat. Cell Biol. 18, 579–586 (2016).
Huang, X. & Dixit, V.M. Drugging the undruggables: exploring the ubiquitin system for drug development. Cell Res. 26, 484–498 (2016).
Zheng, Q. et al. Dysregulation of ubiquitin-proteasome system in neurodegenerative diseases. Front. Aging Neurosci. 8, 303 (2016).
Zhang, W. & Sidhu, S.S. Development of inhibitors in the ubiquitination cascade. FEBS Lett. 588, 356–367 (2014).
Merin, N.M. & Kelly, K.R. Clinical use of proteasome inhibitors in the treatment of multiple myeloma. Pharmaceuticals (Basel) 8, 1–20 (2014).
Sheridan, C. Drug makers target ubiquitin proteasome pathway anew. Nat. Biotechnol. 33, 1115–1117 (2015).
Abdul Rehman, S.A. et al. MINDY-1 is a Member of an evolutionarily conserved and structurally distinct new family of deubiquitinating enzymes. Mol. Cell 63, 146–155 (2016).
Clague, M.J. et al. Deubiquitylases from genes to organism. Physiol. Rev. 93, 1289–1315 (2013).
Ye, Y., Scheel, H., Hofmann, K. & Komander, D. Dissection of USP catalytic domains reveals five common insertion points. Mol. Biosyst. 5, 1797–1808 (2009).
Pal, A., Young, M.A. & Donato, N.J. Emerging potential of therapeutic targeting of ubiquitin-specific proteases in the treatment of cancer. Cancer Res. 74, 4955–4966 (2014).
Heideker, J. & Wertz, I.E. DUBs, the regulation of cell identity and disease. Biochem. J. 465, 1–26 (2015).
D'Arcy, P., Wang, X. & Linder, S. Deubiquitinase inhibition as a cancer therapeutic strategy. Pharmacol. Ther. 147, 32–54 (2015).
Ratia, K. et al. A noncovalent class of papain-like protease/deubiquitinase inhibitors blocks SARS virus replication. Proc. Natl. Acad. Sci. USA 105, 16119–16124 (2008).
Nicholson, B. & Suresh Kumar, K.G. The multifaceted roles of USP7: new therapeutic opportunities. Cell Biochem. Biophys. 60, 61–68 (2011).
Cummins, J.M. et al. Tumour suppression: disruption of HAUSP gene stabilizes p53. Nature 428, 1, 486 10.1038/nature02501 (2004).
Kon, N. et al. Inactivation of HAUSP in vivo modulates p53 function. Oncogene 29, 1270–1279 (2010).
Tavana, O. & Gu, W. Modulation of the p53/MDM2 interplay by HAUSP inhibitors. J. Mol. Cell Biol. 9, 45–52 (2017).
Song, M.S. et al. The deubiquitinylation and localization of PTEN are regulated by a HAUSP-PML network. Nature 455, 813–817 (2008).
Wu, H.T. et al. K63-polyubiquitinated HAUSP deubiquitinates HIF-1α and dictates H3K56 acetylation promoting hypoxia-induced tumour progression. Nat. Commun. 7, 13644 (2016).
Tavana, O. et al. HAUSP deubiquitinates and stabilizes N-Myc in neuroblastoma. Nat. Med. 22, 1180–1186 (2016).
Wang, Q. et al. Stabilization of histone demethylase PHF8 by USP7 promotes breast carcinogenesis. J. Clin. Invest. 126, 2205–2220 (2016).
Zhou, Z. et al. Deubiquitination of Ci/Gli by Usp7/HAUSP Regulates Hedgehog Signaling. Dev. Cell 34, 58–72 (2015).
van der Horst, A. et al. FOXO4 transcriptional activity is regulated by monoubiquitination and USP7/HAUSP. Nat. Cell Biol. 8, 1064–1073 (2006).
van Loosdregt, J. et al. Stabilization of the transcription factor Foxp3 by the deubiquitinase USP7 increases Treg-cell-suppressive capacity. Immunity 39, 259–271 (2013).
Hall, J.A., Tabata, M., Rodgers, J.T. & Puigserver, P. USP7 attenuates hepatic gluconeogenesis through modulation of FoxO1 gene promoter occupancy. Mol. Endocrinol. 28, 912–924 (2014).
Jäger, W. et al. The ubiquitin-specific protease USP7 modulates the replication of Kaposi's sarcoma-associated herpesvirus latent episomal DNA. J. Virol. 86, 6745–6757 (2012).
Meredith, M., Orr, A. & Everett, R. Herpes simplex virus type 1 immediate-early protein Vmw110 binds strongly and specifically to a 135-kDa cellular protein. Virology 200, 457–469 (1994).
Holowaty, M.N. et al. Protein profiling with Epstein-Barr nuclear antigen-1 reveals an interaction with the herpesvirus-associated ubiquitin-specific protease HAUSP/USP7. J. Biol. Chem. 278, 29987–29994 (2003).
Kemp, M. Recent Advances in the Discovery of Deubiquitinating Enzyme Inhibitors. Prog. Med. Chem. 55, 149–192 (2016).
Chen, C. et al. Synthesis and biological evaluation of thiazole derivatives as novel USP7 inhibitors. Bioorg. Med. Chem. Lett. 27, 845–849 (2017).
Weinstock, J. et al. Selective dual Inhibitors of the cancer-related deubiquitylating proteases USP7 and USP47. ACS Med. Chem. Lett. 3, 789–792 (2012).
Reverdy, C. et al. Discovery of specific inhibitors of human USP7/HAUSP deubiquitinating enzyme. Chem. Biol. 19, 467–477 (2012).
