Abstract
Archaea represent a prevalent and abundant microbial group and are believed to play a crucial role in marine ecosystems. Archaeal distribution remains poorly resolved in the western subarctic Pacific (WSP). We investigated the relative abundance, vertical distribution, and phylogenetic composition of Archaea in WSP. The abundance of Euryarchaeota marine group II (Poseidoniales) was the highest (> 77%) in the epipelagic layer and decreased with depth, the abundance of Thaumarchaeota increased with depth, and Euryarchaeota marine group III (Pontarchaea) was absent in the epipelagic layer but was highly abundant in mesopelagic and bathypelagic layers. Based on phylogenic analyses of archaeal 16S rRNA gene partial sequences, we noted that archaeal phylotypes in the epipelagic layer were different from those in the mesopelagic and bathypelagic layers. The phylogenetic composition in the archaeal community was strongly segregated along a depth gradient. The Shannon index and the richness values revealed that the mesopelagic layer was the most diverse habitat of the archaeal community. Additionally, the trends of the vertical distribution of Archaea and Thaumarchaeota, elucidated through the qPCR assay, at station K2 were similar to those documented in other ocean regions, although their absolute quantity was low. Moreover, the unexpected bias in qPCR was noted to be caused by the commonly used Thaumarchaeota-specific primer MCGI391F, which mismatched some sequences derived from the epipelagic water.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Archaea were long believed to exclusively inhabit extreme environments on Earth, until Fuhrman et al. (1992) and DeLong (1992) reported about nonthermophilic planktonic Archaea in the marine water column. Since then, archaeal distribution and diversity have been investigated in various oceanic regions (Church et al., 2003; Herndl et al., 2005; Karner et al., 2001; Kirchman et al., 2007; Tamburini et al., 2009). These studies have revealed high Archaea abundance in the surface, mesopelagic, and bathypelagic layers of various oceanic ecosystems, which accounted for approximately one-third of all prokaryotic cells in the global ocean. This suggests that Archaea plays a crucial role in marine ecosystems. Most oceanic Archaea are unculturable. As research has progressed, the methods of analysis have also evolved from the 16S rRNA sequencing-based initial systematic classification to the application of metagenomic analysis and, more recently, single-cell sequencing. Archaeal classification has progressed from the two phyla (Crenarchaea and Euryarchaeota) named after their discovery in 1977 to more than two dozen phyla (Galand et al., 2009), although the classification of some individual phyla is controversial. New research conducted from 2011 to 2017 has led to the creation of three superphyla: TACK, DPANN, and Asgard. The superphylum Asgard comprises several phyla, including Thaumarchaeota, Aigarchaeota, Crenarchaeota, and Korarchaeota. Euryarchaeota does not fall within a superphylum. Euryarchaeota marine group II (MGII) and MGIII are now referred to as Poseidoniales and Pontarchaea ( Baker et al., 2020; Rinke et al., 2019).
The community structure, diversity, and spatial distribution of Archaea in the pelagic ocean have been described in ongoing research. According to earlier studies, MGI Thaumarchaeota was the most abundant in the deep ocean, whereas Euryarchaeota was more abundant in the upper layer (Herndl et al., 2005; Karner et al., 2001; Massana et al., 2000). In the eastern part of the North Pacific Ocean Gyre, MGI Thaumarchaeota accounts for more than 30% of all microbial cells from 200- to 3000-m depth (Karner et al., 2001). The distribution patterns of major marine Archaea groups have also been mapped based on oceanic circulation (Galand et al., 2009), latitude (Varela et al., 2008), and characteristics of ocean water masses (Tamburini et al., 2009). Differences in the physicochemical and biological parameters of seawater masses possibly influence archaeal biogeography (Herfort et al., 2007).
The western subarctic Pacific (WSP) is considered among the most productive areas in the ocean. WSP is characterized as a high-nutrient, low-chlorophyll area to which cool waters from the Bering Sea are brought by the Oyashio current. Thus, WSP is remarkably different from the eastern North Pacific in terms of bioelement concentrations, primary production, bacterial production, export carbon flux and export rate, and carbon sedimentation rate (Komuro et al., 2005; Longhurst et al., 1995; Sambrotto et al., 1984). For instance, because of deeper winter mixing and distinct hydrography, WSP has higher primary production than the eastern North Pacific, where a greater diversity of communities is observed (Brown et al., 2009; DeLong et al., 2006; Karner et al., 2001; Massana et al., 1997).
Moreover, few studies of microbial diversity and population structure have been conducted at stations KNOT and K2 in WSP. Seasonal variability in the picoplankton community structure and in heterotrophic bacteria abundance has been reported at these time-series stations (Komuro et al., 2005; Liu et al., 2002; Suzuki et al., 2002), but knowledge about archaeal diversity or distribution at these stations is very limited. To obtain some insights into the potential role of Archaea in global marine ecology, in the present study, the phylogenetic composition of Archaea assemblages was investigated in WSP by using a 16S rRNA gene clone library approach. The relative abundance and distribution of Archaea and MGI Thaumarchaeota were precisely characterized through quantitative PCR.
2 Materials and methods
2.1 Sampling sites and collection
Sampling was conducted aboard the R/V Mirai of the Japan Agency for Marine-Earth Science and Technology (JAMSTEC) during the MR07-05 cruise (September 4 to October 2, 2007). The samples collected from station K2 (47°N, 160°E) were used for investigating the vertical distribution of Archaea. Water samples were collected using a conductivity–temperature–depth–optical (CTDO) system at multiple depths (5, 50, 500, 1000, 4000, and 5000 m) in 36 of the 12-L Niskin-X water samplers. All samples were immediately placed at 4 °C. Then, microbial biomass in 2-L seawater samples was trapped through sequential filtering by using quartz fiber filters (Whatman QM-A, diameter: 47 mm) and polycarbonate filters (0.2-µm Whatman, diameter: 47 mm) on board within 2 h. These filters were frozen at −20 °C until after the cruise. For analyzing the total microbial cell number in the future, 10 mL seawater samples were fixed with formaldehyde (4% final concentration) and stored at 4 °C in the dark until analysis.
