Abstract
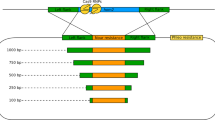
To improve the analysis of unknown flanking DNA sequences adjacent to known sequences in nuclear genomes of photoautotrophic eukaryotic organisms, we established the technique of ligation-mediated suppression-PCR (LMS-PCR) in the green alga Chlamydomonas reinhardtii for (1) walking from a specific nuclear insertion fragment of random knockout mutants into the unknown flanking DNA sequence to identify and analyse disrupted genomic DNA regions and for (2) walking from highly conserved DNA regions derived from known gene iso-forms into flanking DNA sequences to identify new members of protein families. The feasibility of LMS-PCR for these applications was successfully demonstrated in two different approaches. The first resulted in the identification of a genomic DNA fragment flanking a nuclear insertion vector in a random knockout mutant whose phenotype was characterised by its inability to perform functional LHC state transitions. The second approach targeted the cab gene family. An oligonucleotide of a cabII gene, derived from a highly conserved region, was used to identify potential cab gene regions in the nuclear genome of Chlamydomonas. LMS-PCR combined with 3′ rapid amplification of cDNA ends (3′ RACE) and a PCR-based screening of a cDNA library resulted in the identification of the new cabII gene lhcb4. Both results clearly indicate that LMS-PCR is a powerful tool for the identification of flanking DNA sequences in the nuclear genome of Chlamydomonas reinhardtii.
Similar content being viewed by others
References
Allen DK and Staehelin LA (1994) Polypeptide composition, assembly and phosphorylation patterns of the Photosystem II antenna system of Chlamydomonas reinhardtii. Planta 194: 42–54
Asamizu E, Nakamura Y, Sato S, Fukuzawa H and Tabata S (1999) A large scale structural analysis of cDNAs in a unicellular green alga, Chlamydomonas reinhardtii. I. Generation of 3433 non-redundant expressed sequence tags. DNA Res 6: 369–373
Bassi R and Wollman FA (1991) The chlorophyll-a/b proteins of Photosystem II in Chlamydomonas reinhardtii. Planta 183: 423–433
Debuchy R, Purton S and Rochaix JD (1989) The argininosuccinate lyase gene of Chlamydomonas reinhardtii: An important tool for nuclear transformation and for correlating the genetic and molecular maps of the ARG7 locus. EMBO J 8: 2803–2809
Fleischmann MM, Ravanel S, Delosme R, Olive J, Zito F, Wollman FA and Rochaix JD (1999) Isolation and characterisation of photoautotrophic mutants of Chlamydomonas reinhardtii deficient in state transition. J Biol Chem 274: 30987–30994
Frohmann MA and Martin GR (1989) Rapid amplification of cDNA ends using nested primers. Technique 1: 165–170
Goodenough UW (1992) Green yeast. Cell 70: 533–538
Gumpel NJ and Purton S (1994) Playing tag with Chlamydomonas. Trends Cell Biol 4: 299–301
Harris EH (1989) The Chlamydomonas Sourcebook. A Comprehensive Guide to Biology and Laboratory Use. Academic Press, San Diego, California
Hippler M, Redding K and Rochaix JD (1998) Chlamydomonas genetics, a tool for the study of bioenergetic pathways. Biochim Biophys Acta 1367: 1–62
Hobe S, Forster R, Klingler J and Paulsen H (1995) N-proximal sequence motif in light-harvesting chlorophyll a/b-binding protein is essential for the trimerization of light-harvesting chlorophyll a/b complex. Biochemistry 34: 10224–10228
Imbault P, Wittemer C, Johanningmeier U, Jacobs JD and Howell SH (1988) Structure of the Chlamydomonas reinhardtii cabII-1 gene encoding a chlorophyll-a/b-binding protein. Gene 73: 397–407
Jansson S (1999) A guide to the Lhc genes and their relatives in Arabidopsis. Trends Plant Sci 4: 236–240
Kindle KL (1990) High-frequency nuclear transformation of Chlamydomonas reinhardtii. Proc Natl Acad Sci USA 87: 1228–1232
Kindle KL, Schnell RA, Fernández E and Lefebvre PA (1989) Stable nuclear transformation of Chlamydomonas using the Chlamydomonas gene for nitrate reductase. J Cell Biol 109: 2589–2601
Kruse O, Nixon PJ, Schmid GH and Mullineaux CW (1999) Isolation of state transition mutants of Chlamydomonas reinhardtii by fluorescence video imaging. Photosynth Res 61: 43–51
LaRoche J, Bennett J and Falkowski PG (1990) Characterization of a cDNA encoding for the 28.5-kDa LHC II apoprotein from the unicellular marine chlorophyte, Dunaliella tertiolecta. Gene 95: 165–171
Levine RP and Ebershold WT (1960) The genetics and cytology of Chlamydomonas. Annu Rev Microbiol 14: 197–216
Nicholas KB and Nicholas KBJr (1997) Gene Doc: A tool for editing and annotating multiple sequence alignments. Distributed by the author
O'Connor C, Somanchi A, Handley ER and Moroney JV (1998a) Chlamydomonas reinhardtii Lhcb2 mRNA. Published in GenBank_, Acc No AF104630
O'Connor C, Somanchi A, Handley ER and Moroney JV (1998b) Chlamydomonas reinhardtii Lhcb3 mRNA. Published in GenBank_, Acc No AF104631
Purton S and Rochaix JD (1995) Characterisation of the ARG7 gene of Chlamydomonas reinhardtii and its application to nuclear transformation. Eur J Phycol 30: 141–148
Rochaix JD and Van Dillewijn J (1982) Transformation of the green alga Chlamydomonas reinhardtii with yeast DNA. Nature 296: 70–73
Rochaix JD (1995) Chlamydomonas reinhardtii as the photosynthetic yeast. Annu Rev Genet 29: 209–230
Rochaix JD, Goldschmidt-Clermont M and Merchant S (1998) The molecular biology of chloroplasts and mitochondria in Chlamydomonas. Kluwer Academic Publishers, Dodrecht, The Netherlands
Siebert PD, Chenchik A, Kellogg DE, Lukyanov KA and Lukyanov SA (1995) An improved PCR method for walking in uncloned genomic DNA. Nucleic Acids Res 23: 1087–1088
Stevens DR, Rochaix JD and Purton S (1996) The bacterial phleomycin resistance gene ble as a dominant selectable marker in Chlamydomonas. Mol Gen Genet 251: 23–30
Tam LW and Lefebvre PA (1993) Cloning of flagellar genes in Chlamydomonas reinhardtii by DNA insertional mutagenesis. Genetics 135: 375–384
The Arabidopsis Genome Initiative (2000) Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature 408: 796–815
Zhang H, Herman P and Weeks DP (1994) Gene isolation through genomic complementation using an indexed library of Chlamydomonas reinhardtii DNA. Plant Mol Biol 24: 663–672
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Strauss, C., Mussgnug, J.H. & Kruse, O. Ligation-mediated suppression-PCR as a powerful tool to analyse nuclear gene sequences in the green alga Chlamydomonas reinhardtii . Photosynthesis Research 70, 311–320 (2001). https://doi.org/10.1023/A:1014713612509
Issue Date:
DOI: https://doi.org/10.1023/A:1014713612509