Abstract
Gain-of-function mutations in the mouse nicotinamide mononucleotide adenylyltransferase type 1 (Nmnat1) produce two remarkable phenotypes: protection against traumatic axonal degeneration and reduced hypoxic brain injury. Despite intensive efforts, the mechanism of Nmnat1 cytoprotection remains elusive. To develop a new model to define this mechanism, we heterologously expressed a mouse Nmnat1 non-nuclear-localized gain-of-function mutant gene (m-nonN-Nmnat1) in the nematode Caenorhabditis elegans and show that it provides protection from both hypoxia-induced animal death and taxol-induced axonal pathology. Additionally, we find that m-nonN-Nmnat1 significantly lengthens C. elegans lifespan. Using the hypoxia-protective phenotype in C. elegans, we performed a candidate screen for genetic suppressors of m-nonN-Nmnat1 cytoprotection. Loss of function in two genes, haf-1 and dve-1, encoding mitochondrial unfolded protein response (mitoUPR) factors were identified as suppressors. M-nonN-Nmnat1 induced a transcriptional reporter of the mitoUPR gene hsp-6 and provided protection from the mitochondrial proteostasis toxin ethidium bromide. M-nonN-Nmnat1 was also protective against axonal degeneration in C. elegans induced by the chemotherapy drug taxol. Taxol markedly reduced basal expression of a mitoUPR reporter; the expression was restored by m-nonN-Nmnat1. Taken together, these data implicate the mitoUPR as a mechanism whereby Nmnat1 protects from hypoxic and axonal injury.
Similar content being viewed by others
Main
About 25 years ago, Lunn et al.1 reported a remarkable mouse strain (later named wlds) where Wallerian axonal degeneration after traumatic nerve injury was markedly reduced. The responsible genetic lesion in the wlds strain was found to be a translocation and triplication that produced a mutant fusion protein consisting of a segment of an E4-ubiquitin ligase-like protein and full-length Nmnat1, a primarily nuclear-localized nicotinamide adenine dinucleotide (NAD) biosynthetic enzyme.2, 3, 4 Subsequently, overexpression of Nmnat1 or a non-nuclear-localized mutant form of Nmnat1 (nonN-Nmnat1) was shown to be sufficient to prevent axonal degeneration in mice.5 The mechanism whereby Nmnat1 is axonal protective has remained elusive, although substantial evidence points to a mitochondrial site of action.6, 7, 8 Recently, a transgenic mouse with the expression of nonN-Nmnat1 was found to have reduced brain injury in a model of neonatal hypoxic/ischemic stroke.9 Although the reduced stroke volume might be secondary to a reduced axonal degeneration, this result suggests the hypothesis that nonN-Nmnat1 may have broader cytoprotective properties.
Caenorhabditis elegans has become an increasingly important model for the study of both hypoxic and axonal injury.10, 11, 12, 13, 14, 15, 16 In this regard, the strengths of the model lie in its genetic tractability, fully defined anatomy and cellular identity, and the ability to directly observe cell pathology in live animals. Screens in C. elegans for genes that control hypoxic sensitivity have implicated multiple distinct pathways as determinants of hypoxic death.16, 17, 18, 19 In particular, genetic perturbations that improve cellular proteostasis are generally hypoxia protective.16, 17, 18, 19, 20, 21 These findings argue that hypoxia perturbs protein folding and that this perturbation contributes to cell death; indeed, substantial evidence in C. elegans and in other models indicates that hypoxia/ischemia disrupts protein folding homeostasis.16, 17, 19, 22, 23
In this work, we set out to answer two outstanding questions about Nmnat1. First, is hypoxia protection a general feature of Nmnat1 expression or is it peculiar to the mouse transgenic model previously tested? Second, what is the mechanism whereby Nmnat1 protects from hypoxia? For the first question, we expressed the mouse non-nuclear-localized-Nmnat1 (m-nonN-Nmnat1) in two distinct settings, primary mouse hippocampal neurons and in C. elegans, and asked whether in these contexts m-nonN-Nmnat1 was hypoxia protective. Additionally, we overexpressed the C. elegans homologs of Nmnat1 to determine if other isoforms of Nmnat1 were hypoxia protective. For the second question, we used the C. elegans model to test a variety of candidate pathways for their role in Nmnat1 hypoxia protection.
Results
Mouse nonN-Nmnat1 protects mouse primary hippocampal neuron cultures from hypoxic injury
We first determined whether mouse nonN-Nmnat1 directly protects neurons from hypoxic injury as our primary interest is neuronal hypoxic injury and as the previous study showed Nmnat1 hypoxic protection in a transgenic mouse model where anti-inflammatory effects or genetic background contributions cannot be ruled out.9 Using a lentiviral expression system,24 we expressed m-nonN-Nmnat1 in primary mouse hippocampal neuron cultures essentially devoid of other cell types and found that m-nonN-Nmnat1 indeed provides strong hypoxic protection to neurons (Figure 1).
