Abstract
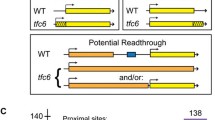
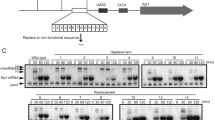
Recent transcriptome analyses using high-density tiling arrays1,2,3 and data from large-scale analyses of full-length complementary DNA libraries by the FANTOM3 consortium4,5 demonstrate that many transcripts are non-coding RNAs (ncRNAs). These transcriptome analyses indicate that many of the non-coding regions, previously thought to be functionally inert, are actually transcriptionally active regions with various features. Furthermore, most relatively large (∼several kilobases) polyadenylated messenger RNA transcripts are transcribed from regions harbouring little coding potential. However, the function of such ncRNAs is mostly unknown and has been a matter of debate2. Here we show that RNA polymerase II (RNAPII) transcription of ncRNAs is required for chromatin remodelling at the fission yeast Schizosaccharomyces pombe fbp1+ locus during transcriptional activation. The chromatin at fbp1+ is progressively converted to an open configuration, as several species of ncRNAs are transcribed through fbp1+. This is coupled with the translocation of RNAPII through the region upstream of the eventual fbp1+ transcriptional start site. Insertion of a transcription terminator into this upstream region abolishes both the cascade of transcription of ncRNAs and the progressive chromatin alteration. Our results demonstrate that transcription through the promoter region is required to make DNA sequences accessible to transcriptional activators and to RNAPII.




Similar content being viewed by others
Accession codes
Primary accessions
ArrayExpress
Data deposits
The raw and analysed data from the genome tiling array experiment are available from ArrayExpress (http://www.ebi.ac.uk/arrayexpress) under the accession numbers E-MEXP-1747.
References
Cawley, S. et al. Unbiased mapping of transcription factor binding sites along human chromosomes 21 and 22 points to widespread regulation of noncoding RNAs. Cell 116, 499–509 (2004)
Cheng, J. et al. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science 308, 1149–1154 (2005)
Wilhelm, B. T. et al. Dynamic repertoire of a eukaryotic transcriptome surveyed at single-nucleotide resolution. Nature 453, 1239–1243 (2008)
The FANTOM Consortium The transcriptional landscape of the mammalian genome. Science 309, 1559–1563 (2005)
Hayashizaki, Y. & Carninci, P. Genome Network and FANTOM3: assessing the complexity of the transcriptome. PLoS Genet. 2, e63 (2006)
Hoffman, C. S. Glucose sensing via the protein kinase A pathway in Schizosaccharomyces pombe . Biochem. Soc. Trans. 33, 257–260 (2005)
Janoo, R. T., Neely, L. A., Braun, B. R., Whitehall, S. K. & Hoffman, C. S. Transcriptional regulators of the Schizosaccharomyces pombe fbp1 gene include two redundant Tup1p-like corepressors and the CCAAT binding factor activation complex. Genetics 157, 1205–1215 (2001)
Neely, L. A. & Hoffman, C. S. Protein kinase A and mitogen-activated protein kinase pathways antagonistically regulate fission yeast fbp1 transcription by employing different modes of action at two upstream activation sites. Mol. Cell. Biol. 20, 6426–6434 (2000)
Wolffe, A. Chromatin: Structure and Function 3rd edn (Academic, 1997)
Grigull, J., Mnaimneh, S., Pootoolal, J., Robinson, M. D. & Hughes, T. R. Genome-wide analysis of mRNA stability using transcription inhibitors and microarrays reveals posttranscriptional control of ribosome biogenesis factors. Mol. Cell. Biol. 24, 5534–5547 (2004)
Higuchi, T., Watanabe, Y. & Yamamoto, M. Protein kinase A regulates sexual development and gluconeogenesis through phosphorylation of the Zn finger transcriptional activator Rst2p in fission yeast. Mol. Cell. Biol. 22, 1–11 (2002)
Hirota, K., Hoffman, C. S. & Ohta, K. Reciprocal nuclear shuttling of two antagonizing Zn finger proteins modulates Tup family corepressor function to repress chromatin remodeling. Eukaryot. Cell 5, 1980–1989 (2006)
