Abstract
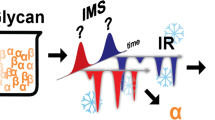
Carbohydrates are ubiquitous biological polymers that are important in a broad range of biological processes1,2,3. However, owing to their branched structures and the presence of stereogenic centres at each glycosidic linkage between monomers, carbohydrates are harder to characterize than are peptides and oligonucleotides4. Methods such as nuclear magnetic resonance spectroscopy can be used to characterize glycosidic linkages, but this technique requires milligram amounts of material and cannot detect small amounts of coexisting isomers5. Mass spectrometry, on the other hand, can provide information on carbohydrate composition and connectivity for even small amounts of sample, but it cannot be used to distinguish between stereoisomers6. Here, we demonstrate that ion mobility–mass spectrometry—a method that separates molecules according to their mass, charge, size, and shape—can unambiguously identify carbohydrate linkage-isomers and stereoisomers. We analysed six synthetic carbohydrate isomers that differ in composition, connectivity, or configuration. Our data show that coexisting carbohydrate isomers can be identified, and relative concentrations of the minor isomer as low as 0.1 per cent can be detected. In addition, the analysis is rapid, and requires no derivatization and only small amounts of sample. These results indicate that ion mobility–mass spectrometry is an effective tool for the analysis of complex carbohydrates. This method could have an impact on the field of carbohydrate synthesis similar to that of the advent of high-performance liquid chromatography on the field of peptide assembly in the late 1970s.



Similar content being viewed by others
References
Dwek, R. A. Glycobiology: toward understanding the function of sugars. Chem. Rev. 96, 683–720 (1996)
Molinari, M. N-glycan structure dictates extension of protein folding or onset of disposal. Nature Chem. Biol. 3, 313–320 (2007)
Varki, A. Sialic acids in human health and disease. Trends Mol. Med. 14, 351–360 (2008)
Bertozzi, C. R. & Rabuka, D. in Essentials of Glycobiology (eds Varki, A. et al.) Ch. 2 (Cold Spring Harbor Laboratory Press, 2009)
Duus, J. Ø., Gotfredsen, C. H. & Bock, K. Carbohydrate structural determination by NMR spectroscopy: modern methods and limitations. Chem. Rev. 100, 4589–4614 (2000)
Dell, A. & Morris, H. R. Glycoprotein structure determination by mass spectrometry. Science 291, 2351–2356 (2001)
Service, R. F. Looking for a sugar rush. Science 338, 321–323 (2012)
Mariño, K., Bones, J., Kattla, J. J. & Rudd, P. M. A systematic approach to protein glycosylation analysis: a path through the maze. Nature Chem. Biol. 6, 713–723 (2010)
Venter, J. C. et al. The sequence of the human genome. Science 291, 1304–1351 (2001)
Domon, B. & Aebersold, R. Mass spectrometry and protein analysis. Science 312, 212–217 (2006)
Plante, O. J., Palmacci, E. R. & Seeberger, P. H. Automated solid-phase synthesis of oligosaccharides. Science 291, 1523–1527 (2001)
Wang, Z. et al. A general strategy for the chemoenzymatic synthesis of asymmetrically branched N-glycans. Science 341, 379–383 (2013)
Boltje, T. J., Buskas, T. & Boons, G.-J. Opportunities and challenges in synthetic oligosaccharide and glycoconjugate research. Nature Chem. 1, 611–622 (2009)
Prien, J. M., Ashline, D. J., Lapadula, A. J., Zhang, H. & Reinhold, V. N. The high mannose glycans from bovine ribonuclease B isomer characterization by ion trap MS. J. Am. Soc. Mass Spectrom. 20, 539–556 (2009)
Daikoku, S., Widmalm, G. & Kanie, O. Analysis of a series of isomeric oligosaccharides by energy-resolved mass spectrometry: a challenge on homobranched trisaccharides. Rapid Commun. Mass Spectrom. 23, 3713–3719 (2009)
Harvey, D. J. Fragmentation of negative ions from carbohydrates: part 1. Use of nitrate and other anionic adducts for the production of negative ion electrospray spectra from N-linked carbohydrates. J. Am. Soc. Mass Spectrom. 16, 622–630 (2005)