2.2 Enumeration of total microbial cell number
Seawater samples (1–5 mL) were vacuum-filtered through a black polycarbonate membrane filter (pore size: 0.20 μm, Advantec, Japan). The filters were stained with 100 μg mL−1 solution of the ethidium bromide fluorescent dye for 5 min, modified from Hobbie et al. (1977), to visualize microbial cells. The cells were counted using an epifluorescence microscope (BX-50, Olympus, Japan).
2.3 DNA extraction and clone library construction
With slight modifications to the method of West and Scanlan (1999), genomic DNA was extracted from the samples collected at three distinct depths at station K2 (epipelagic, 50 m; mesopelagic, 1000 m; and bathypelagic, 4000 m) as follows. Before the nucleic acids were precipitated, 0.1 volume of 3 M sodium acetate and 2.5 volume of 99% ethanol were added to the extracted DNA at room temperature. The extracted DNA was resuspended in TE (50 mM Tris–HCl, pH 8.0; 10 mM EDTA) and stored at − 30 °C for later use. The genomic DNA extracted at station K2 was used to construct the archaeal 16S rRNA gene clone library by using the Archaea-specific primer set A109F1/1492R (Table 1). By using the pCR2.1-TOPO TA vector (Invitrogen, Netherlands), PCR amplification and clone library construction were performed as described previously (Utsumi et al., 2003). The products of four identical PCRs (20 μL) were pooled to create a sufficiently large quantity to serve as a template for constructing the library. For each library, 96 white positive clones were picked randomly and screened through colony PCR with M13 primers. For sequencing, plasmid DNA was retrieved and purified using a Perfectprep Plasmid 96 Vac Kit (Eppendorf, Germany).
2.4 Sequencing, phylogenetic analysis, and diversity calculation
The sequences of insert PCR amplicons, which were selected from the recombinant colonies, were determined using a capillary DNA sequencer (Applied Biosystems 3100, Applied Biosystems, Foster City, CA, USA). The vector-specific primer sets M13F and M13R were employed for sequencing reactions with BigDye Terminator version 3.1 (Applied Biosystems). All obtained sequences of archaeal 16S rRNA genes were verified for chimera formation by using Ribosomal Database Project Chimera Check (Cole et al., 2003). Nonchimeric sequences were aligned using MEGA 4 software (Tamura et al., 2007). The sequences were grouped into different operational taxonomic units (OTUs) by using the 97% similarity level as the cut-off value. Representative sequences from each OTU were aligned using MEGA 6 software. Based on the alignments of partial archaeal nucleotide sequences of 16S rRNA genes, phylogenetic trees were constructed using the neighbor-joining method with MEGA 4. The closest relative for each OTU sequence was identified through a BLASTn search (Altschul et al., 1997). Phylotype diversity in each clone library was estimated using Simpson’s diversity index (Simpson, 1949), the Chao index (Chao, 1984), and the Shannon index and evenness (Margalef, 1958). Phylotypes enumeration and calculators were conducted on the basis of the aforementioned OTU patterns (at a 97% identity threshold between sequences). The 16S rRNA sequences were aligned using the SILVA (Web) Aligner SINA. Phylogenetic trees were estimated using the maximum likelihood method incorporated in the reference (Ref) SSU datasets as of SILVA release 108. These datasets which was contained only high-quality, nearly full-length sequences suitable for in-depth phylogenetic analysis. The data are available in the supplementary material.
2.5 qPCR analyses of environmental DNA templates using different primer sets
The quantified template DNA was created through PCR by using the archaeal 16S rRNA gene primer A109F/A915R and multi-sample DNA templates from several depths at K2. After purification and quantification, the PCR products were diluted to contain 102–106 16S rRNA gene copies. These amplified PCR productions were used as the quantification standard and employed in the qPCR analysis of Archaea in seawater samples conducted using selected primers. All qPCR amplifications were performed on a 7500 Real-time system (Applied Biosystems) in a 20-μL reaction mixture containing 2 μL of template DNA, 0.15 μM of each primer, and 10 μL of Power SYBR® Green PCR Master Mix (TaKaRa, Japan). For the qPCR analysis of Thaumarchaeota, the standard template DNA was generated through PCR by using correlated same specific primers. The DNA of the quantification standard underwent all qPCR reactions along with the samples. Reliable exponential patterns were yielded through the amplifications, with template amounts ranging from 102 to 106 16S rRNA gene copies. The data were employed to fit standard curves relating cycle threshold values to the amount of template DNA added. Linear plots were created over four orders of magnitude with high correlation coefficients (Supporting Information, Table S1). The melting curve analyses were performed to confirm PCR product specificity following amplification by measuring fluorescence continuously while increasing the temperature from 60 °C to 95 °C. Cycle thresholds were established automatically using 7500 system software (version 1.3). PCR reactions were performed with an initial denaturation step at 95 °C for 4 min, followed by 45 cycles of denaturation at 94 °C for 30 s, primer annealing (annealing temperatures for different primer sets are listed in Supporting Information, Table S1) for 40 s, and extension at 72 °C for 40 s. To ensure stringent product quantification, the fluorescent signal was read after each cycle at 78 °C for 35 s. Each reaction was performed in a white 96-well qPCR plate (Applied Biosystems) capped with an optical ultra-clear seal. The reactions were carried out in triplicate by using dilutions of template DNA from each sample. Three reactions were also performed without DNA to control for contamination.
3 Results
Table 2 presents sampling site locations and physicochemical and biological characteristics of the water samples. The hydrographic data at station K2 were cited from the JAMSTEC website (http://www.jamstec.go.jp).
3.1 Nucleotide sequence accession numbers
In total, 231 16S rRNA gene sequences obtained were submitted to the GenBank public database and assigned accession numbers GU144021 to GU144254.
3.2 Composition of the archaeal community at different depths at station K2
The archaeal community structure was characterized by analyzing 16S rRNA genes in the water samples derived from different depths (epipelagic, 50 m; mesopelagic, 1000 m; and bathypelagic, 4000 m) at station K2 in WSP. In total, of the 231 sequences, 97% belonged to one of the three major clusters of Archaea, including Uncultured MGI Thaumarchaeota, Euryarchaeota MGII, and Euryarchaeota MGIII, accounting for 33%, 43%, and 21% of all archaeal sequences, respectively.