m-nonN-Nmnat1 protects primary mouse hippocampal neurons from hypoxic injury. (a) Hypoxic survival of primary mouse hippocampal neurons infected with lentivirus carrying m-nonN-Nmnat1 or control luciferase. Values are mean±S.E.M.; P<0.01 by unpaired T-test of four trials. (b) Representative fluorescent photomicrographs of hypoxia-exposed hippocampal neurons with and without m-nonN-Nmat1 infection. Neurons were visualized with Tuj1 antibody. Scale bar, 10 μm
Mouse nonN-Nmnat1 protects C. elegans from hypoxic injury
Having demonstrated definitively that m-nonN-Nmnat1 protected mouse neurons from hypoxia, we wanted to know if this phenotype extended across phyla and therefore is likely a general property of Nmnat1. Towards this end, we expressed m-nonN-Nmnat1 heterologously in the nematode C. elegans (Table 1). As in mouse, m-nonN-Nmnat1 expression in C. elegans was strongly protective against hypoxic injury. Strains with ubiquitous expression in all C. elegans cells (ub-m-nonN-Nmnat1) were hypoxia resistant with even a single copy insertion conferring significant hypoxic protection (Figure 2a). Given the primarily neuronal phenotype in mouse, we asked whether m-nonN-Nmnat1 expression in neurons alone (neuro-m-nonN-Nmnat1) is sufficient to protect the organism from hypoxia and found that neuronal expression did indeed provide strong protection (Figure 2b). We examined the relative level of expression of the various transgenes, in particular to determine why one of six neuro-m-nonN-Nmnat1 strains had only a low, nonstatistically significant level of protection. The level of protection correlated for neuronal transgenes with the m-nonN-Nmnat1 transcript levels consistent with its gain-of-function activity (Supplementary Figure S1). We next asked whether the m-nonN-Nmnat1 transgene was functioning like the native C. elegans Nmnat1 and found that ub-m-nonN-Nmnat1 could rescue the sterility phenotype of a loss-of-function mutant of nmat-2, one of the two C. elegans Nmnat1 homologs (Figure 2c).25 Similarly, ubiquitous overexpression of nmat-2 and the other C. elegans Nmnat1 homolog nmat-1 both provided significant hypoxic protection (Figure 2d). Thus, hypoxia protection is a conserved phenotype of both worm and mouse Nmnat1 gain-of-function.
m-nonN-Nmnat1 protects worms from hypoxia. (a) Hypoxic survival of ubiquitous-expressing ub-m-nonN-Nmnat1 strains. High copy – gcIs40 and single copy – gcSi1 integrants of the Prpl-28::mCherry::m-nonN-Nmnat1 transgene were tested. (***P<0.0001, **P<0.01, NS, not significant – unpaired T-test, mean±S.E.M. shown, n=15 trials for N2(wild type) and gcIs40, n=6 for gcSi1). Inset: gcIs40 worms (top – DIC; bottom – mCherry filter). (b) Hypoxic survival of pan-neuronal-expressing neuro-m-nonN-Nmnat1 strains (Prab3::mCherry::m-nonN-Nmnat1) (trials n=5–20). Inset: gcIs30 worms (top – DIC; bottom – mCherry filter; scale bars for (a) and (b)=100 μm). (c) The infertility of nmat-2(tm2905) could be rescued by ubiquitous C. elegans nmat-1 overexpression (gcIs37) and m-nonN-Nmnat1 expression (gcSi1) (each data point represents the total brood from one animal). (d) Hypoxic survival of C. elegans nmat-1 and nmat-2 overexpression. gcEx145 contains an extrachromosomal nmat-1 array [Prpl-28::nmat-1::mCherry]; gcIs29 and gcIs37 contain integrated nmat-2 arrays (Prpl-28::nmat-2::mCherry) (trials: n=12, 7, 12 and 3 for N2, gcIs29, gcIs37 and gcEx154, respectively)
M-nonN-Nmnat1 does not affect global oxygen consumption in C. elegans
Given that Nmnat1 is a biosynthetic enzyme for NAD, a general mechanism of Nmnat1 hypoxia protection that we considered was that the m-nonN-Nmnat1 expression reduces oxygen consumption and thereby reduces the duration of cellular hypoxia and severity of injury. However, oxygen consumption was similar to wild type in both m-nonN-Nmnat1 and nmat-2 overexpression animals (Supplementary Figure S2). Thus, the observed Nmnat1 hypoxia protection is not because of a simple oxygen conservation mechanism.
Mitochondrial unfolded protein response genes are required for m-nonN-Nmnat1 hypoxia resistance
We tested a variety of mutant genes that might plausibly have a phenotypic interaction with m-nonN-Nmnat1. Most genes that were tested had no effect on the hypoxia-protective phenotype (Supplementary Figure S3). However, we found that loss-of-function mutations in the gene haf-1, which encodes a mitochondrial outer membrane ABC transporter previously shown to function as an activator of the mitochondrial unfolded protein response (mitoUPR),26 significantly suppressed the hypoxia resistance of both ub-m-nonN-Nmnat1 and neuro-m-nonN-Nmnat1 transgenes (Figures 3a and b). The genes dve-1 and atfs-1 also function in the C. elegans mitoUPR, encoding transcription factors that promote the expression of the mitochondrial-specific chaperones HSP-6 and HSP-60.26,27,28,29 Similar to haf-1(lf) mutants, dve-1 double-stranded RNA-mediated interference (RNAi) suppressed the m-nonN-Nmnat1 hypoxia-resistant phenotype (Figure 3c). Interestingly, neither loss-of-function mutations nor RNAi knockdown of atfs-1, a gene believed to be required for mitoUPR induction under most conditions, suppressed m-nonN-Nmnat1-mediated hypoxia