Abstract
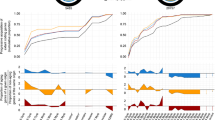
The Red Queen hypothesis proposes that coevolution of interacting species (such as hosts and parasites) should drive molecular evolution through continual natural selection for adaptation and counter-adaptation1,2,3. Although the divergence observed at some host-resistance4,5,6 and parasite-infectivity7,8,9 genes is consistent with this, the long time periods typically required to study coevolution have so far prevented any direct empirical test. Here we show, using experimental populations of the bacterium Pseudomonas fluorescens SBW25 and its viral parasite, phage Φ2 (refs 10, 11), that the rate of molecular evolution in the phage was far higher when both bacterium and phage coevolved with each other than when phage evolved against a constant host genotype. Coevolution also resulted in far greater genetic divergence between replicate populations, which was correlated with the range of hosts that coevolved phage were able to infect. Consistent with this, the most rapidly evolving phage genes under coevolution were those involved in host infection. These results demonstrate, at both the genomic and phenotypic level, that antagonistic coevolution is a cause of rapid and divergent evolution, and is likely to be a major driver of evolutionary change within species.


Similar content being viewed by others
References
Van Valen, L. A new evolutionary law. Evol. Theory 1, 1–30 (1973)
Van Valen, L. Molecular evolution as predicted by natural selection. J. Mol. Evol. 3, 89–101 (1974)
Stenseth, N. C. & Smith, J. M. Coevolution in ecosystems: Red Queen evolution or stasis? Evolution 38, 870–880 (1984)
Clark, A. G. et al. Evolution of genes and genomes on the Drosophila phylogeny. Nature 450, 203–218 (2007)
Hedrick, P. W. Evolutionary genetics of the major histocompatibility complex. Am. Nat. 143, 945–964 (1994)
Obbard, D. J., Jiggins, F. M., Halligan, D. L. & Little, T. J. Natural selection drives extremely rapid evolution in antiviral RNAi genes. Curr. Biol. 16, 580–585 (2006)
Blanc, G. et al. Molecular evolution of Rickettsia surface antigens: evidence of positive selection. Mol. Biol. Evol. 22, 2073–2083 (2005)
Mu, J. et al. Genome-wide variation and identification of vaccine targets in the Plasmodium falciparum genome. Nature Genet. 39, 126–130 (2006)
Barrett, L. G. et al. Diversity and evolution of effector loci in natural populations of the plant pathogen Melampsora lini . Mol. Biol. Evol. 26, 2499–2513 (2009)
Buckling, A. & Rainey, P. B. Antagonistic coevolution between a bacterium and a bacteriophage. Proc. R. Soc. Lond. B 269, 931–936 (2002)
Brockhurst, M. A., Morgan, A. D., Fenton, A. & Buckling, A. Experimental coevolution with bacteria and phage: the Pseudomonas fluorescens–Φ2 model system. Infect. Genet. Evol. 7, 547–552 (2007)
Woolhouse, M. E., Webster, J. P., Domingo, E., Charlesworth, B. & Levin, B. R. Biological and biomedical implications of the co-evolution of pathogens and their hosts. Nature Genet. 32, 569–577 (2002)
Brockhurst, M. A., Morgan, A. D., Rainey, P. B. & Buckling, A. Population mixing accelerates coevolution. Ecol. Lett. 6, 975–979 (2003)
Morgan, A., Gandon, S. & Buckling, A. The effect of migration on local adaptation in a coevolving host–parasite system. Nature 437, 253–256 (2005)
Morgan, A. & Buckling, A. Relative number of generations of hosts and parasites does not influence parasite local adaptation in coevolving populations of bacteria and phages. J. Evol. Biol. 19, 1956–1963 (2006)
Poullain, V., Gandon, S., Brockhurst, M. A., Buckling, A. & Hochberg, M. E. The evolution of specificity in evolving and coevolving antagonistic interactions between a bacteria and its phage. Evolution 62, 1–11 (2008)