Bohrer, B. C., Merenbloom, S. I., Koeniger, S. L., Hilderbrand, A. E. & Clemmer, D. E. Biomolecule analysis by ion mobility spectrometry. Annu. Rev. Anal. Chem. 1, 293–327 (2008)
Uetrecht, C., Rose, R. J., van Duijn, E., Lorenzen, K. & Heck, A. J. R. Ion mobility mass spectrometry of proteins and protein assemblies. Chem. Soc. Rev. 39, 1633–1655 (2010)
Ruotolo, B. T. et al. Evidence for macromolecular protein rings in the absence of bulk water. Science 310, 1658–1661 (2005)
Bleiholder, C., Dupuis, N. F., Wyttenbach, T. & Bowers, M. T. Ion mobility–mass spectrometry reveals a conformational conversion from random assembly to β-sheet in amyloid fibril formation. Nature Chem. 3, 172–177 (2011)
Gabryelski, W. & Froese, K. L. Rapid and sensitive differentiation of anomers, linkage, and position isomers of disaccharides using high-field asymmetric waveform ion mobility spectrometry (FAIMS). J. Am. Soc. Mass Spectrom. 14, 265–277 (2003)
Plasencia, M. D., Isailovic, D., Merenbloom, S. I., Mechref, Y. & Clemmer, D. E. Resolving and assigning N-linked glycan structural isomers from ovalbumin by IMS-MS. J. Am. Soc. Mass Spectrom. 19, 1706–1715 (2008)
Zhu, M., Bendiak, B., Clowers, B. & Hill, H. H., Jr Ion mobility-mass spectrometry analysis of isomeric carbohydrate precursor ions. Anal. Bioanal. Chem. 394, 1853–1867 (2009)
Williams, J. P. et al. Characterization of simple isomeric oligosaccharides and the rapid separation of glycan mixtures by ion mobility mass spectrometry. Int. J. Mass Spectrom. 298, 119–127 (2010)
Fenn, L. S. & McLean, J. A. Structural resolution of carbohydrate positional and structural isomers based on gas-phase ion mobility-mass spectrometry. Phys. Chem. Chem. Phys. 13, 2196–2205 (2011)
Both, P. et al. Discrimination of epimeric glycans and glycopeptides using IM–MS and its potential for carbohydrate sequencing. Nature Chem. 6, 65–74 (2014)
Kröck, L. et al. Streamlined access to conjugation-ready glycans by automated synthesis. Chem. Sci. 3, 1617–1622 (2012)
Pringle, S. D. et al. An investigation of the mobility separation of some peptide and protein ions using a new hybrid quadrupole/travelling wave IMS/oa-ToF instrument. Int. J. Mass Spectrom. 261, 1–12 (2007)
Pagel, K. & Harvey, D. J. Ion mobility-mass spectrometry of complex carbohydrates—collision cross sections of sodiated N-linked glycans. Anal. Chem. 85, 5138–5145 (2013)
Hofmann, J. et al. Estimating collision cross sections of negatively charged N-glycans using traveling wave ion mobility-mass spectrometry. Anal. Chem. 86, 10789–10795 (2014)
Eller, S., Collot, M., Yin, J., Hahm, H. S. & Seeberger, P. H. Automated solid-phase synthesis of chondroitin sulfate glycosaminoglycans. Angew. Chem. Int. Ed. 52, 5858–5861 (2013)
Martin, C. E., Weishaupt, M. W. & Seeberger, P. H. Progress toward developing a carbohydrate-conjugate vaccine against Clostridium difficile ribotype 027: synthesis of the cell-surface polysaccharide PS-I repeating unit. Chem. Commun. 47, 10260–10262 (2011)
Werz, D. B., Carstagner, B. & Seeberger, P. H. Automated synthesis of the tumor-asssociated carbohydrate antigens Gb-3 and Globo-H: incorporation of α-galactosidic linkages. J. Am. Chem. Soc. 129, 2770–2771 (2007)
Acknowledgements
We thank the Free University Berlin and the Max Planck Society for financial support. J.H. and K.P. thank G. von Helden, J.L.P. Benesch and W.B. Struwe for comments.
Author information
Authors and Affiliations
Contributions
P.H.S. and K.P. designed the research; J.H. and H.S.H. performed the research. All authors analysed data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Automated synthesis of oligosaccharides 20–25.
Ar, 2-methyl-5-tert-butylphenyl; Bn, benzyl; Bz, benzoyl; Cbz, carboxybenzyl; Et, ethyl; Fmoc, fluorenylmethyloxycarbonyl.
Extended Data Figure 2 Automated synthesis of oligosaccharides 26–28.
Ac, acetyl; Bn, benzyl; Bu, butyl; Bz, benzoyl; Cbz, carboxybenzyl; Et, ethyl; Fmoc, fluorenylmethyloxycarbonyl; lev, levulinoyl; TCA, trichloroacetimidate; UV, ultraviolet.
Extended Data Figure 3 Drift-time distributions of trisaccharides 1–6 as different species in positive- and negative-ion mode.
