Abstract
Background:
Ovarian clear cell carcinoma (OCCC) is mostly resistant to standard chemotherapy that results in poor patient survival. To understand the genetic background of these tumours, we performed whole-genome sequencing of OCCC tumours.
Methods:
Tumour tissue samples and matched blood samples were obtained from 55 Japanese women diagnosed with OCCC. Whole-genome sequencing was performed using the Illumina HiSeq platform according to standard protocols.
Results:
Alterations to the switch/sucrose non-fermentable (SWI/SNF) subunit, the phosphatidylinositol-3-kinase (PI3K)/Akt signalling pathway, and the receptor tyrosine kinase (RTK)/Ras signalling pathway were found in 51%, 42%, and 29% of OCCC tumours, respectively. The 3-year overall survival (OS) rate for patients with an activated PI3K/Akt signalling pathway was significantly higher than that for those with inactive pathway (91 vs 40%, hazard ratio 0.24 (95% confidence interval (CI) 0.10–0.56), P=0.0010). Similarly, the OS was significantly higher in patients with the activated RTK/Ras signalling pathway than in those with the inactive pathway (91 vs 53%, hazard ratio 0.35 (95% CI 0.13–0.94), P=0.0373). Multivariable analysis revealed that activation of the PI3K/Akt and RTK/Ras signalling pathways was an independent prognostic factor for patients with OCCC.
Conclusions:
The PI3K/Akt and RTK/Ras signalling pathways may be potential prognostic biomarkers for OCCC patients. Furthermore, our whole-genome sequencing data highlight important pathways for molecular and biological characterisations and potential therapeutic targeting in OCCC.
Similar content being viewed by others

Main
Ovarian clear cell carcinoma (OCCC) is recognised in the World Health Organisation classification of ovarian tumours as a distinct histological entity that demonstrates a markedly unique clinical behaviour from the other epithelial ovarian cancers (Scully, 1975; Itamochi et al, 2008). The OCCC constitutes ∼4–12% of all epithelial ovarian cancers in western countries and >20% in Japan (Itamochi et al, 2008). The OCCC was diagnosed twice as frequently (11.1%) among Asian women living in the United States that that among Caucasians (4.8%) (Chan et al, 2008). However, the reason for the ethnic differences of OCCC prevalence remains unknown. The poor prognosis of patients with advanced disease (median survival time 12.7 months) may reflect the resistance of CCC to conventional platinum- and taxane-based chemotherapy (Sugiyama et al, 2000; Itamochi et al, 2008). Recently, randomised phase III clinical trial of irinotecan plus cisplatin (CPT-P) compared with paclitaxel plus carboplatin (TC) in treating patients with CCC was conducted by the Japanese Gynecologic Oncology Group (JGOG) in collaboration with the Gynecologic Cancer Intergroup (GCIG; JGOG3017/GCIG Trial) (Sugiyama et al, 2016). However, no significant survival benefit was observed for CPT-P. Therefore, effective and novel treatment strategies (e.g., incorporating molecular targeted agents) are required to improve outcomes for women with advanced OCCC.
Previous molecular studies have shown that CCC had a variety of genetic alterations, such as frequent mutations of the AT-rich interactive domain 1A (ARID1A, 40–57%) and phosphatidylinositol-4,5-bisphosphate 3-kinase catalytic subunit α (PIK3CA, 33–50%) genes, amplifications of AKT2 (14%) and protein phosphatase, Mg2+/Mn2+ dependent 1D (PPM1D, 10%) genes, and loss of mismatch repair genes (7–18%) (Wilson and Roberts, 2011; Tan et al, 2013; Itamochi et al, 2015; Friedlander et al, 2016). Loss of phosphatase and tensin homolog (PTEN, 40–51%) expression and amplification and overexpression of Erb-b2 receptor tyrosine kinase 2 (ERBB2, 9.3–14%) has also been reported (Tan et al, 2013; Friedlander et al, 2016). Most of these studies have analysed only targeted genes, although ovarian cancers, including OCCC, have heterogeneous gene alterations (Bast et al, 2009; Tan et al, 2011).
Recently, high-throughput sequencing of DNA has been successfully applied to several cancers, enabling the discovery of cancer genes and network-attacking mutations that can possibly translate into advances in cancer diagnosis and treatment (Garraway and Lander, 2013; Creixell et al, 2015). In the present study, whole-genome sequencing of 55 fresh-frozen surgically resected OCCC tumours was performed to identify novel molecular drivers and molecular features that may be predictive of clinical behaviour and enable the development of novel therapeutic strategies for individual patients. Here we provide novel evidence that OCCC tumours have a high incidence of mutations of neuroblastoma breakpoint family (NBPF) member genes. We also found that activations of the phosphatidylinositol-3-kinase (PI3K)/Akt signalling pathway and the receptor tyrosine kinase (RTK)/Ras signalling pathway were independent prognostic factors for patients with OCCC.