According to the phylogenic analyses of the sequences, obvious distinctions were observed in the archaeal community along a depth gradient (Fig. 1). Euryarchaeota MGII were abundant in the epipelagic layer (77% of the Archaea sequence abundance at 50 m), and the abundance declined with depth. The abundance of MGI Thaumarchaeota sequences was 23% in the epipelagic layer, and the abundance increased with depth, reaching up to 30% and 47% at 1000 and 4000 m, respectively. Our results were consistent with previous results (DeLong et al., 2006; Herndl et al., 2005; Karner et al., 2001). Moreover, no Euryarchaeota MGIII sequences were detected in the pelagic layer, and the abundance of Euryarchaeota MGIII sequences was 30% and 16% of the archaeal sequences, respectively, in mesopelagic and bathypelagic layers.
The depth-stratified archaeal communities were also revealed through phylogenetic analysis by using MEGA 4 software. In total, 24 phylotypes (OTUs, at 97% sequence identity threshold) were identified at all depths through clustering analysis. All of the 8 OTUs detected at the epipelagic layer were unique to the other two deep layers. Of the 8 OTUs, 7 OTUs belonged to Euryarchaeota MGII, and one belonged to MGI Thaumarchaeota. By contrast, the archaeal community in the mesopelagic layer exhibited high phylogenetic similarity to that in the bathypelagic layer by sharing seven overlap OTUs.
Based on their OTU patterns, the archaeal community diversity at the three depths at station K2 was assessed using the Shannon index of diversity, evenness, richness, and the reciprocal of Simpson’s diversity index (Table 3). Among the three depths, Archaea in the mesopelagic waters exhibited the highest phylotype diversity, followed by that in the epipelagic and bathypelagic water masses. At all depths, OTU numbers and the Shannon and Chao1 richness of Euryarchaeota were generally higher than those of Thaumarchaeota, whereas the OTU-based archaeal phylotype richness estimates were higher in the mesopelagic waters than in the epipelagic and bathypelagic waters. Our results are consistent with those of Brown et al. (2009), which investigated archaeal diversity over a depth profile at the Hawaii Ocean Time-Series Station, ALOHA. The results also revealed that the mesopelagic layer had considerably greater archaeal community richness.
3.3 Phylogenetic dissection of the community structure for each archaeal marine group
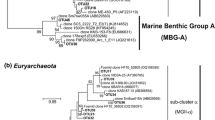
According to the phylogenetic analysis, 4 OTUs composed of 77 archaeal 16S rRNA gene sequences were classified into the MGI Thaumarchaeota group. Two clusters were clearly noted in the phylogenetic tree of MGI Thaumarchaeota 16S rRNA sequences (Fig. 2). One cluster was the ‘ammonia-oxidizing Thaumarchaeota isolate cluster’ (Agogué et al., 2008) composed of OTU3 (20 sequences from the epipelagic layer) and OTU4 (4 sequences from the bathypelagic layer). The other two OTUs (OTU1 and OTU2), composed of 53 sequences from the mesopelagic and bathypelagic layers, were grouped into another cluster. OTU1 contained 51 sequences and was the most abundant crenarchaeal phylotype in our study. OTU1 was phylogenetically close to 1 OTU (A3-75-A2-14), affiliated to a subtropical and equatorial cluster. It was also the most abundant OTU of crenarchaeal 16S rRNA sequences recovered from the North Atlantic Ocean at different stations and depths (Agogué et al., 2008). The sequences retrieved from all four OTUs of MGI Thaumarchaeota in our study exhibited high sequence identity with those reported in various oceans (Agogué et al., 2008; DeLong et al., 2006; Galand et al., 2009). The cosmopolitan phylotypes of MGI Thaumarchaeota were speculated to majorly contribute to the N cycle by participating in chemoautotrophic ammonia oxidation, thereby highlighting the common ecological role of these phylotypes throughout the ocean water. Another Thaumarchaeota phylotype, Crenarchaeota group 1A (DeLong et al., 2006), related to a hot spring clade (pSL12), was reported as abundant below the euphotic zone in the North Pacific Subtropical Gyre (Mincer et al., 2007), North Atlantic Ocean (Agogué et al., 2008), Ionian Sea, and Adriatic Sea (Martin-Cuadrado et al., 2008). However, it was absent in our 16S rRNA sequences. The absence of Crenarchaeota group 1A was also reported in the 500 m-deep Polar Front cosmid library (DeepAnt) and the 1,000-m-deep South Atlantic library (SAT1000) planktonic metagenomic libraries. Furthermore, Galand et al. (2009) reported this group to be present in low abundance in the deep Arctic, as revealed by the massively parallel tag sequencing method. Genes involved in nitrogen fixation were recently found in the Crenarchaeota 1A group, which implied that they may play a role in the ocean’s N cycle (Mincer et al., 2007). The reason for the low abundance or absence of the Crenarchaeota group 1A cluster remains unclear because little is known about environmental factors influencing the distribution of this cluster in different oceanic areas.
Phylogenetic tree of Thaumarchaeota-related 16S rRNA gene partial sequences from the station K2 in the WSP. One representative of OTU is indicated in bold, and the total number of clones is represented by a sequence, while the water depth where they were found is indicated in parentheses. Bar indicates point mutations per nucleotide. Depths: ★, 50 m; ■, 1000 m; ◆, 4000 m
The phylogenetic tree of Euryarchaeota MGII included 102 sequences and was classified into 13 OTUs (Fig. 3). Two-thirds of the sequences were derived from the epipelagic layer and formed into 7 OTUs. A distinct epipelagic cluster consisted of five epipelagic OTUs (OTU11–OTU15) (Fig. 3), and the other two epipelagic branches consisted of 2 OTUs (OTU5 and OTU6). The distribution of mesopelagic or/and bathypelagic OTUs (OTU6–OTU10, OTU16, OTU17) was scattered in the phylogenetic tree. OTU9 (16 sequences) and OTU17 (9 sequences) were constituted by 16S rRNA sequences derived from the mesopelagic and bathypelagic layers. Moreover, five low-abundant OTUs, namely OTU8, OTU10, OTU14, OTU15, and OTU16, which only contained one or two sequences, were found in this group. Only very few sequences in GenBank were similar to those of these low-abundant OTUs, except OTU8, which has six similar sequences in GenBank at 97% identity threshold. Additionally, OTU16 was novel and formed a relatively separate branch with a low (91%) identity with published sequences. In the present study, phylogenetically, Euryarchaeota MGII exhibited high diversity at station K2 and greatly differed from that at ALOHA and in other oceanic areas. It attributed to some disparate ‘Novel’ OTUs accounting for 42% of all Euryarchaeota MGII clones (43 of the 102 sequences) (Fig. 3). The 16S rRNA sequences of Euryarchaeota MGII were more diverse than those of MGI Thaumarchaeota in WSP, which is consistent with previous results (Bano et al., 2004; Massana et al., 2000). Although Euryarchaeota MGII plays a limited role in marine microbial ecology, Frigaard et al. (2006) revealed some proteorhodopsin gene-containing Euryarchaeota MGII in the ocean’s upper water column. The abundance of Euryarchaeota 16S rRNA sequences was approximately 10% in the photic zone. However, no 16S rRNA gene related to the proteorhodopsin gene-containing Euryarchaeote MGII group was detected in our samples.
