Abstract
The human body harbors a diverse microbiota across various anatomical sites, including the lungs, which play a crucial role in maintaining respiratory health and influencing disease outcomes. This review synthesizes current research on the lung microbiota's impact on pulmonary diseases, emphasizing its interactions with microbiota from other body regions, such as the gut and oral cavity. Emerging evidence suggests that lung microbiota dysbiosis is intricately linked to the pathogenesis of various pulmonary diseases, including idiopathic pulmonary fibrosis, lung cancer, pneumonia, asthma, and chronic obstructive pulmonary disease (COPD). The review highlights the significance of understanding these microbial interactions to advance diagnostic, preventive, and therapeutic strategies for respiratory diseases. By elucidating the mechanisms through which microbiota influence lung health, this review underscores the potential of microbiome-based diagnostics and therapeutics in revolutionizing the management of pulmonary diseases, paving the way for personalized medicine and innovative treatment approaches.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The surfaces and orifices of the human body contain a variety of microorganisms such as in the oral cavity, nasal cavity, respiratory system, digestive system, and urogenital tract. Collectively, these microorganisms are known as the microbiota. They coexist with the human body and have a symbiotic relationship with it, benefiting each other. The microbiota is composed of a diverse set of bacteria in a balanced proportion. Each bacterial species relies on and constrains one another, creating an ecological equilibrium in both quality and quantity. This dynamic balance is important for diverse physiological functions, such as nutrition [1], immunity [2] [3], anti-aging [4, 5] and anti-tumor effects [6].
Researchers have become increasingly interested in studying the impact of microbiota on human health and disease. This is mainly due to technological advancements that have enabled the analysis of complex microbial communities. Meta-omics techniques, such as metagenomics, metatranscriptomics, metaproteomics, and metabolomics, help identify the composition and activities of microbial communities. By using techniques like shotgun metagenomic sequencing and 16S rRNA gene sequencing, researchers can gain a deeper understanding of the molecular mechanisms that underlie the relationship between the microbiome and various diseases [7]. There are new concepts and approaches that can connect microbial ecology and molecular mechanisms [8].
The study of microbiota and its impact on human health has garnered significant attention in recent years, particularly in understanding the role of microbiota in various diseases. The human body is home to diverse microbial communities that reside in different anatomical sites, including the gut, oral cavity, and lungs. These microbial communities play crucial roles in maintaining homeostasis and influencing disease processes. In particular, the lung microbiota, though less studied compared to gut microbiota, has been increasingly recognized as a key player in respiratory health and disease. The intricate interactions between lung microbiota and other microbial communities in the body are thought to contribute to the pathogenesis of several pulmonary diseases, including chronic obstructive pulmonary disease (COPD), asthma, and idiopathic pulmonary fibrosis (IPF).
Previous research has provided foundational insights into the characteristics and physiological roles of various bacteria that could influence respiratory health. For instance, Abbasli studied the morpho-cultural and physiological characteristics of bacteria isolated from thermal springs in Azerbaijan, highlighting the diversity and adaptability of these microorganisms to different environments [9]. Such studies are crucial as they offer a broader understanding of the potential sources of bacteria that may colonize the human body, including the lungs. Moreover, natural products and their interactions with human health have also been explored, as seen in the work of Miryusifova [10], who examined the effects of saffron on experimental epilepsy, and Karadağ and Omarova [11], who discussed the use of Prunus armeniaca seed oil in health and cosmetic products. These studies underscore the therapeutic potential of natural substances, which aligns with the growing interest in microbiome-based interventions for treating pulmonary diseases. By exploring these connections, our review aims to bridge the gap between environmental microbial sources, their colonization of the human body, and their impact on lung health, ultimately contributing to the development of novel diagnostic and therapeutic approaches.
Alterations in the composition and function of the human microbiota can lead to dysbiosis. This can cause diseases, and it is important to understand the causes and consequences of bacterial dysbiosis [12]. The gut microbiome, which refers to the microorganisms living in the digestive tract, undergoes changes in both composition and function as people age. These changes are affected by a variety of factors, including biological, behavioral, and environmental factors. This alteration plays a crucial role in the development of frailty, a condition associated with age-related decline in physical and cognitive function. To gain a comprehensive understanding of the effects of age-related changes in the gut microbiome on frailty, it is important to objectively evaluate these factors [13].
Microbial communities may affect the pathogenicity and viability of Vibrio cholerae in the host, playing a crucial role in the pathogenesis of cholera [14]. It is noteworthy to investigate how microbiota can enhance or diminish carcinogenesis, response to cancer treatment, and cancer-related complications [15]. The microbiota is expected to provide new insights for the development and management of cancer in humans.
The gut microbiota has received considerable attention. Emerging research has shed light on the crucial role of the lung microbiota in maintaining respiratory health and its involvement in various pulmonary diseases. It is of paramount importance to understand the intricate relationship between the lung microbiota and pulmonary diseases advance our knowledge, diagnosis and treatment strategies. This review provides a comprehensive analysis of the intricate relationship between microbiota and pulmonary diseases, emphasizing the potential for microbiome-based diagnostics and therapeutics. By elucidating the mechanisms through which microbiota influence lung health, our work offers valuable insights that could lead to the development of novel, targeted treatments for various pulmonary conditions. Furthermore, the review underscores the promise of personalized medicine, where microbiome profiles could be used to tailor treatments to individual patients, thereby improving outcomes. The potential applications of this work extend to early disease detection, monitoring disease progression, and optimizing therapeutic interventions, making it highly relevant for advancing biomedical research and clinical practice.
2 Overview the relationship between microbiota in various sites and pulmonary diseases
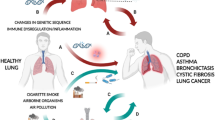
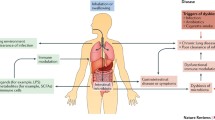
Recent studies have underscored the intricate connections between the lung microbiota and microbiota in other anatomical sites. Firstly, the composition of the lung microbiota is intricately tied to processes such as the inhalation of oral microorganisms, microaspiration of microorganisms from the upper respiratory tract, and the translocation of intestinal microorganisms through the mucosal barrier. Secondly, when dysbiosis occurs in microbiota elsewhere in the body, it can perturb the equilibrium of the lung microbiota, potentially giving rise to lung dysbiosis and subsequent pulmonary disorders (Fig. 1).
The left part of the image shows the lung microbiome is characterized by low biomass, dynamic community composition, and balance between microbial immigration and microbial elimination. immigration is mediated via inhalation, microaspiration and mucosal dispersion; elimination proceeds through cough, mucociliary transport and immune mechanisms. The right part of the image shows the association of each departmental microbiota with lung microbiota and the association of each departmental microbiota with pulmonary diseases. Created with BioRender.com.
2.1 Microbiota in various parts and pulmonary disease
A mounting body of evidence indicates a strong correlation between pulmonary dysbiosis and a spectrum of pulmonary diseases. It's noteworthy that pulmonary dysbiosis is frequently intertwined with alterations in the microbiota of the oral, nasal, and intestinal tracts, emphasizing the interconnected nature of these microbial shifts.
2.1.1 Gut microbiota and pulmonary disease
2.1.1.1 The gut-lung axis: insights from early life and COPD
The gut microbiota has gained recognition as a pivotal regulator of human health and disease. Researchers have delved into the interplay between the gut microbiota and the lungs, aiming to decipher the underlying pathogenesis of specific pulmonary diseases and explore potential therapeutic avenues [16, 17]. Importantly, an intriguing discovery has emerged from studies examining the maturation of the gut microbiome within the first year of life, particularly in a farm environment. This phenomenon appears to confer protection against asthma, underscoring the existence of a compelling “gut-lung axis” in humans, bridging the gut and the lungs [18].
2.1.1.2 Modulating gut microbiota for pulmonary disease management
The investigations conducted in mice have unveiled the substantial impact of the gut microbiome on the development of obstructive pulmonary disease (COPD). Notably, with the 16S rRNA gene sequencing, the identification of a symbiotic bacterium, Parabacteroides goldsteinii, has demonstrated its potential to alleviate COPD symptoms. These findings underscore the profound connection between the gut microbiota and metabolite composition in pulmonary diseases. This highlights the critical role of the gut-lung axis in modulating immune responses and inflammation, paving the way for manipulating gut microbiota and metabolites as a promising treatment strategy for pulmonary diseases [19, 20]. Crucially, these extensive interactions between microbiota and immune responses emphasize the significance of interventions like dietary adjustments, probiotics, vitamin supplementation, fecal microbiota transplantation, and emerging therapeutic modalities. These interventions hold the potential to provide preventive therapies for obstructive pulmonary disease (COPD) and advance the field of pulmonary disease management.
2.1.1.3 Linking gut bacteria to acute lung injury
The impact of microbiota extends to acute pulmonary diseases as well. A comprehensive investigation, integrating microbial profiling with physiological analyses, has revealed a substantial link between the gut bacterial metabolite succinate and acute lung injury induced by intestinal ischemia–reperfusion (I/R). The study has shed light on how intestinal ischemia–reperfusion (I/R) can lead to gut microbiome dysbiosis, resulting in an excessive production of succinate. This accumulated succinate subsequently infiltrates the lungs, triggering the polarization of alveolar macrophages via succinate receptor 1. Ultimately, this cascade of events culminates in lung injury by promoting the apoptosis of alveolar epithelial cells [21]. Moreover, with the NGS, studies have unveiled alterations in the gut microbiota of children afflicted with respiratory infections. These findings have sparked investigations into the connection between a child's microbiota, their immune function, and the development of diseases, emphasizing the pivotal role of the microbiota in these aspects [22].
2.1.1.4 The gut microbiome for idiopathic pulmonary fibrosis and lung function
Some study investigates the causal relationship between gut microbiota and idiopathic pulmonary fibrosis (IPF) as well as lung function using a two-sample Mendelian randomization approach with 16S fecal microbiome data from 18,340 participants across 24 cohorts. The findings suggest that certain gut microbes, like the Bifidobacteriales order and Ruminococcaceae genus, have protective effects against IPF, while others, like the Coprococcus genus, may promote its development. Additionally, lung function impacts the abundance of specific gut microbiota, indicating a bidirectional relationship [23]. Another study demonstrates that gut microbiota significantly influences the severity of lung fibrosis following acute lung injury in mice with metagenomic analyses. Specifically, a lower diversity of gut microbiota exacerbates lung fibrosis, while certain microbes, such as Lactobacillus, may protect against it by reducing pro-inflammatory responses [24]. These findings highlight the potential influence of microbial communities on lung inflammation and fibrosis, suggesting that targeting the lung microbiome could be a promising therapeutic approach for managing IPF [25].
2.1.1.5 The gut microbiome’s role in COVID-19
A recent study has emphasized the significant changes that SARS-CoV-2 infection can induce in the gut microbiome with 16S rRNA gene sequencing. These alterations have demonstrated a robust correlation with the severity of infection, suggesting a potential role for gut microbes in the pathogenesis and progression of COVID-19. Building upon these findings, researchers have proposed interventions aimed at regulating the gut microbiota as a strategy for managing COVID-19 [26].
These intricate connections and communication between the gut microbiota and pulmonary diseases present a compelling area for further exploration. Advancing our understanding of how gut microbiota influence lung function and the immune environment holds the potential to shed light on the origins of pulmonary diseases and cultivate innovative therapeutic approaches.
