Abstract
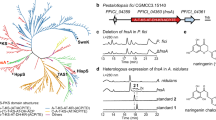
Chalcone synthases, which biosynthesize chalcones (the starting materials for many flavonoids1,2), have been believed to be specific to plants. However, the rppA gene from the Gram-positive, soil-living filamentous bacterium Streptomyces griseus encodes a 372-amino-acid protein that shows significant similarity to chalcone synthases3. Several rppA-like genes are known, but their functions and catalytic properties have not been described. Here we show that a homodimer of RppA catalyses polyketide synthesis: it selects malonyl-coenzyme-A as the starter, carries out four successive extensions and releases the resulting pentaketide to cyclize to 1,3,6,8-tetrahydroxynaphthalene (THN). Site-directed mutagenesis revealed that, as in other chalcone synthases4,5, a cysteine residue is essential for enzyme activity. Disruption of the chromosomal rppA gene in S. griseus abolished melanin production in hyphae, resulting in ‘albino’ mycelium. THN was readily oxidized to form 2,5,7-trihydroxy-1,4-naphthoquinone (flaviolin), which then randomly polymerized to form various coloured compounds. THN formed by RppA appears to be an intermediate in the biosynthetic pathways for not only melanins but also various secondary metabolites containing a naphthoquinone ring. Therefore, RppA is a chalcone-synthase-related synthase that synthesizes polyketides and is found in the Streptomyces and other bacteria.




Similar content being viewed by others
References
Schröder, J. Afamily of plant-specific polyketide synthases: facts and predictions. Trends Plant Sci. 2, 373–378 (1997).
Schröder, J. Comprehensive Natural Products Chemistry Vol. 1(ed. Sankawa, U.) 749–771 (Elsevier, Amsterdam, 1999).
Ueda, K., Kim, K.-M., Beppu, T. & Horinouchi, S. Overexpression of a gene cluster encoding a chalcone synthase-like protein confers redbrown pigment production in Streptomyces griseus. J. Antibiot. 48, 638–646 (1995).
Tropf, S., Kärcher, B., Schröder, G. & Schröder, J. Reaction mechanisms of homodimeric plant polyketide synthases (stilbene and chalcone synthase). J. Biol. Chem. 270, 7922–7928 (1995).
Lanz, T. et al. The role of cysteine in polyketide synthases. Site-directed mutagenesis of resveratrol and chalcone synthases, two key enzymes in different plant-specific pathways. J. Biol. Chem. 266, 9971–9976 (1991).
Fujii, I. et al. Heterologous expression and product identification of Colletotrichium lagenarium polyketide synthase encoded by the PKS1 gene involved in melanin biosynthesis. Biosci. Biotechnol. Biochem. 63, 1445–1452 (1999).
Eckermann, S. et al. New pathway to polyketides in plants. Nature 396, 387–390 (1998).
Dimroth, P., Ringelmann, E. & Lynen, F. 6-Methylsalicylic acid synthase from Penicillium patulum. Some catalytic properties of the enzyme and its relation to fatty acid synthase. Eur. J. Biochem. 68, 591–596 (1976).
Hopwood, D. A. Genetic contributions to understanding polyketide synthases. Chem. Rev. 97, 2465–2497 (1997).
Hutchinson, C. R. & Fujii, I. Polyketide synthase gene manipulation: a structure–function approach in engineering novel antibiotics. Annu. Rev. Microbiol. 49, 201–238 (1995).
Katz, L. & Donadio, S. Polyketide synthesis: prospects for hybrid antibiotics. Annu. Rev. Microbiol. 47, 875–912 (1993).
Bangera, M. G. & Thomashow, L. S. Characterization of a genomic locus required for synthesis of the antibiotic 2,4-diacetylphloroglucinol by the biological control agent Pseudomonas fluorescensQ2-87. Mol. Plant–Microbe Interact. 9, 83–90 (1996).
van Wageningen, A. M. A. et al. Sequencing and analysis of genes involved in the biosynthesis of a vancomycin group antibiotic. Chem. Biol. 5, 155–162 (1998).