Phylogenetic tree of Euryarchaeota marine group II-related 16S rRNA gene partial sequences from station K2. One representative of OTU is indicated in bold, and the total number of clones is represented by a sequence, while the water depth where they were found is indicated in parentheses. Bar indicates point mutations per nucleotide. Depths: ★, 50 m; ■, 1000 m; ◆, 4000 m
In total, 5 OTUs (Fig. 4) were affiliated with Euryarchaeota MGIII and were composed of the three clusters. One cluster comprised three OTUs (OTU18, OTU20, and OTU21) that composed the OTU D cluster (Galand et al., 2009; Martin-Cuadrado et al., 2008). Each of the other two clusters, OTU22 and OTU19, consisted of a single branch. OTU22 exhibited the highest similarity (98%) to the representative sequence of an MGIII cluster, ‘DH148-W24’ (Galand et al., 2009). By contrast, the representative OTU19 sequence was most closely related (95% sequence identity) to the fosmid 16S rRNA gene sequence HF130_43E12, which was recovered from 130-m depth in the eastern North Pacific Ocean (DeLong et al., 2006). Most Euryarchaeota MGIII sequences were retrieved from the mesopelagic or bathypelagic samples and have rarely been found in the coastal water or epipelagic layer (Bano et al., 2004; Galand et al., 2006, 2009). Based on some investigations on the archaeal community in the eastern Mediterranean Sea, Moeseneder et al. (2001) reported on the abundance of Euryarchaeota MGIII (12%) at 10-m seawater. Gland and colleagues (Galand et al., 2009) found that Euryarchaeota MGIII was more abundant in deep-water masses and represented the largest archaeal group in the deep Atlantic layer of the central Arctic Ocean. Our results concur with the findings of Galand et al. (2009) because we also found Euryarchaeota MGIII to be abundant in the mesopelagic and bathypelagic layers at station K2 in WSP. Brown et al. (2009) retrieved Euryarchaeota MGIII from North Pacific Ocean depths and reported on their low abundance throughout the water column at ALOHA.
Phylogenetic tree of Euryarchaeota marine group III-related sequences and other sequences unaffiliated with any of the three main Archaea groups, based on 16S rRNA gene partial sequences from station K2. The numbers at the nodes are bootstrap values out of 1000 replications. One representative OTU is indicated in bold, and the total number of clones is represented by a sequence, while the water depth where they were found is depicted in parentheses. Closely related database and cultivated Archaea sequences (with sequence ID numbers in parentheses) are shown in plain font. The bar indicates 0.01 fixed point mutations per nucleotide. Depths: ★, 50 m; ■, 1000 m; ◆, 4000 m
In addition, 2 OTUs (OTU23, 2 sequences; OTU24, 4 sequences) were not phylogenetically affiliated with the three uncultured Archaea groups. OTU23 comprised an exceptional branch with few closely related sequences in the 16S rRNA gene database. It showed 96% similarity with only one uncultured archaeon, PaswB60, which was observed in hydrothermal areas in the southern Mariana trough. OTU24 was closest to a cluster of the culturable Euryarchaeota Methanosarcina mazei. A similar result (Brown et al., 2009) was observed in the low abundance of methanogen sequences in the mesopelagic (800 m) and bathypelagic (4,400 m) layers in the North Pacific Ocean.
3.4 Archaeal distribution in WSP determined through qPCR analysis
The vertical distribution of Archaea and Thaumarchaeota in WSP was investigated based on the relevant specific 16S rRNA gene copy numbers quantified through qPCR by using seawater samples from different layers at station K2. The results (Table 4) revealed that the Archaea and Thaumarchaeota 16S rRNA gene copy numbers on the surface were 7.6 × 103 and 15 copies mL−1, respectively. The abundance of Archaea increased with depth, and the maximum Archaea 16S rRNA gene copy number at 50 m depth was 1.05 × 105 copies mL−1. This number decreased with depth, and the Archaea 16S rRNA gene copy number dropped to 0.9 × 103 copies mL−1 at 4000 m. A slight rise in copy number (1.4 × 103 copies mL−1) was found at 5000 m. The variation trend of MGI Thaumarchaeota was slightly different from that of Archaea at a depth below 50 m. The maximal Thaumarchaeota 16S rRNA gene copy number (3.2 × 103 copies mL−1) was observed at 500 m, and then, the gene copy number slowly decreased to 2.1 × 103 copies m L−1 at 1000 m. The gene copy number decreased to 0.73 × 103 copies mL−1 at 4,000 m and then slightly increased to 0.91 × 103 copies mL−1 at 5,000 m.
The contribution of Archaea to the total microbial cell number was the highest at 50 m (26.6%) and decreased to 20.7% at 500 m and 12.7% at 1,000 m. At 4,000 and 5,000 m depth, the contribution of Archaea to the total microbial cell number was 18.0% and 12.6%, respectively. The variation trend of relative contributions of MGI Thaumarchaeota to the total microbial cell number was similar to that of Archaea to the total microbial cell number despite their abundance being lower in both the surface and epipelagic layers. However, the contribution of MGI Thaumarchaeota to the total microbial cell count increased quickly with depth, reaching 8.1% at 500 m, peaking at 11.8% at 1,000 m, and then decreasing slightly to approximately 7% relative to the total microbial cell number in the bathypelagic water. The relative abundance of crenarchaeal 16S rRNA copy numbers to that of total archaeal 16S rRNA copy numbers was very low at the surface, whereas it increased sharply with depth, reaching up to 30% at 500 m and up to 89% at 1,000 m, and then declining from 1,000 m to 5,000 m, but still maintaining a high value.
