Abstract
Abiotic stresses are major factors constraining the growth, development and productivity of tomato (Solanum lycopersicum), the most cultivated vegetable crop worldwide. Uridine diphosphate glycosyltransferases (UDPGTs or UGTs) are essential enzymes that utilize 5-uridine diphosphate as a glycosyl donor molecule to facilitate the catalysis of glycosylation reactions across diverse substrates, thereby playing a pivotal role in conferring abiotic stress tolerance. Currently, there is a limited understanding of the structure and functions of the UDPGT gene family in tomato. In this work, 106 members of the SlUDPGT gene family were identified through in silico analysis, besides, their protein sequence properties, phylogenetic relationships, gene structure, chromosomal distribution, cis-acting elements, tissue expression and hormone- and stress-induced expression were comprehensively investigated. The expression of representative SlUDPGTs under abiotic stress and exogenous hormone treatments, including salt, polyethylene glycol, methyl viologen, gibberellic acid, jasmonic acid, abscisic acid and brassinolide, was investigated through qRT‒PCR analysis. Numerous cis-acting elements linked to stress and hormone signaling were present in the promoter regions of SlUDPGTs. According to microarray data, most SlUDPGT genes were responsive to hormones and abiotic stresses, while certain SlUDPGTs were specifically differentially expressed under Botrytis cinerea and tomato spotted wilt virus infection. Additionally, diverse expression profiles of SlUDPGTs were observed in various tissues and developmental stages. Furthermore, CRISPR/Cas9-mediated knockout of SlUDPGT52 led to enhanced drought tolerance due to enhanced reactive oxygen species (ROS) scavenging. These findings lay the foundations for the future functional characterization of specific UDPGT gene family members, assisting the biotechnology-mediated improvement of tomato and other horticultural crops.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glycosylation is a vital biochemical process that involves the modification of various receptor molecules within and on the surface of the cells, thereby facilitating the maintenance of cellular homeostasis and the execution of specific functions. Glycosylation involves the attachment of one or more carbohydrate units to the target substrate. Protein glycosylation can modify their physicochemical properties, regulate the catalytic activity of enzymes, hormone signaling and fine-tune immune responses and other processes (Lis and Sharon 1993). Regarding lipids, glycolipids are vital to the membrane and viral and bacterial receptors. They can interact with lectins, toxins, hormones and other biological response modifiers (Curatolo 1987).
Glycosylation is catalyzed by glycosyltransferases (GTs) that are widely present in humans, animals, plants and microorganisms. According to the CAZY database (Cantarel et al. 2009), GTs are classified into 114 categories based on their catalytic properties and target substrates, amino acid sequence similarity and other factors. GT-1 is the most prominent family among GTs, whose members use 5-uridine diphosphate as a donor molecule to catalyze glycosylation by transferring glycosyl groups from activated donor molecules to receptor molecules (Yonekura-Sakakibara and Hanada 2011). Therefore, it is also referred to as the uridine diphosphate glycosyltransferase (UDPGT or UGT, UDP-glycosyltransferase) family. While the UDPGT sequences are less conserved, their C-terminal region features a conserved motif known as plant secondary product glycosyltransferase (PSPG) (Mackenzie et al. 1997). The PSPG motif consists of 44 amino acids (aa), a conserved region among all plant UDPGTs (Vogt and Jones 2000). The binding of the UDP moiety of the nucleotide sugar is thought to occur in this region (Mackenzie et al. 1997). The N-terminus of the UDPGT protein is highly diversified, considered to be the driver for the wide range of UDPGT substrate specificities (Wang 2009; Lairson et al. 2008). Thus far, the crystal structure of UDPGTs has been obtained in various species, such as in humans (Fujiwara et al. 2016) and Medicago truncatula (Modolo et al. 2009; Shao et al. 2005). Although their sequence similarity is relatively low, all UDPGTs have two β/α/β Rossmann-like domains, which comprise a GT-B fold (Yonekura-Sakakibara and Hanada 2011).