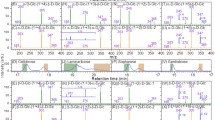
The CCS difference between the most compact and the most extended isomer of each set is given as a percentage. Small CCS differences are observed in positive-ion mode (a, b), which makes an unambiguous identification of the trisaccharides difficult. The largest CCS differences are observed using deprotonated ions (c), allowing the identification of linkage isomers (for example, 3 + 6) and stereoisomers (for example, 2 + 3). A clear identification of regioisomers with a terminal 1→3 or 1→4 glycosidic bond can be obtained for chloride adducts (d).
Extended Data Figure 4 Comparison of drift times and CCSs of structurally similar precursor ions and fragments.
a, Mass spectra of trisaccharides 5 and 6, as well as a tandem MS spectrum of 7 (β-Gal-(1→3)–β-GlcNAc-(1→3)–α-Gal-(1→4)–β-Gal-(1→4)–β-Glc-L; L = C5H10NH2) in negative-ion mode. The pentasaccharide 7 has the same core structure as the trisaccharide 6. Collision-induced dissociation of deprotonated 7 consequently produces a fragment with the same mass as the deprotonated precursor ion of 6. b, Drift-time distributions of [M-H]− = 588 ions. The collision-induced dissociation fragment arising from deprotonated 7 exhibits an drift time and CCS identical to those of the intact deprotonated trisaccharide 6. This indicates that glycans and glycan fragments with identical structures also exhibit identical CCSs. Seen from a broader perspective, this highlights the exceptional potential of negative-ion CCSs to be used as a diagnostic parameter for glycan sequencing.
Extended Data Figure 5 IM–MS differentiation and identification of the hexasaccharides 8 (black) and 9 (red).
As deprotonated ions, 8 and 9 show almost identical drift times and therefore cannot be distinguished. However, smaller collision-induced dissociation fragments containing five, four, or three monosaccharide building blocks (m/z 1,017, 832, and 652, respectively) exhibit highly diagnostic drift times. At m/z 832, a double peak is observed for the branched oligosaccharide 8 (inset, black trace), because two isomeric fragments are formed. Both fragments can be detected simultaneously using IM–MS, with cleavage at the 3-antenna being clearly preferred. The disaccharide fragments at m/z 467 and 364 are identical for 8 and 9 and consequently exhibit identical drift times.
Extended Data Figure 6 Alternative synthesis of oligosaccharide 5 and corresponding IM–MS analysis.
a, An alternative route for synthesizing 5 uses building block 11 instead of 12, which results in a mixture of the disaccharides 31 and 32 and subsequently in a mixture of trisaccharides 5 and 30. Neither the fully protected trisaccharides 29 and 29-by-product, nor the deprotected sugars 5 and 30, can be separated by HPLC. The formation of 29-by-product can be detected using NMR analysis, but a clear structural assignment is not possible owing to the low relative concentration. b, [M-H]− = 588 and c, [M+Cl]− = 624 drift-time distributions of trisaccharides 1–6 compared to the drift time of the crude mixture consisting of 5 and 30 clearly reveal a content of about 5% by-product 30. In particular, the drift time of the chloride adduct of 30 is very diagnostic, because it differs considerably from the drift times of all other trisaccharides investigated here.
Extended Data Figure 7 Correlation between signal intensity and ion mobility peak width in mixtures of 2 and 4.
a, Drift-time distributions of [M + H]+ and [M + Na]+ ions at high (upper panels) and low (lower panels) signal intensity. The given average ion count per second corresponds to the signal detected for the major isotope peak. High signal intensities result in peak broadening and reduced ion mobility resolution, whereas a considerably improved separation is achieved at lower intensity. b, Drift-time distributions of [M-H]− ions from a mixture of <1% 4 and >99% 2. Measurements at high signal intensity can be used to qualitatively detect 4. At low intensity, however, 4 is discriminated and no signal can be detected.
Supplementary information
Supplementary Information
This file contains Supplementary Text and Data, Supplementary Figures 1-27, Supplementary Tables 1-4 and additional references. (PDF 9856 kb)
Rights and permissions
About this article
Cite this article
Hofmann, J., Hahm, H., Seeberger, P. et al. Identification of carbohydrate anomers using ion mobility–mass spectrometry. Nature 526, 241–244 (2015). https://doi.org/10.1038/nature15388
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature15388
- Springer Nature Limited
This article is cited by
-
High-resolution separation of bioisomers using ion cloud profiling
Nature Communications (2023)
-
Carbohydrates and carbohydrate degradation gene abundance and transcription in Atlantic waters of the Arctic
ISME Communications (2023)
-
Advances in single-molecule junctions as tools for chemical and biochemical analysis
Nature Chemistry (2023)
-
Understanding of protomers/deprotomers by combining mass spectrometry and computation
Analytical and Bioanalytical Chemistry (2023)
-
Dextran as internal calibrant for N-glycan analysis by liquid chromatography coupled to ion mobility-mass spectrometry
Analytical and Bioanalytical Chemistry (2022)