Abstract
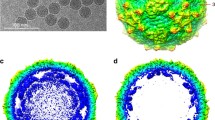
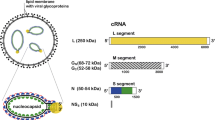
Rotavirus, the major cause of life-threatening infantile gastroenteritis, is a member of the Reoviridae1. Although the structures of rotavirus2 and other members of the Reoviridae3,4 have been extensively studied, little is known about the structures of virus-encoded non-structural proteins that are essential for genome replication and packaging. The non-structural protein NSP2 of rotavirus, which exhibits nucleoside triphosphatase, single-stranded RNA binding5, and nucleic-acid helix-destabilizing6 activities, is a major component of viral replicase complexes7,8. We present here the X-ray structure of the functional octamer9 of NSP2 determined to a resolution of 2.6 Å. The NSP2 monomer has two distinct domains. The amino-terminal domain has a new fold. The carboxy-terminal domain resembles the ubiquitous cellular histidine triad (HIT) group of nucleotidyl hydrolases10. This structural similarity suggests that the nucleotide-binding site is located inside the cleft between the two domains. Prominent grooves that run diagonally across the doughnut-shaped octamer are probable locations for RNA binding. Several RNA binding sites, resulting from the quaternary organization of NSP2 monomers, may be required for the helix destabilizing activity of NSP2 and its function during genome replication and packaging.



Similar content being viewed by others
References
Estes, M. K. et al. in Virology (ed. Fields, B. N.) 1625–1655 (Raven, New York, 1996)
Prasad, B. V. V. et al. Visualization of ordered genomic RNA and localization of transcriptional complexes in rotavirus. Nature 382, 471–473 (1996)
Reinisch, K. M., Nibert, M. L. & Harrison, S. C. Structure of the reovirus core at 3.6-Å resolution. Nature 404, 960–967 (2000)
Grimes, J. M. et al. The atomic structure of the bluetongue virus core. Nature 395, 470–478 (1998)
Taraporewala, Z., Chen, D. & Patton, J. Multimers formed by the rotavirus nonstructural protein NSP2 bind to RNA and have nucleoside triphosphatase activity. J. Virol. 73, 9934–9943 (1999)
Taraporewala, Z. & Patton, J. Identification and characterization of the helix-destabilizing activity of rotavirus nonstructural protein NSP2. J. Virol. 75, 4519–4527 (2001)
Aponte, C., Poncet, D. & Cohen, J. Recovery and characterization of a replicase complex in rotavirus-infected cells by using a monoclonal antibody against NSP2. J. Virol. 70, 985–991 (1996)
Gallegos, C. & Patton, J. Characterization of rotavirus replication intermediates: a model for the assembly of single-shelled particles. Virology 172, 616–627 (1989)
Schuck, P., Taraporewala, Z., McPhie, P. & Patton, J. T. Rotavirus nonstructural protein NSP2 self-assembles into octamers that undergo ligand-induced conformational changes. J. Biol. Chem. 276, 9679–9687 (2001)
Lima, C. D., Klein, M. G. & Hendrickson, W. A. Structure-based analysis of catalysis and substrate definition in the HIT protein family. Science 278, 286–290 (1997)
Petrie, B. L., Greenberg, H. B., Graham, D. Y. & Estes, M. K. Ultrastructural localization of rotavirus antigens using colloidal gold. Virus Res. 1, 133–152 (1984)
Kattoura, M., Chen, X. & Patton, J. The rotavirus RNA-binding protein NS35 (NSP2) forms 10S multimers and interacts with the viral RNA polymerase. Virology 202, 803–813 (1994)
Ramig, R. & Petrie, B. L. Characterization of temperature-sensitive mutants of simian rotavirus SA11:protein synthesis and morphogenesis. J. Virol. 49, 665–673 (1984)
Chen, D., Gombold, J. L. & Ramig, R. F. Intracellular RNA synthesis directed by temperature-sensitive mutants of simian rotavirus SA11. Virology 178, 143–151 (1990)