It has been proposed that UDPGTs are insignificantly associated with plant responses to abiotic stresses. Overexpression of UGT79B2/B3 in Arabidopsis enhanced cold, drought and salt tolerance by regulating anthocyanin accumulation, while ugt79b2/b3 double mutants were more sensitive to adverse environmental conditions (Li et al. 2017b). Ectopic expression of UGT76E11 in Arabidopsis increased flavonoid accumulation and further enhanced abiotic tolerance by upregulating stress-associated genes (Li et al. 2018b). Moreover, Arabidopsis UGT87A2 was shown to be induced by ABA, drought and salt (Li et al. 2017a), and UGT87E7, a salicylic acid carboxyl glucosyltransferase, regulated disease resistance in Camellia sinensis (Hu et al. 2022). In tomato, UDPGTs can catalyze the glycosylation of ABA and regulate the dynamic equilibrium of ABA levels. SlUGT75C1-RNAi lines exhibited improved drought tolerance and accelerated fruit ripening as a result of increased ABA levels and the earlier induction of ethylene release (Sun et al. 2017). SlUGT5 was highly expressed in the flowers and the ripening fruit, and the recombinant SlUGT5 protein influenced the activity of guaiacol and eugenol, benzyl alcohol, and methyl salicylate (Louveau et al. 2011).
Tomato is one of the most widely consumed vegetables, with an annual global production of 189 million tons (Food and Agriculture Organization of the United Nations, 2021). Moreover, its fresh produce and processed products have a high economic value. Tomato significance extends to being a crucial model organism for scientific research due to its exceptional genetic characteristics. Although genome-wide identification and expression analysis of the UDPGT gene family has been performed in other plant species, such as soybean (Mamoon Rehman et al. 2016), its structural and functional properties remain unexplored in Solanum lycopersicum. In this study, we identified a total of 106 SlUDPGTs using bioinformatics approaches. We comprehensively analyzed their structure and function, including their physicochemical properties, subcellular localization, exon-intron structure, protein tertiary structures, phylogenetic relationships, gene duplication events, chromosome distribution patterns and the presence of cis-acting elements. Additionally, the expression profiles of SlUDPGTs were investigated under various stress conditions and hormone treatments. Selected representative genes were further validated through qPCR analysis. SlUDPGT52 was shown to negatively regulate drought tolerance by enhancing reactive oxygen species (ROS) scavenging. This comprehensive investigation provides a thorough perspective regarding the functional characteristics and structural attributes of SlUDPGTs, establishing a solid foundation for future studies to unravel their biological significance.
Materials and methods
Plant materials
Tomato (S. lycopersicum cv. AC) wild-type (WT) plants were cultivated in a greenhouse under an 8 h dark/16 h light photoperiod. Six-leaf-stage tomato WT seedlings were subjected to hormone and stress treatments. Tomato leaves were sprayed with 100 µM Gibberellic acid (GA), 100 µM Methyl Jasmonate (MeJA), 100 µM Abscisic Acid(ABA), 1 µM Brassinolide (BR), and water until uniform coverage was achieved. Soil concentrations of 100 mM NaCl, 100 mM polyethylene glycol (PEG), and 100 µM methyl viologen (MV) were used for abiotic stress treatments (Li et al. 2018a), while deionized water water treatment was used as the control. To avoid the effects of the circadian clock on gene expression differences, untreated plants were used as the control. After 0, 0.25, 0.5, 1, 2, 6, 12 and 24 h of treatment, leaves were immediately sampled from the plants, placed and frozen in liquid nitrogen, and stored at −80°C. Three biological replicates were performed for each treatment.
