Abstract
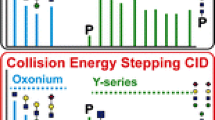
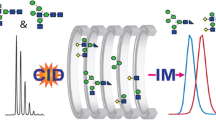
Combining source collision-induced dissociation (CID) and tandem mass spectral acquisition in a pseudo-MS3 experiment using a linear ion trap results in a highly selective and sensitive approach to identifying glycopeptide elution from a protein digest. The increased sensitivity is partially attributed to the nonselective nature of source CID, which allows simultaneous activation of all charge states and coeluting glycoforms generating greater ion abundance for the mass-to-charge (m/z) 204 and/or 366 oxonium ions. Unlike source CID alone, a pseudo-MS3 approach adds selectivity while improving sensitivity by eliminating chemical noise during the tandem mass spectral acquisition of the oxonium ions in the linear ion trap. Performing the experiments in the hybrid linear ion trap/Fourier transform-ion cyclotron resonance (FT-ICR) enables subsequent high-resolution/high-mass accuracy full-scan mass spectra (MS) and parallel acquisition of MS/MS in the linear ion trap to be completed in 2 s directly following the pseudo-MS3 scan to collate identification and characterization of glycopeptides in one experimental scan cycle. Analysis of bovine fetuin digest using the combined pseudo-MS3, high-resolution MS, and data-dependent MS/MS events resulted in identification of four N-linked and two O-linked glycopeptides without enzymatic cleavage of the sugar moiety or release of the sialic acids before analysis. In addition, over 95% of the total protein sequence was identified in one analytical run.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
An, H. J.; Peavy, T. R.; Hedrick, J. L.; Lebrilla, C. B. Determination of N-Glycosylation Sites and Site Heterogeneity in Glycoproteins. Anal. Chem. 2003, 75, 5628–5637.
Jebanathirajah, J.; Steen, H.; Roepstorff, P. Using Optimized Collision Energies and High Resolution, High Accuracy Fragment Ion Selection to Improve Glycopeptide Detection by Precursor Ion Scanning. J. Am. Soc. Mass Spectrom. 2003, 14, 777–784.
Sandra, K.; Devreese, B.; Van Beeumen, J.; Stals, I.; Claeyssens, M. The Q-Trap Mass Spectrometer, A Novel Tool in the Study of Protein Glycosylation. J. Am. Soc. Mass Spectrom. 2004, 15, 413–423.
Wuhrer, M.; Koeleman, C. A. M.; Hokke, C. H.; Deelder, A. M. Protein Glycosylation Analyzed by Normal-Phase Nano-Liquid Chromatography-Mass Spectrometry of Glycopeptides. Anal. Chem. 2005, 77, 886–894.
Morelle, W.; Michalski, J. C. The Mass Spectrometric Analysis of Glycoproteins and Their Glycan Structures. Current Anal. Chem. 2005, 1, 29–57.
Medzihradszky, K. F.; Gillece-Castro, B. L.; Townsend, R. R.; Burlingame, A. L.; Hardy, M. R. Structural Elucidation of O-Linked Glycopeptides by High Energy Collision-Induced Dissociation. J. Am. Soc. Mass Spectrom. 1996, 7, 319–328.
Weiskopf, A. S.; Vouros, P.; Harvey, D. J. Characterization of Oligosaccharide Composition and Structure by Quadrupole Ion Trap Mass Spectrometry. Rapid Commun. Mass Spectrom. 1997, 11, 1493–1504.
Schulz, B. L.; Packer, N. H.; Karlsson, N. G. Small-Scale Analysis of O-Linked Oligosaccharides from Glycoproteins and Mucins Separated by Gel Electrophoresis. Anal. Chem. 2002, 74, 6088–6097.
Harvey, D. J. Identification of Protein-Bound Carbohydrates by Mass Spectrometry. Proteomics 2001, 1, 311–328.
Huddleston, M. J.; Bean, M. F.; Carr, S. A. Collisional Fragmentation of Glycopeptides by Electrospray Ionization LC/MS and LC/MS/MS: Methods for Selective Detection of Glycopeptides in Protein Digests. Anal. Chem. 1993, 65, 877–884.
Carr, S. A.; Huddleston, M. J.; Bean, M. F. Selective Identification and Differentiation of N- and O-Linked Oligosaccharides in Glycoproteins by Liquid Chromatography-Mass Spectrometry. Protein Sci. 1993, 2, 183–196.
Ritchie, M. A.; Gill, A. C.; Deery, M. J.; Lilley, K. Precursor Ion Scanning for Detection and Structural Characterization of Heterogeneous Glycopeptide Mixtures. J. Am. Soc. Mass Spectrom. 2002, 13, 1065–1077.
Carr, S. A.; Roberts, G. D. Carbohydrate Mapping by Mass Spectrometry: A Novel Method for Identifying Attachment Sites of Asn-linked Sugars in Glycoproteins. Anal. Biochem. 1986, 157, 396–406.