Uitenweerde, J. M., Theron, J., Stoltz, M. A. & Huismans, H. The multimeric nonstructural NS2 proteins of bluetongue virus, African horsesickness virus, and epizootic hemorrhagic disease virus differ in their single-stranded RNA-binding ability. Virology 209, 624–632 (1995)
Taraporewala, Z., Chen, D. & Patton, J. Multimers of the bluetongue virus nonstructural protein, NS2, possess nucleotidyl phosphatase activity: similarities between NS2 and rotavirus NSP2. Virology 280, 221–231 (2001)
Gillian, A. L., Schmechel, S. C., Livny, J., Schiff, L. A. & Nibert, M. L. Reovirus protein sigma NS binds in multiple copies to single-stranded RNA and shares properties with single-stranded DNA binding proteins. J. Virol. 74, 5939–5948 (2000)
Cohen, J. Rethinking a vaccine risk. Science 293, 1576–1577 (2001)
Holm, L. & Sander, C. Protein folds and families: sequence and structure alignments. Nucleic Acid Res. 27, 244–247 (1999)
Lima, C. D., Klein, M. G., Weinstein, I. B. & Hendrickson, W. A. Three-dimensional structure of human protein kinase C interacting protein 1, a member of the HIT family of proteins. Proc. Natl. Acad. Sci. USA 28, 5357–5362 (1996)
Brenner, C., Biegonowski, P., Pace, H. C. & Huebner, K. The histidine superfamily of nucleotide-binding proteins. J. Cell. Phys. 181, 179–187 (1999)
Raghunathan, S., Kozlov, A. G., Lohman, T. M. & Waksman, G. Structure of the DNA binding domain of E. coli SSB bound to ssDNA. Nature Struct. Biol. 7, 648–652 (2000)
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Collaborative Computational Project, Number 4. The CCP4 Suite: Programs for Protein Crystallography. Acta Crystallogr. D 50, 760–763 (1994)
Brunger, A. T. et al. Crystallography and NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54, 905–921 (1998)
Jones, T. A., Zou, J.-Y., Cowan, S. W. & Kjeldgaard, M. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991)
Brünger, A. T. Free R value: a novel statistical quantity for assessing the accuracy of crystal structures. Nature 355, 472–475 (1992)
Kraulis, P. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991)
Merritt, E. A. & Bacon, D. J. Raster3D: photorealistic molecular graphics. Methods Enzymol. 277, 505–524 (1997)
Nicholls, A., Sharp, K. A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991)
Acknowledgements
We thank A. Nickitenko, B. Bowman, W. Meador and the National Synchroton Light Source beamline (X8C) staff for help with data collection, and F.A. Quiocho for allowing us to use X-ray diffraction facilities at the Baylor College of Medicine. We also thank M. K. Estes and R. F. Ramig, and M. Sowa, for helpful discussions. This work was supported by grants from the NIH and R. Welch foundation to B.V.V.P and an NIH intramural grant to J.P.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Jayaram, H., Taraporewala, Z., Patton, J. et al. Rotavirus protein involved in genome replication and packaging exhibits a HIT-like fold. Nature 417, 311–315 (2002). https://doi.org/10.1038/417311a
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/417311a
- Springer Nature Limited
This article is cited by
-
Reassessing the mechanism of genome packaging in plant viruses with lessons from ATPase fold
Australasian Plant Pathology (2021)
-
Genetic determinants restricting the reassortment of heterologous NSP2 genes into the simian rotavirus SA11 genome
Scientific Reports (2017)
-
Structural insights into the coupling of virion assembly and rotavirus replication
Nature Reviews Microbiology (2012)
-
Conventional and unconventional mechanisms for capping viral mRNA
Nature Reviews Microbiology (2012)
-
Parallels among positive-strand RNA viruses, reverse-transcribing viruses and double-stranded RNA viruses
Nature Reviews Microbiology (2006)