Identification of SlUDPGT genes
Hmmsearch (Finn et al. 2011) V3.1b1 was used to query the tomato genome (ITAG 2.4 release) obtained from the SGN (http://solgenomics.net/) (Mueller et al. 2005) tomato database and identify the SlUDPGT gene family members. A hidden Markov model of the UDPGT domain (PF00201) was downloaded from Pfam release 32 (http://pfam.xfam.org/) (El-Gebali et al. 2019). The sequences whose e-values were lower than 1e-19 were considered to belong to SlUDPGT gene family members. All the protein and conserved domain sequences from the result file were collected and were used for BLAST comparisons against all protein sequences using Diamond to confirm the result of Hmmsearch (Buchfink et al. 2015). SMART (http://smart.embl-heidelberg.de/) (Letunic and Bork 2018) and Pfam (El-Gebali et al. 2019) were utilized to confirm the UDPGT domain and identify the signal peptide and transmembrane domains. The SlUDPGT enzyme physicochemical properties and subcellular localization were identified with ExPASy (Artimo et al. 2012) and WoLF PSORT (https://wolfpsort.hgc.jp/) (Horton et al. 2007).
Evolutionary analysis
The protein sequences of the SlUDPGT gene family members were extracted from the ITAG 2.4 genome annotation release from the SGN database (Mueller et al. 2005). MUSCLE (Larkin et al. 2007) was used to conduct multiple sequence alignment. The phylogenetic tree was then built using the neighbor-joining (NJ) method with 1000 bootstraps in MEGA 7.0.26 software (Kumar et al. 2016). The Blast program was used to identify tandemly duplicated genes (Johnson et al. 2008). Tandemly duplicate gene pairs were assigned when the identity of two genes was greater than 75%, and the alignment coverage of the longer sequence was greater than 75%. The Ka/Ks of all tandemly duplicated gene pairs were calculated using KaKs_Calculator (Wang et al. 2010), and their relationships were displayed via Circos (Krzywinski et al. 2009).
Chromosomal location, gene structure, sequence alignment and protein tertiary structure
The chromosome localization map was drawn using MG2C (http://mg2c.iask.in/mg2c_v2.1/) by collecting the chromosome positions of each gene from the annotated gff3 file (Chao et al. 2015). The CDS and gDNA sequences from the ITAG 2.4 annotation were aligned with the Gene Structure Display Server 2.0 (GSDS: http://gsds.cbi.pku.edu.cn) (Guo 2007) to identify the exon and intron structures. The expression of five genes was evaluated with qRT-PCR. Other genes of interest were used to predict the tertiary structure of the SlUDPGT proteins using the Alphafold2 software (Jumper et al. 2021). The following parameters are used: db_preset=full_dbs, model_preset=monomer. After calculation, PDB 3D viewer (https://www.rcsb.org/3d-view) (Sussman et al. 1998) was used to visualize the protein tertiary structure.
Prediction of cis-acting elements in the gene promoters
The promoter sequences (1.5 kb upstream of the 5ʹ UTR) were retrieved from the ITAG 2.4 genome annotation based on the location and chromosome number of SlUDPGT genes obtained from the gff3 file. All promoter sequences were uploaded to the PlantCARE database to predict cis-elements (Rombauts et al. 1999), and the results were visualized with GSDS 2.0.
RNA extraction and expression analysis
RNA was isolated from all the samples using an RNAiso Plus kit (Takara, Japan) following the manufacturer’s instructions. Then, the RNA was reverse transcribed into cDNA by using a reverse transcription kit (Takara, Japan). The primers used for quantitative real-time PCR (qRT‒PCR) are listed in Supplementary Table S2. Each 10 μL of the PCR reaction mixture contained 5 μl of Ultra SYBR Mixture (CWBIO, Beijing), 40 ng of cDNA, and 0.5 μM of each primer. The actin gene (Solyc11g005330.1.1) was used as the internal control to normalize target gene expression. The following program was used for qRT‒PCR in an Analytik Jena (Germany) q-Tower: 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 59°C (actin, UDPGT050)/60°C (UDPGT054, UDPGT077)/55°C (UDPGT091)/57.5°C (UDPGT094) for 30 s.
The microarray expression profiles of SlUDPGTs under different stress conditions were obtained through the TFGD database (http://ted.bti.cornell.edu/) (Fei et al. 2011), which provides genome-wide microarray data for various environmental stresses, including drought, salt, heat, Botrytis cinerea infection and tomato spotted wilt virus (TSWV). Microarray data from the TOM2 oligo array and Affymetrix genome array platforms were used. The probe sets of SlUDPGTs were identified through the BlastN program. If many probes were detected per gene, their average value was used. Then, log2 logarithmic transformation was performed on the expression data.