2.1.2 Oral cavity microbiota and pulmonary disease
2.1.2.1 The oral microbiome as a reservoir for respiratory pathogens
The oral cavity harbors a diverse microbial population that, when in a state of equilibrium, contributes to maintaining oral health. However, it is crucial to recognize that oral microorganisms can also serve as a significant reservoir for potential respiratory pathogens that may infiltrate the lungs through microaspiration, potentially leading to infections or immune activation [27]. With the 16S rRNA gene, some studies have indicated that microaspiration of oral commensals can instigate dysbiosis in the lower airways, ultimately promoting an IL-17-driven inflammatory phenotype that has the potential to impact lung tumorigenesis [28].
2.1.2.2 Oral microaspiration and its influence on cancer and COPD
In a study exploring the lung and oral microbiomes of individuals with lung cancer, it was observed that the microorganisms found in the bronchoalveolar lavage fluid (BALF) of the majority of lung cancer patients were also prevalent in their oral samples [29]. This underscores the significance of oral microaspiration as a mechanism through which oral microorganisms can influence lung health by entering the lower respiratory tract and lungs from the oral cavity.
Disruption in the oral microbiota and the influx of bacteria into the lungs via microaspiration have detrimental effects on lung function and the progression of chronic obstructive pulmonary disease (COPD). This exacerbates lung microbiota dysbiosis and contributes to lung injury [30].
2.1.2.3 Respiratory infections and oral microbiota
Furthermore, respiratory infections can impact the composition of the oral microbiota. For example, the composition of the oral microbiome varies depending on an individual’s smoking habits and HIV status. Among HIV-infected patients, changes in the oral microbiota were associated with abnormal lung function. An increased relative abundance of Veillonella, Streptococcus, and Lactobacillus suggested that the oral microbiome might serve as a biomarker for lung function in HIV patients [31].
SARS-CoV-2 infections have been shown to induce alterations in the oral microbiota, which can exacerbate pulmonary diseases. This highlights a close association between the oral microbiome and SARS-CoV-2 co-infections in the lungs, emphasizing the importance of effective oral health care measures in reducing these infections [32].
In summary, a robust and interconnected relationship exists between the oral and lung microbiomes, significantly impacting respiratory health and diseases. Manipulating the oral microbiome may hold therapeutic potential for pulmonary illnesses and reduce the likelihood of secondary infections in vulnerable populations.
2.1.2.4 Oral microbiota and idiopathic pulmonary fibrosis
A recent study investigates the relationship between oral microbiota and disease severity in idiopathic pulmonary fibrosis (IPF) using buccal swabs from 511 patients follow by 16S rRNA gene sequencing. It found that greater microbial diversity in the mouth is associated with worse lung function and an increased risk of death, while a higher proportion of the oral commensal Streptococcus mitis is linked to better lung function and improved survival. This suggests that the composition of oral microbiota may influence the progression and outcomes of IPF[33].
2.1.3 Dysbiosis in upper respiratory tract microbiota and pulmonary diseases
2.1.3.1 Advanced pulmonary disease and oropharyngeal dysbiosis
Recent research has unveiled compelling connections between upper respiratory tract (URT) microbiota dysbiosis and various pulmonary diseases. Individuals with advanced pulmonary disease, specifically lung transplant recipients six months post-transplantation, exhibited substantial dysbiosis in their oropharyngeal microbiota compared to healthy controls. This association suggests that advanced pulmonary disease is linked to microecological dysbiosis in the URT, potentially rendering the lungs susceptible to microbial abnormalities and heightened infection risk [34].
2.1.3.2 Pulmonary hypertension and respiratory microbiota alterations
Furthermore, a study on patients with pulmonary hypertension (PH) shed light on the link between PH and alterations in the upper respiratory microbiota. PH patients displayed increased respiratory microbiota abundance and reduced community diversity compared to healthy individuals. Notably, higher proportions of Streptococcus, Lautropia, and Ralstonia were observed in PH patients, accompanied by altered gene expression patterns in associated pathogenic bacteria [35].
2.1.3.3 Nasal microbiota variations in asthma
Additionally, significant differences in nasal microbiota were noted in asthma patients when compared to healthy controls with 16S rRNA gene sequencing. Subjects with both exacerbated and non-exacerbated asthma exhibited enriched taxa of Proteobacteria and Bacteroidetes [36].
2.1.3.4 Dysbiosis and its role in pneumonia development
The role of microbiota in pneumonia development has been extensively studied, revealing that dysbiosis plays a pivotal role in its pathogenesis. Among senior patients with pneumonia, reduced microbial species richness in the oropharynx and increased bacterial density were observed compared to healthy individuals. This bacterial overgrowth was directly linked to disease severity and is thought to result from disturbances in the oropharyngeal microbiota, reducing colonization resistance and the ability to restrain potential pathogens. Consequently, pathogens are transmitted to the lower respiratory tract, leading to pneumonia [37].
2.1.3.5 Nasopharyngeal microbiota and lower respiratory tract infections
In children with lower respiratory tract infections (LRTI), a strong concordance was discovered between viral and bacterial composition in nasopharyngeal and deep tracheal samples from the lower respiratory tract 16S rRNA gene sequencing. This suggests that nasopharyngeal samples can serve as surrogates for pulmonary microbiota during LRTI [38].
2.1.3.6 Nasopharyngeal microbiota and invasive pneumococcal disease
Furthermore, investigations into the nasopharyngeal microbiota's role in invasive pneumococcal disease revealed that colonization of the nasopharynx with S. pneumoniae, H. influenzae, or M. catarrhalis during childhood is associated with a higher likelihood of respiratory infections and allergies during the same period [39].
2.1.3.7 Immunocompromised patients and oropharyngeal dysregulation
It is worth noting that the development of pneumonia in immunocompromised patients may not be directly caused by lung microbiome dysregulation but rather by the inhalation of oropharyngeal pathogens into the lungs [40]. Studies in immunocompromised patients have shown significant dysregulation in the oropharyngeal microbiome, correlating with the severity of pulmonary disease. Low IgA levels were also associated with respiratory inflammatory responses and lung injury [41].
In conclusion, this body of research underscores the potential contribution of URT microbiota imbalance to pulmonary diseases. However, the precise pathogenesis of these diseases requires further elucidation. It is imperative to explore the intricate relationship between upper respiratory system microbiota and pulmonary diseases using a multi-omics and multi-site approach for comprehensive insights and potential therapeutic interventions.
2.1.4 Lung microbiota and pulmonary disease
2.1.4.1 Lung microbiota: a hidden community
With the arrival and advancement of high-throughput sequencing technologies, scientists have verified the presence of a multifaceted microbial community within a healthy lung. Due to their anatomical structure and physiological function, lung microorganisms exhibit a low biomass, and their community composition is dynamically balanced between microbial immigration (driven by inhalation, microaspiration, and mucosal dispersion) and microbial elimination (driven by cough, mucociliary transport, and immune mechanisms) [42].
2.1.4.2 Lung microbiota and pulmonary diseases
Respiratory health is closely linked to lung microbiota, where microecological dysbiosis contributes to the development and progression of chronic pulmonary diseases. The intricate correlations between pulmonary microbial communities and pulmonary diseases have been extensively discussed [43, 44]. The alteration of the lung microbiota can influence the risk of malignancy in various ways, and research has shown a link between the pulmonary microbiome and the development of lung cancer as well as the spread of cancer from other primary sites to the lungs[45].
2.1.4.3 Dysbiosis, inflammation, and disease progression
The worsening of chronic pulmonary disease is a result of dysbiosis, and dysregulated host immune response is triggered by microbial dysbiosis, which alters microbial growth and promotes further dysbiosis, perpetuating a cycle of inflammation and microbiota disruption [46]. And not only that, dysbiosis of the lung microbiota has been demonstrated in numerous pulmonary diseases not traditionally believed to be caused by microorganisms[47].
2.1.4.4 Predictive value of lung microbiota
In an evaluation of the impact of lung microbiome characteristics on subsequent chronic lung allograft dysfunction (CLAD)-free survival in healthy lung transplant recipients, the composition of the lung bacterial community in patients who developed CLAD or died was significantly different from those who survived and remained CLAD-free. This suggests that increased lung bacterial burden in asymptomatic lung transplant recipients one year after transplantation predicted chronic rejection and death [48].
2.1.4.5 Biomarkers and disease control
The survey indicates that the lung microbiota diversity decreased in patients with both pulmonary tuberculosis (TB, representing infectious pulmonary disease) and interstitial pneumonia (IP, representing non-infectious pulmonary disease) compared to healthy volunteers. Meanwhile, Mycobacterium's abundance increased in the communities of TB patients, and Firmicutes' abundance decreased in IP. However, there was a significant difference between TB and pneumonia patients: the abundance of Fusobacterium was noticeably lower in TB patients [49]. These findings indicate that alterations in lung microbiota may serve as biomarkers for pulmonary diseases, and monitoring and managing microecology could result in new disease control strategies. In the following section, we will delve into the relationship between lung microbiota and various pulmonary diseases.
The lung microbiome has a close relationship with immune function and respiratory health. Diseases are associated with both lung microbiota dysbiosis and diversity loss. An understanding of lung microbiome ecology and its disease interactions may facilitate microbiome-based diagnostics and therapeutics.
2.1.5 Tumor microbiota and pulmonary disease
2.1.5.1 Unveiling tumor microbiota
Tumor microbiota pertains to the microbial community found within tumor tissue. With the advancement and usage of next-generation sequencing technologies, mounting evidence suggests that tumor microbiota significantly contributes to cancer development, progression, and treatment [50].
2.1.5.2 The enigmatic relationship: cancer cells and microbes
There is no definitive evidence regarding the correlation between tumor microbiota and cancer cells. A study proposes several potential connections between the two: (1) In an obligate mutualistic relationship, cancer cells and bacterial cells depend entirely on each other, so the loss of one results in the loss of the other; (2) In a facultative mutualistic relationship, cancer cells and bacterial cells can coexist and benefit each other, but either can survive without the other; (3) In a commensalistic relationship, one party benefits while the other suffers neither harm nor benefit[51].
2.1.5.3 Tumor microbiota diversity and therapeutic impact
Recent evidence suggests that each tumor type exhibits a distinctive microbiota profile, influencing the effectiveness of cancer therapies. The impact of tumor microbiota depends on various factors, including tumor type, tissue type, and tumor status [52].
2.1.5.4 Mechanisms of tumorigenesis and immune suppression
Tumor microbiota can contribute to tumorigenesis and progression through multiple mechanisms, including DNA mutations, oncogenic signaling pathway activation, chronic inflammation, complement system activation, and promoting metastasis. Conversely, they can impair anti-tumor immune functions, increase reactive oxygen species levels, create anti-inflammatory environments, inhibit T-cell activity, and weaken cellular immunity [53].
2.1.5.5 Tumor suppression and immune enhancement
On the other hand, these microorganisms can also impair the anti-tumor monitoring and clearance functions of the body through various indirect pathways, such as increasing the level of reactive oxygen species (ROS) leading to damage of immune cells, promoting an anti-inflammatory environment and thus suppressing the immune response, inhibiting T-cell activity and weakening cellular immunity, and generally leading to a decrease in the functioning of the immune system [54].
Concerning tumor suppression, tumor microbiota may enhance anti-tumor immunity through mechanisms such as activating STING signaling, activating T and NK cells, producing tumor-associated lymphoid structures, and presenting microbial-derived antigens.