4 Discussion
Studies on the archaeal community structure and distribution in different ocean regions have developed rapidly (Agogué et al., 2008; Brown et al., 2009; DeLong et al., 2006; Galand et al., 2009; Hu et al., 2011a, 2011b; Ionescu et al., 2009; Karl et al., 2001; Massana et al., 1997, 2000; Mincer et al., 2007). Depth-related change in the archaeal community was noted in different oceanic areas (Bano et al., 2004; DeLong et al., 2006; Massana et al., 1997, 2000). Euryarchaeota was dominant in the photic zone, and Thaumarchaeota was dominant in deep water (Bano et al., 2004). Consistent with previous works (Bano et al., 2004; DeLong et al., 2006; Massana et al., 1997, 2000), a distinct partitioning of the marine water column was found between Thaumarchaeota and Euryarchaeota in the present study, based on the relative abundance of their sequences in the 16S rRNA gene clone libraries derived from different layers in WSP. Furthermore, the depth-related separation of the ‘archaeal community structure’ was observed between the different phylotypes of certain archaeal groups. For example, the Thaumarchaeota phylotype (OTU3) occurred only in the epipelagic layer, whereas the different Thaumarchaeota phylotypes (OTU1, OTU2) dominated the mesopelagic and bathypelagic layers. The distinct depth-related distribution of Thaumarchaeota phylotypes has also been reported in other studies (Agogue et al., 2008; Hu et al., 2011a, 2011b; Massana et al., 1997, 2000). Some studies (Hu et al., 2011a, 2011b) have suggested that niche partitioning may occur between the shallow and deep marine Thaumarchaeota phylotypes. Moreover, qPCR analyses indicated that Thaumarchaeota amoA (Agogué et al., 2008; Beman et al., 2008; Francis et al., 2005; Hu et al., 2011a, b; Mincer et al., 2007; Yakimov et al., 2011) and accA (Hu et al., 2011a, b) gene abundances varied similarly with Thaumarchaeota 16S rRNA gene abundances in the epipelagic, mesopelagic, and bathypelagic zones. The segregation of the phylogenetic composition of Thaumarchaeota assemblages between the epipelagic water and mesopelagic or bathypelagic waters implies the different explicit functional and ecological roles.
Quantitative real-time PCR (qPCR) has been widely applied to investigate Thaumarchaeota abundance in various environments. The Thaumarchaeota-specific 16S rRNA gene primer pair MCGI391F/MCGI554R was commonly used in previous studies (Auguet et al., 2011; Coolen et al., 2004; De Corte et al., 2009; Galand et al., 2009; Herfort et al., 2007; Pitcher et al., 2011; Pouliot et al., 2009; Takai et al., 2000; Wuchter et al., 2006) for quantifying the abundance of Thaumarchaeota. When we determined the specificity of the primers for Thaumarchaeota in the Silva 16S rRNA database (Pruesse et al., 2007), we found that the forward primer MCGI391F was not sufficient to cover all MGI Thaumarchaeota 16S rRNA sequences deposited in the database, especially the primer MCGI391F mismatched with the target 16S RNA sequences of MGI Thaumarchaeota. Most of these mismatched sequences were phylogenetically close to the representative of the phylotype OTU3, which was derived from the epipelagic seawater in the present study. Bias possibly occurred in qPCR using the primer MCGI391F, especially when the samples were collected from the surface or upper water column, where the ‘shallow layer’ phylotype OTU3 dominated. For testifying this assumption, a new qPCR primer set Arc519F/Cren770R was used here to quantify the abundance of Thaumarchaeota at different depths of seawater by using the same samples as used for the primer pair MCGI391F/MCGI554R. Cren770R was a newly developed reverse primer that has high coverage and is specific to most MGI Thaumarchaeota Expectedly, the qPCR results with the new primer set Arc519F/Cren770R revealed that the abundance of Thaumarchaeota was higher than that obtained with the primer set MCGI391F/MCGI554R in the upper water samples (< 500 m).
Although the primer MCGI391F only mismatched with some sequences of MGI Thaumarchaeota by one base at terminal 3′, Simsek and Adnan (2000) reported that a single mismatch at or near the terminal 3′ base of a primer affects PCR results more dramatically. The primer MCGI391F was first designed by Takai et al. (2004). It was complementary to the conserved regions of Thaumarchaeota MGI 16S rRNA sequences presented in the original database, which contained a small number of sequences. However, it was not found to be complementary to 701 sequences in the Silva 108 ‘SSU REF’ database released (Pruesse et al., 2007).
In the present study, the vertical distribution of Archaea was investigated through qPCR. The results revealed a lower abundance of Archaea in WSP than in other oceanic areas, which were determined through the FISH or CARD-FISH method (Herfort et al., 2007; Herndl et al., 2005; Karner et al., 2001; Massana et al., 1997). As determined through qPCR, the 16S rRNA gene copy number of Thaumarchaeota was very low level in the surface of WSP. In general, Archaea and Thaumarchaeota exhibited similar variation trends with depth despite their low contribution (Fig. 5) to the microbial abundance in bathypelagic waters (1000–5000 m depth). Moreover, the maximum copy number and highest abundance of Thaumarchaeota 16S sequences were noted at 500 and 1000 m at station K2, where a pronounced oxygen minimum zone also existed (Fig. 5). This result is also consistent with those of other studies conducted in the Arabian Sea, the Black Sea, the subtropical North Atlantic, and the upwelling area (Coolen et al., 2007; Francis et al., 2005; Lam et al., 2007; Sinninghe Damsté et al., 2002; Varela et al., 2008). In contrast, the low contribution of Thaumarchaeota to the total microbial cell number and their high contribution to the total Archaea (> 80%) suggested that some bias (e.g., caused by the sampling method and low DNA recovery from the samples) in qPCR accounts for this discrepancy.
Although the current investigation provides an interesting insight into the distribution of archaeal diversity in WSP, some microbes escaped the QM-A filter and were later detected in the 0.2-µm polycarbonate filter. However, the numbers were very low and accounted for only 1%–7% (data not shown) of the total Archaea count after the samples collected from the same sea area were reanalyzed using the 0.2-µm polycarbonate filter and QM-A filter in 2009. Further analysis on this topic is underway.