RNA-seq data from the platform Tomato Functional Genomics Database (http://ted.bti.cornell.edu/) were used to assess the expression patterns of SlUDPGTs in the leaves, roots, flower buds, fully opened flowers and 1, 2 and 3 cm, mature green, breaker, and breaker + 10 days stage fruits of the tomato cultivar Heinz (Fei et al. 2011). The gene expression levels were determined using their normalized expression values, which corresponded to reads per kilobases per million (RPKM) for each tissue/stage. The RPKM values were log2 logarithmically transformed, and heatmaps were plotted to examine the SlUDPGTs expression levels.
Vector construction and genetic transformation
The specific single guide RNAs (sgRNAs) for CRISPR/Cas9 were designed using CRISPR 2.0 (http://cbi.hzau.edu.cn/CRISPR2/) and are shown in Supplementary Table S2. The fragment was ligated to the expression vector PTX (Song et al. 2022) by homologous recombination using T4 DNA ligase (NEB, M0202T). Then, the ligated product was transferred to Escherichia coli (DH5α). The positive plasmids were extracted after picking single clones, which were subsequently cultured for sequence verification. The plasmids with the correctly arranged and precise sequences were transferred into Agrobacterium (LBA4404). The Agrobacterium-mediated leaf disc transformation method into the drought-sensitive tomato cultivar AC was carried out to generate SlUDPGT52 transgenic tomato plants.
To verify the CRISPR/Cas9 mutations in the transgenic tomato plants, DNA was extracted using the CTAB method, and the target fragments were assessed by PCR, while WT plants were used as controls. The target fragments were amplified by PCR using gene-specific primers, and the PCR products were sequenced.
Drought tolerance assay and measurement of physiological indicators
Seeds from the WT and SlUDPGT52 CRISPR/Cas9 (CR) knockout lines (CR-1, CR-2 and CR-3) were sown in MS medium for vertical culture. After sterilization, the seeds with uniform germination were selected and transferred to MS medium in three biological replicates for each line in each treatment group. MS medium without any other compounds added was used as the control, while MS medium containing 200 mmol/L mannitol was used to simulate drought stress. After sowing the seeds on the corresponding medium, the plants were grown at 25℃ to observe potential growth differences among the treatments.
Regarding the soil-based experiments, uniform WT and transgenic knockout line seedlings were transplanted into pots with 150 g weight of substrate (peat soil:perlite:vermiculite = 3:1:1). Tomato plants of uniform size with five leaves were selected for the drought treatment. A control group was assigned with normal watering and a drought treatment group with strict water control, with three biological replicates (three plants per line) in each group. Before the drought treatment, each treatment group was watered uniformly. Then, phenotypic observations were made, and the plants were photographed and measured at specific time points. To further determine the physiological and biochemical changes in the transgenic plants affected by the SlUDPGT52 knockout during drought stress compared to WT, the physiological and biochemical parameters were measured on the fourth day of drought treatment, including proline (Pro), malondialdehyde (MDA), electrical conductivity, and various antioxidant enzyme activities. Diaminobenzidine (DAB) and nitroblue tetrazolium (NBT) staining were performed to detect oxidative stress in plants under drought stress. The physiological indicators were measured as previously described (Wang et al., 2023).
Statistical analysis
All data were analyzed using the GraphPad 8.0 software. Student’s t-test was used to determine statistically significant differences between the two datasets (* P<0.05, ** P<0.01, *** P<0.001, **** P<0.0001).
Results
Identification and characterization of the UDPGT gene family in tomato
In this study, a total of 106 SlUDPGT genes were identified in the tomato reference genome (Supplementary Table S1). The genes were numbered from UDPGT001 to UDPGT106 based on their distributions on chromosomes. The protein length, chromosome location, molecular weight, isoelectric point, hydrophilicity coefficient and subcellular localization were assessed. As shown in Supplementary Table S1, the protein length of SlUDPGTs was greater than 183 aa, and the average was 449 aa. Most proteins (86.79%) ranged between 400 and 500 aa in length. The average molecular weight was 50656.44, and their isoelectric points ranged from 4.87 to 9.67. The subcellular localization prediction indicated that more than half of the SlUDPGTs were localized in the chloroplast (52.83%). The number of proteins located in the nucleus was equal to those located in the cytosol. Only a few SlUDPGTs were found to be localized in the extracellular environment (1.89%), mitochondria (0.94%), and endoplasmic reticulum (1.89%).