2.1.5.6 Microbial characterization in lung cancer
Recent studies on microbial characterization in lung cancer tissue have yielded inconsistent findings with 16S rRNA gene sequencing and metagenomic analyses. While some observed lower abundance and diversity of lung tumor microbiota compared to normal tissue, others found elevated prevalence and variety of certain pathogenic species [55, 56]. However, some studies have reached the opposite conclusion, indicating that the prevalence and variety of certain conditionally pathogenic species linked to the development and progression of cancer were elevated in lung cancer tissues compared to standard controls [57]. Differing conclusions suggest a potential link between changes in lung microbiota composition and lung cancer development [58, 59]. Further research is needed to clarify the mechanism. Some studies have identified microbial species strongly correlated with specific lung cancer subtypes, enhancing the understanding of tissue microbial alterations in lung cancer [60]. Future initiatives may focus on characterizing lung cancer microbiomes for diagnosis, treatment, and prognosis. The Cancer Microbiome Atlas (TCMA) offers valuable resources for investigating microbiota's role in various cancer types and predicting microbes as biomarkers [61].
2.1.5.7 Exploring fungal microbiota
In addition to bacteria, recent studies have endeavored to analyze the role of fungi in the microbiome of lung cancer. The findings indicate that despite their low absolute abundance, fungi are not uncommon in all common types of human lung cancer. Furthermore, specific groups of fungi can predict the survival of patients with tumors [62]. These results are a crucial step in initiating research into the hypothesized driving mechanisms for the role of fungal flora in various cancers.
Although the precise mechanism of action between tumor cells and microorganisms is not clear, there are specificities in the microbial composition of different tumor types that can bi-directionally regulate tumor progression. The conclusions of existing studies on tumor microbial profiles are divergent, but some microbial species have been shown to be highly associated with specific tumors. Future studies could focus on exploring the specificity of the tumor microbiome, which could help to improve the diagnosis and treatment of tumors.
2.2 Correlation between lung microbiota and other microbiota
There exists an intricate association between the microbiota of various regions of the human body and their reciprocal regulation. As a crucial organ of the respiratory system, variations in the indigenous microbiota inside the lung correspond to shifts in other areas of the body, like the oral cavity, upper respiratory tract, and intestinal tract. These changes can impact the lung microbiota in multiple ways [63, 64].
2.2.1 Gut microbiota and lung microbiota
In recent years, it has been found that there exists an important connection and interaction between the gut microbiota and lung microbiota in different parts of the human body. A complex network of interactions has been established between the two through direct microbial transmission and indirect communication of immune cells, which is very important for the maintenance of normal physiological functions of the human body. There are several key pathways for the establishment of this interaction.
Firstly, from the perspective of microbial transmission, the epithelial surfaces of the gut and respiratory tract are exposed to a variety of microorganisms. Externally ingested microorganisms can enter both areas simultaneously, and some members of the gut microbiota can directly enter the respiratory tract and lungs through pharyngeal reflux or microinhalation. Therefore, there is a certain degree of similarity and overlap between the two in terms of microbiota composition [65].
Secondly, from the perspective of metabolites, gut microbiota can produce a variety of small-molecule metabolites, which can enter the lymphatic and blood circulatory systems shared by the gut and lung, and ultimately reach the lungs through blood transportation or lymphatic transport, and have an impact on the lung microenvironment and immune status [66]. It has been found that short-chain fatty acids (SCFA) produced by gut microbiota regulate hematopoiesis through the gut-bone marrow-lung axis to activate the immune system to fight against lung infections and allergic reactions [67].
Thirdly, from the perspective of immune cells, the gut microbiota can stimulate the activation and proliferation of related immune cells, which migrate from the gut to the lungs through the common mucosal immune system (MIS) [68]. The migration of immune cells from the gut to the lungs can enhance the body's ability to resist infection, but it may also lead to pathological changes in the lungs under disease conditions. The gut and the lungs regulate the immune response of the lungs in normal and disease states through bidirectional interactions between immune cell migration and inflammatory mediators [66].
Fourthly, the interaction between the gut and lung microbiota is bidirectional. On the one hand, changes in gut microbiota can affect the composition and function of lung microbiota, and lead to changes in lung immune response. On the other hand, lung microbiota and pulmonary diseases can also in turn affect the balance of gut microbiota. This interaction occurs both through direct microbial contact and indirect effects through stimulation of immune cells [69, 70].
In summary, a close biological and immunological connection exists between the microorganisms in the gut and lungs. They can interact not only directly, but also remotely by modulating the host immune system and signaling pathways. This complex network constitutes the gut-lung axis [71]
2.2.2 Oral cavity microbiota and lung microbiota
A burgeoning body of epidemiological, microbiological, and molecular biology research has revealed a strong association between the oral microbiota and the lung microbiota. The movement of microbes within the oral cavity is believed to be a crucial means by which microbiomes populate the lungs during states of wellbeing. In healthy lungs, bacterial enclaves overlap with those found in the oral cavity, albeit at reduced quantities, with less diverse membership, and a distinct configuration [72]. Further studies have revealed that the oral and nasal cavities can act as external sources of lung microorganisms, and microorganisms from these two sites can contribute to the structure and composition of the lung microbiota by colonizing distinct ecological niches in the lungs. This highlights the significance of oral samples, such as saliva, as a means of monitoring and assessing lung microbial status in a clinical setting [73]. Collectively, it is apparent that the oral microbiome has a significant impact on the composition of both healthy and diseased individuals' lung microbiomes, contributing significantly to lung microbial diversity [74].
However, recent studies have also shown that the oral microbiome does not fully account for all the constituents of the lung microbiome. For instance, a neutral model comparison revealed that while most lung bacteria also exist in the oral cavity, certain bacteria, such as Enterobacteriaceae, Haemophilus, Methylobacterium, and Ralstonia species, were overrepresented in the lungs compared to values predicted by the neutral model. Additionally, Tropheryma is only present in the lungs and not in the oral cavity, indicating that not all lung microorganisms originate from the oral cavity [75]. In a study examining the composition of the oral and lower respiratory microbiota in young, healthy individuals, distinctions were observed between the oral wash and bronchoalveolar lavage samples. Notably, significant differences were found between the oral and lung microbiota [76].
Overall, the oral microbiota significantly impacts the lung microbiota in both healthy and diseased states. Certain oral microorganisms migrate to the lungs through microrespiration, subsequently contributing to the formation of the lung microecology. However, the unique structure and function of the lungs in comparison to the oral cavity means that the oral microbiota alone cannot fully dictate the overall composition of lung microorganisms. Additionally, the lung microbiota is subject to other influences, resulting in its distinct characteristics.
2.2.3 Upper respiratory tract microbiota and lung microbiota
2.2.3.1 Respiratory tract continuum
The upper respiratory tract serves as the gateway to the respiratory system, intricately connected to the lower respiratory tract both anatomically and microbially. This close linkage gives rise to a highly related microbial composition between the upper and lower respiratory tracts. Within this continuum, the lungs, as vital organs of the lower respiratory tract, host a diverse microbial community. In states of health, the lung microbiota closely resembles the oropharyngeal microbiota. At the phylum level, the predominant bacterial groups include Firmicutes, Bacteroidetes, and Proteobacteria. Notably, Streptococcus, Prevotella, and Veillonella emerge as the dominant genera. Conversely, pathogenic Aspergillus, such as Haemophilus, are found in significantly lower abundance [77].
2.2.3.2 Microbial diversity in the upper respiratory tract
Exploring the microbiome composition of the upper respiratory tract in healthy military personnel reveals distinct patterns. Elevated levels of Staphylococcus, Corynebacterium, and Propionibacterium are observed in the nasal cavity and nasopharynx. In contrast, Streptococcus reigns as the dominant bacterial genus within the oropharynx, albeit with greater diversity than the nasal cavity or nasopharynx. An analysis of diversity underscores a noticeable overlap between nasal and nasopharyngeal samples, with oropharyngeal samples forming a distinct cluster. This reveals unique microbial profiles within these areas [78].
2.2.3.3 Consistency across respiratory systems
Further investigation into the respiratory systems of healthy individuals unveils a consistent microbial composition despite variations in biomass between the upper and lower respiratory tracts. This consistency suggests a robust intrinsic link between the upper and lower respiratory tracts [79]. Intriguingly, a combined sampling approach, encompassing both oropharyngeal and nasopharyngeal microbiota, better reflects the bronchoalveolar lavage microbiota for most individuals, surpassing the analysis of either upper respiratory tract location alone[80].
2.2.3.4 Disease states and microbiota interaction
The interplay between the upper respiratory tract and lung microbiota extends beyond health and into disease states. For instance, in cystic fibrosis (CF) patients, research highlights a close relationship between their respiratory tract microbiota, despite individual differences. Moreover, evidence suggests that sinuses may serve as a significant source of lower respiratory tract infections, with the sinus and lung microbiota playing crucial roles in modulating the presence and abundance of typical CF pathogens [81].
2.2.3.5 Th17 immune activation and microbiota
An intriguing study delving into the lung microbiota through bronchoalveolar lavage identified a subset of healthy subjects with lung microbiota resembling common oral microbiota components, referred to as pneumotype-SPT. Pneumotype-SPT was associated with an augmented lung inflammatory phenotype, characterized by increased lymphocytes, Th17 cells, immune-characterized free fatty acids, elevated Th17 chemokines, heightened mRNA expression of inflammatory pathways, and reduced TLR4 responses. This observation suggests a correlation between Th17 immune activation in the lower respiratory tract mucosa and the constitution of the lung microbiota derived from the upper respiratory tract’s microrespiratory microbiota [82].
2.2.3.6 Mutual influence of upper and lower respiratory tract microbiota
In summary, the microbiota of the upper respiratory tract and lungs engage in a dynamic interaction, mutually influencing each other. Understanding the mechanisms underlying this interaction is imperative for predicting and comprehending pulmonary disease development. This intricate relationship underscores the need for further research in this evolving field of study, shedding light on the profound implications for respiratory health and disease management.
3 Association of lung microbiota with various types of pulmonary disease
A growing statistical data and experimental evidence shows that changes in the lung microbiota are highly correlated with the development of pulmonary disease. Treatments targeting specific pathogenic microorganisms have also shown significant efficacy [83, 84]. These findings emphasize the need for an in-depth study of the interrelationship between the two. By analyzing the mechanisms of microbe-host interactions, it may be possible to identify the causes of disease and provide a theoretical basis for future preventive, diagnostic and therapeutic strategies. Therefore, an in-depth study of the relationship between lung microorganisms and lung cancer, pneumonia, asthma, chronic obstructive pulmonary disease and other common pulmonary diseases will help to understand the pathogenesis, provide reference for diagnosis and treatment, and have important theoretical and practical significance for disease prevention and treatment. This is a necessary and valuable research topic (Fig. 2).
3.1 Idiopathic pulmonary fibrosis
Idiopathic pulmonary fibrosis (IPF) is the most common and severe type of idiopathic interstitial pneumonia (IIP). Conventional wisdom suggests that IPF is not associated with the lung microbiota; however, recent studies have found that the lung lung microbiota may have an impact on the pathogenesis and disease progression of IPF [85].