Materials and methods
Clinical samples
Ovarian tumour tissue samples and matched blood samples were obtained from 55 Japanese women diagnosed with OCCC and treated surgically at Iwate Medical University, Tottori University, Niigata University, Tokai University, Tohoku University, Juntendo University, Shikoku Cancer Center, Kurume University, Hokkaido University, Keio University, Osaka University, Kagoshima City, or Hirosaki University Hospitals from 2003 to 2012. The study protocol was approved by the institutional review boards of each institution and all patients submitted written informed consent before collection of specimens, in accordance with institutional guidelines. These patients underwent complete surgical staging, including intraperitoneal cytology, bilateral salpingo-oophorectomy, hysterectomy, omentectomy, pelvic-/paraaortic lymphadenectomy, and aggressive cytoreductive surgery, followed by platinum-based chemotherapy. Thirty-five (64%) patients received postoperative chemotherapy, consisting of paclitaxel at 175 mg m−2 plus an area under the curve of 6.0 mg ml−1 of carboplatin on day 1, every 3 weeks for up to six cycles. The remaining 20 (36%) patients received 60 mg m−2 of irinotecan on days 1, 8, and 15, plus 60 mg m−2 of cisplatin on day 1, every 4 weeks for up to six cycles. There were 28 patients with stage I disease, 5 with stage II, 19 with stage III, and 3 with stage IV according to the surgicopathological staging guidelines of the International Federation of Gynaecology and Obstetrics (FIGO).
Tumour tissue samples (∼1 g each) were collected during surgery and then immediately frozen in liquid nitrogen and stored at −80 °C until assayed. Blood was withdrawn and fractionated as a pretreatment to separate the buffy coat, comprising white blood cells and platelets, from erythrocytes and plasma. The buffy coat was cryopreserved and was used for normal genomic DNA matching to the tumour specimens. Before whole-exome sequencing, germline single-nucleotide polymorphism analysis was performed on tumour-blood pairs to confirm identities, showing matching of all tumour samples to the respective blood samples.
Massively parallel sequencing
Genomic DNA from tumour and blood was extracted using the DNeasy Blood & Tissue DNA kit (QIAGEN, Hilden, Germany). Exome capture was performed using the Agilent SureSelect BTA XT AUTO Human All Exon V5+IncRNA Platform (Agilent Technologies, Santa Clara, CA, USA) according to the manufacturer’s instructions. Whole-genome sequencing was performed on the Illumina HiSeq platform (Illumina, Inc., San Diego, CA, USA) according to standard protocols (Imielinski et al, 2012). Base calling and quality scoring was conducted using Real-Time Analysis Software, then de-multiplexing and generation of FASTQ files were performed using the CASAVA package provided by Illumina (Illumina, Inc.). The data set analysed here (the data set ID: JGAS00000000076) is available at the National Bioscience Database Center (NBDC) website (https://humandbs.biosciencedbc.jp/en/hum0067-v1) in controlled access.
Detection of somatic single-nucleotide variants (SNVs) and short insertion and deletion (INDEL) variants
Mapping of genomes of raw sequence data (FASTQ files) was performed using the Burrows–Wheeler alignment algorithm (Li and Durbin, 2009). Then, realignment, recalibration, and generation of binary sequence alignment/map files were performed using the GATK toolkit and SAMtools (Li et al, 2009; McKenna et al, 2010). The SNVs and INDELs were detected by comparing sequence data from tumour-blood pairs using the MuTect, VarScan, and SomaticIndelDetector GATK algorithms (Koboldt et al, 2012; Cibulskis et al, 2013). Then, the ANNOVAR tool was used to annotate all SNVs and INDELs (Wang et al, 2010).
Somatic copy number variation (CNV) detection
Exome CNV and Control-FREEC were used to analyse the copy number (CN) in sequence data from tumour-blood pairs (Sathirapongsasuti et al, 2011; Boeva et al, 2012). A CN of ⩾4 was considered as a CN gain and the size of regions of ⩽3 Mb were defined as focal amplifications.
Statistical analyses
Statistical analyses were performed with JMP, version 12, software (SAS Institute Inc., Cary, NC, USA) and GraphPad PRISM, version 7, software (GraphPad Software, Inc., La Jolla, CA, USA). The χ2 test, Fisher’s exact test, and unpaired t-test were used for statistical analysis. Cluster analyses were performed by hierarchical clustering with Ward’s minimum variance method. Survival distributions were calculated using the Kaplan–Meier method and the significance of apparent differences in survival distribution between groups was tested using the log-rank test. In addition, the Cox proportional hazards model for multivariable analysis was applied. A probability (P) value of <0.05 was considered statistically significant.