Conserved domain and phylogenetic analyses of SlUDPGTs
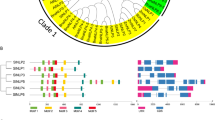
The conserved domain structure exhibited limited variability among the UDPGT gene family in tomato, with all members containing a highly conserved UDPGT domain (Fig. 1). Additionally, UDPGT031, UDPGT048 and UDPGT062 possessed an additional Glyco_tran_28_C domain, while UDPGT002 and other UDPGT proteins carried transmembrane domains. These findings suggest the potential involvement of SlUDPGTs in transmembrane transport and protein synthesis.
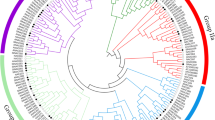
Due to the high divergence of UDPGTs, a phylogenetic analysis of SlUDPGTs was conducted to gain further insights into their phylogenetic relationships. Based on the results, the lowest overall mean distance (0.704) was observed for the p-distance, and the distances based on the other models were above 1. Based on the phylogenetic tree, the SlUDPGTs were divided into five subfamilies labeled from A to E (Fig. 2a). Subfamily E was the largest subfamily, with 39 members. Three members with Glyco_tran_28_C domains were presented in subfamilies A, D and E. Subfamily C was the smallest, with five members.
Phylogenetic analysis of the uridine diphosphate glycosyltransferase (UDPGT) gene family in tomato and analysis of tandemly duplicated genes. (a) Phylogenetic analysis. Different colors indicate different subfamilies based on sequence similarity annotation analysis. Red denotes the SlUDPGT genes selected for subsequent experiments. (b) Analysis of tandemly duplicated genes. The red lines represent the tandemly duplicated gene pairs located in a single chromosome. The green lines indicate the tandemly duplicated gene pairs located in different chromosomes. The chromosome number and tandemly duplicated gene pairs are marked. The ch01-12 represent 12 chromosomes in the tomato genome
A total of 28 pairs of tandemly duplicated genes were identified in the SlUDPGT gene family (Fig. 2b). The Ka, Ks and Ka/Ks of these 28 pairs of genes were calculated (Supplementary Table S3). A Ka/Ks value greater than 1.2 indicated positive selection within a gene pair, a value less than 0.5 suggested purifying selection and a value between 0.5 and 1.2 revealed neutral selection (Betran et al. 2002; Emerson et al. 2004). The results showed a conspicuous distribution of Ka/Ks. Overall, 85.7% of gene pairs were influenced by selection evolutionary forces, indicating that the SlUDPGT gene family is actively evolving. Furthermore, most gene pairs (71.4%) have undergone purifying selection. Only two gene pairs had Ka/Ks values greater than 1.2. Despite their low occurrence, these two gene pairs had Ka/Ks values that were much higher than 1, reaching 6.0–7.0, which suggested that they were largely affected by positive selection. Four genes were under neutral selection. Notably, the Ka/Ks values within each of the three selection modes were concentrated toward extreme values. For example, in gene pairs undergoing purifying selection, their Ka/Ks values were much lower than 0.5, ranging from 0.1 to 0.3. A similar pattern was observed in gene pairs under neutral and positive selection pressure.
Chromosomal localization, gene structure, and protein tertiary structure of SlUDPGTs
SlUDPGTs were distributed in a relatively non-random manner in chromosomes and tended to be present as gene clusters. The 106 SlUDPGT genes were mapped to the tomato reference genome based on the ITAG 2.4 annotation (Fig. 3). The SlUDPGT genes were distributed across all 12 chromosomes. SlUDPGT genes predominantly exhibited a distal distribution from the centromere. Regions of high gene density were observed on chromosomes 01, 03, 04, 07, 08, 10, 11, and 12, while SlUDPGT genes were more evenly distributed on chromosomes 02 and 06.