It was noted that reduced lung microbial diversity may be associated with disease activity in IPF and that the diversity of the lung microbiota may serve as a predictor of IPF pathogenesis [86]. Honeycombing is a major histopathologic feature of IPF and is closely associated with changes in the composition of the lung microbial community. This honeycombing phenomenon may cause ecological changes in the distal airways, thereby altering the composition of the microbial community and exacerbating lung tissue damage. This phenomenon suggests a dynamic bidirectional interaction between the lung microbiota and anatomical destruction in IPF [87]. Microbiota characteristics in bronchoalveolar lavage fluid from patients with early IPF were significantly correlated with their disease progression, and higher bacterial loads were not only an unfavorable factor for early prognosis, but also profoundly influenced the clinical manifestations of advanced IPF [88]. Altered microbiome burden, diversity, and composition may contribute to disease onset, progression, acute exacerbation, and death in IPF. If a mechanistic link between dysbiosis of the lung microbiome and the onset and progression of IPF is established, microbiome manipulation aimed at restoring “healthy” microbiome communities will soon become a potential therapeutic intervention for IPF [89]. As the pathogenesis of interstitial lung disease (ILD) continues to be investigated, the relationship between pulmonary fibrosis and the lung microbiota as well as immunomodulatory mechanisms has received increasing attention from medical researchers [90]. Patients diagnosed with idiopathic pulmonary fibrosis (IPF) had significantly increased intra-alveolar concentrations of pro-inflammatory fibrotic cytokines and growth factors when bacteria diversity in their lungs was reduced. The aforementioned points elucidate a corrective relationship between lung microbiota and inflammation, indicating that lung bacterial load prognosticates disease progression in IPF patients [91].
Despite the growing body of evidence supporting the link between lung microbiota and IPF, several points of controversy remain in the field. For example, some studies have had difficulty detecting microbiota in lung explants from patients with IPF, although this may be related to a variety of factors such as sample collection, processing, and analysis methods [92]. Therefore, whether microbiota dysbiosis directly causes the development of IPF needs to be verified by further scientific studies through more rigorous experimental designs.
3.2 Lung cancer
Lung cancer is one of the most rapidly increasing malignant tumors in terms of incidence rate and mortality, posing an imminent threat to human health and life. The occurrence of lung cancer is linked to a multitude of environmental and lifestyle factors, including smoking, radiation exposure, and genetic susceptibility. Recent studies have shown a correlation between changes in lung microbiota and cancer[93].
3.2.1 Altered lung microbiota in cancer patients
Recent research has highlighted substantial differences in the composition and diversity of lung microbiota between cancer patients and healthy individuals. In a study comparing lung tumor tissues with adjacent normal lung tissues, several genera exhibited significant variations. Notably, Phormidium, Propionibacterium, Rhodobacter, Finegoldia, and Microbacterium were found to be markedly reduced in tumor tissues, while Modestobacter showed enrichment in tumor tissues. This shift in microbial abundance suggests a decrease in potentially pro-inflammatory microbial genera within tumors [94].
3.2.2 Characteristics of lung tissue microbiota
An analysis of lung tissue microbiota in both tumor and non-tumor patients identified the predominant constituents as Proteobacteria, Actinobacteria, Firmicutes, and Bacteroidetes, with Proteobacteria being the most prevalent in both types of tissues. Notably, specific genera, including Massilia, Phenylobacterium, and Pseudoxanthomonas, were more abundant in tumor tissues, while others like Brevibacillus, Cupriavidus, and Anaerococcus were enriched in non-tumor tissue samples. Variations in abundance were also observed in genera such as Brevundimonas, Ruminococcus, and Polaromonas, which differed between squamous cell carcinoma and adenocarcinoma[95].
3.2.3 Lung microbiota and cancer progression
Emerging evidence suggests that alterations in lung microbiota may play a role in cancer progression. Normally, lung microbiota maintain a stable symbiotic relationship with their hosts, contributing to lung health. However, dysregulation in the composition and structure of lung microbiota, leading to a disrupted microbe-host balance, may be implicated in the development of lung cancer through various mechanisms [96]. These findings suggest that changes in lung microbiota could serve as an etiological factor in lung cancer development and influence its progression and response to therapy [97].
3.2.4 Role of specific microorganisms in lung cancer
In a comparative study of bronchoalveolar lavage fluid from patients with non-small cell lung cancer (NSCLC) and those with benign lung disease, distinct microbial patterns emerged. The NSCLC group exhibited significant enrichment at the phylum level of Firmicutes, Bacteroidetes, and Fusobacteriota, as well as at the genus level of Streptococcus, Prevotella, and Veillonella. Particularly, Veillonella (Parvula) was identified as an oncogenic bacterium that promotes lung cancer progression [98]. These findings underscore the relevance of specific microbial signatures in lung cancer and open avenues for further research into their diagnostic and therapeutic implications.
3.2.5 Microbial correlations in non-small cell lung cancer (NSCLC)
Recent studies have unveiled the potential of certain microorganisms in predicting and treating cancer [84]. An investigation involving 87 untreated non-small cell lung cancer (NSCLC) patients at various stages and 34 healthy volunteers shed light on the significant correlations within microbial data. The analysis of fecal and sputum microbiota data revealed a robust association between alterations in sputum microbiota and NSCLC, particularly distant metastasis (DM). This suggests that the lung microbiota might play a more pivotal role in NSCLC development than the gut microbiota. Furthermore, the study identified commonly shared microbial biomarkers across different disease stages, implying the possibility of early detection of DM. Notably, Pseudomonas aeruginosa emerged as a significant microbial marker for brain metastases [99].
3.2.6 Non-invasive diagnosis and biomarkers
These findings offer compelling support for non-invasive diagnosis of non-small cell lung cancer, even from a distance. They also highlight the potential to distinguish it from benign pulmonary diseases based on pronounced differences in lung microbiota. Notably, elevated levels of bacteria like Capnocytophaga were observed in lung cancer patients compared to those with benign pulmonary diseases [100]. Furthermore, lung cancer patients often exhibit high levels of Aspergillus spp. in their lungs, with greater diversity in squamous cell carcinomas, particularly among heavy-smoking men [101]. These discoveries emphasize the potential of lung microbiota as bacterial biomarkers, tools for patient stratification, future prognostic indicators, or even therapeutic targets, especially when combined with clinical tumor markers. Such a combination holds promise for enhanced cancer prediction to some extent, warranting further research and exploration [102].
It is unclear what the specific mechanism is, although the results provide a potential link between the lung microbiota and cancer mechanisms. Further discussion is necessary to determine how changes in microbial composition and function can promote cancer cell proliferation. Identifying key microorganisms or metabolites could offer new insights for disease diagnosis and treatment. In conclusion, comprehending the interplay between the human microbiome and cancer can aid in clarifying the pathogenesis of cancer and in creating novel diagnostic and therapeutic approaches.
3.3 Pneumonia
Pneumonia can be classified into viral pneumonia, bacterial pneumonia, and other forms based on the pathogen causing the infection, with each type showcasing distinct clinical presentations, severity, and prognosis. The lung microbiota exhibit varying roles in the development and advancement of the different types of pneumonia [103].
COVID‑19 SARS-CoV-2 is the pathogen responsible for causing novel coronavirus pneumonia. The composition of the microbial community in the BAL samples differed significantly between the COVID-19 and non-COVID-19 groups, with lower microbial diversity in the former. Staphylococcus aureus, Streptococcus anginosus, and Olsenella were significantly more abundant among microorganisms in the COVID-19 group as compared to the non-COVID-19 group of patients with Staphylococcus aureus ventilator-associated pneumonia (SA-VAP). These findings suggest that COVID-19 significantly affects the microbiological and clinical features of SA-VAP and is linked to a distinct lung microbiota composition [104]. In SARS patients, ACE2 expression is down-regulated during infection, which decreases antimicrobial peptide secretion, leading to increased intestinal pathogens and intestinal flora imbalance. Therefore, the regulation of intestinal flora by ACE2 may be a cause of diarrhea in COVID-19 patients. In addition, the microbial characteristics of lungs in COVID-19 patients may also predict the occurrence and prognosis of ARD [105]. Another study found changes in the lung microbiota in COVID-19 patients and that enrichment of the lung microbiota with bacteria from the gut was associated with the onset and long-term outcome of acute respiratory distress syndrome. and thatthe lung microbiota may affect the host immune response, with some specific bacterial and fungal species linked to decreased anti-inflammatory IL-4 signaling and increased signaling of IL-2, IL-3, IL-5, IL-6, IL-7, IL-10, IL-20, IL-22, and IL-23, as well as the IFNα and TGFβ signaling transduction pathways, inflammatory pathway, and immune dysregulation, which impact the severity and outcome of COVID-19 [106]. The composition of the lung microbiome was found to correlate with successful extubation in BAL samples from 114 COVID-19 patients receiving mechanical ventilation. Patients with higher lung bacterial and fungal loads had increased mortality rates, while higher bacterial loads in BAL samples positively correlated with concentrations of alveolar pro-inflammatory cytokines such as TNF-α and IL-1ß. This confirms the significance of the lung microbiome in both COVID-19 and ARDS [107]. A study discovered that RS-5645 (4-(thiophen-3-yl)-1-(p-tolyl)-1H-pyrrol-3-yl) (3,4,5-trimethoxyphenyl)methanone) lessened the effects of SARS-CoV-2 spike protein and LPS by adjusting the lung microbiota. Furthermore, it also reduced lung inflammatory cell infiltration, as well as the inflammatory storm caused by SARS-CoV-2 spike protein and LPS by regulating the pulmonary microbiota [108].
These studies indicate that COVID-19 is closely linked to the lung microbiota, and that modifying host microbes could be a promising new target for treating COVID-19. Deepening our understanding of the intricate mechanisms of the microbiota-host interaction will aid in the exploration of microecological treatment options for SARS-CoV-2 infection.
Influenza pneumonia Influenza virus infection may cause influenza pneumonia, which in severe cases can advance to acute respiratory distress syndrome and become life-threatening. Numerous studies have established a link between the lung microbiota and the development of influenza pneumonia. A recent study of mice infected with influenza A virus (IAV) discovered that the lung microbiota of healthy mice was primarily dominated by Lactobacillus spp., whereas the lung microbiota of IAV-infected mice was primarily dominated by Streptococcus. Further studies have indicated a decreased abundance of Lactobacillus in the oropharyngeal, nasopharyngeal, lung, and intestinal microbiota, which are vital for antiviral immunity. This suggests that the reduced abundance of L. murinus strains may be significant in IAV infection. Thus, L. murinus has been identified as a biomarker for IAV infection and a crucial target for intervention in post-influenza bacterial superinfection [109].
The lung microbiome has a significant impact on the development and operation of the pulmonary immune system in early stages of life. Changes in the microbial community could compromise the organism’s vulnerability to viral pneumonia. The microbiota composition was identified as a crucial factor in regulating the immune response of CD4 + and CD8 + T cells and antibodies post-respiratory influenza virus infection. The presence of neomycin-sensitive bacteria elicited a lung immune response, hinting at the microbiota’s critical role in managing adaptive immunity to respiratory virus infection [110]. Microbial-driven interferon (IFN) signature in lung stromal cells constitutes a homeostatic mechanism that maintains epithelial cells in an IFN-initiated state. This mechanism imparts resistance to influenza virus infection in mice by hindering the viral life cycle. The microbiota-driven signaling can operate on several levels. It can activate IFN signaling to induce non-immune cells into an antiviral state which can help in curbing early infections. Additionally, it can enhance the function of immune cells, thereby improving both innate and adaptive immunity during the later stages of infection [111]. These findings highlight the influence of interactions between bacterial and viral exposures on the use of antibiotics.