Availability of data and materials
All data generated or analysed during this study are included in this published article, and the reference 16S rRNA gene dataset used to constructed the phylogenetic tree was prepared by selecting those sequences from the Ref 108 and is available for download on the SILVA website.
References
Agogué, H., Brink, M., Dinasquet, J., & Herndl, G. J. (2008). Major gradients in putatively nitrifying and non-nitrifying Archaea in the deep North Atlantic. Nature, 456(7223), 788–791. https://doi.org/10.1038/nature07535
Altschul, S. F., Madden, T. L., Schäffer, A. A., Zhang, J., Zhang, Z., Miller, W., et al. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Research, 25(17), 3389–3402. https://doi.org/10.1093/nar/25.17.3389
Auguet, J. C., Nomokonova, N., Camarero, L., & Casamayor, E. O. (2011). Seasonal changes of freshwater ammonia-oxidizing archaeal assemblages and nitrogen species in oligotrophic alpine lakes. Applied and Environmental Microbiology, 77(6), 1937–1945. https://doi.org/10.1128/AEM.01213-10
Baker, B. J., De Anda, V., Seitz, K. W., Dombrowski, N., Santoro, A. E., & Lloyd, K. G. (2020). Diversity, ecology and evolution of Archaea. Nature Microbiology, 5(7), 887–900. https://doi.org/10.1038/s41564-020-0715-z
Bano, N., Ruffin, S., Ransom, B., & Hollibaugh, J. T. (2004). Phylogenetic composition of Arctic Ocean archaeal assemblages and comparison with Antarctic assemblages. Applied and Environmental Microbiology, 70(2), 781–789. https://doi.org/10.1128/AEM.70.2.781-789.2004
Beman, J. M., Popp, B. N., & Francis, C. A. (2008). Molecular and biogeochemical evidence for ammonia oxidation by marine Crenarchaeota in the Gulf of California. The ISME Journal, 2(4), 429–441. https://doi.org/10.1038/ismej.2007.118
Brown, M. V., Philip, G. K., Bunge, J. A., Smith, M. C., Bissett, A., Lauro, F. M., et al. (2009). Microbial community structure in the North Pacific ocean. The ISME Journal, 3(12), 1374–1386. https://doi.org/10.1038/ismej.2009.86
Chao, A. (1984). Nonparametric estimation of the number of classes in a population. Scandinavian Journal of Statistics, 11, 265–270.
Church, M. J., DeLong, E. F., Ducklow, H. W., Karner, M. B., Preston, C. M., & Karl, D. M. (2003). Abundance and distribution of planktonic Archaea and Bacteria in the waters west of the Antarctic Peninsula. Limnology and Oceanography, 48(5), 1893–1902. https://doi.org/10.4319/lo.2003.48.5.1893
Cole, J. R., Chai, B., Marsh, T. L., Farris, R. J., Wang, Q., Kulam, S. A., et al. (2003). The Ribosomal Database Project (RDP-II): Previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Research, 31(1), 442–443. https://doi.org/10.1093/nar/gkg039
Coolen, M. J., Hopmans, E., Rijpstra, W. I., Muyzer, G., Schouten, S., Volkman, J. K., et al. (2004). Evolution of the methane cycle in Ace Lake (Antarctica) during the Holocene: Response of methanogens and methanotrophs to environmental change. Organic Geochemistry, 35, 1151–1167.
Coolen, M. J., Abbas, B., van Bleijswijk, J., Hopmans, E. C., Kuypers, M. M., Wakeham, S. G., et al. (2007). Putative ammonia-oxidizing Crenarchaeota in suboxic waters of the Black Sea: A basin-wide ecological study using 16S ribosomal and functional genes and membrane lipids. Environmental Microbiology, 9(4), 1001–1016. https://doi.org/10.1111/j.1462-2920.2006.01227.x
De Corte, D., Yokokawa, T., Varela, M. M., Agogué, H., & Herndl, G. J. (2009). Spatial distribution of Bacteria and Archaea and amoA gene copy numbers throughout the water column of the Eastern Mediterranean Sea. The ISME Journal, 3(2), 147–158. https://doi.org/10.1038/ismej.2008.94
DeLong, E. F. (1992). Archaea in coastal marine environments. Proceedings of the National Academy of Sciences of the United States of America, 89(12), 5685–5689. https://doi.org/10.1073/pnas.89.12.5685
DeLong, E. F., Preston, C. M., Mincer, T., Rich, V., Hallam, S. J., Frigaard, N. U., et al. (2006). Community genomics among stratified microbial assemblages in the ocean’s interior. Science (New York, N.Y.), 311(5760), 496–503. https://doi.org/10.1126/science.1120250
Francis, C. A., Roberts, K. J., Beman, J. M., Santoro, A. E., & Oakley, B. B. (2005). Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. Proceedings of the National Academy of Sciences of the United States of America, 102(41), 14683–14688. https://doi.org/10.1073/pnas.0506625102
Frigaard, N. U., Martinez, A., Mincer, T. J., & DeLong, E. F. (2006). Proteorhodopsin lateral gene transfer between marine planktonic Bacteria and Archaea. Nature, 439(7078), 847–850. https://doi.org/10.1038/nature04435
Fuhrman, J. A., McCallum, K., & Davis, A. A. (1992). Novel major archaebacterial group from marine plankton. Nature, 356(6365), 148–149. https://doi.org/10.1038/356148a0
Galand, P. E., Lovejoy, C., & Vincent, W. F. (2006). Remarkably diverse and contrasting archaeal communities in a large arctic river and the coastal Arctic Ocean. Aquatic Microbial Ecology, 44, 115–126.