Results
Identification of gene mutations
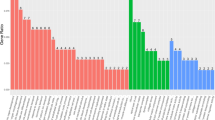
A total of 4792 genomic alterations were identified in the 55 tumours with a median of 62 alterations per tumour (range 25–2676), although no germline mutations and CNVs were correlated with these genomic alterations. Of these, 37 genes were found to be mutated in at least 10 (18%) of the tumours, with the most common alterations being NBPF members 20 (67% of tumours), 10 (60%), and 14 (60%) (Supplementary Table 1). A total of 51 (93%) cases had a mutation in NBPF20, 10, and 14 genes, and these mutations were all nonsynonymous SNVs. The relationship between nonsynonymous SNVs of these genes and clinicopathological factors and prognostic significance in OCCC were not observed (Supplementary Table 2). Mutations of ARID1A were found in 23 (42%) tumours. Univariate analysis revealed that mutations in PIK3CA (35% of tumours), major histocompatibility complex, class II, DR β1 (HLA-DRB1) (25%), mucin 4, cell surface associated (MUC4) (22%), zinc finger protein 717 (ZNF717) (22%), or ARID1B (18%) of these 37 genes were correlated with better overall survival (OS) of patients with OCCC. In contrast, mutations of TBC1 domain family member 3 (TBC1D3) (24%) or ataxin 1 (ATXN1) (18%) were correlated with poor OS of these patients.
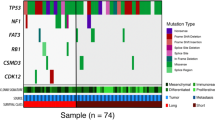
Unsupervised hierarchical clustering by these 37 genes grouped the tumours into four clusters (Figure 1A). Cluster 1 tumours had frequent mutations of NBPF20 (88% of these tumours) and NBPF10 (88%). Cluster 2 tumours were mostly lacking mutations of ARID1A (0.5% of these tumours), PIK3CA (0.5%), and ARID1B (0%). Patients in clusters 2 and 4 tended to have worse OS compared with those in clusters 1 and 3, with the 3-year survival rates being 52 and 75% (hazard ratio 2.24 (95% confidence interval (CI) 0.96–5.24), P=0.0623), respectively (Figure 1B).
Somatic mutations in ovarian clear cell carcinomas. (A) Tumours were hierarchically clustered into four groups based on somatic mutations. The heat map shows somatic mutations in each tumour (horizontal axis) plotted by genes (vertical axis). (B) Kaplan–Meier curves of overall survival (OS) for each mutation cluster. There was no significant difference in OS between patients in each cluster.
Pathways influencing patient’s outcome
Although the relationship between molecular alterations of genes and clinicopathological variables and prognostic significance in several cancers has been reported recently, it is unclear whether gene alterations affect the clinical behaviour of OCCC. Therefore, we investigated whether the alteration status could be a prognostic biomarker for this disease.
Somatic mutations of ARID1A, which encodes a member of the switch/sucrose non-fermentable (SWI/SNF) family protein, were identified in approximately half of OCCC cases (Wilson and Roberts, 2011; Itamochi et al, 2015). We therefore examined mutations of the SWI/SNIF subunit genes. A total of 28 (51%) cases had a mutation in ARID1A (42%), ARID1B (18%), B-cell CLL/Lymphoma 11A (BCL11A) (2%), double PHD fingers 1 (DPF1) (2%), SWI/SNF related, matrix associated, actin dependent regulator of chromatin (SMARC) A1 (2%), SMARCA2 (2%), SMARCA4 (5%), or SMARCC1 (2%) (Figure 2A). These mutations occurred more frequently in patients with FIGO stage I/II disease compared with those with stage III/IV (Table 1). Univariate analysis demonstrated that mutations of the SWI/SNF subunit genes tend to correlate with better OS among patients with OCCC, with the 3-year survival rates being 74 and 47% (hazard ratio 0.46 (95% CI 0.20–1.07), P=0.0711), respectively (Figure 2B). However, the OS rate was not significantly higher in patients with FIGO stage I or II OCCC with mutations of the SWI/SNF subunit genes, compared with those with the wild-type phenotype of these genes, with 3-year survival rates of 85 and 67% (hazard ratio 0.37 (95% CI 0.08–1.75), P=0.2082), respectively. Similarly, these mutations did not influence OS of patients with FIGO stage III or IV OCCC (hazard ratio 1.12 (95% CI 0.37–3.34), P=0.8464).
Mutations of the switch/sucrose non-fermentable (SWI/SNF) family subunit genes and outcome prediction in ovarian clear cell carcinoma of the ovary (OCCC). (A) The patterns of gene mutations of the SWI/SNF family subunit. Mutations of the SWI/SNF family subunit gene were observed in 51% of cases. (B) Kaplan–Meier analysis of OS for mutation status of SWI/SNF family subunit genes. No significant difference in OS was observed according to the mutation status of these genes.
