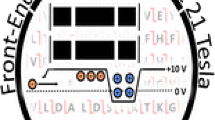
Abstract
We have developed and implemented a novel mass spectrometry (MS) platform combining the advantages of high mass accuracy and resolving power of Fourier transform ion cyclotron resonance (FTICR) with the economy and speed of multiple ion traps for tandem mass spectrometry. The instruments are integrated using novel algorithms and software and work in concert as one system. Using chromatographic time compression, a single expensive FTICR mass spectrometer can match the throughput of multiple relatively inexpensive ion trap instruments. Liquid chromatography (LC)-mass spectrometry data from the two types of spectrometers are aligned and combined to hybrid datasets, from which peptides are identified using accurate mass from the FTICR data and tandem mass spectra from the ion trap data. In addition, the high resolving power and dynamic range of a 12 tesla FTICR also allows precise label-free quantitation. Using two ion traps in parallel with one LC allows simultaneous MS/MS experiments and optimal application of collision induced dissociation and electrontransfer dissociation throughout the chromatographic separation for increased proteome coverage, characterization of post-translational modifications and/or simultaneous measurement in positive and negative ionization mode. An FTICR-ion trap cluster can achieve similar performance and sample throughput as multiple hybrid ion trap-FTICR instruments, but at a lower cost. We here describe the first such FTICR-ion trap cluster, its performance and the idea of chromatographic compression.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
March, R. E. An Introduction to Quadrupole Ion Trap Mass Spectrometry. J. Mass Spectrom. 1997, 32, 351–369.
Morris, H. R.; Paxton, T.; Dell, A.; Langhorne, J.; Berg, M.; Bordoli, R. S.; Hoyes, J.; Bateman, R. H. High Sensitivity Collisionally-Activated Decomposition Tandem Mass Spectrometry on a Novel Quadrupole/ Orthogonal-Acceleration Time-of-Flight Mass Spectrometer. Rapid Commun. Mass Spectrom. 1996, 10, 889–896.
Makarov, A.; Denisov, E.; Kholomeev, A.; Balschun, W.; Lange, O.; Strupat, K.; Horning, S. Performance Evaluation of a Hybrid Linear Ion Trap/Orbitrap Mass Spectrometer. Anal. Chem. 2006, 78, 2113–2120.
Syka, J. E.; Marto, J. A.; Bai, D. L.; Horning, S.; Senko, M. W.; Schwartz, J. C.; Ueberheide, B.; Garcia, B.; Busby, S.; Muratore, T.; Shabanowitz, J.; Hunt, D. F. Novel Linear Quadrupole Ion Trap/FT Mass Spectrometer: Performance Characterization and Use in the Comparative Analysis of Histone H3 Post-Translational Modifications. J. Proteome Res. 2004, 3, 621–626.
O’Connor, P. B.; Pittman, J. L.; Thomson, B. A.; Budnik, B. A.; Cournoyer, J. C.; Jebanathirajah, J.; Lin, C.; Moyer, S.; Zhao, C. A New Hybrid Electrospray Fourier Transform Mass Spectrometer: Design and Performance Characteristics. Rapid Commun. Mass Spectrom. 2006, 20, 259–266.
Pasa-Tolic, L.; Masselon, C.; Barry, R. C.; Shen, Y.; Smith, R. D. Proteomic Analyses Using an Accurate Mass and Time Tag Strategy. Biotechniques 2004, 37, 621–624, 626–633, 636 passim.
Petritis, K.; Kangas, L. J.; Ferguson, P. L.; Anderson, G. A.; Pasa-Tolic, L.; Lipton, M. S.; Auberry, K. J.; Strittmatter, E. F.; Shen, Y.; Zhao, R.; Smith, R. D. Use of Artificial Neural Networks for the Accurate Prediction of Peptide Liquid Chromatography Elution Times in Proteome Analyses. Anal. Chem. 2003, 75, 1039–1048.
Petritis, K.; Kangas, L. J.; Yan, B.; Monroe, M. E.; Strittmatter, E. F.; Qian, W. J.; Adkins, J. N.; Moore, R. J.; Xu, Y.; Lipton, M. S.; Camp, D. G. II; Smith, R. D. Improved Peptide Elution Time Prediction for Reversed-Phase Liquid Chromatography-MS by Incorporating Peptide Sequence Information. Anal. Chem. 2006, 78, 5026–5039.