In addition, research on the gut-lung axis microbiota and the pathogenesis of influenza pneumonia has revealed that certain drugs may treat influenza pneumonia by influencing the microenvironment of the gut-lung axis. For instance, during pathological conditions, fritillary polysaccharide (HCP) interacts with the intestinal microbiota to decrease the migration of CCR6 + Th17/CCR6 + Treg cells from the intestinal mucosa-associated lymphoid tissues (GALT) to the lungs. Hence, this regulates the intestinal-lung Th17/Treg balance and mitigates the acute lung injury induced by influenza viruses. Please note: technical abbreviation HCP has been defined upon first use [112]. The six active ingredients in Lianhua Qingdian capsules have anti-inflammatory effects by regulating the microenvironment of the intestinal-lung axis and inhibiting the TLR/NF-κB pathway in the lungs. As a result, they alleviate lung and intestinal injuries, reduce lung viral load, improve survival rates, and suppress inflammatory responses [113]. Thus, the development of influenza pneumonia is closely related to microorganisms in the lungs and intestines. The microbial community plays a role in regulating immune function and inflammatory response, affecting viral clearance and causing lung injury.
Other pneumonias It was discovered that discrepancies existed among the lung microbial communities of patients with varying pneumonia types. The analysis of BALF samples showed that children with Mycoplasma pneumoniae pneumonia (MPP) exhibited lower bacterial α-diversity in their lungs in comparison to those with adenovirus pneumonia (AVP). At the phylum level, Tenericutes were the dominant bacteria in children with MPP while Firmicutes were the prominent bacteria in children with AVP. At the genus level, mycoplasma prevailed in children with MPP whereas streptococcus prevailed in children with AVP. The structure of the pulmonary microbial community significantly differed between the two types of pneumonia patients at the taxonomic level. This indicates that the dynamics of the pulmonary microbial community are closely associated with the emergence of various types of pneumonia [103]. High-throughput sequencing analysis revealed that community-acquired pneumonia (CAP) may alter the lung microbiota composition with a notable increase in the relative abundance of Proteobacteria, Bacteroidetes, Euryarchaeota, Firmicutes, and Spirochaetes. Klebsiella pneumoniae and Bacillus cereus were identified as potential microbial markers for CAP diagnosis and treatment [114]. In a study of 67 patients with severe community-acquired pneumonia (SCAP) admitted to the ICU, a higher α-diversity of lung microorganisms and a greater relative abundance of Prevotellaceae and Actinomycetaceae were found to be positively linked to symptomatic improvement in SCAP patients. The study suggests that tracking microbial changes in the lungs can add value to the evaluation of SCAP and the guidance of treatment [115]. The mechanism behind the development of ventilator-associated pneumonia (VAP) remains inadequately explained. The detection of lung microbiota has called into question the conventional pathological theory of VAP. As a result, molecular microbial detection techniques have become increasingly necessary for accurate VAP diagnosis. This may facilitate the use and advancement of immunomodulatory drugs and probiotic agents in the treatment of VAP [116].
These studies indicate that different types of pneumonia were significantly associated with changes in the composition and structure of lung microorganisms.
3.4 Asthma
The development and advancement of asthma are linked to a microbial imbalance within the lungs. Alterations in the microbial composition affect the asthma phenotype and severity, often manifesting as an elevation in roteobacteria, particularly those of the Haemophilus genus [117]. In addition, patients suffering from severe asthma may encounter a wider range of dysbiosis in the lung microbiota, resulting in a substantial increase in Actinobacteria compared to patients with mild-to-moderate asthma or healthy controls, with the most significant variation in Klebsiella [118]. Nonetheless, variations in research outcomes suggest that the heterogeneity of dysbiosis warrants further exploration to resolve discrepancies among studies.
The intricate interplay between gut microbiota and the development of the host's immune system during infancy is a pivotal factor impacting lung microbiota through the gut-lung axis. Early exposure to environmental microbes can potentially influence susceptibility to asthma later in life, primarily by modulating microbial metabolites associated with the gut-lung axis, such as short-chain fatty acids [119]. Disruption of the bidirectional communication of the gut-lung axis due to improper use of antibiotics, anti-ulcer drugs, or other medications can diminish the diversity of both intestinal and lung microbiota, increasing the risk of hypersensitivity reactions to respiratory and food allergens [120]. This implies that changes in the gut microbiome or the presence of specific pathogenic bacteria in the lungs during early life stages are linked to an elevated risk of allergic asthma. Notably, a study identified a significant increase in sputum-specific Gram-negative bacteria in corticosteroid-resistant asthmatics compared to corticosteroid-sensitive asthmatics and healthy controls. This microbial shift activated the TAK1/MAPK signaling pathway, resulting in corticosteroid resistance. However, the suppression of TAK1 restored cellular sensitivity to corticosteroids [121].
While the mechanisms underlying the interaction between asthma and lung microbiota are becoming clearer, research in this field is still in its nascent stages. As an integral component of asthma pathogenesis, many unknown mechanisms await in-depth exploration. A more comprehensive understanding necessitates larger-scale, scientifically designed studies employing multi-omics and bioinformatics approaches to delve into this complex relationship.
3.5 Chronic obstructive pulmonary disease (COPD)
3.5.1 Lung microbial composition in COPD
Chronic obstructive pulmonary disease (COPD), a prevalent respiratory ailment, has long been a subject of extensive research to decipher its precise pathogenesis. Recent studies have shed light on the substantial impact of microbial composition alterations in the lungs of COPD patients on disease progression. In individuals with mild to moderate COPD, the bronchial and lung regions harbor microorganisms such as Streptococcus, Corynebacterium, Alloiococcus, Prevotella, Veillonella, and Rothia, predominantly inhaled from the upper respiratory tract. These microbes engage with the host, inciting inflammatory reactions in the airways and lungs, thereby contributing to disease progression [122].
3.5.2 Microbial dynamics during copd exacerbations
A yearlong study delving into the lung microbiota of COPD patients during periods of stability and exacerbation revealed that patients experiencing frequent COPD exacerbations and deteriorations exhibited declining microbiome stability. Moreover, recurrent bacterial and eosinophilic exacerbations were more likely in such cases. Haemophilus and Moraxella emerged as strongly associated with disease severity, exacerbation events, and bronchodilation [123]. The heightened prevalence of Haemophilus spp. and other Proteobacteria species significantly disrupts the microbial community composition, leading to lung microflora dysbiosis and an inflammatory host response. This underscores the potential of Haemophilus spp. and similar bacteria as critical “keystone species” for lung microflora dysbiosis and potential therapeutic targets [124].
3.5.3 Lung microbiome and metabolites in COPD
Data pertaining to the lung microbiota and metabolites in smokers demonstrate a high correlation with disease severity and clinical indicators in COPD patients. This correlation not only reflects the strong association between the lung microbiome and clinical indicators but also suggests a synergistic effect between microbial composition and metabolites. These findings offer a novel perspective on the role of the microbial-host metabolic axis in the pathogenesis of pulmonary disease [125].
3.5.4 Fungal involvement in AECOPD
In addition to bacteria, fungal infections may also be linked to acute exacerbations of COPD (AECOPD). A study analyzing lung fungal communities in patients experiencing AECOPD found a negative correlation between alpha diversity and severity in non-survivors, patients requiring invasive mechanical ventilation, and those with acidemia AECOPD. This suggests a potential link between the alpha diversity of lung fungal communities and the severity, progression, and prognosis of AECOPD [126].
3.5.5 Exploring COPD pathogenesis
The pathogenesis of COPD is a multifaceted process involving various factors, with dysbiosis playing a significant role. Animal model studies have illuminated how microorganisms in the airway can activate multiple immune pathways, ultimately fostering COPD progression and leading to persistent inflammation and tissue damage in immune cells and airway epithelial cells [127]. Recent investigations into the mechanisms of lung function decline in COPD patients have revealed intriguing correlations. The presence of opportunistic pathogens at baseline airway dysbiosis in COPD patients was linked to a rapid decline in forced expiratory volume over two years. This suggests that Staphylococcus aureus colonization in the airways triggers homocysteine production, contributing to pulmonary function decline in emphysema [128].
COPD microbiology research continues to delve into the alterations and activities of microorganisms within the respiratory tract and lungs of COPD patients. The primary goals include uncovering the precise molecular mechanisms through which microorganisms impact COPD pathogenesis, assessing their diagnostic potential, and devising safe and effective microbial treatments. The aim is to swiftly translate the emerging diagnostic and therapeutic technologies targeting respiratory microorganisms from research laboratories to clinical settings, ultimately benefiting COPD patients.
3.6 Other lung diseases
Lung microbes are closely correlated with several other respiratory diseases. Patients diagnosed with idiopathic pulmonary fibrosis (IPF) had significantly increased intra-alveolar concentrations of pro-inflammatory fibrotic cytokines and growth factors when bacteria diversity in their lungs was reduced. The aforementioned points elucidate a corrective relationship between lung microbiota and inflammation, indicating that lung bacterial load prognosticates disease progression in IPF patients [91].
Analysis of the microbial composition of the lungs of patients with primary pulmonary tuberculosis (PTB), lung cancer (LC), and community-acquired pneumonia (CAP) revealed that, although all types of patients exhibited increased levels of microorganisms such as Mycobacterium, Selenomonas, Leptotrichia, Campylobacter, and Lactobacillus. Dysbiosis of lung microbiota in PTB patients promotes lung inflammation, which leads to higher levels of inflammatory factors in BALF of PTB patients than LC and CAP patients. In addition, bacterial α-diversity and abundance were higher in LC patients than in PTB and CAP patients, and microbiota alterations in CAP patients were significantly different between the PTB and LC groups, and these results reveal a pattern of differences in microbial and related immune mechanisms in the pathogenesis of different respiratory diseases [129].
In patients diagnosed with cystic fibrosis (CF), the α-diversity of the lung microbial community decreased significantly as the disease progressed. Meanwhile, P. melaninogenica, among others, became more and more predominant in patients with better lung function. The results indicate that a comprehensive analysis of changes in the microbial composition and dominant flora of CF patients' lungs can function as a reference index to evaluate the severity of CF disease [83].
Patients with connective tissue disease-associated interstitial pulmonary disease (CTD-ILD) who are colonized by Pseudomonas aeruginosa, Haemophilus influenzae, and non-tuberculous mycobacteria experience faster disease progression and greater impairment of lung function. This finding suggests that lung microbial colonization is associated with the severity of CTD-ILD and its mortality rate [130].
Understanding the microbial characteristics of the lungs of patients with various pulmonary diseases can aid in comprehending the pathogenesis of the diseases and creating novel diagnostic and therapeutic methods involving microorganisms. As research advances, fresh drugs and individualized therapeutic regimens, centered on the modulation of lung microorganisms, will surface, bringing new advantages to patients with respiratory illnesses.
4 Commentary and discussion
4.1 Clinical significance
Microbiota research has uncovered a profound link between microorganisms and human health, providing a new perspective on physiological mechanisms and disease pathogenesis. This has opened up new opportunities for disease diagnosis and treatment. In particular, the study of pulmonary flora has gained significant attention in recent years due to its crucial role in regulating lung immunity, the pathogenesis of respiratory diseases, and therapeutic efficacy. This article discusses the clinical significance of pulmonary flora research from three perspectives (Fig. 3).
