Abstract
Ca2+ antagonist drugs are widely used in therapy of cardiovascular disorders1,2. Three chemical classes of drugs bind to three separate, but allosterically interacting, receptor sites on CaV1.2 channels, the most prominent voltage-gated Ca2+ (CaV) channel type in myocytes in cardiac and vascular smooth muscle3,4,5,6,7,8,9. The 1,4-dihydropyridines are used primarily for treatment of hypertension and angina pectoris and are thought to act as allosteric modulators of voltage-dependent Ca2+ channel activation, whereas phenylalkylamines and benzothiazepines are used primarily for treatment of cardiac arrhythmias and are thought to physically block the pore1,2. The structural basis for the different binding, action, and therapeutic uses of these drugs remains unknown. Here we present crystallographic and functional analyses of drug binding to the bacterial homotetrameric model CaV channel CaVAb, which is inhibited by dihydropyridines and phenylalkylamines with nanomolar affinity in a state-dependent manner. The binding site for amlodipine and other dihydropyridines is located on the external, lipid-facing surface of the pore module, positioned at the interface of two subunits. Dihydropyridine binding allosterically induces an asymmetric conformation of the selectivity filter, in which partially dehydrated Ca2+ interacts directly with one subunit and blocks the pore. In contrast, the phenylalkylamine Br-verapamil binds in the central cavity of the pore on the intracellular side of the selectivity filter, physically blocking the ion-conducting pathway. Structure-based mutations of key amino-acid residues confirm drug binding at both sites. Our results define the structural basis for binding of dihydropyridines and phenylalkylamines at their distinct receptor sites on CaV channels and offer key insights into their fundamental mechanisms of action and differential therapeutic uses in cardiovascular diseases.




Similar content being viewed by others
Accession codes
Primary accessions
Protein Data Bank
Data deposits
The coordinates and structure factors have been deposited in the Protein Data Bank with the following accession codes 5KLB (CavAb, 5 mM Ca2+ 2.7Å; 5KLG (CavAb-W195Y-UK-59811, 5 mM Ca2+); 5KLS (CavAb-UK-59811, 5 mM Ca2+); 5KMD (CavAb-W195Y-amlodipine, 5 mM Ca2+); 5KMF (CavAb-W195Y nimodipine, 5 mM Ca2+); and 5KMH (CavAb-Br-verapamil, 5 mM Ca2+).
References
Zamponi, G. W., Striessnig, J., Koschak, A. & Dolphin, A. C. The physiology, pathology, and pharmacology of voltage-gated calcium channels and their future therapeutic potential. Pharmacol. Rev. 67, 821–870 (2015)
Hondeghem, L. M. & Katzung, B. G. Antiarrhythmic agents: the modulated receptor mechanism of action of sodium and calcium channel-blocking drugs. Annu. Rev. Pharmacol. Toxicol. 24, 387–423 (1984)
Murphy, K. M., Gould, R. J., Largent, B. L. & Snyder, S. H. A unitary mechanism of calcium antagonist drug action. Proc. Natl Acad. Sci. USA 80, 860–864 (1983)
Catterall, W. A. & Striessnig, J. Receptor sites for Ca2+ channel antagonists. Trends Pharmacol. Sci. 13, 256–262 (1992)
Hockerman, G. H., Peterson, B. Z., Johnson, B. D. & Catterall, W. A. Molecular determinants of drug binding and action on L-type calcium channels. Annu. Rev. Pharmacol. Toxicol. 37, 361–396 (1997)
Striessnig, J. Pharmacology, structure and function of cardiac L-type Ca2+ channels. Cell. Physiol. Biochem. 9, 242–269 (1999)
Hofmann, F., Lacinová, L. & Klugbauer, N. Voltage-dependent calcium channels: from structure to function. Rev. Physiol. Biochem. Pharmacol. 139, 33–87 (1999)
Cheng, R. C., Tikhonov, D. B. & Zhorov, B. S. Structural model for phenylalkylamine binding to L-type calcium channels. J. Biol. Chem. 284, 28332–28342 (2009)
Tikhonov, D. B. & Zhorov, B. S. Structural model for dihydropyridine binding to L-type calcium channels. J. Biol. Chem. 284, 19006–19017 (2009)