protection (Figures 3b and c), demonstrating that m-nonN-Nmnat1 is activating the mitoUPR in a non-atfs-1-dependent manner. Several mitochondrial-resident proteases are thought to function upstream of the peptide transporter HAF-1.27, 29 Mutations in these genes greatly compromised worm development and rendered data from the double mutations difficult to interpret. Nevertheless, in cases where the double mutant animals were healthy enough for hypoxia testing, none of these mutations suppressed m-nonN-Nmnat1 hypoxia protection (Supplementary Figure S4a). Thus, the suppression of the hypoxia-resistant phenotype of m-nonN-Nmnat1 is not a general property of mitoUPR mutants but is relatively specific to haf-1 and dve-1. Mutations in ABC transporters for other organelles such as haf-2 and haf-3 did not suppress m-nonN-Nmnat1 protection against hypoxia (Supplementary Figure S4b). Additionally, the suppression by the haf-1 is specific to Nmnat1 hypoxia resistance in that haf-1(lf) did not suppress the hypoxic protection produced by a previously identified hypoxia-resistant allele of the arginyl tRNA synthetase gene – rars-1(gc47) (Figure 3d).17
Mitochondrial UPR components are required for m-nonN-Nmnat1 phenotypes. (a and b) Loss-of-function mutations of haf-1 but not atfs-1 suppressed the hypoxia resistance of Ub-Nmnat1(gcIs40[ub-m-nonN-Nmnat1]) (a) and Neuro-Nmnat1(gcIs30[neuro-m-nonN-Nmnat1] (b) (mean±S.E.M., unpaired T-test, n=5–10 trials). (c) RNAi against dve-1 but not atfs-1 suppressed Ub-Nmnat1(gcIs40[Ub-m-nonN-Nmnat1]) hypoxia resistance. Hypoxic treatment was for 18 h (mean±S.E.M., unpaired T-test, n=3 trials). (d) Loss of function of haf-1 did not suppress rars-1(gc47) hypoxia resistance; hypoxic treatment was for 30 h (mean±S.E.M., unpaired T-test, n=3 trials). (e) Mutation of haf-1 rescued the low fertility of Ub-Nmnat1[gcIs40(Ub-m-nonN-Nmnat1]) (n=14–49, unpaired T-test, N2 versus Ub-Nmnat1 – P<0.0001; N2 versus Neuro-Nmnat1(gcIs30[Neuro-m-nonN-Nmnat1] – P=0.13. (f) Worm lifespan was significantly extended by Ub-Nmnat1(gcIs40[ub-m-nonN-Nmnat1] (P<0.0001, N2 versus gcIs40, log-rank test); the lifespan extension was not significantly suppressed by haf-1(tm843)
M-nonN-Nmnat1 worms are long lived
While generally characterizing the m-nonN-Nmnat1 strains, we found that the strongly expressing ub-m-nonN-Nmnat1 strain gcIs40 had two additional phenotypes, reduced fertility and long lifespan (Supplementary Table S1). We asked whether these Nmnat1 phenotypes were regulated by the same pathways as hypoxic injury. Both haf-1 mutants strongly suppressed the infertility phenotype of ub-m-nonN-Nmnat1 (Figure 3e and Supplementary Table S2). However, surprisingly, haf-1(tm843) did not suppress lifespan extension (Figure 3f). It is important to point out that infertility was not causally related to the hypoxia resistance phenotype of m-nonN-Nmnat1 in that gcSi1, a single copy insertion allele of ub-m-nonN-Nmnat1, is hypoxia resistant but has normal fertility (Figures 2a and c), and several of the hypoxia-resistant neuro-m-nonN-Nmnat1 strains have normal fertility (Figures 2b and Supplementary Table S1). The lack of concordance of the suppression of the lifespan extension and infertility phenotypes in the haf-1(lf);gcIs40 mutants argues that the underlying mechanisms of hypoxia resistance and lifespan extension by Nmnat1 are also distinct.
M-nonN-Nmnat1 protects from mitochondrial proteostasis stress
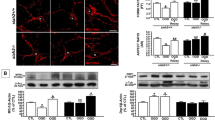
The suppression phenotypes by haf-1 and dve-1 loss-of-function suggests that m-nonN-Nmnat1 may ameliorate mitochondrial proteostasis stress. To test this hypothesis, we measured the sensitivity of m-nonN-Nmnat1 to ethidium bromide (EtBr), which produces mitochondrial protein folding stress.30 Both neuronal and ubiquitous m-nonN-Nmnat1 strains were highly protective against the toxic effects of EtBr (Figure 4a). Consistent with amelioration of mitochondrial folding stress rather than other effects of EtBr on mitochondria such as suppressing transcription, neuro-m-nonN-Nmnat1 significantly reduced the induction by EtBr of the mitoUPR transcriptional reporter, Phsp-6::GFP(zcIs13) (Figure 4b).30 Surprisingly, in the absence of EtBr, both neuro- and ub-m-nonN-Nmnat1 mildly but significantly increased the basal level of activation of Phsp-6::GFP, and neuro-m-nonN-Nmnat1 increased Phsp-60::GFP (Figures 4c and d). The results suggest that the protective effect of m-nonN-Nmnat1 may be hormetic in nature, by producing a low level of mitochondrial proteostasis stress that induces a mitoUPR-dependent protective response. Consistent with this hypothesis, basal mitoUPR induction by m-nonN-Nmnat1 was significantly suppressed by haf-1(lf) (Figure 4e). Moreover, expression of m-nonN-Nmnat1 in mouse primary hippocampal neurons could induce the expression of mitochondrial chaperone hspa9 (closest hsp-6 homolog in mouse); however, no induction of hspd1 was observed (closest hsp-60 homolog) (Supplementary Figure S5). An enzymatic activity compromised version of m-nonN-Nmnat1 (H24A)24 had not effect on the expression of hspd1, demonstrating that enzymatic activity of nonN-Nmnat1 is required for induction in mouse neurons.