Clinical Implications of Lung Microbiota Studies. A This schem discusses the effects of lung microbiota on the pathogenesis of pulmonary diseases, including impaired microbreathing, impaired airway clearance, and synergistic effects of lung microbiota and metabolites. B The scheme explores how the study of lung microbiota can be applied to the diagnosis, prevention, and treatment of diseases. Created using BioRender.com
4.1.1 Research results explain the mechanism of pulmonary disease
Changes in the microbial composition of the lungs and their physiological functions are intricately connected with the development of various respiratory diseases. Notably, patients afflicted with pulmonary hypertension (PH) manifest elevated respiratory microbiota richness, reduced community diversity, and marked increases in Streptococcus, Raucheria, and Lachnospira when compared to healthy individuals. These observations hint at a potential association between abnormalities in the respiratory microbiota and the pathogenesis of PH [35]. Additionally, respiratory colonization by Streptococcus pneumoniae, H. influenzae, and M. catarrhalis has been linked to respiratory infections and allergic predisposition in childhood [39]. Furthermore, colonization by Staphylococcus aureus has been shown to exacerbate emphysema through the involvement of homocysteine [128]. The New Coronary Pneumonia Study has revealed an active lung microbial community's association with the mechanisms of SARS-CoV-2 infection and its complications [131].
As the exploration of the relationship between lung microbes and lung diseases advances, a growing body of data is offering crucial insights into understanding the pathogenesis of various lung disorders. The available data suggest that the effects of microorganisms on the lung can be categorized into direct and indirect effects.
4.1.2 Direct effects
-
1.
In patients with mild to moderate COPD, microorganisms like Streptococcus, Corynebacterium, Alloiococcus, Prevotella, Veillonella, and Rothia, commonly found in the upper respiratory tract, infiltrate the lungs through microbial respiration. This intrusion triggers inflammatory responses in the airways and lungs, contributing to the progression of COPD [122].
-
2.
Impaired airway clearance may also play a role in the pathogenesis of lung cancer. An enrichment of Veillonella parvula, a pathogenic bacterium originating from the oral cavity, has been observed in the microbiota of lung cancer patients, and this enrichment has been correlated with a poor prognosis and reduced survival in early-stage patients [28].
-
3.
Synergistic interactions between microorganisms and metabolites may also be involved in the disease process. The gut microbiota and its metabolites in asthma patients may influence disease development by regulating ACE2 receptors, among other pathways [132]. Furthermore, microbes and metabolites can impact the course of COPD [125]
4.1.3 Indirect effects
Indirect effects primarily stem from lung microbial communities’ potential involvement in disease development by modulating the body's immune function and inflammatory responses. For example, an increase in Haemophilus spp. can lead to shifts in microbial community dynamics, resulting in dysbiosis of the lung microflora and provoking pro-inflammatory responses in the host [124]. Animal studies have also demonstrated that the microbial community structure governs the immune response of CD4 + and CD8 + T cells, as well as antibodies, in the context of respiratory influenza virus infection [110]. These findings imply that microbial communities might be integral to the body's defense against respiratory pathogens. Moreover, it is worth noting that microinhalation of oropharyngeal microbes has been associated with the reprogramming of host immune responses in the lung, possibly by promoting Th17 immune responses and inhibiting macrophage-mediated anti-inflammatory TLR4 signaling [82].
In conclusion, research into the interplay between lung microbes and respiratory diseases has illuminated fresh perspectives for understanding pathogenesis and is advancing towards personalized precision medicine. It is anticipated that with the continued development of technology, this field will witness further breakthroughs, ushering in a new era of diagnosis and treatment for respiratory diseases.
4.1.4 Application of research results in disease diagnosis and prediction
4.1.4.1 The cancer microbiome atlas: a pioneering resource
Recent advancements in microbiota analysis have revealed its substantial potential in diagnosing and predicting various diseases, including cancer and respiratory conditions. A groundbreaking development in this arena is The Cancer Microbiome Atlas (TCMA), serving as an unparalleled resource for investigating the role of tissue-associated microbiota across diverse cancer types. TCMA also stands as a predictive microbial biomarker for cancer diagnosis and treatment [61].
4.1.4.2 Lung microbiome insights in lung cancer
Examination of the lung microbiome via 16S rRNA sequencing of bronchoalveolar lavage fluid (BALF) samples has uncovered significant enrichments of bacteria such as Capnocytophaga in lung cancer patients [100]. Combining these microbial markers with clinical tumor indicators holds promise for enhancing lung cancer prediction. Further investigations have revealed distinct bacterial profiles in different subtypes of lung cancer, with proteobacteria showing higher abundance in squamous cell carcinoma, especially among male heavy smokers, compared to adenocarcinoma patients [101]. This discovery suggests the potential use of bacteria as biomarkers for stratifying lung cancer patients.
4.1.4.3 Microbial markers across various diseases
Studies exploring microbial markers have provided valuable insights across a spectrum of diseases:
-
Streptococci exhibited notable increases in the microbiome of central lung cancer patients compared to controls [133].
-
Acidovorax displayed higher abundance in cases of squamous cell carcinoma associated with TP53 mutations, a correlation not observed in adenocarcinoma [60].
-
In non-small cell lung cancer, significant alterations in the gut microbiota were detected, with Pseudomonas aeruginosa emerging as a potential microbial biomarker for brain metastasis [99].
-
Mouse models of influenza A virus infection revealed the reduction of Lactobacillus murinus during early infection, establishing it as a biomarker for influenza A virus infection [109].
-
Community-acquired pneumonia (CAP) profoundly impacted pulmonary microbial diversity, with Klebsiella pneumoniae and Bacillus cereus identified as potential diagnostic biomarkers [114].
-
Microbiota analysis in bronchoalveolar lavage fluid (BALF) samples from children with adenovirus pneumonia (AVP) or Mycoplasma pneumoniae pneumonia (MPP) suggested its potential in distinguishing between different pneumonia types [103].
-
In asthma, Actinobacteria enrichment in severe asthma patients compared to those with mild to moderate asthma or healthy individuals, with Klebsiella species exhibiting significant differences, has implications for asthma diagnosis and stratification [118].
-
For COPD patients, reduced microbial community stability in worsening COPD cases was observed, with Haemophilus and Moraxella identified as microbial markers enhancing COPD classification precision [123].
4.1.4.4 Harnessing microbiota analysis for disease insights
In summary, these findings underscore the vast potential of microbiota analysis in disease diagnosis and prediction. The identified microbial markers provide novel insights into understanding disease mechanisms, monitoring changes in health status, and guiding treatment strategies. This burgeoning field holds promise for revolutionizing disease management and precision medicine.
4.1.5 Application of research results in the treatment of pulmonary diseases
Exploring the intricate world of lung microbiota has emerged as a crucial endeavor in the diagnosis, treatment, and management of pulmonary diseases. In-depth investigations into the underlying mechanisms of pulmonary conditions have unveiled the pivotal role played by lung microbiota in disease pathogenesis and the regulation of pulmonary immunity.
4.1.5.1 Modifying microbiota for improved clinical outcomes
Significant progress has been made in understanding how lung microbiota can be modified to benefit patients. One notable study revealed a positive correlation between increased alpha diversity in lung microbiota and the enrichment of Prevotellaceae and Actinomycetaceae in individuals suffering from severe community-acquired pneumonia. These findings were associated with improved clinical symptoms, suggesting that the microbiota may be modifiable through interventions like probiotics or bacterial supplements [115].
4.1.5.2 Targeting microbes for COPD treatment
In the realm of Chronic Obstructive Pulmonary Disease (COPD), specific bacteria, such as Haemophilus spp., have emerged as potential treatment targets. During the progression of COPD, Haemophilus spp. and other Aspergillus spp. exhibit increased abundance, triggering a pro-inflammatory response in the host. Targeting airway microorganisms like Haemophilus spp. could provide a novel approach to slowing down COPD progression [124].
4.1.5.3 Restoring lung function through microbiome interventions
Studies have unveiled the significant impact of Staphylococcus aureus colonization in the airways of COPD patients, contributing to emphysema and diminished lung function. Interestingly, depleting Staphylococcus aureus through phagocytosis has shown promise in restoring lung function in mice with emphysema [128]. These findings underscore the potential of pulmonary microbiome studies in revolutionizing the treatment landscape for lung diseases. Understanding microbial variations can aid in the selection of microbial-based therapeutic strategies and provide a theoretical foundation for new drug development.
4.1.5.4 Personalized microbial modulation
The intricate interplay between individual genotypes and lifestyles leads to personalized microbial communities. This personalized microbial modulation approach holds promise, enabling real-time monitoring of individual microecological changes and tailored interventions. This personalized approach is poised to usher in a new paradigm in future medical treatments, where treatments are fine-tuned to individual microbiota profiles.
In summary, the exploration of lung microbiota offers a transformative potential in reshaping the landscape of pulmonary disease management. These insights not only provide new avenues for treatment but also pave the way for personalized and more effective medical interventions in the future.
4.2 Challenges and prospects
4.2.1 Challenges
Considerable progress has been made in the examination of the relationship between lung microbiota and diseases, yet there are still significant areas that require further attention. Firstly, the composition and origin of lung microbiota exert a profound influence on the occurrence and progression of lung diseases. Numerous studies have attempted to dissect the species composition and origins of lung microbiota. However, due to the intricate and individualized nature of these microbial communities, research results often bear a degree of uncertainty, necessitating additional investigations.
Secondly, delving into the mechanisms underlying the interplay between lung microbiota and disease onset and progression represents a crucial avenue in lung microbiome research. While many studies have explored this connection, limitations in research methods and techniques have predominantly led to descriptive and correlational studies. Functional investigations and clinical trials remain relatively scarce. Descriptive studies alone cannot conclusively establish microbial involvement in disease development, and correlational studies are insufficient to determine causation. Therefore, there is a pressing need for functional studies, along with experimental research and clinical trials of microbial therapies.
Lastly, the diagnosis and treatment of lung microbiota-related diseases are at the forefront of microbiological research. Numerous studies have been conducted in this realm, yet the development of microbial diagnostic and therapeutic products faces challenges related to technical and clinical validation. The microbial markers identified in existing studies have not yet been directly applied to clinical diagnosis, requiring rigorous methodological validation, large-scale clinical testing, and product development. This critical stage must be traversed to facilitate clinical translation.
4.2.2 Prospects
Looking ahead, the investigation of lung microorganisms and their connection to lung diseases must prioritize multidisciplinary collaboration, the development of advanced research methods and technologies intensified examination of lung microbial communities, exploration of the intricate mechanisms linking lung microbiota to lung diseases, and the pursuit of more effective diagnostic and therapeutic approaches. These endeavors will provide a more scientifically grounded foundation for the prevention and treatment of lung diseases and pave the way for novel breakthroughs in microbial therapies.
Our review highlights the emerging evidence that microbiome manipulation, such as the use of probiotics, prebiotics, and fecal microbiota transplants, could offer a more personalized and targeted approach to treating pulmonary diseases. These methods have the potential to address the underlying microbial dysbiosis that traditional therapies might overlook. Additionally, microbiome-based interventions could complement existing treatments by enhancing their efficacy or reducing side effects.
However, we also acknowledge that while the potential is significant, more research is needed to fully establish these approaches as reliable alternatives. This includes large-scale clinical trials and further exploration of the long-term effects of microbiome-based therapies. By incorporating this discussion, we aim to provide a balanced perspective on the current and future role of microbiome-based strategies in the therapeutic landscape for pulmonary diseases.