Next we analysed alterations to PI3K/Akt signalling genes. A total of 23 (42%) cases had alteration in these genes, including mutations of PIK3CA (35%), PIK3R1 (7%), and PTEN (2%), and amplifications of PIK3R2 (5%), AKT1 (4%), and AKT2 (9%) (Figure 3A and B). These gene alterations were frequently found in patients with FIGO stage I/II OCCC, compared with those with stage III/IV (Table 1), and were significantly more common in tumours with mutations of the SWI/SNIF subunit genes (82%, P<0.0001). Univariate analysis demonstrated that activation of the PI3K/Akt signalling pathway was correlated with better OS of patients with OCCC, with 3-year survival rates of 91% and 40% (hazard ratio 0.24 (95% CI 0.10–0.56), P=0.0010), respectively (Figure 3C). After adjustment for patient age, FIGO stage, and treatment regimen, activation of this pathway was also correlated with better OS of patients with OCCC (hazard ratio 0.27 (95% CI 0.06–0.86), P=0.0249). Interestingly, the OS rate was also significantly higher in patients with FIGO stage I or II OCCC with activated pathway compared with not activated, with 3-year survival rates of 94% and 61% (hazard ratio 0.19 (95% CI 0.04–0.83), P=0.0276), respectively.
Alterations to the phosphatidylinositol-3-kinase (PI3K)/Akt signalling pathway and outcome prediction in ovarian clear cell carcinoma (OCCC). (A) The PI3K/Akt signalling pathway is altered through several mechanisms. Alteration frequencies are expressed as the percentage of all cases. Activated genes are shown in red and inactivated genes in blue. (B) Patterns of gene alterations in the PI3K/Akt signalling pathway. Alterations to the PI3K/Akt signalling pathway genes were observed in 42% of cases. (C) Kaplan–Meier analysis of overall survival (OS) for the activation status of the PI3K/Akt signalling pathway. Overall survival was significantly higher in patients with activation of the PI3K/Akt signalling pathway compared with those with no activation of this pathway.
Alterations of the RTK/Ras signalling pathways genes were found in a total of 16 (29%) cases, including amplifications of ERBB2 (11%) and ERBB3 (5%), and mutations of ERBB2 (4%), ERBB3 (7%), KRAS (9%), and BRAF (2%) (Figure 4A and B). Of these alterations, the RTK/Ras signalling pathway was activated in 11 (20%) cases. Univariate analysis revealed that activation of this signalling pathway was correlated with better OS of patients with OCCC, with 3-year survival rates of 91% and 53% (hazard ratio 0.35 (95% CI 0.13–0.94), P=0.0373), respectively (Figure 4C). After adjustment for patient age, FIGO stage, and treatment regimen, activation of this pathway was also correlated with better OS of patients with OCCC (hazard ratio 0.11 (95% CI 0.01–0.54), P=0.0034).
Pathway alterations of receptor tyrosine kinase (RTK)/Ras signalling and outcome prediction in ovarian clear cell carcinoma (OCCC). (A) The RTK/Ras signalling pathway is altered through several mechanisms. Alteration frequencies are expressed as the percentage of all cases. Activated genes are shown in red and inactivated genes in blue. (B) Patterns of gene alterations to the RTK/Ras signalling pathway. Alterations to the RTK/Ras signalling pathway genes were observed in 29% of cases. (C) Kaplan–Meier analysis of overall survival (OS) for the activation status of the RTK/Ras signalling pathway. Overall survival was significantly higher in patients with activation of the RTK/Ras signalling pathway compared with those with no activation of this pathway.
Multivariable analysis of age, FIGO stage, and genes alterations was performed that found that FIGO stage, activation of PI3K/Akt signalling, and activation of RTK/Ras signalling were independent prognostic factors of OCCC (Table 2).
Discussion
Analysis of whole-genome sequencing data from 55 OCCC tumours revealed that the ARID1A and PIK3CA genes, which are known to be altered in OCCC, were frequently altered in this study. We also found frequent genetic alterations in NBPF member genes. To our knowledge, this is the first study to report the comprehensive landscape of somatic genome alterations in OCCC.
The ARID1A encodes a member of the SWI/SNF family protein BAF250a (ARID1A) and is the most frequently mutated of the SWI/SNF subunit genes in solid cancers, including OCCC (Wilson and Roberts, 2011). The SWI/SNF family proteins, which form SWI/SNF complexes, are involved in the regulation of many cellular processes, including development, differentiation, proliferation, DNA repair, and tumour suppression. Indeed, we found mutations in the SWI/SNF subunit genes in 51% (28/55) of cases and the most frequently mutated genes were ARID1A (42%) followed by ARID1B (18%) and SMARCA4 (5%). Eight of the 10 cases (80%) with an ARID1B mutation also had an ARID1A mutation. In contrast, 2 of the 3 cases (67%) with an SMARCA4 mutation harboured no mutations in ARID1A or ARID1B. Similarly, an SMARCA4 mutation was observed in 1 of 8 OCCC cases (13%) that did not have mutations of ARID1A (5 of these tumours) or ARID1B (one) (Shain and Pollack, 2013). Both the ARID1A and ARID1B subunits contain an ARID domain that binds DNA in a sequence-nonspecific manner, whereas the SMARCA4 protein, also known as BRG1, is a catalytic ATPase subunit of the SWI/SNF complex (Wilsker et al, 2004; Wilson and Roberts, 2011). These observations suggest that one of these mutations may be sufficient for deterioration of the tumour suppressor role of this complex in OCCC.