The exon-intron structure is a significant gene structural property from which the existence of transcript isoforms can be inferred. The exon and intron structures of SlUDPGTs were obtained by comparing the cDNA and gDNA sequences. Among all UDPGTs, only UDPGT003 and UDPGT072 exhibited a three-exon structure, representing 1.9% of the family members. Notably, 52 genes lacked introns, accounting for 49.1% of all UDPGTs. Furthermore, members within the same subfamily displayed structural similarities. UDPGT056, UDPGT057, UDPGT059 and UDPGT060 were located on the same branch of the phylogenetic tree, and upstream or downstream sequences were found at the beginning and end of these genes. Their coding sequence was split due to the presence of an intron in the middle of their genomic sequence. UDPGT085, UDPGT086, UDPGT087, UDPGT088 and UDPGT089 were classified in the same subfamily with similar sequence lengths and intron exon distribution patterns (Supplementary Fig. S1).
The protein tertiary structures were obtained based on the top five scores from different models for each protein, among which the model with the highest score was used for visualization (Supplementary Fig. S2). The results indicated that each protein contained two or more β/α/β Rossmann-like domains, and most of the proteins exhibited alternating structures of several β-sheets and α-helices. For example, UDPGT054 contains Rossmann-like domains composed of 5 β-lamellar and 5 α-helices. Besides, the β-sheets in Rossmann-like domains tended to congregate to form a curved surface, and no β-turns were observed. These patterns were observed in all 11 protein tertiary structures, indicating that these proteins share the same structural pattern.
Cis‑acting elements in the promoters of SlUDPGT genes
Cis-acting elements are present in the promoter sequence, integrating the developmental and environmental signals in different tissues and growth stages. Detailed information on the cis-acting elements in each gene promoter is shown in Supplementary Fig. S3 and Supplementary Table S2. Based on our results, 48 cis-acting elements were identified in more than 20 SlUDPGT genes (Supplementary Table S5). Among the 48 cis-acting elements, 21 were annotated by the PlantCARE database. TATA box and CAAT box motifs were identified in all SlUDPGT promoters, proving that the promoter sequence cis-acting elements were predicted with high accuracy. Most promoters contained the Box 4 motif, which is a light-responsive element. In addition to the TATA box and CAAT box motifs, 20 cis-acting elements were divided into six classes. The Box 4, GT1-motif, TCT-motif, G-box, GATA-motif, G-Box, I-box and MRE motifs are involved in light responsiveness. ABRE, CGTCA-motif, TGACG-motif and P-box are involved in hormone signaling induction. WUN-motif, TC-rich repeats and LTR are abiotic and biotic stress-inducible. The O2 site and CAT box are involved in growth and development. In addition, an element called Unnamed_1, which is a 60K protein binding site, and a regulatory element named the A-box were identified.