5 Literature selection process in this review
We conducted a comprehensive search of relevant literature using major databases, including PubMed, Scopus, and Web of Science. Our search focused on publications from the last 5 years (few 10 years) to capture the most current and relevant findings in the field. We used specific keywords related to pulmonary diseases, microbiota, and associated mechanisms to ensure a thorough and targeted search. We selected studies based on their relevance, quality, and contributions to the understanding of the interplay between microbiota and pulmonary health. This approach allowed us to provide a well-rounded and up-to-date review of the topic.
Data availability
No datasets were generated or analysed during the current study.
References
Shanahan F, et al. Feeding the microbiota: transducer of nutrient signals for the host. Gut. 2017;66(9):1709–17.
Donald K, Finlay BB. Early-life interactions between the microbiota and immune system: impact on immune system development and atopic disease. Nat Rev Immunol. 2023. https://doi.org/10.1038/s41577-023-00874-w.
Bouskra D, et al. Lymphoid tissue genesis induced by commensals through NOD1 regulates intestinal homeostasis. Nature. 2008;456(7221):507–10.
Zhao X, et al. Parishin from gastrodia elata ameliorates aging phenotype in mice in a gut microbiota-related manner. Front Microbiol. 2022;13: 877099.
Li M, et al. Theaflavins in black tea mitigate aging-associated cognitive dysfunction via the microbiota-gut-brain axis. J Agric Food Chem. 2023;71(5):2356–69.
Viaud S, et al. The intestinal microbiota modulates the anticancer immune effects of cyclophosphamide. Science. 2013;342(6161):971–6.
Joris BR, Gloor GB. Unaccounted risk of cardiovascular disease: the role of the microbiome in lipid metabolism. Curr Opin Lipidol. 2019;30(2):125–33.
Alexander M, Turnbaugh PJ. Deconstructing mechanisms of diet-microbiome-immune interactions. Immunity. 2020;53(2):264–76.
Abbasli G. Morpho-cultural and physiological characteristics of bacteria isolated from some thermal springs of the republic of azerbaijan. Adv Biol Earth Sci. 2024;9(1):124–33.
Khuraman Miryusifova GM. Arzu Allahverdiyeva, Nigar Huseynova, Aysun Umudlu, the saffron effects on the dynamics of experimental epilepsy. Adv Biol Earth Sci. 2024;9(1):196–202.
Karadağ M. Use of prunus armeniaca l seed oil and pulp in health and cosmetic products. Adv Biol Earth Sci. 2024;9:105–10.
Levy M, et al. Dysbiosis and the immune system. Nat Rev Immunol. 2017;17(4):219–32.
Piggott DA, Tuddenham S. The gut microbiome and frailty. Transl Res. 2020;221:23–43.
Hsiao A, Zhu J. Pathogenicity and virulence regulation of Vibrio cholerae at the interface of host-gut microbiome interactions. Virulence. 2020;11(1):1582–99.
Garrett WS. Cancer and the microbiota. Science. 2015;348(6230):80–6.
Zhou A, et al. Gut microbiota: the emerging link to lung homeostasis and disease. J Bacteriol. 2021;203:4.
Chunxi L, et al. The gut microbiota and respiratory diseases: new evidence. J Immunol Res. 2020;2020:2340670.
Depner M, et al. Maturation of the gut microbiome during the first year of life contributes to the protective farm effect on childhood asthma. Nat Med. 2020;26(11):1766–75.
Lai HC, et al. Gut microbiota modulates COPD pathogenesis: role of anti-inflammatory Parabacteroides goldsteinii lipopolysaccharide. Gut. 2022;71(2):309–21.
Zhang D, et al. The cross-talk between gut microbiota and lungs in common lung diseases. Front Microbiol. 2020;11:301.
Wang YH, et al. Gut microbiota-derived succinate aggravates acute lung injury after intestinal ischaemia/reperfusion in mice. Eur Respir J. 2023;61:2.
Zhu W, et al. Gut-lung axis: microbial crosstalk in pediatric respiratory tract infections. Front Immunol. 2021;12: 741233.
Ren Y, et al. Genetic liability of gut microbiota for idiopathic pulmonary fibrosis and lung function: a two-sample Mendelian randomization study. Front Cell Infect Microbiol. 2024;14:1348685.
Chioma OS, et al. Gut microbiota modulates lung fibrosis severity following acute lung injury in mice. Commun Biol. 2022;5(1):1401.
Fabbrizzi A, et al. Microbiota and IPF: hidden and detected relationships. Sarcoidosis Vasc Diffuse Lung Dis. 2021;38(3): e2021028.
Zhang F, et al. Gut microbiota in COVID-19: key microbial changes, potential mechanisms and clinical applications. Nat Rev Gastroenterol Hepatol. 2023;20(5):323–37.
Dong J, et al. Relationships between oral microecosystem and respiratory diseases. Front Mol Biosci. 2021;8: 718222.
Tsay JJ, et al. Lower airway dysbiosis affects lung cancer progression. Cancer Discov. 2021;11(2):293–307.
Sun Y, et al. Characterization of lung and oral microbiomes in lung cancer patients using culturomics and 16S rRNA gene sequencing. Microbiol Spectr. 2023. https://doi.org/10.1128/spectrum.00314-23.
Wen S, et al. The role of oral microbiota in chronic obstructive pulmonary disease. Respiration. 2022;101(9):859–68.
Yang L, et al. Alterations in oral microbiota in HIV are related to decreased pulmonary function. Am J Respir Crit Care Med. 2020;201(4):445–57.
Bao L, et al. Oral microbiome and SARS-CoV-2: beware of lung co-infection. Front Microbiol. 2020;11:1840.
O’Dwyer DN, et al. Commensal oral microbiota, disease severity, and mortality in fibrotic lung disease. Am J Respir Crit Care Med. 2024;209(9):1101–10.
Simon-Soro A, et al. Upper respiratory dysbiosis with a facultative-dominated ecotype in advanced lung disease and dynamic change after lung transplant. Ann Am Thorac Soc. 2019;16(11):1383–91.
Zhang C, et al. Altered airway microbiota composition in patients with pulmonary hypertension. Hypertension. 2020;76(5):1589–99.
Fazlollahi M, et al. The nasal microbiome in asthma. J Allergy Clin Immunol. 2018;142(3):834–43.
de Steenhuijsen Piters WA, et al. Dysbiosis of upper respiratory tract microbiota in elderly pneumonia patients. ISME J. 2016;10(1):97–108.
Man WH, et al. Bacterial and viral respiratory tract microbiota and host characteristics in children with lower respiratory tract infections: a matched case-control study. Lancet Respir Med. 2019;7(5):417–26.
Dietl B, et al. Related factors to streptococcus pneumoniae invasive infection and clinical manifestations: the potential role of nasopharyngeal microbiome. Front Med (Lausanne). 2021;8: 650271.
Dong H, et al. The effects of immunosuppression on the lung microbiome and metabolites in rats. Front Microbiol. 2022;13: 817159.
Berbers RM, et al. Low IgA associated with oropharyngeal microbiota changes and lung disease in primary antibody deficiency. Front Immunol. 2020;11:1245.
Natalini JG, Singh S, Segal LN. The dynamic lung microbiome in health and disease. Nat Rev Microbiol. 2023;21(4):222–35.
Shukla SD, et al. Microbiome effects on immunity, health and disease in the lung. Clin Transl Immunology. 2017;6(3): e133.
Xiang L, Meng X. Emerging cellular and molecular interactions between the lung microbiota and lung diseases. Crit Rev Microbiol. 2022;48(5):577–610.
Ramirez-Labrada AG, et al. The influence of lung microbiota on lung carcinogenesis, immunity, and immunotherapy. Trends Cancer. 2020;6(2):86–97.
Dickson RP, Martinez FJ, Huffnagle GB. The role of the microbiome in exacerbations of chronic lung diseases. Lancet. 2014;384(9944):691–702.
Whiteside SA, McGinniss JE, Collman RG. The lung microbiome: progress and promise. J Clin Invest. 2021;131:15.
Combs MP, et al. Lung microbiota predict chronic rejection in healthy lung transplant recipients: a prospective cohort study. Lancet Respir Med. 2021;9(6):601–12.
Vazquez-Perez JA, et al. Alveolar microbiota profile in patients with human pulmonary tuberculosis and interstitial pneumonia. Microb Pathog. 2020;139: 103851.
Bahuguna A, Dubey SK. Relevance of tumor microbiome in cancer incidence, prognosis, and its clinical implications in therapeutics. Reviews Cancer. 2023;1878:5.
Bullman S. The intratumoral microbiota: From microniches to single cells. Cell. 2023;186(8):1532–4.
Ferrari V, Rescigno M. The intratumoral microbiota: friend or foe? Trends Cancer. 2023;9(6):472–9.
Wong-Rolle A, et al. Unexpected guests in the tumor microenvironment: microbiome in cancer. Protein Cell. 2020;12(5):426–35.
Xie Y, et al. Microbiota in tumors: from understanding to application. Adv Sci. 2022;9:21.
Fu A, et al. Emerging roles of intratumor microbiota in cancer metastasis. Trends Cell Biol. 2022. https://doi.org/10.1016/j.tcb.2022.11.007.
Peters BA, et al. The microbiome in lung cancer tissue and recurrence-free survival. Cancer Epidemiol Biomarkers Prev. 2019;28(4):731–40.
Ekanayake A, et al. Respiratory bacterial microbiota and individual bacterial variability in lung cancer and bronchiectasis patients. Indian J Microbiol. 2020;60(2):196–205.
Xu N, et al. Microbiota dysbiosis in lung cancer: evidence of association and potential mechanisms. Translational Lung Cancer Res. 2020;9(4):1554–68.
Mao Q, et al. Differential flora in the microenvironment of lung tumor and paired adjacent normal tissues. Carcinogenesis. 2020;41(8):1094–103.
Greathouse KL, et al. Interaction between the microbiome and TP53 in human lung cancer. Genome Biol. 2018;19(1):123.
Dohlman AB, et al. The cancer microbiome atlas: a pan-cancer comparative analysis to distinguish tissue-resident microbiota from contaminants. Cell Host Microbe. 2021;29(2):281–98.
Li X, Saxena D. The tumor mycobiome: a paradigm shift in cancer pathogenesis. Cell. 2022;185(20):3648–51.
Nicola I, et al. Characterization of the upper and lower respiratory tract microbiota in Piedmontese calves. Microbiome. 2017;5:1.
Vientós-Plotts AI, Ericsson AC, Reinero CR. The respiratory microbiota and its impact on health and disease in dogs and cats: a one Health perspective. J Vet Intern Med. 2023;37(5):1641–55.
Budden KF, et al. Emerging pathogenic links between microbiota and the gut-lung axis. Nat Rev Microbiol. 2017;15(1):55–63.
Wypych TP, Wickramasinghe LC, Marsland BJ. The influence of the microbiome on respiratory health. Nat Immunol. 2019;20(10):1279–90.
Dang AT, Marsland BJ. Microbes, metabolites, and the gut–lung axis. Mucosal Immunol. 2019;12(4):843–50.
Anand S, Mande SS. Diet microbiota and gut-lung connection. Front Microbiol. 2018. https://doi.org/10.3389/fmicb.2018.02147.