The impact of ARID1A status on the treatment outcome of patients with OCCC has been evaluated. Several studies failed to find a correlation between negative expression of the ARID1A protein or mutations of the gene and OS of patients with OCCC (Itamochi et al, 2015). However, we previously found that the 5-year OS rate for FIGO stage I or II OCCC patients with negative tumour expression of ARID1A was lower than that of patients with positive tumour expression of ARID1A (74% vs 91% respectively). Another study showed that the loss of ARID1A expression was significantly correlated with shorter progression-free survival for patients with OCCC, but not OS (Katagiri et al, 2012). In contrast, Abou-Taleb et al (2016) reported that the loss of expression of one or multiple SWI/SNF subunit proteins demonstrated aggressive behaviour and poor prognosis of OCCC. In this study however, we observed no significant difference in OS between patients with mutations of the SWI/SNF subunit genes and those with the wild-type phenotypes of these genes, regardless of FIGO stage. Therefore, further studies are needed to elucidate the prognostic significance of SWI/SNF subunit alterations in OCCC.
The relationship between the loss of ARID1A expression and activation of the PI3K/Akt pathway has been reported in various cancers, including OCCC, suggesting a collaboration in tumourigenesis (Yamamoto et al, 2012; Zang et al, 2012; Bosse et al, 2013; Huang et al, 2014). Yamamoto et al (2012) reported that mutations of PIK3CA (which encodes the catalytic subunit p110α of PI3K) were detected in 40% (17/42) of OCCC tumours and a majority (71%) of these were found in ARID1A-deficient carcinomas. Another study also showed that the loss of ARID1A was more frequent in OCCC tumours with an activated PI3K/Akt pathway (PIK3CA mutations or loss of PTEN expression) (54%) than those without alterations to the PI3K/Akt pathway (30%) (Huang et al, 2014). Indeed, in the present study, 82% of tumours with activation of the PI3K/Akt pathway were observed in tumours with mutations of the SWI/SNIF subunit genes. Interestingly, clustering analysis revealed that tumours in cluster 2 were almost lacking mutations of ARID1A, PIK3CA, and ARID1B, and patients in this cluster tended to have worse OS compared with those in clusters 1 and 3. Furthermore, univariate and multivariable analyses revealed that activation of the PI3K/Akt signalling pathway, but not mutations of the SWI/SNIF subunit genes, was correlated with better OS of patients with OCCC. Several studies have shown that PIK3CA mutations or overexpression were correlated with improved OS of patients with OCCC (Rahman et al, 2012; Abe et al, 2013). Other studies, however, reported that PIK3CA status was not a prognostic factor for these patients (Huang et al, 2014; Ye et al, 2016). Although the prognostic significance of PIK3CA alterations remains controversial, these novel findings suggest that activation of the PI3K/Akt signalling pathway may have some effect on the treatment outcome of patients with OCCC.
Amplification and overexpression of ERBB2, also known as human epidermal growth factor receptor 2 (HER2), and mutations of KRAS have been reported in 9.3–14% and 4.7–13% of OCCC cases respectively (Tan et al, 2011, 2013; Friedlander et al, 2016; Zannoni et al, 2016). Overexpression of HER2 is associated with poor sensitivity to conventional anticancer agents and poor prognosis in several types of tumours, including ovarian cancer (Kim et al, 1998; Rolitsky et al, 1999; Itamochi et al, 2008). On the contrary, Nodin et al (2013) reported that a KRAS mutation was associated with a significant improvement in cancer-specific survival in patients with ovarian endometrioid carcinoma. However, the prognostic significance of these signalling pathways for OCCC remain largely unknown. Here, we found that the activation of the RTK/Ras signalling pathway was observed in 20% of tumours and was a favourable prognostic factor for OCCC. Moreover, the RTK/Ras pathway and its downstream signalling pathway are thought to be potential targets for cancer therapy (Mandal et al, 2016). We previously reported that inhibition of mitogen-activated protein kinase (MEK) 1/2, which is the downstream signalling cascade of RTK/Ras, by selumetinib reduced growth of OCCC cell lines and suppressed tumour growth in a OCCC xenograft model (Bartholomeusz et al, 2012). These findings suggest that the RTK/Ras signalling pathway may be an important prognostic biomarker for patients with OCCC and a potential therapeutic target for OCCC.