m-nonN-Nmnat1 protects from mitochondrial folding stress and activates the mitoUPR. (a) m-nonN-Nmnat1 protected worms from EtBR-induced developmental arrest. Eggs were laid on plates with or without EtBr (40 μg/ml) and percent adults was scored 4 days later (mean±S.E.M., unpaired T-test; n=4 trials). (b) The induction of the mitoUPR reporter Phsp-6::GFP (zcIs13) by EtBr was significantly reduced by Neuro-Nmnat1(gcIs41[Neuro-m-nonN-Nmnat1]) (mean±S.E.M., unpaired T-test; n=28–50 animals per condition); note gcIs41 heterozygotes were used because of difficulty constructing zcIs13;gcIs41 homozygotes. (c and d) Under basal conditions, both Ub-Nmnat1(gcSi6;gcSi7[7X-Ub-m-nonN-Nmnat1] and Neuro-Nmnat1(gcIs41[Neuro-m-nonN-Nmat1]) activate mitoUPR reporter zcIs13[Phsp-6::GFP]; Neuro-Nmnat1(gcIs41[Neuro-m-nonN-Nmat1]) also activates zcIs9[Phsp-60::GFP]. (e) haf-1(tm843) suppressed Neuro-Nmnat1(gcIs41[neuro-m-nonN-Nmnat1])-induced mitoUPR (for (b–e) mean±S.E.M., unpaired T-test; three trials with at least 15 worms imaged for each genotype/condition)
Taxol-induced axonal degeneration ameliorated by m-nonN-Nmnat1
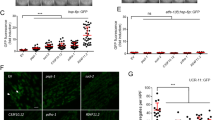
Finally, we asked whether m-nonN-Nmnat1 protected against axonal degeneration in C. elegans, the original phenotype of the wlds mouse. Worm axon degeneration has been reported in spectrin unc-70 mutants and in a model of necrosis-induced neuron degeneration,25, 31 and overexpression of worm nmat-2 was shown to be protective against degeneration of mechanosensory neurons.25 To examine the role m-nonN-Nmnat1 in axon degeneration, we used taxol, a chemotherapeutic agent, previously shown to induce axonal degeneration in cancer patients and in experimental models.32, 33, 34 Worms hatched from taxol-containing plates had significant growth retardation that could be rescued by ub-m-nonN-Nmnat1 (Figure 5a). To observe axonal degeneration directly, GFP-labeled mechanosensory neurons were examined after taxol treatment. In elderly animals, mechanosensory axonal pathology manifesting as axonal beading and gaps, particularly in the PLM neurons in the tail of the animals, was seen (Figure 5b) and was significantly increased in taxol-treated animals (Figure 5c). The taxol-induced axonal degeneration was significantly reduced by the neuro-m-nonN-Nmnat1 transgene gcIs35 (Figure 5c); the low expressing gcIs35 transgene was used because it did not extend lifespan, thereby avoiding a potential confounding factor (Supplementary Figure S6 and Supplementary Table S2). Despite initial developmental growth retardation, taxol did not alter adult lifespan in wild-type animals (Supplementary Figure S6 and Supplementary Table S2). However, taxol was lethally toxic to elderly haf-1 but not atfs-1 loss-of-function animals (Figure 5d). These data suggest the hypotheses that taxol produces mitochondrial folding stress or that it affects the haf-1-dependent induction of the mitoUPR in response to mitochondrial proteostatic stress. Consistent with the latter hypothesis, taxol dose dependently reduced the basal expression of the Phsp-6::GFP mitoUPR reporter and this reduction of the expression of the mitoUPR reporter was blocked by neuronal m-nonN-Nmnat1 (Figures 5e and f). The analogous ER unfolded protein response reporter Phsp-4::GFP35 was unaffected by taxol, indicating relative specificity for the mitoUPR (Figure 5g).
m-nonN-Nmnat1 reduces axonal degeneration induced by taxol. (a) Taxol-mediated C. elegans developmental arrest is rescued by Ub-Nmnat1(gcIs40[ub-m-nonN-Nmnat1]) (mean±S.E.M., unpaired T-test, three trials). Eggs were laid on taxol plates and four days later the number of adults was scored. (b) Worm mechanosensory neurons and axons labeled by zdIs5[mec-4::GFP] (scale bar: 100 μm; scale bar in inset: 10 μm). Representative images from the tail region comparing DMSO buffer versus taxol. Note fragmented (arrow) and beaded (arrowhead) axons in the taxol condition. (c) Taxol-induced axon degeneration was rescued by Neuro-Nmnat1[gcIs35(neuro-m-nonN-Nmnat1]) (Fisher's exact test, 141–494 axons were scored in five independent trials and pooled together). (d) haf-1(tm843) reduced survival of NeuroNmnat1 animals (gcIs35[neuro-m-nonN-Nmnat1]) in the presence of chronic taxol but atfs-1(tm4919) did not. Eggs were laid on taxol (3 μM) plates and at 15 days after adulthood, surviving worms were scored (Fisher's exact test; data were pooled from four independent trials, n=239–314). (e) Taxol acutely reduced basal mitoUPR reporter zcIs13[Phsp-6::GFP] activity). Inset: worms under DIC (top) and GFP filter (bottom, scale bar: 100 μm). (f) Heterozygous Neuro-Nmnat1(gcIs41[Neuro-m-nonN-Nmnat1]) preserved mitoUPR reporter activity following taxol (3 μm) treatment (mean±S.E.M., unpaired T-test; three independent trials). (g) Taxol did not alter the expression of the ER-UPR reporter activity zcIs4 [Phsp-4::GFP] (mean±S.E.M., unpaired T-test; three independent trials)
Discussion
Nmnat1 has been found to protect from axonal degeneration secondary to a variety of insults.36, 37, 38, 39 Most recently, a transgenic mouse strain expressing the same non-nuclear-localized Nmnat1 protein used in this study was found to have a reduced stroke volume following a hypoxic ischemic insult.9 Unlike previous work, the Verghese et al.9 study suggests that nonN-Nmnat1 protects neuronal cell bodies from acute injury, not just axons from delayed degeneration. Our results in both mouse primary neurons and in C. elegans definitively confirm the hypoxia-protective activity of non-nuclear-localized Nmnat1. Our data also show that hypoxia protection is not peculiar to m-nonN-Nmnat1 but rather is a general feature of Nmnats since overexpressions of two C. elegans Nmnat homologs also improve hypoxic survival. Additionally, our work uncovered a novel nicotinamide mononucleotide adenylyltransferase (Nmnat) stress resistant phenotype – prolonged lifespan. Our data show that m-nonN-Nmnat1 activates the mitoUPR. Given that the mitoUPR has been implicated in lifespan extension in C. elegans and other animals,40 it is not particularly surprising that m-nonN-Nmnat1 extends lifespan in C. elegans. Further, the lifespan extension phenotype shows that nonN-Nmnat1 is not only protective against the acute high level stress of hypoxia and traumatic nerve injury but also against the indolent stress of aging.