Enaud R, et al. The gut-lung axis in health and respiratory diseases: a place for inter-organ and inter-kingdom crosstalks. Front Cell Infect Microbiol. 2020;10:9.
Bingula R, et al. Desired turbulence? gut-lung axis, immunity, and lung cancer. J Oncol. 2017;2017:1–15.
He Y, et al. Gut-lung axis: the microbial contributions and clinical implications. Crit Rev Microbiol. 2017;43(1):81–95.
Bassis CM, et al. Analysis of the upper respiratory tract microbiotas as the source of the lung and gastric microbiotas in healthy individuals. Mbio. 2015;6(2):e00037.
Zhang J, et al. Differential oral microbial input determines two microbiota pneumo-types associated with health status. Adv Sci (Weinh). 2022;9(32): e2203115.
Mammen MJ, Scannapieco FA, Sethi S. Oral-lung microbiome interactions in lung diseases. Periodontol. 2020;83(1):234–41.
Morris A, et al. Comparison of the respiratory microbiome in healthy nonsmokers and smokers. Am J Respir Crit Care Med. 2013;187(10):1067–75.
Leitao Filho FS, et al. Characterization of the lower airways and oral microbiota in healthy young persons in the community. Biomedicines. 2023;11:3.
Faner R, et al. The microbiome in respiratory medicine: current challenges and future perspectives. Eur Respir J. 2017;49:4.
Hang J, et al. Composition and variation of respiratory microbiota in healthy military personnel. PLoS ONE. 2017;12(12): e0188461.
Charlson ES, et al. Topographical continuity of bacterial populations in the healthy human respiratory tract. Am J Respir Crit Care Med. 2011;184(8):957–63.
Marsh RL, et al. The microbiota in bronchoalveolar lavage from young children with chronic lung disease includes taxa present in both the oropharynx and nasopharynx. Microbiome. 2016;4(1):37.
Lucas SK, et al. 16S rRNA gene sequencing reveals site-specific signatures of the upper and lower airways of cystic fibrosis patients. J Cyst Fibros. 2018;17(2):204–12.
Segal LN, et al. Enrichment of the lung microbiome with oral taxa is associated with lung inflammation of a Th17 phenotype. Nat Microbiol. 2016;1:16031.
Cuthbertson L, et al. Lung function and microbiota diversity in cystic fibrosis. Microbiome. 2020;8(1):45.
Yang D, et al. The impact of lung microbiota dysbiosis on inflammation. Immunology. 2020;159(2):156–66.
Amati F, et al. Lung microbiome in idiopathic pulmonary fibrosis and other interstitial lung diseases. Int J Mol Sci. 2022;23:2.
Takahashi Y, et al. Impaired diversity of the lung microbiome predicts progression of idiopathic pulmonary fibrosis. Respiratory Res. 2018;19:1.
Dickson RP, et al. Radiographic honeycombing and altered lung microbiota in patients with idiopathic pulmonary fibrosis. Am J Respir Crit Care Med. 2019;200(12):1544–7.
Valenzi, E.B., et al., 2020.
Spagnolo P, et al. The role of the lung’s microbiome in the pathogenesis and progression of idiopathic pulmonary fibrosis. Int J Mol Sci. 2019;20:22.
Zhang T, et al. Potential targeted therapy based on deep insight into the relationship between the pulmonary microbiota and immune regulation in lung fibrosis. Front Immunol. 2023. https://doi.org/10.3389/fimmu.2023.1032355.
O’Dwyer DN, et al. Lung microbiota contribute to pulmonary inflammation and disease progression in pulmonary fibrosis. Am J Respir Crit Care Med. 2019;199(9):1127–38.
Kitsios GD, et al. Microbiome in lung explants of idiopathic pulmonary fibrosis: a case–control study in patients with end-stage fibrosis. Thorax. 2018;73(5):481–4.
Huynh M, Crane MJ, Jamieson AM. The lung, the niche, and the microbe: Exploring the lung microbiome in cancer and immunity. Front Immunol. 2023. https://doi.org/10.3389/fimmu.2022.1094110/full.
Mao Q. Comparison between the microbiota in lung tumor and adjacent normal tissues in lung cancer patients. J Clin Oncol. 2018;36:e20548–e20548.
Dong H, et al. Convergent alteration of lung tissue microbiota and tumor cells in lung cancer. Iscience. 2022;25(1):103638.
Khan FH, et al. Microbiome dysbiosis and epigenetic modulations in lung cancer: From pathogenesis to therapy. Semin Cancer Biol. 2022;86:732–42.
Sommariva M, et al. The lung microbiota: role in maintaining pulmonary immune homeostasis and its implications in cancer development and therapy. Cell Mol Life Sci. 2020;77(14):2739–49.
Zeng W, et al. Alterations of lung microbiota in patients with non-small cell lung cancer. Bioengineered. 2022;13(3):6665–77.
Lu H, et al. Alterations of the human lung and gut microbiomes in non-small cell lung carcinomas and distant metastasis. Microbiol Spectr. 2021;9(3): e0080221.
Cheng C, et al. Characterization of the lung microbiome and exploration of potential bacterial biomarkers for lung cancer. Transl Lung Cancer Res. 2020;9(3):693–704.
Gomes S, et al. Profiling of lung microbiota discloses differences in adenocarcinoma and squamous cell carcinoma. Sci Rep. 2019;9(1):12838.
Weinberg F, et al. The Lung Microbiome: a central mediator of host inflammation and metabolism in lung cancer patients? Cancers. 2020;13:1.
Wang H, et al. Lung microbiota and pulmonary inflammatory cytokines expression vary in children with tracheomalacia and adenoviral or mycoplasma pneumoniae pneumonia. Front Pediatr. 2019;7:265.
De Pascale G, et al. Staphylococcus aureus ventilator-associated pneumonia in patients with COVID-19: clinical features and potential inference with lung dysbiosis. Crit Care. 2021;25(1):197.
He Y, et al. Main clinical features of COVID-19 and potential prognostic and therapeutic value of the microbiota in SARS-CoV-2 Infections. Front Microbiol. 2020;11:1302.
Dereschuk K, et al. Identification of lung and blood microbiota implicated in covid-19 prognosis. Cells. 2021;10:6.
Kullberg RFJ, et al. Lung microbiota of critically ill patients with covid-19 are associated with nonresolving acute respiratory distress syndrome. Am J Respir Crit Care Med. 2022;206(7):846–56.
Liu T, et al. RS-5645 attenuates inflammatory cytokine storm induced by SARS-CoV-2 spike protein and LPS by modulating pulmonary microbiota. Int J Biol Sci. 2021;17(13):3305–19.
Chen Q, et al. Topography of respiratory tract and gut microbiota in mice with influenza a virus infection. Front Microbiol. 2023;14:1129690.
Ichinohe T, et al. Microbiota regulates immune defense against respiratory tract influenza A virus infection. Proc Natl Acad Sci U S A. 2011;108(13):5354–9.
Bradley KC, et al. Microbiota-driven tonic interferon signals in lung stromal cells protect from influenza virus infection. Cell Rep. 2019;28:1.
Shi C, et al. Intestinal microbiota metabolizing Houttuynia cordata polysaccharides in H1N1 induced pneumonia mice contributed to Th17/Treg rebalance in gut-lung axis. Int J Biol Macromol. 2022;221:288–302.
Xu P, et al. Intestinal microbiota analysis and network pharmacology reveal the mechanism by which Lianhua Qingwen capsule improves the immune function of mice infected with influenza A virus. Front Microbiol. 2022;13:1035941.
Hong L, Chen Y, Ye L. Characteristics of the lung microbiota in lower respiratory tract infections with and without history of pneumonia. Bioengineered. 2021;12(2):10480–90.
Du S, et al. Clinical factors associated with composition of lung microbiota and important taxa predicting clinical prognosis in patients with severe community-acquired pneumonia. Front Med. 2022;16(3):389–402.
Fernandez-Barat L, Lopez-Aladid R, Torres A. Reconsidering ventilator-associated pneumonia from a new dimension of the lung microbiome. EBioMedicine. 2020;60: 102995.
Chung KF. Airway microbial dysbiosis in asthmatic patients: A target for prevention and treatment? J Allergy Clin Immunol. 2017;139(4):1071–81.
Huang YJ, et al. The airway microbiome in patients with severe asthma: associations with disease features and severity. J Allergy Clin Immunol. 2015;136(4):874–84.
Loverdos K, et al. Lung microbiome in asthma: current perspectives. J Clin Med. 2019;8:11.
Hufnagl K, et al. Dysbiosis of the gut and lung microbiome has a role in asthma. Semin Immunopathol. 2020;42(1):75–93.
Goleva E, et al. The effects of airway microbiome on corticosteroid responsiveness in asthma. Am J Respir Crit Care Med. 2013;188(10):1193–201.
Pragman AA, et al. The lung tissue microbiota of mild and moderate chronic obstructive pulmonary disease. Microbiome. 2018;6(1):7.
Mayhew D, et al. Longitudinal profiling of the lung microbiome in the AERIS study demonstrates repeatability of bacterial and eosinophilic COPD exacerbations. Thorax. 2018;73(5):422–30.
Wang Z, et al. Lung microbiome dynamics in COPD exacerbations. Eur Respir J. 2016;47(4):1082–92.
Madapoosi SS, et al. Lung microbiota and metabolites collectively associate with clinical outcomes in milder stage chronic obstructive pulmonary disease. Am J Respir Crit Care Med. 2022;206(4):427–39.
Enaud R, et al. Lung mycobiota α-diversity is linked to severity in critically ill patients with acute exacerbation of chronic obstructive pulmonary disease. Microbiol Spectrum. 2023;11:2.
Kayongo A, et al. Airway microbiome-immune crosstalk in chronic obstructive pulmonary disease. Front Immunol. 2022;13:1085551.
Liang W, et al. Airway dysbiosis accelerates lung function decline in chronic obstructive pulmonary disease. Cell Host Microbe. 2023;31:6.
Xia X, et al. Comparative analysis of the lung microbiota in patients with respiratory infections, tuberculosis, and lung cancer: A preliminary study. Front Cell Infect Microbiol. 2022;12:1024867.
Ricci A, et al. The role of lung colonization in connective tissue disease-associated interstitial lung disease. Microorganisms. 2021;9:5.
Han Y, et al. The active lung microbiota landscape of COVID-19 patients through the metatranscriptome data analysis. Bioimpacts. 2022;12(2):139–46.
Barcik W, et al. The role of lung and gut microbiota in the pathology of asthma. Immunity. 2020;52(2):241–55.
Bello S, et al. Core microbiota in central lung cancer with streptococcal enrichment as a possible diagnostic marker. Arch Bronconeumol. 2021;57(11):681–9.
Acknowledgements
This research was supported by Natural Science Foundation of Xuzhou [KC21056]
Funding
Natural Science Foundation of Xuzhou, KC21056
Author information
Authors and Affiliations
Contributions
Y.S., J.Y. and F.G.: Conceptualization, literature review, writing – original draft, writing – review & editing. P.L., S.Z., H.C.: Literature search, writing – review & editing. Y.Y. and Y.M.: Literature search, writing, and editing for the revision.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Yu, J., Li, P., Yu, Y. et al. A comprehensive insight of complex interplay of microbiota in pulmonary diseases. Discov Med 1, 48 (2024). https://doi.org/10.1007/s44337-024-00063-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44337-024-00063-1