Colland, F. et al. Small-molecule inhibitor of USP7/HAUSP ubiquitin protease stabilizes and activates p53 in cells. Mol. Cancer Ther. 8, 2286–2295 (2009).
Altun, M. et al. Activity-based chemical proteomics accelerates inhibitor development for deubiquitylating enzymes. Chem. Biol. 18, 1401–1412 (2011).
Tian, X. et al. Characterization of selective ubiquitin and ubiquitin-like protease inhibitors using a fluorescence-based multiplex assay format. Assay Drug Dev. Technol. 9, 165–173 (2011).
Wu, J. et al. Chemical approaches to intervening in ubiquitin specific protease 7 (USP7) function for oncology and immune oncology therapies. J. Med. Chem. http://dx.doi.org/10.1021/acs.jmedchem.7b00498 (2017).
Ndubaku, C. & Tsui, V. Inhibiting the deubiquitinating enzymes (DUBs). J. Med. Chem. 58, 1581–1595 (2015).
Lee, J.G., Baek, K., Soetandyo, N. & Ye, Y. Reversible inactivation of deubiquitinases by reactive oxygen species in vitro and in cells. Nat. Commun. 4, 1568 (2013).
Wrigley, J.D. et al. Enzymatic characterisation of USP7 deubiquitinating activity and inhibition. Cell Biochem. Biophys. 60, 99–111 (2011).
Winter, A. et al. Biophysical and computational fragment-based approaches to targeting protein-protein interactions: applications in structure-guided drug discovery. Q. Rev. Biophys. 45, 383–426 (2012).
Kessler, B.M. Selective and reversible inhibitors of ubiquitin-specific protease 7: a patent evaluation (WO2013030218). Expert Opin. Ther. Pat. 24, 597–602 (2014).
Lor, L.A. et al. A simple assay for detection of small-molecule redox activity. J. Biomol. Screen. 12, 881–890 (2007).
Ritorto, M.S. et al. Screening of DUB activity and specificity by MALDI-TOF mass spectrometry. Nat. Commun. 5, 4763 (2014).
Tovar, C. et al. MDM2 small-molecule antagonist RG7112 activates p53 signaling and regresses human tumors in preclinical cancer models. Cancer Res. 73, 2587–2597 (2013).
Wang, S. et al. SAR405838: an optimized inhibitor of MDM2-p53 interaction that induces complete and durable tumor regression. Cancer Res. 74, 5855–5865 (2014).
Cummins, J.M. & Vogelstein, B. HAUSP is required for p53 destabilization. Cell Cycle 3, 689–692 (2004).
Kategaya, L. et al. USP7 small-molecule inhibitors interfere with ubiquitin binding. Nature 550, 534–538 (2017).
Turnbull, A.P. et al. Molecular basis of USP7 inhibition by selective small-molecule inhibitors. Nature 550, 481–486 (2017).
Lamberto, I. et al. Structure-guided development of a potent and selective non-covalent active-site inhibitor of USP7. Cell Chem. Biol. http://dx.doi.org/10.1016/j.chembiol.2017.09.003 (2017).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Bochevarov, A.D. et al. Jaguar: A high-performance quantum chemistry software program with strengths in life and materials sciences. Int. J. Quantum Chem. 113, 2110–2142 (2013).
Acknowledgements
The authors thank collaborators at Ubiquigent (Dundee, UK), Boston Biochem (Boston, US) and Beactica (Uppsala, Sweden) for their contributions, as well as Crelux GmbH (Martinsried, Germany) for solving the crystal structures and C. Scott and J. Burrows (Queen's University, Belfast) for helpful discussions and for providing plasmids, respectively. This study was supported by the Almac Group, the European Regional Development Fund and Invest Northern Ireland (Grant RD1010668).
Author information
Authors and Affiliations
Contributions
T.H. conceived the concept and directed the research. C.R.O'D. and G.G. helped develop the concept and designed and supervised medicinal chemistry and biology experiments. M.D.H., E.A., J.F. and H.M. carried out the design, synthesis and characterization of compounds. A.D. performed surface plasmon resonance experiments. C.H., K.M., E.O., E.C., A.D. and N.P. carried out compound screening, target validation and biochemical and cellular profiling studies. O.B. carried out computational modeling and structural analysis. T.H. and G.G. wrote the manuscript with input from other authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–3 and Supplementary Figures 1–16 (PDF 3747 kb)
Supplementary Note
Synthetic procedures (PDF 877 kb)
Rights and permissions
About this article
Cite this article
Gavory, G., O'Dowd, C., Helm, M. et al. Discovery and characterization of highly potent and selective allosteric USP7 inhibitors. Nat Chem Biol 14, 118–125 (2018). https://doi.org/10.1038/nchembio.2528
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2528
- Springer Nature America, Inc.
This article is cited by
-
Rolapitant treats lung cancer by targeting deubiquitinase OTUD3
Cell Communication and Signaling (2024)
-
Structural basis for the bi-specificity of USP25 and USP28 inhibitors
EMBO Reports (2024)
-
Pharmaceutical targeting of OTUB2 sensitizes tumors to cytotoxic T cells via degradation of PD-L1
Nature Communications (2024)
-
Backbone 1H, 13C and 15N resonance assignment of the ubiquitin specific protease 7 catalytic domain (residues 208–554) in complex with a small molecule ligand
Biomolecular NMR Assignments (2024)
-
Discovery of an OTUD3 inhibitor for the treatment of non-small cell lung cancer
Cell Death & Disease (2023)