Despite considerable effort, the mechanism whereby nonN-Nmnat1 protects from axonal degeneration, much less the mechanism of any of its more recently defined phenotypes, has not been defined. However, mitochondria have been repeatedly implicated as having a role. The mitochondrially localized mammalian isoform Nmnat3 has been shown to phenocopy the axonal degeneration-protective activity of the Nmnat1 wlds mutant, whereas simple overexpression of the nuclear-localized Nmnat1 or the golgi-localized Nmnat2 does not protect from axonal degeneration.8, 41 The axonal-protective phenotype correlates with nonN-Nmnat1 expression in the mitochondria matrix and mitochondrial ATP content has been found to be higher in Nmnat3 and wlds mice compared to wild-type mice.8 Nmnat3 heterologous expression in Drosophila was also protective against axotomy-induced axonal degeneration, and as in mammalian cells, Nmnat3 localizes to fly mitochondria.6, 42 Knockdown of endogenous Drosophila nicotinamide mononucleotide adenylyltransferase (dNmnat) in fly results in a rapid loss of axonal mitochondria, suggesting that dNmnat is required for mitochondrial health.43 Expression of the wlds protein in fly results in an increase in mitochondrial calcium buffering capacity resulting in improved axonal mitochondrial motility.42 Whether improved mitochondrial calcium buffering capacity, improved mitochondrial proteostasis or a combination of the two is responsible for Nmnat-mediated stress protection is unclear. However, we recently reported that hypoxia disrupts mitochondrial proteostasis and that induction of the mitoUPR by genetic or pharmacological agents was hypoxia protective in C. elegans.44 Thus, we favor improved mitochondrial proteostasis by Nmnat as a primary mediator of hypoxic survival.
The enzymatic product of Nmnat1, NAD, has been shown to be axonal-protective5 and to activate the mitoUPR to maintain mitochondrial proteostasis.45 Further NAD and its precursors have been found to extend lifespan in C. elegans and multiple other models.46 Even though overall NAD levels have been found to be unchanged in nonN-Nmnat1 transgenic mice,24 the enzymatic activity of Nmnat1 is required for its axonal-protective phenotype in mouse and for its hypoxia-protective phenotype in mouse primary neuron cultures (Supplementary Figure S5). Mammalian mitochondrial NAD has to be synthesized in situ by the mitochondrial-specific isoform Nmnat3;47 perhaps, nonN-Nmnat1 similarly increases mitochondrial NAD levels without affecting overall cellular NAD. Therefore, our working hypothesis based on these data is that m-nonN-Nmnat1 maintains mitochondrial NAD levels and thereby supports mitoUPR function and mitochondrial proteostasis in injured neurons and presumably other cell types to prevent degeneration and cell death.
Materials and Methods
Strains
C. elegans strains were obtained from the Caenorhabditis Genetics Center and Japan National BioResource Project and outcrossed three times before testing. Mutations were confirmed after outcrossing by PCR. All strains were maintained at 20 °C on nematode growth media agar seeded with OP50 bacteria as described previously.48 Information for all alleles used is available at www.wormbase.org, and all mutations were consistent with the published information at Wormbase. New transgenic strains are listed in Table 1. Double and triple mutant strains were generated by standard genetic methods,49 and genotypes were confirmed by phenotype and/or PCR. RNAi experiments were performed as described previously.18
Transgenic animals
nmat-1 and nmat-2 cDNA was amplified from wild-type N2 and confirmed by sequencing. The rpl-28 promoter (1.4 kb) was subcloned from the Andy Fire Vector Kit (pPD129.57; Addgene, Cambridge, MA, USA). An expression construct with the rab-3 promoter was a gift from Mike Nonet (Washington University School of Medicine, St Louis, MO, USA).50 Mouse nonN-Nmnat1 cDNA was a gift from Jeffery Milbrandt (Washington University School of Medicine) and was tagged with mCherry and 6X-His.24 Prpl-28::mCherry::m-nonN-Nmnat1, Prpl-28::nmat-1::mCherry, Prpl-28::nmat-2::mCherry and Prab3::m-nonN-Nmnat1::mCherry were injected into N2 gonads along with the coinjection marker pPHGFP (except gcIs41, which was without coinjection marker).51 Transgenic animals were selected by the expression of fluorescent markers. To make integrated lines, 30–40 young adult worms carrying transgenes were UV irradiated as described previously.51 At least 100 F1s with expression from the transgene were cloned and lines where 100% of animals stably expressed the transgene were kept and outcrossed at least 10 times.
Single copy chromosomal insertions were produced as described previously.52 Single copy (gcSi1), two tandem copy (gcSi3) and four tandem copy (gcSi6) of Prpl-28::m-nonN-Nmnat1 was inserted into ttTi5605 II site by the MosSCI method, whereas three tandem copies (gcSi7) was inserted into the cxTi10882 IV site.52 Insertion sites and integrity of the integrated sequences were verified by PCR and restriction mapping.
C. elegans hypoxia assays
Hypoxic incubation and scoring of C. elegans was performed as described previously.18 For each trial, three plates of young adult worms (one day after L4 worms, 30–60 worms per plate) were used. After hypoxic treatment, worms from these three plates were scored after a 24-h recovery as alive or dead, and the values from the three plates were pooled as a single trial. Hypoxic incubation was for 20 h, unless otherwise stated. Numbers of trials are listed in the individual figure legends.
Fertility tests
L4 worms (14–20 for each genotype) were placed in plates individually. The worms were transferred to new plates and eggs were counted daily until no eggs were laid for two consecutive days. Worms burrowing, bagging or desiccated were censored from analysis.
Lifespan analysis
Lifespan assays were conducted at 20 °C as described previously.53 After the L4 molting, animals were transferred to plates containing 100 μM 5-fluoro-2′-deoxyuridine (FUDR; Sigma). We used the L4 molt as t=0 for lifespan analysis. Strains were grown at 20 °C for at least five generations before lifespan determination. Exploded, bagged and desiccated worms were censored.
EtBr and taxol developmental assay
EtBr (40 μg/ml) or taxol (3 μM) were added to plates 24 h before synchronization. Eggs (30–50/plate, three plates/genotype/trial) were allowed to hatch on EtBr or taxol plates. The adult animals were counted 4 days after hatching and divided by total eggs to obtain percent of adults.
