Abstract
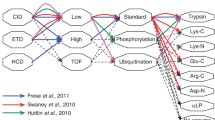
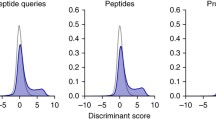
Proteomic analyses via tandem mass spectrometry have been greatly enhanced by the recent development of fast, highly accurate instrumentation. However, successful application of these developments to high-throughput experiments requires careful optimization of many variables which adversely affect each other, such as mass accuracy and data collection speed. We examined the performance of three shotgun-style acquisition methods ranging in their data collection speed and use of mass accuracy in identifying proteins from yeast-derived complex peptide and phosphopeptide-enriched mixtures. We find that the combination of highly accurate precursor masses generated from one survey scan in the FT-ICR cell, coupled with ten data-dependent tandem MS scans in a lower-resolution linear ion trap, provides more identifications in both mixtures than the other examined methods. For phosphopeptide identifications in particular, this method identified over twice as many unique phosphopeptides as the second-ranked, lower-resolution method from triplicate 90-min analyses (744 ± 50 vs. 308 ± 50, respectively). We also examined the performance of four popular peptide assignment algorithms (Mascot, Sequest, OMSSA, and Tandem) in analyzing the results from both high-and low-resolution data. When compared in the context of a false positive rate of approximately 1%, the performance differences between algorithms were much larger for phosphopeptide analyses than for an unenriched, complex mixture. Based upon these findings, acquisition speed, mass accuracy, and the choice of assignment algorithm all largely affect the number of peptides and proteins identified in high-throughput studies.






Similar content being viewed by others
References
Syka JE, Marto JA, Bai DL, Horning S, Senko MW, Schwartz JC, Ueberheide B, Garcia B, Busby S, Muratore T, Shabanowitz J, Hunt DF (2004) J Proteome Res 3:621–626
Florens L, Washburn MP, Raine JD, Anthony RM, Grainger M, Haynes JD, Moch JK, Muster N, Sacci JB, Tabb DL, Witney AA, Wolters D, Wu Y, Gardner MJ, Holder AA, Sinden RE, Yates JR, Carucci DJ (2002) Nature 419:520–526
Everley PA, Bakalarski CE, Elias JE, Waghorne CG, Beausoleil SA, Gerber SA, Faherty BK, Zetter BR, Gygi SP (2006) J Proteome Res 5:1224–1231
Lasonder E, Ishihama Y, Andersen JS, Vermunt AM, Pain A, Sauerwein RW, Eling WM, Hall N, Waters AP, Stunnenberg HG, Mann M (2002) Nature 419:537–542
Dieguez-Acuna FJ, Gerber SA, Kodama S, Elias JE, Beausoleil SA, Faustman D, Gygi SP (2005) Mol Cell Proteomics 4:1459–1470
Beausoleil SA, Jedrychowski M, Schwartz D, Elias JE, Villen J, Li J, Cohn MA, Cantley LC, Gygi SP (2004) Proc Natl Acad Sci USA 101:12130–12135
Beausoleil SA, Villen J, Gerber SA, Rush J, Gygi SP (2006) Nat Biotechnol 24:1285–1292
Villen J, Beausoleil SA, Gerber SA, Gygi SP (2007) Proc Natl Acad Sci USA 104:1488–1493
Li X, Gerber SA, Rudner AD, Beausoleil SA, Haas W, Villen J, Elias JE, Gygi SP (2007) J Proteome Res 6:1190–1197
Olsen JV, Blagoev B, Gnad F, Macek B, Kumar C, Mortensen P, Mann M (2006) Cell 127:635–648
Olsen JV, Ong SE, Mann M (2004) Mol Cell Proteomics 3:608–614
Mayya V, Rezaul K, Cong YS, Han D (2005) Mol Cell Proteomics 4:214–223
Haas W, Faherty BK, Gerber SA, Elias JE, Beausoleil SA, Bakalarski CE, Li X, Villen J, Gygi SP (2006) Mol Cell Proteomics 5:1326–1337
Perkins DN, Pappin DJ, Creasy DM, Cottrell JS (1999) Electrophoresis 20:3551–3567
Geer LY, Markey SP, Kowalak JA, Wagner L, Xu M, Maynard DM, Yang X, Shi W, Bryant SH (2004) J Proteome Res 3:958–964
Eng JK, McCormack AL, Yates I, John R (1994) J Am Soc Mass Spectrom 5:976–989
Craig R, Beavis RC (2004) Bioinformatics 20:1466–1467
Guthrie C, Fink GR (1991) Guide to yeast genetics and molecular biology. Academic, San Diego
Mortimer RK, Johnston JR (1986) Genetics 113:35–43
Shevchenko A, Wilm M, Vorm O, Mann M (1996) Anal Chem 68:850–858
Pedrioli PG, Eng JK, Hubley R, Vogelzang M, Deutsch EW, Raught B, Pratt B, Nilsson E, Angeletti RH, Apweiler R, Cheung K, Costello CE, Hermjakob H, Huang S, Julian RK, Kapp E, McComb ME, Oliver SG, Omenn G, Paton NW, Simpson R, Smith R, Taylor CF, Zhu W, Aebersold R (2004) Nat Biotechnol 22:1459–1466
Elias JE, Gygi SP (2007) Nat Methods 4:207–214
Elias JE, Haas W, Faherty BK, Gygi SP (2005) Nat Methods 2:667–675
Liu H, Sadygov RG, Yates JR 3rd (2004) Anal Chem 76:4193–4201
Acknowledgements
This work was supported in part by National Institutes of Health Grants GM67945 and HG3456 (to S. P. G.). We would like to thank J. Elias, J. Mintseris, J. Villen, and S. Beausoleil for helpful advice and discussions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 476 kb)
Rights and permissions
About this article
Cite this article
Bakalarski, C.E., Haas, W., Dephoure, N.E. et al. The effects of mass accuracy, data acquisition speed, and search algorithm choice on peptide identification rates in phosphoproteomics. Anal Bioanal Chem 389, 1409–1419 (2007). https://doi.org/10.1007/s00216-007-1563-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-007-1563-x