The genomic alterations have also been examined in high-grade serous ovarian cancer (HGS-OvCa), which is the most common histological subtype, by the Cancer Genome Atlas (TCGA) study (Cancer Genome Atlas Research Network, 2011). These TCGA analyses reveal that the BRCA1 and BRCA2 mutations in 20% HGS-OvCa samples triggered aberrations in DNA damage repair by homologous recombination. Patients with germline BRCA1/2 mutation are thought to have clinical benefit from using PARP (poly ADP-ribose polymerase) inhibitors, such as olaparib (Sato and Itamochi, 2015). However, in this study of OCCC, only 3 cases (5%) had somatic BRCA2 mutations (2 nonsynonymous SNV and 1 frameshift insertion) (data not shown). Similarly, almost all HGS-OvCa tumours (96%) had mutations in TP53, but 3 OCCC tumours (5%) had mutations (1 stopgain, 1 frameshift deletion, and 1 nonsynonymous SNV) in this gene (data not shown). On the contrary, ARID1A and PIK3CA mutations were more common in OCCC cases compared with that in HGS-OvCa. These differences between ovarian cancer subtypes suggested that subtype-specific treatment strategies might be needed to improve ovarian cancer outcomes.
The limitations of this study are that we have analysed only DNA sequencing data in a relatively small number of OCCC samples, and it lacks independent validation of the prognostic association. Therefore, the sequencing of larger OCCC cohorts will be necessary to determine a more comprehensive genomic landscape for OCCC. Furthermore, precise prevalence frequency and prognostic significance for the mutated genes detected in our analysis also need to be validated by DNA sequencing as well as by multiple-assay platform. Recently, Friedlander et al (2016) examined the results of a multiplatform profiling panel, such as DNA sequencing, immunohistochemistry, fluorescent or chromogenic in situ hybridisation, and RNA fragment analysis, in OCCC to identify the potential therapeutic targets. Consistent with our results, they confirmed that the PIK3CA/Akt/mTOR pathway was altered in 61% OCCC tumours.
In summary, our data showed that mutations of ARID1A and PIK3CA were frequently observed in OCCC tumours. We also found that the activation of PI3K/Akt and RTK/Ras signalling pathways may be a favourable prognostic marker for patients with OCCC. We believe that these whole-genome sequencing data will be valuable and useful for further analysis of the molecular and biological characteristics of OCCC, and may lead to the establishment of novel treatment strategies to improve survival of patients with OCCC.
References
Abe A, Minaguchi T, Ochi H, Onuki M, Okada S, Matsumoto K, Satoh T, Oki A, Yoshikawa H (2013) PIK3CA overexpression is a possible prognostic factor for favorable survival in ovarian clear cell carcinoma. Hum Pathol 44: 199–207.
Abou-Taleb H, Yamaguchi K, Matsumura N, Murakami R, Nakai H, Higasa K, Amano Y, Abiko K, Yoshioka Y, Hamanishi J, Koshiyama M, Baba T, Yamada R, Matsuda F, Konishi I, Mandai M (2016) Comprehensive assessment of the expression of the SWI/SNF complex defines two distinct prognostic subtypes of ovarian clear cell carcinoma. Oncotarget 7: 54758–54770.
Bartholomeusz C, Oishi T, Saso H, Akar U, Liu P, Kondo K, Kazansky A, Krishnamurthy S, Lee J, Esteva FJ, Kigawa J, Ueno NT (2012) MEK1/2 inhibitor selumetinib (AZD6244) inhibits growth of ovarian clear cell carcinoma in a PEA-15-dependent manner in a mouse xenograft model. Mol Cancer Ther 11: 360–369.
Bast RC Jr, Hennessy B, Mills GB (2009) The biology of ovarian cancer: new opportunities for translation. Nat Rev Cancer 9: 415–428.
Boeva V, Popova T, Bleakley K, Chiche P, Cappo J, Schleiermacher G, Janoueix-Lerosey I, Delattre O, Barillot E (2012) Control-FREEC: a tool for assessing copy number and allelic content using next-generation sequencing data. Bioinformatics 28: 423–425.
Bosse T, ter Haar NT, Seeber LM, v Diest PJ, Hes FJ, Vasen HF, Nout RA, Creutzberg CL, Morreau H, Smit VT (2013) Loss of ARID1A expression and its relationship with PI3K-Akt pathway alterations, TP53 and microsatellite instability in endometrial cancer. Mod Pathol 26: 1525–1535.
Cancer Genome Atlas Research Network (2011) Integrated genomic analyses of ovarian carcinoma. Nature 474: 609–615.
Chan JK, Teoh D, Hu JM, Shin JY, Osann K, Kapp DS (2008) Do clear cell ovarian carcinomas have poorer prognosis compared to other epithelial cell types? A study of 1411 clear cell ovarian cancers. Gynecol Oncol 109: 370–376.
Cibulskis K, Lawrence MS, Carter SL, Sivachenko A, Jaffe D, Sougnez C, Gabriel S, Meyerson M, Lander ES, Getz G (2013) Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat Biotechnol 31: 213–219.
Creixell P, Schoof EM, Simpson CD, Longden J, Miller CJ, Lou HJ, Perryman L, Cox TR, Zivanovic N, Palmeri A, Wesolowska-Andersen A, Helmer-Citterich M, Ferkinghoff-Borg J, Itamochi H, Bodenmiller B, Erler JT, Turk BE, Linding R (2015) Kinome-wide decoding of network-attacking mutations rewiring cancer signaling. Cell 163: 202–217.