O2 consumption
Oxygen consumption assays were performed as reported previously.16 At least 3000 young adults were used for each test for each genotype and the experiments were repeated at least three times.
GFP reporter assays
Worms were mounted on 2% agarose pads and immobilized using 10 μM levamisole in M9. All images were taken using a Zeiss Axioskop 2 microscope (Carl Zeiss Microscopy, Jena, Germany) with the x10 objective and a Retiga EXi Fast1394 digital camera (QImaging, Surrey, BC, Canada). At least 15 animals were used for each trial, with at least three trials for each genotype/treatment. Fluorescent intensities were measured and analyzed with ImageJ software (NIH, Bethesda, MD, USA) as described previously.17
Taxol and axon degeneration
Paclitaxel/taxol was purchased from Sigma (T7402) and dissolved in dimethyl sulfoxide (DMSO; Sigma D2650) to make a 10 mM stock solution and stored at −80 °C. The stock solution was diluted immediately before use into DMSO to make 0.6 mM intermediate solution, 50 μl of which was added to NGM plates (10 ml) the night before use to make final 3 μM taxol assay plates. Control plates contained 50 μl DMSO. The 0.3 and 1 μm plates were made similarly. Worm populations were synchronized on these plates and 100 μM mitotic inhibitor FUDR (F0503; Sigma) was added into plates once worms grew into young adults. When the food became scarce, worms were picked onto new plates with the same conditions. All surviving day 15 adults (L4=day 0) were picked to examine axon phenotypes under a compound microscope (Zeiss Axioskop2 plus, Zeiss North America, Thornwood, NY, USA). Pictures of all PLM posterior processes were taken by using a x63 objective and Photoshop software (Adobe Systems, San Jose, CA, USA) was used for further image processing such as rotating and adding pseudocolor. An axon was scored as pathological when clear gaps, truncation and beading were present. The images were scored blinded to condition. Only the posterior processes of PLM neurons were found to have reproducible axon degeneration. Sporadic degenerations at the anterior of PLM and in the ALM neurons was rarely observed.
RT-qPCR
The detailed protocol has been described previously.19 Total RNA was extracted from worms or primary neurons with TRIzol LS reagent (Life Technologies, Carlsbad, CA, USA; no. 10296-010). Primers for m-nonN-Nmnat1 (all nonN-Nmnat1 constructs were mCherry and 6 × his tagged): forward, 5′-ACTGGAAAAGCCTGGGGCGGCGGCGGC-3′ and reverse, 5′-ATGGTGATGGTGATGCAAAGTGGAATGG-3′; β-actin: forward, 5′-GACATGGAGAAGATCTGGCA-3′ and reverse, 5′-GGTCTCAAACATGATCTGGGT-3′; hspa9: forward, 5′-CAGGAAGAAGGAACGTGTTG-3′ and reverse, 5′-GCTTGTTGCACTCATCAGCAG-3′; hspd1: forward, 5′-CCTGTGACAACCCCTGAAG-3′ and reverse, 5′-CACTCAACAAGACATAGGCA-3′. qPCR (SYBR green) was run at 95 °C for 15 s and 60 °C for 30 s and analyzed using the ΔΔCT method.
Hippocampal neurons and hypoxic treatment
Primary P0 mouse hippocampal neurons were established in poly-d-lysine-coated 4-well dishes with Neurobasal medium supplemented with B27 (Life Technologies; 17504-044), l-glutamine and FUDR to prevent glial growth. After 5 days in vitro, the neurons were infected with lentivirus containing sequence encoding the desired proteins.24 Five days after infection, nonN-Nmnat1 expression was confirmed by checking for the coexpressing GFP signal. Cultures were either harvested for RNA extraction or treated with hypoxia (<0.3% O2) for 7 h at 37 °C, followed by overnight recovery under normal culture condition. To score for cell death, cultures were stained by ethidium homodimer (Life Technologies; E1169) before fixation. Cells were then stained with Tuj1 (Abcam, Cambridge, UK, AB14545) antibody to identify neurons (green signal shown in Supplementary Figure S1) and with DAPI to identify all cells. At least 10 high-power fields were taken for each infection, and survival values from these fields were pooled as one trial.
Statistical analysis
GraphPad Prism 6 software (La Jolla, CA, USA) was used for all statistical analysis. Statistical significance was determined by tests indicated in individual figure legends. Paired T-tests were used when all trials had concurrent controls; unpaired T-tests were used for all other experiments except lifespan comparisons where log-rank tests were performed.
Abbreviations
- C. elegans :
-
Caenorhabditis elegans
- DMSO:
-
dimethyl sulfoxide
- EtBr:
-
ethidium bromide
- GFP:
-
green fluorescent protein
- mitoUPR:
-
mitochondrial unfolded protein response
- NAD:
-
nicotinamide adenine dinucleotide
- Nmnat:
-
nicotinamide mononucleotide adenylyltransferase
- Nmnat1:
-
mouse nicotinamide mononucleotide adenylyltransferase type 1
- dNmnat:
-
Drosophila nicotinamide mononucleotide adenylyltransferase
- m-nonN-Nmnat1:
-
mouse non-nuclear NMNAT1
- neuro-m-nonN-Nmnat1:
-
pan-neuronally expressed mouse non-nuclear NMNAT1
- ub-m-nonN-Nmnat1:
-
ubiquitously expressed mouse non-nuclear NMNAT1
- RNAi:
-
double-stranded RNA-mediated interference
References
Lunn ER, Perry VH, Brown MC, Rosen H, Gordon S . Absence of Wallerian degeneration does not hinder regeneration in peripheral nerve. Eur J Neurosci 1989; 1: 27–33.
Coleman MP, Conforti L, Buckmaster EA, Tarlton A, Ewing RM, Brown MC et al. An 85-kb tandem triplication in the slow Wallerian degeneration (Wlds) mouse. Proc Natl Acad Sci USA 1998; 95: 9985–9990.