Takahashi, M., Seagar, M. J., Jones, J. F., Reber, B. F. & Catterall, W. A. Subunit structure of dihydropyridine-sensitive calcium channels from skeletal muscle. Proc. Natl Acad. Sci. USA 84, 5478–5482 (1987)
Tanabe, T. et al. Primary structure of the receptor for calcium channel blockers from skeletal muscle. Nature 328, 313–318 (1987)
Mikami, A. et al. Primary structure and functional expression of the cardiac dihydropyridine-sensitive calcium channel. Nature 340, 230–233 (1989)
Wu, J. et al. Structure of the voltage-gated calcium channel Cav1.1 complex. Science 350, aad2395 (2015)
Ren, D. et al. A prokaryotic voltage-gated sodium channel. Science 294, 2372–2375 (2001)
Catterall, W. A. & Zheng, N. Deciphering voltage-gated Na+ and Ca2+ channels by studying prokaryotic ancestors. Trends Biochem. Sci. 40, 526–534 (2015)
Payandeh, J., Scheuer, T., Zheng, N. & Catterall, W. A. The crystal structure of a voltage-gated sodium channel. Nature 475, 353–358 (2011)
Payandeh, J., Gamal El-Din, T. M., Scheuer, T., Zheng, N. & Catterall, W. A. Crystal structure of a voltage-gated sodium channel in two potentially inactivated states. Nature 486, 135–139 (2012)
Zhang, X. et al. Crystal structure of an orthologue of the NaChBac voltage-gated sodium channel. Nature 486, 130–134 (2012)
Tang, L. et al. Structural basis for Ca2+ selectivity of a voltage-gated calcium channel. Nature 505, 56–61 (2014)
Catterall, W. A., Perez-Reyes, E., Snutch, T. P. & Striessnig, J. Voltage-gated calcium channels: introduction. IUPHAR/BPS Guide to Pharmacology (http://guidetopharmacology.org/GRAC/FamilyIntroductionForward?familyId=80) (2011)
Striessnig, J., Murphy, B. J. & Catterall, W. A. Dihydropyridine receptor of L-type Ca2+ channels: identification of binding domains for [3H](+)-PN200-110 and [3H]azidopine within the α1 subunit. Proc. Natl Acad. Sci. USA 88, 10769–10773 (1991)
Yamaguchi, S. et al. Key roles of Phe1112 and Ser1115 in the pore-forming IIIS5-S6 linker of L-type Ca2+ channel α1C subunit (CaV 1.2) in binding of dihydropyridines and action of Ca2+ channel agonists. Mol. Pharmacol. 64, 235–248 (2003)
Hescheler, J., Pelzer, D., Trube, G. & Trautwein, W. Does the organic calcium channel blocker D600 act from inside or outside on the cardiac cell membrane? Pflügers Archiv 393, 287–291 (1982)
Striessnig, J., Glossmann, H. & Catterall, W. A. Identification of a phenylalkylamine binding region within the α1 subunit of skeletal muscle Ca2+ channels. Proc. Natl Acad. Sci. USA 87, 9108–9112 (1990)
Hockerman, G. H., Johnson, B. D., Scheuer, T. & Catterall, W. A. Molecular determinants of high affinity phenylalkylamine block of L-type calcium channels. J. Biol. Chem. 270, 22119–22122 (1995)
Yatani, A. & Brown, A. M. The calcium channel blocker nitrendipine blocks sodium channels in neonatal rat cardiac myocytes. Circ. Res. 56, 868–875 (1985)
Kass, R. S., Arena, J. P. & Chin, S. Block of L-type calcium channels by charged dihydropyridines. Sensitivity to side of application and calcium. J. Gen. Physiol. 98, 63–75 (1991)
Bangalore, R., Baindur, N., Rutledge, A., Triggle, D. J. & Kass, R. S. L-type calcium channels: asymmetrical intramembrane binding domain revealed by variable length, permanently charged 1,4-dihydropyridines. Mol. Pharmacol. 46, 660–666 (1994)
Peterson, B. Z. & Catterall, W. A. Calcium binding in the pore of L-type calcium channels modulates high affinity dihydropyridine binding. J. Biol. Chem. 270, 18201–18204 (1995)
Glossmann, H., Ferry, D. R., Goll, A., Striessnig, J. & Zernig, G. Calcium channels and calcium channel drugs: recent biochemical and biophysical findings. Arzneimittelforschung 35, 1917–1935 (1985)
Gamal El-Din, T. M., Martinez, G. Q., Payandeh, J., Scheuer, T. & Catterall, W. A. A gating charge interaction required for late slow inactivation of the bacterial sodium channel NavAb. J. Gen. Physiol. 142, 181–190 (2013)