Galand, P. E., Casamayor, E. O., Kirchman, D. L., Potvin, M., & Lovejoy, C. (2009). Unique archaeal assemblages in the Arctic Ocean unveiled by massively parallel tag sequencing. The ISME Journal, 3(7), 860–869. https://doi.org/10.1038/ismej.2009.23
Herfort, L., Schouten, S., Abbas, B., Veldhuis, M. J., Coolen, M. J., Wuchter, C., et al. (2007). Variations in spatial and temporal distribution of Archaea in the North Sea in relation to environmental variables. FEMS Microbiology Ecology, 62(3), 242–257. https://doi.org/10.1111/j.1574-6941.2007.00397.x
Herndl, G. J., Reinthaler, T., Teira, E., van Aken, H., Veth, C., Pernthaler, A., et al. (2005). Contribution of Archaea to total prokaryotic production in the deep Atlantic Ocean. Applied and Environmental Microbiology, 71(5), 2303–2309. https://doi.org/10.1128/AEM.71.5.2303-2309.2005
Hobbie, J. E., Daley, R. J., & Jasper, S. (1977). Use of nuclepore filters for counting bacteria by fluorescence microscopy. Applied and Environmental Microbiology, 33(5), 1225–1228. https://doi.org/10.1128/aem.33.5.1225-1228.1977
Hu, A., Jiao, N., & Zhang, C. L. (2011a). Community structure and function of planktonic Crenarchaeota: Changes with depth in the South China Sea. Microbial Ecology, 62(3), 549–563. https://doi.org/10.1007/s00248-011-9866-z
Hu, A., Jiao, N., Zhang, R., & Yang, Z. (2011b). Niche partitioning of marine group I Crenarchaeota in the euphotic and upper mesopelagic zones of the East China Sea. Applied and Environmental Microbiology, 77(21), 7469–7478. https://doi.org/10.1128/AEM.00294-11
Ionescu, D., Penno, S., Haimovich, M., Rihtman, B., Goodwin, A., Schwartz, D., et al. (2009). Archaea in the Gulf of Aqaba. FEMS Microbiology Ecology, 69(3), 425–438. https://doi.org/10.1111/j.1574-6941.2009.00721.x
Karner, M. B., DeLong, E. F., & Karl, D. M. (2001). Archaeal dominance in the mesopelagic zone of the Pacific Ocean. Nature, 409(6819), 507–510. https://doi.org/10.1038/35054051
Kirchman, D. L., Elifantz, H., Dittel, A. I., Malmstrom, R. R., & Cottrell, M. T. (2007). Standing stocks and activity of Archaea and bacteria in the western Arctic ocean. Limnology and Oceanography, 52(2), 495–507. https://doi.org/10.4319/lo.2007.52.2.0495
Komuro, C., Narita, H., Imai, K., Nojiri, Y., & Jordan, R. W. (2005). Microplankton assemblages at Station KNOT in the subarctic western Pacific, 1999–2000. Deep-Sea Research Part Ii-Topical Studies in Oceanography, 52, 2206–2217.
Lam, P., Jensen, M. M., Lavik, G., McGinnis, D. F., Müller, B., Schubert, C. J., et al. (2007). Linking crenarchaeal and bacterial nitrification to anammox in the Black Sea. Proceedings of the National Academy of Sciences of the United States of America, 104(17), 7104–7109. https://doi.org/10.1073/pnas.0611081104
Lane, D. J. (1991). 16S/23S rRNA sequencing. In E. Stackebrandt & M. Goodfellow (Eds.), Nucleic Acid Techniques in Bacterial Systematics (pp. 115–148). Wiley.
Liu, H., Imai, K., Suzuki, K., Nojiri, Y., Tsurushima, N., & Saino, T. (2002). Seasonal variability of picophytoplankton and bacteria in the western subarctic Pacific Ocean at station KNOT. Deep-Sea Research Part Ii-Topical Studies in Oceanography, 49, 5409–5420. https://doi.org/10.1016/S0967-0645(02)00199-6
Longhurst, A. R., Sathyendranath, S., Platt, T., & Caverhill, C. (1995). An estimate of global primary production in the ocean from satellite radiometer data. Journal of Plankton Research, 17, 1245–1271.
Margalef, R. (1958). Information theory in ecology. International Journal of General Systems, 3, 36–71.
Martin-Cuadrado, A. B., Rodriguez-Valera, F., Moreira, D., Alba, J. C., Ivars-Martínez, E., Henn, M. R., Talla, E., et al. (2008). Hindsight in the relative abundance, metabolic potential and genome dynamics of uncultivated marine archaea from comparative metagenomic analyses of bathypelagic plankton of different oceanic regions. The ISME Journal, 2(8), 865–886. https://doi.org/10.1038/ismej.2008.40
Massana, R., Murray, A. E., Preston, C. M., & DeLong, E. F. (1997). Vertical distribution and phylogenetic characterization of marine planktonic Archaea in the Santa Barbara Channel. Applied and Environmental Microbiology, 63(1), 50–56. https://doi.org/10.1128/aem.63.1.50-56.1997
Massana, R., DeLong, E. F., & Pedrós-Alió, C. (2000). A few cosmopolitan phylotypes dominate planktonic archaeal assemblages in widely different oceanic provinces. Applied and Environmental Microbiology, 66(5), 1777–1787. https://doi.org/10.1128/AEM.66.5.1777-1787.2000
Mincer, T. J., Church, M. J., Taylor, L. T., Preston, C., Karl, D. M., & DeLong, E. F. (2007). Quantitative distribution of presumptive archaeal and bacterial nitrifiers in Monterey Bay and the North Pacific Subtropical Gyre. Environmental Microbiology, 9, 1162–1175. https://doi.org/10.1111/j.1462-2920.2007.01239.x
Moeseneder, M. M., Winter, C., Arrieta, J. M., & Herndl, G. J. (2001). Terminal-restriction fragment length polymorphism (T-RFLP) screening of a marine archaeal clone library to determine the different phylotypes. Journal of Microbiological Methods, 44(2), 159–172. https://doi.org/10.1016/s0167-7012(00)00247-5
Ovreås, L., Forney, L., Daae, F. L., & Torsvik, V. (1997). Distribution of bacterioplankton in meromictic Lake Saelenvannet, as determined by denaturing gradient gel electrophoresis of PCR-amplified gene fragments coding for 16S rRNA. Applied and Environmental Microbiology, 63(9), 3367–3373. https://doi.org/10.1128/aem.63.9.3367-3373.1997
Pitcher, A., Villanueva, L., Hopmans, E. C., Schouten, S., Reichart, G. J., & Sinninghe Damsté, J. S. (2011). Niche segregation of ammonia-oxidizing archaea and anammox bacteria in the Arabian Sea oxygen minimum zone. The ISME Journal, 5(12), 1896–1904. https://doi.org/10.1038/ismej.2011.60
Pouliot, J., Galand, P. E., Lovejoy, C., & Vincent, W. F. (2009). Vertical structure of archaeal communities and the distribution of ammonia monooxygenase A gene variants in two meromictic High Arctic lakes. Environmental Microbiology, 11(3), 687–699. https://doi.org/10.1111/j.1462-2920.2008.01846.x
Pruesse, E., Quast, C., Knittel, K., Fuchs, B. M., Ludwig, W., Peplies, J., et al. (2007). SILVA: A comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Research, 35(21), 7188–7196. https://doi.org/10.1093/nar/gkm864
Rinke, C., Rubino, F., Messer, L. F., Youssef, N., Parks, D. H., Chuvochina, M., et al. (2019). A phylogenomic and ecological analysis of the globally abundant Marine Group II archaea (Ca. Poseidoniales ord. nov.). The ISME Journal, 13(3), 663–675. https://doi.org/10.1038/s41396-018-0282-y
Sambrotto, R. N., Goering, J. J., & McRoy, C. P. (1984). Large yearly production of phytoplankton in the Western bering strait. Science (New York, N.Y.), 225(4667), 1147–1150. https://doi.org/10.1126/science.225.4667.1147
Simpson, E. (1949). Measurement of Diversity. Nature, 163, 688. https://doi.org/10.1038/163688a0
Simsek, M., & Adnan, H. (2000). Effect of single mismatches at 3’-end of primers on polymerase chain reaction. Journal for Scientific Research. Medical Sciences, 2(1), 11–14.