Jaitly, N.; Monroe, M. E.; Petyuk, V. A.; Clauss, T. R.; Adkins, J. N.; Smith, R. D. Robust Algorithm for Alignment of Liquid Chromatography-Mass Spectrometry Analyses in an Accurate Mass and Time Tag Data Analysis Pipeline. Anal. Chem. 2006, 78, 7397–7409.
Palmblad, M.; Mills, D. J.; Bindschedler, L. V.; Cramer, R. Chromatographic Alignment of LC-MS and LC-MS/MS Datasets by Genetic Algorithm Feature Extraction. J. Am. Soc. Mass Spectrom. 2007, 18, 1835–1843.
Jacob, F.; Monod, J. Genetic Regulatory Mechanisms in the Synthesis of Proteins. J. Mol. Biol. 1961, 3, 318–356.
Traxler, M. F.; Chang, D. E.; Conway, T. Guanosine 3′,5′-Bispyrophosphate Coordinates Global Gene Expression During Glucose-Lactose Diauxie in Escherichia coli. Proc. Natl. Acad. Sci. U.S.A. 2006, 103, 2374–2379.
Craig, R.; Beavis, R. C. TANDEM: Matching Proteins with Tandem Mass Spectra. Bioinformatics. 2004, 20, 1466–1467.
Palmblad, M.; Bindschedler, L. V.; Gibson, T. M.; Cramer, R. Automatic Internal Calibration in Liquid Chromatography/Fourier Transform Ion Cyclotron Resonance Mass Spectrometry of Protein Digests. Rapid Commun. Mass Spectrom. 2006, 20, 3076–3080.
www.matrixscience.com.
Colinge, J.; Masselot, A.; Giron, M.; Dessingy, T.; Magnin, J. OLAV: Towards High-Throughput Tandem Mass Spectrometry Data Identification. Proteomics 2003, 3, 1454–1463.
Syka, J. E.; Coon, J. J.; Schroeder, M. J.; Shabanowitz, J.; Hunt, D. F. Peptide and Protein Sequence Analysis by Electron Transfer Dissociation Mass Spectrometry. Proc. Natl. Acad. Sci. U.S.A. 2004, 101, 9528–9533.
Leinenbach, A.; Hartmer, R.; Lubeck, M.; Kneissl, B.; Elnakady, Y. A.; Baessmann, C.; Muller, R.; Huber, C. G. Proteome Analysis of Sorangium cellulosum employing 2D-HPLC-MS/MS and Improved Database Searching Strategies for CID and ETD Fragment Spectra. J. Proteome Res. 2009, 8, 4350–4361.
Haas, W.; Faherty, B. K.; Gerber, S. A.; Elias, J. E.; Beausoleil, S. A.; Bakalarski, C. E.; Li, X.; Villen, J.; Gygi, S. P. Optimization and Use of Peptide Mass Measurement Accuracy in Shotgun Proteomics. Mol. Cell. Proteom. 2006, 5, 1326–1337.
Bakalarski, C. E.; Haas, W.; Dephoure, N. E.; Gygi, S. P. The Effects of Mass Accuracy, Data Acquisition Speed, and Search Algorithm Choice on Peptide Identification Rates in Phosphoproteomics. Anal. Bioanal. Chem. 2007, 389, 1409–1419.
Ishihama, Y.; Oda, Y.; Tabata, T.; Sato, T.; Nagasu, T.; Rappsilber, J.; Mann, M. Exponentially Modified Protein Abundance Index (emPAI) for Estimation of Absolute Protein Amount in Proteomics by the Number of Sequenced Peptides Per Protein. Mol. Cell. Proteom. 2005, 4, 1265–1272.
Palmblad, M.; Mills, D. J.; Bindschedler, L. V. Heat-Shock Response in Arabidopsis thaliana Explored by Multiplexed Quantitative Proteomics Using Differential Metabolic Labeling. J. Proteome Res. 2008, 7, 780–785.