Friedlander ML, Russell K, Millis S, Gatalica Z, Bender R, Voss A (2016) Molecular profiling of clear cell ovarian cancers: identifying potential treatment targets for clinical trials. Int J Gynecol Cancer 26: 648–654.
Garraway LA, Lander ES (2013) Lessons from the cancer genome. Cell 153: 17–37.
Huang HN, Lin MC, Huang WC, Chiang YC, Kuo KT (2014) Loss of ARID1A expression and its relationship with PI3K-Akt pathway alterations and ZNF217 amplification in ovarian clear cell carcinoma. Mod Pathol 27: 983–990.
Imielinski M, Berger AH, Hammerman PS, Hernandez B, Pugh TJ, Hodis E, Cho J, Suh J, Capelletti M, Sivachenko A, Sougnez C, Auclair D, Lawrence MS, Stojanov P, Cibulskis K, Choi K, de Waal L, Sharifnia T, Brooks A, Greulich H, Banerji S, Zander T, Seidel D, Leenders F, Ansen S, Ludwig C, Engel-Riedel W, Stoelben E, Wolf J, Goparju C, Thompson K, Winckler W, Kwiatkowski D, Johnson BE, Janne PA, Miller VA, Pao W, Travis WD, Pass HI, Gabriel SB, Lander ES, Thomas RK, Garraway LA, Getz G, Meyerson M (2012) Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150: 1107–1120.
Itamochi H, Kigawa J, Terakawa N (2008) Mechanisms of chemoresistance and poor prognosis in ovarian clear cell carcinoma. Cancer Sci 99: 653–658.
Itamochi H, Oumi N, Oishi T, Shoji T, Fujiwara H, Sugiyama T, Suzuki M, Kigawa J, Harada T (2015) Loss of ARID1A expression is associated with poor prognosis in patients with stage I/II clear cell carcinoma of the ovary. Int J Clin Oncol 20: 967–973.
Katagiri A, Nakayama K, Rahman MT, Rahman M, Katagiri H, Nakayama N, Ishikawa M, Ishibashi T, Iida K, Kobayashi H, Otsuki Y, Nakayama S, Miyazaki K (2012) Loss of ARID1A expression is related to shorter progression-free survival and chemoresistance in ovarian clear cell carcinoma. Mod Pathol 25: 282–288.
Kim YC, Park KO, Kern JA, Park CS, Lim SC, Jang AS, Yang JB (1998) The interactive effect of Ras, HER2, P53 and Bcl-2 expression in predicting the survival of non-small cell lung cancer patients. Lung Cancer 22: 181–190.
Koboldt DC, Zhang Q, Larson DE, Shen D, McLellan MD, Lin L, Miller CA, Mardis ER, Ding L, Wilson RK (2012) VarScan 2: somatic mutation and copy number alteration discovery in cancer by exome sequencing. Genome Res 22: 568–576.
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25: 1754–1760.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R 1000 Genome Project Data Processing Subgroup (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25: 2078–2079.
Mandal R, Becker S, Strebhardt K (2016) Stamping out RAF and MEK1/2 to inhibit the ERK1/2 pathway: an emerging threat to anticancer therapy. Oncogene 35: 2547–2561.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20: 1297–1303.
Nodin B, Zendehrokh N, Sundstrom M, Jirstrom K (2013) Clinicopathological correlates and prognostic significance of KRAS mutation status in a pooled prospective cohort of epithelial ovarian cancer. Diagn Pathol 8: 106.
Rahman M, Nakayama K, Rahman MT, Nakayama N, Ishikawa M, Katagiri A, Iida K, Nakayama S, Otsuki Y, Shih Ie M, Miyazaki K (2012) Clinicopathologic and biological analysis of PIK3CA mutation in ovarian clear cell carcinoma. Hum Pathol 43: 2197–2206.
Rolitsky CD, Theil KS, McGaughy VR, Copeland LJ, Niemann TH (1999) HER-2/neu amplification and overexpression in endometrial carcinoma. Int J Gynecol Pathol 18: 138–143.
Sathirapongsasuti JF, Lee H, Horst BA, Brunner G, Cochran AJ, Binder S, Quackenbush J, Nelson SF (2011) Exome sequencing-based copy-number variation and loss of heterozygosity detection: ExomeCNV. Bioinformatics 27: 2648–2654.
Sato S, Itamochi H (2015) DNA Repair and Chemotherapy. InTech.
Scully RE (1975) World Health Organization classification and nomenclature of ovarian cancer. Natl Cancer Inst Monogr 42: 5–7.
Shain AH, Pollack JR (2013) The spectrum of SWI/SNF mutations, ubiquitous in human cancers. PLoS One 8: e55119.
Sugiyama T, Kamura T, Kigawa J, Terakawa N, Kikuchi Y, Kita T, Suzuki M, Sato I, Taguchi K (2000) Clinical characteristics of clear cell carcinoma of the ovary: a distinct histologic type with poor prognosis and resistance to platinum-based chemotherapy. Cancer 88: 2584–2589.