Expression patterns of SlUDPGT genes
The data from the TOM2 oligo array and the Affymetrix genome array were retrieved from the TFGD database (Supplementary Table S6) and drawn into a heatmap to explore the expression patterns of SlUDPGTs under various abiotic stresses (Fig. 4a-c) and biotic stresses (Fig. 4d-e). No corresponding probes were identified for ten genes (9.43%). The tomato experimental varieties CO-3 and EC-520061 are sensitive and resistant to abiotic stress, respectively (Mishra et al. 2016). The expression data from the two varieties were analyzed and compared to explore the effect of abiotic stress on gene expression during flowering (Fig. 4a). During the flowering stage, under drought stress treatment, a significant upregulation of genes was observed in the drought tolerant (DT) variety compared to the drought sensitive (DS) variety. This finding suggests that SlUDPGTs may be crucial in conferring drought tolerance during the tomato flowering stage. Furthermore, UDPGT074, UDPGT023, UDPGT042, UDPGT071, UDPGT072, UDPGT073, UDPGT075, UDPGT091 and UDPGT092 were expressed at very low levels in DT varieties, but their expression levels in DS varieties were nearly similar to that of non-stress control (CK) varieties. When exposed to heat stress, the two varieties exhibited no significant differences in response to this stress condition. However, the expression of 29 SlUDPGT genes, including UDPGT089, was observed to be downregulated under heat treatments in susceptible varieties. The tomato variety PI365967 exhibits salt tolerance, while Moneymaker is a salt-sensitive cultivar. The transcriptome data analysis of the SlUDPGT gene family members in response to salt stress in these two tomato varieties are shown in Fig. 4b. 20 SlUDPGT genes (UDPGT008, UDPGT061, UDPGT078, UDPGT104, UDPGT089, UDPGT033, UDPGT034, UDPGT058, UDPGT102, UDPGT057, UDPGT056, UDPGT002, UDPGT014, UDPGT005, UDPGT059, UDPGT062, UDPGT060, UDPGT093, UDPGT038 and UDPGT101) were significantly upregulated under salt-stress conditions in both varieties (Fig. 4b). IL 2-5 and IL 9-1 are drought-resistant introgression lines, and M82 is their recurrent parent (Gong et al. 2010). Upon exposure to drought stress, UDPGT015 showed significant upregulation, while the expression of 20 SlUDPGT genes was notably downregulated. Conversely, UDPGT078, UDPGT066, UDPGT001 and UDPGT014 exhibited very low expression levels overall (Fig. 4c).
Expression patterns of Solanum lycopersicum uridine diphosphate glycosyltransferase (SlUDPGT) genes under various stresses. (a) Expression profiles of SlUDPGTs in tomato under high temperature and drought stress at the flowering stage. CK, non-stress control; DS, drought susceptibility; DT, drought tolerance; HS, heat susceptibility; HT, heat tolerance. (b) Expression profiles of SlUDPGTs under salt stress. MM, Moneymaker; PI, PI365967. (c) Expression profiles of SlUDPGTs under drought stress. (d) Expression profiles of SlUDPGTs in tomato fruits infected with Botrytis cinerea. MW, wounded mature green fruits; MH, healthy mature green fruits; MB, B. cinerea-infected mature green fruit; RH, healthy red ripe fruit; RW, wounded red ripe fruit; RB, B. cinerea-infected red ripe fruit. (e) Expression profiles of SlUDPGTs after tomato infection with tomato spotted wilt virus (TSWV). IR, infected roots; MR, mock-inoculated roots; IL, infected leaves; ML, mock-inoculated leaves; Red indicates upregulated expression, and green indicates downregulated expression; a gray box indicates that no reading was detected. The scale represents the expression levels
The AC and Moneymaker tomato varieties were assessed to explore the expression levels of genes under B. cinerea and TSWV infection, respectively (Cantu et al. 2009; Catoni et al. 2009). As shown in Fig. 4d, the expression of more than half of SlUDPGTs was higher in the healthy red ripe fruit (RH) stage than in the healthy mature green fruit (MH) stage. When the mature green fruit was wounded, the expression levels of most SlUDPGTs were slightly upregulated. These genes were continuously slightly upregulated when the varieties were wound-inoculated with B. cinerea. On the other hand, UDPGT086, UDPGT015, UDPGT084, UDPGT091, UDPGT092, UDPGT023, UDPGT042, UDPGT071, UDPGT072, UDPGT073, UDPGT075, UDPGT070 and UDPGT076 were downregulated in both varieties after inoculation with B. cinerea. During the red fruit stage, SlUDPGT gene expression exhibited significant differences after wounding and inoculation with B. cinerea. UDPGT081, UDPGT079 and UDPGT080 were weakly expressed, whereas UDPGT005 and UDPGT059 showed the highest expression levels in healthy plants. When infected by TSWV, the varieties exhibited remarkably different SlUDPGT expression profiles between the roots and leaves (Fig. 4e). UDPGT006, UDPGT067 and UDPGT105 demonstrated extremely high expression levels in leaves but low expression levels in roots. On the other hand, no expression could be detected for UDPGT053 in the leaves and UDPGT010 in the roots.