Faham, S. & Bowie, J. U. Bicelle crystallization: a new method for crystallizing membrane proteins yields a monomeric bacteriorhodopsin structure. J. Mol. Biol. 316, 1–6 (2002)
Faham, S. et al. Crystallization of bacteriorhodopsin from bicelle formulations at room temperature. Protein Sci. 14, 836–840 (2005)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Brünger, A. T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Murshudov, G. N., Vagin, A. A. & Dodson, E. J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004)
Laskowski, R. A., Moss, D. S. & Thornton, J. M. Main-chain bond lengths and bond angles in protein structures. J. Mol. Biol. 231, 1049–1067 (1993)
DeLano, W. L. PyMol molecular viewer (V.1.2r3pre) (http://www.pymol.org) (2002)
Acknowledgements
We are grateful to the beamline staff at the Advanced Light Source (BL8.2.1 and BL8.2.2) for their assistance during data collection. Research reported in this publication was supported by the National Heart, Lung, and Blood Institute (NHLBI) of the National Institutes of Health under award number R01 HL112808 (W.A.C. and N.Z.), and a National Research Service Award from training grant T32 GM008268 (T.M.S.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. This work was also supported by research grants from the Howard Hughes Medical Institute (N.Z.) and by the National Institute of Neurological Disorders and Stroke (NINDS) of the National Institutes of Health under award number R01 NS26254 (W.A.C.). D.C.P. acknowledges support from Neusentis, Pfizer Inc., Cambridge, UK during the course of this work.
Author information
Authors and Affiliations
Contributions
L.T., T.M.G., T.M.S., T.S., D.C.P., N.Z., and W.A.C. designed the experiments. D.C.P. provided compound UK-59811. L.T. conducted protein purification, crystallization, and X-ray diffraction experiments for dihydropyridines. L.T. and T.M.S. conducted protein purification, crystallization, and X-ray diffraction experiments for Br-verapamil. L.T. and T.M.S. determined the structures and analysed the structural results with input from T.M.G. and N.Z. T.M.G. designed and analysed mutants that block drug binding and performed all of the electrophysiological studies. T.M.G., T.S., and W.A.C. analysed the electrophysiological results. All authors contributed to the interpretation of the structures in light of the physiological data. L.T., N.Z., and W.A.C. wrote the manuscript with input from all co-authors.
Corresponding authors
Extended data figures and tables
Extended Data Figure 1 Biophysical characterization of CaVAb I199S.
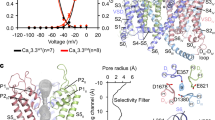
a, Ba2+ currents recorded from a holding potential of −120 mV to test potentials from −60 mV to 20 mV in 10 mV steps for I199S. b, G–V curves of CaVAb and CaVAb I199S derived from peak I–V relationships. The voltages for half-maximal activation and slopes are: CaVAb: V1/2 = −18.8 ± 0.3, k = 3.68 ± 0.43, n = 7; CaVAb I199S: V1/2 = −18.8 ± 0.3, k = 3.88 ± 0.47 (n = 5). c, Repetitive depolarization to 0 mV at 1 Hz from a holding potential of −120 mV (n = 5). d, Steady-state inactivation of CaVAb and CaVAb I199S. Two pulses were applied: a 300-ms conditioning pulse to the indicated potentials followed by 50-ms test pulse to 0 mV (n = 3). e, State-dependent block of CaVAb I199S by 10 nM (green), 100 nM (blue), or 1.5 μM (red) amlodipine during repetitive depolarizations to 0 mV (left, n = 3–5 cells). Ba2+ currents in 100 nM amlodipine for CaVAb I199S (right). f, Concentration-dependent block of CaVAb I199S by nimodipine at 100 nM (blue), 1 μM (red), 5 μM (brown), 10 μM (grey) and 50 μM (black) (left, n = 4–5 cells for each curve). Ba2+ currents in the presence of 5 μM nimodipine for CaVAb I199S (right).