Sinninghe Damsté, J. S., Rijpstra, W. I., Hopmans, E. C., Prahl, F. G., Wakeham, S. G., & Schouten, S. (2002). Distribution of membrane lipids of planktonic Crenarchaeota in the Arabian Sea. Applied and Environmental Microbiology, 68(6), 2997–3002. https://doi.org/10.1128/AEM.68.6.2997-3002.2002
Stahl, D. A., & Amann, R. (1991). Development and application of nucleic acid probes in bacterial systematic. In E. Stackebrandt & M. Goodfellow (Eds.), Nucleic Acid Techniques in Bacterial Systematics (pp. 205–248). John Wiley and Sons.
Suzuki, K., Minami, C., Liu, H., & Saino, T. (2002). Temporal and spatial patterns of chemotaxonomic algal pigments in the subarctic Pacific and the Bering Sea during the early summer of 1999. Deep-Sea Research Part Ii-Topical Studies in Oceanography, 49, 5685–5704.
Takai, K., & Horikoshi, K. (2000). Rapid detection and quantification of members of the archaeal community by quantitative PCR using fluorogenic probes. Applied and Environmental Microbiology, 66(11), 5066–5072. https://doi.org/10.1128/AEM.66.11.5066-5072.2000
Takai, K., Oida, H., Suzuki, Y., Hirayama, H., Nakagawa, S., Nunoura, T., et al. (2004). Spatial distribution of marine crenarchaeota group I in the vicinity of deep-sea hydrothermal systems. Applied and Environmental Microbiology, 70(4), 2404–2413. https://doi.org/10.1128/AEM.70.4.2404-2413.2004
Tamburini, C., Garel, M., Ali, B. A., Mérigot, B., Kriwy, P., Charrière, B., et al. (2009). Distribution and activity of Bacteria and Archaea in the different water masses of the Tyrrhenian Sea. Deep-Sea Research Part Ii-Topical Studies in Oceanography, 56, 700–712.
Tamura, K., Dudley, J., Nei, M., & Kumar, S. (2007). MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Molecular Biology and Evolution, 24(8), 1596–1599. https://doi.org/10.1093/molbev/msm092
Teira, E., Reinthaler, T., Pernthaler, A., Pernthaler, J., & Herndl, G. J. (2004). Combining catalyzed reporter deposition-fluorescence in situ hybridization and microautoradiography to detect substrate utilization by bacteria and Archaea in the deep ocean. Applied and Environmental Microbiology, 70(7), 4411–4414. https://doi.org/10.1128/AEM.70.7.4411-4414.2004
Utsumi, M., Belova, S. E., King, G. M., & Uchiyama, H. (2003). Phylogenetic comparison of methanogen diversity in different wetland soils. The Journal of General and Applied Microbiology, 49(2), 75–83. https://doi.org/10.2323/jgam.49.75
Varela, M. M., van Aken, H. M., Sintes, E., & Herndl, G. J. (2008). Latitudinal trends of Crenarchaeota and Bacteria in the meso- and bathypelagic water masses of the Eastern North Atlantic. Environmental Microbiology, 10(1), 110–124. https://doi.org/10.1111/j.1462-2920.2007.01437.x
West, N. J., & Scanlan, D. J. (1999). Niche-partitioning of Prochlorococcus populations in a stratified water column in the eastern North Atlantic Ocean. Applied and Environmental Microbiology, 65(6), 2585–2591. https://doi.org/10.1128/AEM.65.6.2585-2591.1999
Wuchter, C., Abbas, B., Coolen, M. J., Herfort, L., van Bleijswijk, J., Timmers, P., et al. (2006). Archaeal nitrification in the ocean. Proceedings of the National Academy of Sciences of the United States of America, 103(33), 12317–12322. https://doi.org/10.1073/pnas.0600756103
Yakimov, M. M., Cono, V. L., Smedile, F., DeLuca, T. H., Juárez, S., Ciordia, S., et al. (2011). Contribution of crenarchaeal autotrophic ammonia oxidizers to the dark primary production in Tyrrhenian deep waters (Central Mediterranean Sea). The ISME Journal, 5(6), 945–961. https://doi.org/10.1038/ismej.2010.197
Acknowledgements
We thank all personnel of the R/V Mirai for essential sample collection support in summer cruise MR-0705.
Funding
No funding was received for conducting this study.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by Gang Chen, Takaaki Satake, Norio Sugiura and Motoo Utsumi. The first draft of the manuscript was written by Gang Chen and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, G., Satake, T., Sugiura, N. et al. Vertical distribution and phylogenetic characterization of marine Archaea in the western subarctic Pacific. Mar Dev 2, 5 (2024). https://doi.org/10.1007/s44312-024-00017-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44312-024-00017-2