Jiang, H.; Desaire, H.; Butnev, V. Y.; Bousfield, G. R. Glycoprotein Profiling by Electrospray Mass Spectrometry. J. Am. Soc. Mass Spectrom. 2004, 15, 750–758.
Carr, S. A.; Huddleston, M. J.; Annan, R. S. Selective Detection and Sequencing of Phosphopeptides at the Low Femtomole Level by Mass Spectrometry. Anal. Biochem. 1996, 239, 180–192.
Dziegielewska, K. M.; Brown, W. M.; Casey, S.-J.; Christie, D. L.; Foreman, R. C.; Hill, R. M.; Saunders, N. R. The Complete cDNA and Amino Acid Sequence of Bovine Fetuin. J. Biol. Chem. 1990, 265, 4354–4357.
Zhu, X.; Brorchers, C.; Bienstock, R. J.; Tomer, K. B. Mass Spectrometric Characterization of Glycosylation Pattern of HIV-gp120 Expressed in CHO Cells. Biochemistry 2000, 39, 11194–11204.
Juhasz, P.; Martin, S. A. The Utility of Nonspecific Proteases in the Characterization of Glycoproteins by High-Resolution Time-of-Flight Mass Spectrometry. Int. J. Mass Spectrom. 1997, 169/170, 217–230.
Schwartz, J. C.; Senko, M. W.; Syka, J. E. P. A Two-Dimensional Quadrupole Ion Trap Mass Spectrometer. J. Am. Soc. Mass Spectrom. 2002, 13, 659–669.
Staudacher, E.; Altmann, F.; Glossl, J.; Marz, L.; Schachter, H.; Kamerling, J. P.; Hard, K.; Vliegenthart, F. G. GDP-fucose: β-N-Acetylglucosamine (Fuc to Fucα1-6GlcNAc)-Asn peptide) 1–3-Fucosyltransferase Activity in Honeybee (Apis mellifica) Venom Glands. Eur. J. Biochem. 1991, 199(745), 751.
Kannicht, C. Posttranslational Modifications of Proteins; Humana Press: Totowa, New Jersey, 2002.
Syka, J. E. P.; Marto, J. A.; Bai, D. L.; Horning, S.; Senko, M. W.; Schwartz, J. C.; Ueberheide, B.; Garcia, B.; Busby, S.; Muratore, T.; Shabanowitz, J.; Hunt, D. F. Novel Linear Quadrupole Ion Trap/FT Mass Spectrometer: Performance Characterization and Use in the Comparative Analysis of Histone H3 Posttranslational Modifications. J. Proteome Res. 2004, 2, 5–10.
Tabb, D.; McDonald, W. H.; Yates, J. R. I. DTASelect and Contrast: Tools for Assembling and Comparing Identification from Shotgun Proteomics. J. Proteome Res. 2002, 1, 21–26.
Cooper, C. A.; Gasteiger, E.; Packer, N. H. Glycomod-A Software Tool for Determining Glycosylation Composition from Mass Spectrometer Data. Proteomics 2001, 1, 340–349.
Cooper, C. A.; Gasteiger, E.; Packer, N. H. In Predicting Glycan Composition from Experimental Mass Using GlycoMod; Humana Press: Totowa, NJ, 2003.
Medzihradszky, K. F.; Maltby, D. A.; Hall, S. C.; Settineri, C. A.; Burlingame, A. L. Characterization of Protein N-Glycosylation by Reversed-Phase Microbore Liquid Chromatography/Electrospray Mass Spectrometry, Complementary Mobile Phases, and Sequential Exoglycosidase Digestion. J. Am. Soc. Mass Spectrom. 1994, 5, 350–358.
Sullivan, B.; Addona, T. A.; Carr, S. A. Selective Detection of Glycopeptides on Ion Trap Mass Spectrometers. Anal. Chem. 2004, 76, 3112–3118.
Zhang, S.; Chelius, D. Characterization of Protein Glycosylation Using Chip Based Infusion Nanoelectrospray Linear Ion Trap Tandem Mass Spectrometry. J. Biomol. Tech. 2004, 15, 120–133.
Huang, Y.; Mechref, Y.; Novotny, M. V. Microscale Nonreductive Release of O-Linked Glycans for Subsequent Analysis Through MALDI Mass Spectrometry and Capillary Electrophoresis. Anal. Chem. 2001, 73, 6063–6069.
Author information
Authors and Affiliations
Corresponding author
Additional information
Published online January 10, 2006
Rights and permissions
About this article
Cite this article
Peterman, S.M., Mulholland, J.J. A novel approach for identification and characterization of glycoproteins using a hybrid linear ion trap/FT-ICR mass spectrometer. The official journal of The American Society for Mass Spectrometry 17, 168–179 (2006). https://doi.org/10.1016/j.jasms.2005.10.008
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1016/j.jasms.2005.10.008