Benmayor, R., Hodgson, D. J., Perron, G. G. & Buckling, A. Host mixing and disease emergence. Curr. Biol. 19, 764–767 (2009)
Excoffier, L., Smouse, P. E. & Quattro, J. M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitocondrial DNA restriction data. Genetics 131, 479–491 (1992)
Ceyssens, P. J. et al. Genomic analysis of Pseudomonas aeruginosa phages LKD16 and LKA1: establishment of the ΦKMV subgroup within the T7 supergroup. J. Bacteriol. 188, 6924–6931 (2006)
Calendar, R. L. The Bacteriophages (Oxford Univ. Press, 2005)
Barrick, J. E. et al. Genome evolution and adaptation in a long-term experiment with Escherichia coli . Nature 461, 1243–1247 (2009)
Bull, J. J. et al. Exceptional convergent evolution in a virus. Genetics 147, 1497–1507 (1997)
Wichman, H. A., Badgett, M. R., Scott, L. A., Boulianne, C. M. & Bull, J. J. Different trajectories of parallel evolution during viral adaptation. Science 285, 422–424 (1999)
Thompson, J. N. The Geographic Mosaic of Coevolution (Univ. of Chicago Press, 2005)
Santos, M. A. An improved method for the small scale preparation of bacteriophage DNA based on phage precipitation by zinc chloride. Nucleic Acids Res. 19, 5442 (1991)
Rutherford, K. et al. Artemis: sequence visualization and annotation. Bioinformatics 16, 944–945 (2000)
Barrick, J. E. & Lenski, R. E. Genome-wide mutational diversity in an evolving population of Escherichia coli . Cold Spring Harb. Symp. Quant. Biol. 10.1101/sqb.2009.74.018 (23 September 2009)
Acknowledgements
We are grateful to M. Begon and G. Hurst for comments on previous drafts of this manuscript. We acknowledge funding from the Natural Environment Research Council, the Wellcome Trust, the Leverhulme Trust and the European Research Council. We are grateful for technical assistance from staff at the Centre for Genomic Research, University of Liverpool.
Author Contributions M.A.B. and S.P. conceived and designed the study; T.V. performed selection experiments, infectivity assays and prepared samples for population genomic sequencing; B.L. and M.A.B. performed further assays; R.B. prepared samples for ancestral genome sequencing; N.R.T., M.Q., F.S. and D.W. performed ancestral genome sequencing and assembly; A.J.S. and N.R.T. performed genome annotation; S.P., T.V. and M.A.B. analysed the data; M.A.B., S.P., A.B., A.J.S. and N.R.T. wrote the manuscript; T.V. and N.H. commented on the manuscript; M.A.B., A.F. and S.P. supervised the research; S.P., N.H., A.B. and M.A.B. obtained funding.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figures S1-S2 with Legends, Supplementary Table S1, a Supplementary Discussion and Supplementary References. (PDF 297 kb)
Supplementary Table 2
This table contains the genome annotation for phage Φ2, and the full gene-by-gene statistical analysis of the genome re-sequencing data. (XLS 75 kb)
PowerPoint slides
Rights and permissions
About this article
Cite this article
Paterson, S., Vogwill, T., Buckling, A. et al. Antagonistic coevolution accelerates molecular evolution. Nature 464, 275–278 (2010). https://doi.org/10.1038/nature08798
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature08798
- Springer Nature Limited
This article is cited by
-
Eco-evolutionary dynamics of gut phageome in wild gibbons (Hoolock tianxing) with seasonal diet variations
Nature Communications (2024)
-
Coevolution between marine Aeromonas and phages reveals temporal trade-off patterns of phage resistance and host population fitness
The ISME Journal (2023)
-
Bacteria-phage coevolution with a seed bank
The ISME Journal (2023)
-
Soil viral diversity, ecology and climate change
Nature Reviews Microbiology (2023)
-
Molecular mechanisms of adaptive evolution in wild animals and plants
Science China Life Sciences (2023)