Extended Data Figure 2 Structural comparison of the binding modes of amlodipine, nimodipine, and UK-59811.
a, Superposition of CaVAb in complexes with amlodipine (cyan), nimodipine (yellow), and UK-59811 (magenta) at the dihydropyridine binding site viewed from the side of the pore module. The side chains of dihydropyridine-interacting residues are shown in sticks. b, An Fo–Fc simulated annealing omit map contoured at 2.5σ for nimodipine. c, An Fo–Fc simulated annealing omit map contoured at 2.5σ for UK-59811. d, An Fo–Fc simulated annealing omit map contoured at 2.5σ for DMPC.
Extended Data Figure 3 Biophysical characterization and drug block of CaVAb W195 and CaVAb Y195.
a, Sequence alignment of CaVAb S6 segment and CaV1.1 DIV S6. W195 in CaVAb is equivalent to Y1358 in CaV1.1. b, Ba2+ currents recorded from a holding potential of −120 mV to test potentials from −60 mV to 20 mV in 10 mV steps for CaVAb W195Y. c, G–V curves for CaVAb W195 and CaVAb Y195 derived from peak I–V relationships. The voltages for half-maximal activation and slopes are: CaVAb W195 V1/2 = −18.8 ± 0.3, k = 3.7 ± 0.43, n = 7; CaVAb Y195, V1/2 = −9 ± 0.3, k = 7.4 ± 0.1, n = 5. d, Steady-state inactivation of CaVAb W195 and CaVAb Y195 (n = 3). Two pulses were applied: a 300-ms conditioning pulse followed by 50-ms test pulse to 0 mV. e, State-dependent block of CaVAb W195Y by nimodipine at 100 nM (white), 500 nM (blue), 1 μM (green), 5 μM (red), and control (grey). f, Concentration-dependent block of CaVAb W195Y by nimodipine. IC50 = 508 ± 93 nM (n = 4–5 cells for each point).
Extended Data Figure 4 Biophysical characterization of block by UK-59811.
a, Ba2+ currents for state-dependent block by different concentrations of UK-59811. b, State-dependent block of CaVAb by UK-59811 at 0 nM (black), 100 nM (green), 500 nM (red), 1 μM (blue), and 5 μM (brown). For each curve, n = 4–5 cells. c, Concentration-response curve for UK-59811. Data were fit with a Hill equation assuming a 1:1 binding. IC50 = 194 ± 22 nM, n = 4–5.
Extended Data Figure 5 Evidence for the partially dehydrated Ca2+ binding and carboxyl-carboxylate pairs at the selectivity filter entryway.
a, Top view of an Fo–Fc simulated annealing omit map contoured at 3σ for residues 178 and 181 for the wild-type channel without drug. b, Top view of an Fo–Fc simulated annealing omit map contoured at 3σ for residues 178 and 181 for CaVAb–amlodipine. c, Top view of an Fo–Fc simulated annealing omit map contoured at 2.5σ for residues 178 and 181 for CaVAb–nimodipine. d, Top view of an Fo–Fc simulated annealing omit map contoured at 3σ for residues 178 and 181 for CaVAb–UK-59811. e, Top view of Site 1 with the anomalous difference Fourier map density (red mesh, contoured at 3σ) calculated with diffraction data of crystals collected at 1.75 Å wavelength. Ca2+ is shown as a green sphere. Site 1 residues are shown in sticks. Hydrogen bonds are indicated with dashed lines. f, Top view of Site 2 with the anomalous difference Fourier map density (magenta mesh, contoured at 3σ).