Sugiyama T, Okamoto A, Enomoto T, Hamano T, Aotani E, Terao Y, Suzuki N, Mikami M, Yaegashi N, Kato K, Yoshikawa H, Yokoyama Y, Tanabe H, Nishino K, Nomura H, Kim JW, Kim BG, Pignata S, Alexandre J, Green J, Isonishi S, Terauchi F, Fujiwara K, Aoki D (2016) Randomized phase III trial of irinotecan plus cisplatin compared with paclitaxel plus carboplatin as first-line chemotherapy for ovarian clear cell carcinoma: JGOG3017/GCIG Trial. J Clin Oncol 34: 2881–2887.
Tan DS, Iravani M, McCluggage WG, Lambros MB, Milanezi F, Mackay A, Gourley C, Geyer FC, Vatcheva R, Millar J, Thomas K, Natrajan R, Savage K, Fenwick K, Williams A, Jameson C, El-Bahrawy M, Gore ME, Gabra H, Kaye SB, Ashworth A, Reis-Filho JS (2011) Genomic analysis reveals the molecular heterogeneity of ovarian clear cell carcinomas. Clin Cancer Res 17: 1521–1534.
Tan DS, Miller RE, Kaye SB (2013) New perspectives on molecular targeted therapy in ovarian clear cell carcinoma. Br J Cancer 108: 1553–1559.
Wang K, Li M, Hakonarson H (2010) ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res 38: e164.
Wilsker D, Patsialou A, Zumbrun SD, Kim S, Chen Y, Dallas PB, Moran E (2004) The DNA-binding properties of the ARID-containing subunits of yeast and mammalian SWI/SNF complexes. Nucleic Acids Res 32: 1345–1353.
Wilson BG, Roberts CW (2011) SWI/SNF nucleosome remodellers and cancer. Nat Rev Cancer 11: 481–492.
Yamamoto S, Tsuda H, Takano M, Tamai S, Matsubara O (2012) Loss of ARID1A protein expression occurs as an early event in ovarian clear-cell carcinoma development and frequently coexists with PIK3CA mutations. Mod Pathol 25: 615–624.
Ye S, Yang J, You Y, Cao D, Huang H, Wu M, Chen J, Lang J, Shen K (2016) Clinicopathologic significance of HNF-1beta, AIRD1A, and PIK3CA expression in ovarian clear cell carcinoma: a tissue microarray study of 130 cases. Medicine (Baltimore) 95: e3003.
Zang ZJ, Cutcutache I, Poon SL, Zhang SL, McPherson JR, Tao J, Rajasegaran V, Heng HL, Deng N, Gan A, Lim KH, Ong CK, Huang D, Chin SY, Tan IB, Ng CC, Yu W, Wu Y, Lee M, Wu J, Poh D, Wan WK, Rha SY, So J, Salto-Tellez M, Yeoh KG, Wong WK, Zhu YJ, Futreal PA, Pang B, Ruan Y, Hillmer AM, Bertrand D, Nagarajan N, Rozen S, Teh BT, Tan P (2012) Exome sequencing of gastric adenocarcinoma identifies recurrent somatic mutations in cell adhesion and chromatin remodeling genes. Nat Genet 44: 570–574.
Zannoni GF, Improta G, Pettinato A, Brunelli C, Troncone G, Scambia G, Fraggetta F (2016) Molecular status of PI3KCA, KRAS and BRAF in ovarian clear cell carcinoma: an analysis of 63 patients. J Clin Pathol 69: 1088–1092.
Acknowledgements
This study was supported by the Project for Development of Innovative Research on Cancer Therapeutics, from the Ministry of Education, Culture, Sports, Science and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This work is published under the standard license to publish agreement. After 12 months the work will become freely available and the license terms will switch to a Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License.
Supplementary Information accompanies this paper on British Journal of Cancer website
Supplementary information
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 4.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Itamochi, H., Oishi, T., Oumi, N. et al. Whole-genome sequencing revealed novel prognostic biomarkers and promising targets for therapy of ovarian clear cell carcinoma. Br J Cancer 117, 717–724 (2017). https://doi.org/10.1038/bjc.2017.228
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/bjc.2017.228
- Springer Nature Limited
Keywords
This article is cited by
-
From clinical management to personalized medicine: novel therapeutic approaches for ovarian clear cell cancer
Journal of Ovarian Research (2024)
-
Radiation therapy with phenotypic medicine: towards N-of-1 personalization
British Journal of Cancer (2024)
-
Treatment for ovarian clear cell carcinoma with combined inhibition of WEE1 and ATR
Journal of Ovarian Research (2023)
-
Ovarian clear cell carcinoma with or without endometriosis origin in a single institution cohort
Discover Oncology (2023)
-
Genomic landscapes of ovarian clear cell carcinoma from latin countries reveal aberrations linked to survival and progression
BMC Cancer (2023)