Conforti L, Tarlton A, Mack TG, Mi W, Buckmaster EA, Wagner D et al. A Ufd2/D4Cole1e chimeric protein and overexpression of Rbp7 in the slow Wallerian degeneration (WldS) mouse. Proc Natl Acad Sci USA 2000; 97: 11377–11382.
Mack TG, Reiner M, Beirowski B, Mi W, Emanuelli M, Wagner D et al. Wallerian degeneration of injured axons and synapses is delayed by a Ube4b/Nmnat chimeric gene. Nat Neurosci 2001; 4: 1199–1206.
Araki T, Sasaki Y, Milbrandt J . Increased nuclear NAD biosynthesis and SIRT1 activation prevent axonal degeneration. Science 2004; 305: 1010–1013.
Avery MA, Sheehan AE, Kerr KS, Wang J, Freeman MR . Wld S requires Nmnat1 enzymatic activity and N16-VCP interactions to suppress Wallerian degeneration. J Cell Biol 2009; 184: 501–513.
Court FA, Coleman MP . Mitochondria as a central sensor for axonal degenerative stimuli. Trends Neurosci 2012; 35: 364–372.
Yahata N, Yuasa S, Araki T . Nicotinamide mononucleotide adenylyltransferase expression in mitochondrial matrix delays Wallerian degeneration. J Neurosci 2009; 29: 6276–6284.
Verghese PB, Sasaki Y, Yang D, Stewart F, Sabar F, Finn MB et al. Nicotinamide mononucleotide adenylyl transferase 1 protects against acute neurodegeneration in developing CNS by inhibiting excitotoxic-necrotic cell death. Proc Natl Acad Sci USA 2011; 108: 19054–19059.
Chen L, Chisholm AD . Axon regeneration mechanisms: insights from C. elegans. Trends Cell Biol 2011; 21: 577–584.
Butler JA, Mishur RJ, Bokov AF, Hakala KW, Weintraub ST, Rea SL . Profiling the anaerobic response of C. elegans using GC-MS. PLoS One 2012; 7: e46140.
Fawcett EM, Horsman JW, Miller DL . Creating defined gaseous environments to study the effects of hypoxia on C. elegans. J Vis Exp 2012; 65: e4088.
LaRue BL, Padilla PA . Environmental and genetic preconditioning for long-term anoxia responses requires AMPK in Caenorhabditis elegans. PLoS One 2011; 6: e16790.
Menuz V, Howell KS, Gentina S, Epstein S, Riezman I, Fornallaz-Mulhauser M et al. Protection of C. elegans from anoxia by HYL-2 ceramide synthase. Science 2009; 324: 381–384.
Powell-Coffman JA . Hypoxia signaling and resistance in C. elegans. Trends Endocrinol Metab 2010; 21: 435–440.
Scott B, Sun CL, Mao X, Yu C, Vohra BP, Milbrandt J et al. Role of oxygen consumption in hypoxia protection by translation factor depletion. J Exp Biol 2013; 216: 2283–2292.
Anderson LL, Mao X, Scott BA, Crowder CM . Survival from hypoxia in C. elegans by inactivation of aminoacyl-tRNA synthetases. Science 2009; 323: 630–633.
Mabon ME, Mao X, Jiao Y, Scott BA, Crowder CM . Systematic identification of gene activities promoting hypoxic death. Genetics 2009; 181: 483–496.
Mao XR, Crowder CM . Protein misfolding induces hypoxic preconditioning via a subset of the unfolded protein response machinery. Mol Cell Biol 2010; 30: 5033–5042.
Mabon ME, Scott BA, Crowder CM . Divergent mechanisms controlling hypoxic sensitivity and lifespan by the DAF-2/Insulin/IGF-receptor pathway. PLoS One 2009; 4: e7937.
Scott BA, Avidan MS, Crowder CM . Regulation of hypoxic death in C. elegans by the insulin/IGF receptor homolog DAF-2. Science 2002; 296: 2388–2391.
Badiola N, Penas C, Minano-Molina A, Barneda-Zahonero B, Fado R, Sanchez-Opazo G et al. Induction of ER stress in response to oxygen-glucose deprivation of cortical cultures involves the activation of the PERK and IRE-1 pathways and of caspase-12. Cell Death Dis 2011; 2: e149.
Kim I, Xu W, Reed JC . Cell death and endoplasmic reticulum stress: disease relevance and therapeutic opportunities. Nat Rev Drug Discov 2008; 7: 1013–1030.
Sasaki Y, Vohra BP, Lund FE, Milbrandt J . Nicotinamide mononucleotide adenylyl transferase-mediated axonal protection requires enzymatic activity but not increased levels of neuronal nicotinamide adenine dinucleotide. J Neurosci 2009; 29: 5525–5535.
Calixto A, Jara JS, Court FA . Diapause formation and downregulation of insulin-like signaling via DAF-16/FOXO delays axonal degeneration and neuronal loss. PLoS Genet 2012; 8: e1003141.
Haynes CM, Yang Y, Blais SP, Neubert TA, Ron D . The matrix peptide exporter HAF-1 signals a mitochondrial UPR by activating the transcription factor ZC376.7 in C. elegans. Mol Cell 2010; 37: 529–540.
Haynes CM, Petrova K, Benedetti C, Yang Y, Ron D . ClpP mediates activation of a mitochondrial unfolded protein response in C. elegans. Dev Cell 2007; 13: 467–480.
Nargund AM, Fiorese CJ, Pellegrino MW, Deng P, Haynes CM . Mitochondrial and nuclear accumulation of the transcription factor ATFS-1 promotes OXPHOS recovery during the UPR(mt). Mol Cell 2015; 58: 123–133.
Nargund AM, Pellegrino MW, Fiorese CJ, Baker BM, Haynes CM . Mitochondrial import efficiency of ATFS-1 regulates mitochondrial UPR activation. Science 2012; 337: 587–590.
Yoneda T, Benedetti C, Urano F, Clark SG, Harding HP, Ron D . Compartment-specific perturbation of protein handling activates genes encoding mitochondrial chaperones. J Cell Sci 2004; 117: 4055–4066.