Extended Data Figure 6 Biophysical characterization of verapamil block of CaVAb and functional properties of CaVAb T206S.
a, Chemical structure of verapamil. b, Concentration dependence of verapamil inhibition of CaVAb. The amplitude of the peak Ba2+ current was recorded after applying 20 pulses at a frequency of 1 Hz, where the block reaches steady state. The data were fit by a Hill equation assuming a 1:1 binding ratio. n = 4–7 cells. IC50 = 475 ± 25 nM. c, Ba2+ currents of CaVAb T206S. d, G–V curves. CaVAb (black): V1/2 = 18.8 ± 0.3 mV, k = 3.7 ± 0.43 (n = 5); CaVAb T206S (blue): V1/2 = −15 ± 1.8 mV, k = 6.6 ± 0.4 (n = 5). e, Current traces of CaVAb (black) and CaVAb T206S (blue) during a 1-s depolarizing pulse from a holding potential of –120 mV to −10 mV. f, State-dependent inhibition of CaVAb T206S by Br-verapamil at 10 μM (black), 25 μM (green), 50 μM (red), and 100 μM (blue). For each curve, n = 4–5 cells.
Extended Data Figure 7 Comparison of dihydropyridine binding site in CaVAb and CaV1.2.
The pore domain of CaVAb is illustrated with two subunits in view, one in tan corresponding to domain III of CaV1.2 and one in blue corresponding to domain IV of CaV1.2. The amino acid residues in CaVAb corresponding to those that are important for dihydropyridine binding to CaV1.2 channels are highlighted in red. Bound amlodipine is illustrated with green sticks.
Extended Data Figure 8 Amlodipine binding breaks symmetry.
a, The overall structure of CaVAb in complex with amlodipine (shown in ribbon representation). Measuring the Cα distances of V196 (nearing the amlodipine binding pocket) from the 4 subunits shows the channel is asymmetrical. b, Binding of amlodipine (sticks in red) induces asymmetry and causes rearrangement of the lipid in the central cavity. c–f, The amlodipine binding pocket showing the Cα–Cα distance at two layers (Y195–G164 and I199–F167) horizontally. At layer 1 (Y195–G164), the Cα–Cα distance of its neighbouring sites (11.0 Å in d and 11.0 Å in f) matches the drug binding site (10.9 Å in c), but the diagonal site (e) is too narrow (10.6 Å). At layer 2 (I199–F167), the pocket width of the diagonal site (11.1 Å in e) matches the drug-binding site (11.0 Å in c), but the two diagonal sites are too wide (11.4 Å in d and 11.3 Å in f).
Extended Data Figure 9 Br-verapamil binding breaks symmetry.
a, Alignment of the 4 subunits of CaVAb in complex with Br-verapamil showing the voltage sensor module (VSD) and the ends of S6 are different. b, Measuring the Cα distances between T206 residues in adjacent subunits shows that the channel is indeed asymmetrical with Br-verapamil in the pore.
Supplementary information
Supplementary Information
This file contains a Supplementary Discussion and additional references. (PDF 142 kb)
Rights and permissions
About this article
Cite this article
Tang, L., Gamal El-Din, T., Swanson, T. et al. Structural basis for inhibition of a voltage-gated Ca2+ channel by Ca2+ antagonist drugs. Nature 537, 117–121 (2016). https://doi.org/10.1038/nature19102
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature19102
- Springer Nature Limited
This article is cited by
-
Challenges in the development of novel therapies, vaccines and siRNAs for the treatment of hypertension
Hypertension Research (2023)
-
Quaternary structure independent folding of voltage-gated ion channel pore domain subunits
Nature Structural & Molecular Biology (2022)
-
Synthesis, spectroscopic, DFT, and molecular docking studies on 1,4-dihydropyridine derivative compounds: a combined experimental and theoretical study
Journal of Molecular Modeling (2022)
-
Therapeutic vaccine for chronic diseases after the COVID-19 Era
Hypertension Research (2021)
-
Ligand binding at the protein–lipid interface: strategic considerations for drug design
Nature Reviews Drug Discovery (2021)