Nevedomskaya, E.; Derks, R.; Deelder, A. M.; Mayboroda, O. A.; Palmblad, M. Alignment of Capillary Electrophoresis-Mass Spectrometry Datasets Using Accurate Mass Information. Anal. Bioanal. Chem. 2009, 395, 2527–2533.
Zubarev, R. A.; Kelleher, N. L.; McLafferty, F. W. Electron Capture Dissociation of Multiply Charged Protein Cations. A Nonergodic Process. J. Am. Chem. Soc. 1998, 120, 3265–3266.
Ong, S. E.; Blagoev, B.; Kratchmarova, I.; Kristensen, D. B.; Steen, H.; Pandey, A.; Mann, M. Stable Isotope Labeling by Amino Acids in Cell Culture, SILAC, as a Simple and Accurate Approach to Expression Proteomics. Mol. Cell. Proteom. 2002, 1, 376–386.
Ross, P.; Dey, S.; Pillai, S.; Daniels, S.; Williamson, S.; Guertin, S.; Minkoff, M.; X., C.; Purkayastha, B.; Pappin, D. Proceedings of the 54th ASMS Conference on Mass Spectrometry; Seattle, WA, May 28-June 1, 2006.
Ross, P. L.; Huang, Y. N.; Marchese, J. N.; Williamson, B.; Parker, K.; Hattan, S.; Khainovski, N.; Pillai, S.; Dey, S.; Daniels, S.; Purkayastha, S.; Juhasz, P.; Martin, S.; Bartlet-Jones, M.; He, F.; Jacobson, A.; Pappin, D. J. Multiplexed Protein Quantitation in Saccharomyces cerevisiae Using Amine-Reactive Isobaric Tagging Reagents. Mol. Cell. Proteom. 2004, 3, 1154–1169.
Spengler, B.; Hester, A. Mass-Based Classification (MBC) of Peptides: Highly Accurate Precursor Ion Mass Values Can Be Used to Directly Recognize Peptide Phosphorylation. J. Am. Soc. Mass Spectrom. 2008, 19, 1808–1812.
Horn, D. M.; Zubarev, R. A.; McLafferty, F. W. Automated Reduction and Interpretation of High Resolution Electrospray Mass Spectra of Large Molecules. J. Am. Soc. Mass Spectrom. 2000, 11, 320–332.
Palmblad, M.; Wetterhall, M.; Markides, K.; Håkansson, P.; Bergquist, J. Analysis of Enzymatically Digested Proteins and Protein Mixtures Using a 9.4 Tesla Fourier Transform Ion Cyclotron Mass Spectrometer. Rapid Commun. Mass Spectrom. 2000, 14, 1029–1034.
Palmblad, M.; Drijfhout, J. W.; Deelder, A. M. High Resolution Mass Spectrometry for Rapid Characterization of Combinatorial Peptide Libraries. J. Comb. Chem. 2010, 12, 65–68.
Strittmatter, E. F.; Ferguson, P. L.; Tang, K.; Smith, R. D. Proteome Analyses Using Accurate Mass and Elution Time Peptide Tags with Capillary LC Time-of-Flight Mass Spectrometry. J. Am. Soc. Mass Spectrom. 2003, 14, 980–991.
Zimmer, J. S.; Monroe, M. E.; Qian, W. J.; Smith, R. D. Advances in Proteomics Data Analysis and Display Using an Accurate Mass and Time Tag Approach. Mass Spectrom. Rev. 2006, 25, 450–482.
Old, W. M.; Meyer-Arendt, K.; Aveline-Wolf, L.; Pierce, K. G.; Mendoza, A.; Sevinsky, J. R.; Resing, K. A.; Ahn, N. G. Comparison of Label-Free Methods for Quantifying Human Proteins by Shotgun Proteomics. Mol. Cell. Proteom. 2005, 4, 1487–1502.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Palmblad, M., van der Burgt, Y.E.M., Mostovenko, E. et al. A novel mass spectrometry cluster for high-throughput quantitative proteomics. J Am Soc Mass Spectrom 21, 1002–1011 (2010). https://doi.org/10.1016/j.jasms.2010.02.001
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1016/j.jasms.2010.02.001