Hirota, K., Mizuno, K. I., Shibata, T. & Ohta, K. Distinct chromatin modulators regulate the formation of accessible and repressive chromatin at the fssion yeast recombination hotspot ade6–M26 . Mol. Biol. Cell 19, 1162–1173 (2008)
Martens, J. A. & Winston, F. Recent advances in understanding chromatin remodeling by Swi/Snf complexes. Curr. Opin. Genet. Dev. 13, 136–142 (2003)
Uhler, J. P., Hertel, C. & Svejstrup, J. Q. A role for noncoding transcription in activation of the yeast PHO5 gene. Proc. Natl Acad. Sci. USA 104, 8011–8016 (2007)
Schmitt, S., Prestel, M. & Paro, R. Intergenic transcription through a polycomb group response element counteracts silencing. Genes Dev. 19, 697–708 (2005)
Sanchez-Elsner, T., Gou, D., Kremmer, E. & Sauer, F. Noncoding RNAs of trithorax response elements recruit Drosophila Ash1 to Ultrabithorax. Science 311, 1118–1123 (2006)
Bender, M. A., Bulger, M., Close, J. & Groudine, M. β-globin gene switching and DNase I sensitivity of the endogenous β-globin locus in mice do not require the locus control region. Mol. Cell 5, 387–393 (2000)
Gribnau, J., Diderich, K., Pruzina, S., Calzolari, R. & Fraser, P. Intergenic transcription and developmental remodeling of chromatin subdomains in the human β-globin locus. Mol. Cell 5, 377–386 (2000)
Ashe, H. L., Monks, J., Wijgerde, M., Fraser, P. & Proudfoot, N. J. Intergenic transcription and transinduction of the human β-globin locus. Genes Dev. 11, 2494–2509 (1997)
Miura, F. et al. A large-scale full-length cDNA analysis to explore the budding yeast transcriptome. Proc. Natl Acad. Sci. USA 103, 17846–17851 (2006)
Hirota, K., Hoffman, C. S., Shibata, T. & Ohta, K. Fission yeast Tup1-like repressors repress chromatin remodeling at the fbp1+ promoter and the ade6–M26 recombination hotspot. Genetics 165, 505–515 (2003)
Hirota, K., Steiner, W. W., Shibata, T. & Ohta, K. Multiple modes of chromatin configuration at natural meiotic recombination hot spots in fission yeast. Eukaryot. Cell 6, 2072–2080 (2007)
Takeda, T. & Yamamoto, M. Analysis and in vivo disruption of the gene coding for calmodulin in Schizosaccharomyces pombe . Proc. Natl Acad. Sci. USA 84, 3580–3584 (1987)
Acknowledgements
We thank W. Lin for critically reading this manuscript. This work was supported by grants-in-aid for scientific research in a priority area, the Ministry of Education, Culture, Sports, Science and Technology, Japan.
Author Contributions K.H. and K.O. conceived the project. K.H. planned, performed and analysed most of the experiments. T.M. performed Rst2-ChIP and RACE experiments. T.M. and K.K. performed the genome tiling array experiment. K.H. wrote most of the paper. C.S.H., T.S. and K.O. gave scientific advice and financial support. K.O. supervised the project.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
This file contains Supplementary Figures 1-6 with Legends, Supplementary Table 1 and Supplementary Methods. (PDF 1147 kb)
Rights and permissions
About this article
Cite this article
Hirota, K., Miyoshi, T., Kugou, K. et al. Stepwise chromatin remodelling by a cascade of transcription initiation of non-coding RNAs. Nature 456, 130–134 (2008). https://doi.org/10.1038/nature07348
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07348
- Springer Nature Limited
This article is cited by
-
Gene mapping methodology powered by induced genome rearrangements
Scientific Reports (2022)
-
Biologia futura: combinatorial stress responses in fungi
Biologia Futura (2022)
-
Modular signature of long non-coding RNA association networks as a prognostic biomarker in lung cancer
BMC Medical Genomics (2021)
-
High-resolution analysis of cell-state transitions in yeast suggests widespread transcriptional tuning by alternative starts
Genome Biology (2021)
-
Dissecting the transcriptional regulatory networks of promoter-associated noncoding RNAs in development and cancer
Journal of Experimental & Clinical Cancer Research (2020)