Hammarlund M, Jorgensen EM, Bastiani MJ . Axons break in animals lacking beta-spectrin. J Cell Biol 2007; 176: 269–275.
Bhattacharya MR, Gerdts J, Naylor SA, Royse EX, Ebstein SY, Sasaki Y et al. A model of toxic neuropathy in Drosophila reveals a role for MORN4 in promoting axonal degeneration. J Neurosci 2012; 32: 5054–5061.
Sahenk Z, Barohn R, New P, Mendell JR . Taxol neuropathy. Electrodiagnostic and sural nerve biopsy findings. Arch Neurol 1994; 51: 726–729.
Bennett GJ, Liu GK, Xiao WH, Jin HW, Siau C . Terminal arbor degeneration – a novel lesion produced by the antineoplastic agent paclitaxel. Eur J Neurosci 2011; 33: 1667–1676.
Calfon M, Zeng H, Urano F, Till JH, Hubbard SR, Harding HP et al. IRE1 couples endoplasmic reticulum load to secretory capacity by processing the XBP-1 mRNA. Nature 2002; 415: 92–96.
Ali YO, McCormack R, Darr A, Zhai RG . Nicotinamide mononucleotide adenylyltransferase is a stress response protein regulated by the heat shock factor/hypoxia-inducible factor 1{alpha} pathway. J Biol Chem 2011; 286: 19089–19099.
Coleman MP, Freeman MR . Wallerian degeneration, wld(s), and nmnat. Annu Rev Neurosci 2010; 33: 245–267.
Vohra BP, Sasaki Y, Miller BR, Chang J, DiAntonio A, Milbrandt J . Amyloid precursor protein cleavage-dependent and -independent axonal degeneration programs share a common nicotinamide mononucleotide adenylyltransferase 1-sensitive pathway. J Neurosci 2010; 30: 13729–13738.
Wang JT, Medress ZA, Barres BA . Axon degeneration: molecular mechanisms of a self-destruction pathway. J Cell Biol 2012; 196: 7–18.
Hill S, Van Remmen H . Mitochondrial stress signaling in longevity: a new role for mitochondrial function in aging. Redox Biol 2014; 2: 936–944.
Sasaki Y, Araki T, Milbrandt J . Stimulation of nicotinamide adenine dinucleotide biosynthetic pathways delays axonal degeneration after axotomy. J Neurosci 2006; 26: 8484–8491.
Avery MA, Rooney TM, Pandya JD, Wishart TM, Gillingwater TH, Geddes JW et al. Wld(S) prevents axon degeneration through increased mitochondrial flux and enhanced mitochondrial Ca(2+) buffering. Curr Biol 2012; 22: 596–600.
Fang Y, Soares L, Teng X, Geary M, Bonini NM . A novel Drosophila model of nerve injury reveals an essential role of nmnat in maintaining axonal integrity. Curr Biol 2012; 22: 590–595.
Kaufman DM, Crowder CM . Mitochondrial proteostatic collapse leads to hypoxic injury. Curr Biol 2015; 25: 2171–2176.
Mouchiroud L, Houtkooper RH, Moullan N, Katsyuba E, Ryu D, Canto C et al. The NAD(+)/sirtuin pathway modulates longevity through activation of mitochondrial UPR and FOXO signaling. Cell 2013; 154: 430–441.
Verdin E . The many faces of sirtuins: coupling of NAD metabolism, sirtuins and lifespan. Nat Med 2014; 20: 25–27.
Nikiforov A, Dolle C, Niere M, Ziegler M . Pathways and subcellular compartmentation of NAD biosynthesis in human cells: from entry of extracellular precursors to mitochondrial NAD generation. J Biol Chem 2011; 286: 21767–21778.
Brenner S . The genetics of Caenorhabditis elegans. Genetics 1974; 77: 71–94.
Fay DS . Classical genetic methods (December 30, 2013), WormBook, ed. The C. elegans Research Community, WormBook.
Mahoney MB, Parks AL, Ruddy DA, Tiong SY, Esengil H, Phan AC et al. Presenilin-based genetic screens in Drosophila melanogaster identify novel notch pathway modifiers. Genetics 2006; 172: 2309–2324.
Evans TC (ed). Transformation and microinjection (April 6, 2006), WormBook, ed. The C. elegans Research Community, WormBook, doi/10.1895/ wormbook.1.108.1, doi/10.1895/wormbook.1.165.1.
Frokjaer-Jensen C, Davis MW, Hopkins CE, Newman BJ, Thummel JM, Olesen SP et al. Single-copy insertion of transgenes in Caenorhabditis elegans. Nat Genet 2008; 40: 1375–1383.
Apfeld J, Kenyon C . Regulation of lifespan by sensory perception in Caenorhabditis elegans. Nature 1999; 402: 804–809.
Acknowledgements
We thank members in Crowder laboratory for help and insightful discussions. We thank Michael Nonet and his lab members and Jeffery Milbrandt and Yo Sasaki for reagents, advice and help with technique development. Some strains were provided by the Caenorhabditis Genetics Center, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440). We also thank National BioResource Project, Japan for strains. This work was supported by funding from the National Institute of Neurological Disorders and Stroke (R01 NS045905 and 5R21NS084360-02).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Raschellà
Supplementary Information accompanies this paper on Cell Death and Disease website
Supplementary information
Rights and permissions
Cell Death and Disease is an open-access journal published by Nature Publishing Group. This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Mao, X., Kaufman, D. & Crowder, C. Nicotinamide mononucleotide adenylyltransferase promotes hypoxic survival by activating the mitochondrial unfolded protein response. Cell Death Dis 7, e2113 (2016). https://doi.org/10.1038/cddis.2016.5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2016.5
- Springer Nature Limited
This article is cited by
-
Effect of the mitochondrial unfolded protein response on hypoxic death and mitochondrial protein aggregation
Cell Death & Disease (2021)