Hammond, S. J. et al. On the biosynthesis of the antibiotic vancomycin. J. Chem. Soc. Chem. Commun. 1982, 344–346 (1982).
Shin-ya, K., Furihata, K., Hayakawa, Y. & Seto, H. Biosynthetic studies of naphterpin, a terpenoid metabolite of Streptomyces. Tetrahedron Lett. 31, 6025–6026 (1990).
Funayama, S., Ishibashi, M., Komiyama, K. & Omura, S. Biosynthesis of furaquinocins A and B. J. Org. Chem. 55, 1132–1133 (1990).
Shiomi, K. et al. Structures of new antibiotics napyradiomycins. J. Antibiot. 39, 494–501 (1986).
Pathirana, C., Jensen, P. R. & Fenical, W. Marinone and debromomarinone: antibiotic sesquiterpenoid naphthoquinone of a new structure class from a marine bacterium. Tetrahedron Lett. 33, 7663–7666 (1992).
Blanco, G. et al. Hybridization and DNA sequence analyses suggest an early evolutionary divergence of related biosynthetic gene sets encoding polyketide antibiotics and spore pigments in Streptomyces spp. Gene 130, 107–116 (1993).
Yu, T.-W. et al. Engineered biosynthesis of novel polyketides from Streptomyces spore pigment polyketide synthases. J. Am. Chem. Soc. 120, 7749–7759 (1998).
Bell, A. A. & Wheeler, M. H. Biosynthesis and functions of fungal melanins. Annu. Rev. Phytopathol. 24, 411–451 (1986).
Mayorga, M. E. & Timberlake, W. E. The developmentally regulated Aspergillus nidulans wA gene encodes a polyketide homologous to polypeptide and fatty acid synthases. Mol. Gen. Genet. 235, 205–212 (1992).
Takano, Y. et al. Structural analysis of PKSI, a polyketide synthase gene involved in melanin biosynthesis in Collectotrichum lagenarium. Mol. Gen. Genet. 249, 162–167 (1995).
Maniatis, T., Fritsch, E. F. & Sambrook, J. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1982).
Hopwood, D. A. et al. Genetic Manipulation in Streptomyces: A Laboratory Manual (The John Innes Foundation, Norwich, UK, 1985).
Beck, E. et al. Nucleotide sequence and exact localisation of the neomycin phosphotransferase gene from transposon Tn5. Gene 19, 327–336 (1982).
Oh, S. H. & Chater, K. F. Denaturation of circular or linear DNA facilitates targeted integrative transformation of Streptomyces coelicolor A3(2): possible relevance to other organisms. J. Bacteriol. 179, 122–127 (1997).
Ando, N., Ueda, K. & Horinouchi, S. AStreptomyces griseus gene (sgaA) suppresses the growth disturbance caused by high osmolality and a high concentration of A-factor during early growth. Microbiology 143, 2715–2723 (1997).
Acknowledgements
This work was supported by the Nissan Science Foundation, by the ‘Research for the Future’ Program of JSPS, and by the Bio Design Program from the Ministry of Agriculture, Forestry and Fisheries of Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Funa, N., Ohnishi, Y., Fujii, I. et al. A new pathway for polyketide synthesis in microorganisms. Nature 400, 897–899 (1999). https://doi.org/10.1038/23748
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/23748
- Springer Nature Limited
This article is cited by
-
Melanin biopolymers from microbial world with future perspectives—a review
Archives of Microbiology (2023)
-
Chalcone synthases (CHSs): the symbolic type III polyketide synthases
Planta (2020)
-
Melanin biosynthesis in bacteria, regulation and production perspectives
Applied Microbiology and Biotechnology (2020)
-
Diversity of nonribosomal peptide synthetase and polyketide synthase gene clusters among taxonomically close Streptomyces strains
Scientific Reports (2018)
-
The Combinatorial Biosynthesis of “Unnatural” Products with Polyketides
Transactions of Tianjin University (2018)