We further assessed the expression of SlUDPGTs in various tomato tissues/stages, namely the leaves, roots, flowers, flower buds, 1-, 2- and 3-cm fruits, mature green fruits, breaker fruits and fruits on day 10, using RNA-seq data. The results indicated that SlUDPGTs exhibited different expression levels in different tissues and stages (Supplementary Fig. S4). UDPGT052, UDPGT033, UDPGT034, UDPGT087, UDPGT068, UDPGT053, UDPGT086, UDPGT082, UDPGT046, UDPGT047, UDPGT100, UDPGT013, UDPGT072, UDPGT071, UDPGT001, UDPGT031, UDPGT002 and UDPGT014 exhibited low expression levels in all tissues. UDPGT066 was highly expressed in fully opened flowers, and UDPGT096 was highly expressed in roots. In addition, four genes, including UDPGT094, UDPGT091, UDPGT054 and UDPGT055, were upregulated in the buds, flowers, and 1- and 2-cm fruits but showed decreased expression in breaker-stage fruits. UDPGT067, UDPGT079, UDPGT080, UDPGT081, UDPGT009 and UDPGT011 were highly expressed in breaker stage fruits and +10 days post breaker stage fruits.
qRT‒PCR was performed to more precisely evaluate the expression patterns of 5 randomly selected representative SlUDPGTs, namely, SlUDPGT050, SlUDPGT054, SlUDPGT077, SlUDPGT091 and SlUDPGT094, under different abiotic stresses (Fig. 5) and hormone treatments (Fig. 6). The expression of SlUDPGT054 was significantly upregulated at 0.5 h and 2 h after NaCl treatment. However, SlUDPGT077 did not exhibit expression changes in the first 6 h but increased sharply at 12 h. The expression of SlUDPGT094 increased by 300-fold after 0.5 h of PEG treatment, suggesting that this gene was likely to be involved in drought tolerance. The expression of SlUDPGT091 decreased under treatment with MV, indicating that oxidative stress might inhibit its expression. The SlUDPGT094 gene was initially upregulated but returned to its initial expression level after 24 h of MV treatment, while SlUDPGT091 was substantially upregulated, respectively (Fig. 5). The expression of SlUDPGT050 and SlUDPGT094 was significantly downregulated under ABA treatment (Fig. 6a, e), while the expression of SlUDPGT050 decreased to almost non detectable levels at 24 h after ABA treatment (Fig. 6a). SlUDPGT050 was significantly induced by various stresses and hormones and increased largely within 2 h after the MeJA and BR treatments (Fig. 6a). In plants treated with GA and MeJA , the expression levels of SlUDPGT054 increased initially, peaked and then decreased to normal levels (Fig. 6b). The expression of SlUDPGT077 gradually increased and peaked at 6 h after JA treatment (Fig. 6c). Notably, after 2 h of BR treatment, the expression level of SlUDPGT091 increased by more than 1000-fold, indicating its very high responsivenss to BR (Fig. 6d). In summary, the expression of SlUDPGTs was regulated by various stresses and hormones, suggesting their involvement in the adaptataion and resistance to multiple stresses and their regulation by diverse hormonal signals.
Expression analysis of selected Solanum lycopersicum uridine diphosphate glycosyltransferases (SlUDPGTs) under different stress treatments. Error bars correspond to the ±SE of technical replicates. The relative expression was the ratio of each treatment compared with that of the control group (untreated). The lowercase letters on the column represent significant differences (P < 0.05)
Expression patterns of selected Solanum lycopersicum uridine diphosphate glycosyltransferases (SlUDPGTs) under different hormone treatments. (a)-(e) Relative expression of UDPGT050, UDPGT054, UDPGT077, UDPGT091 and UDPGT094 after gibberellic acid (GA), jasmonic acid (JA), abscisic acid (ABA) and brassinolide (BR) treatments, respectively. The relative expression was the ratio of each treatment compared with that of the control group (untreated). Error bars show ±SE of technical repetition. The lowercase letters on the column represent significant differences (P < 0.05)
SlUDPGT52 knockout enhances drought tolerance in tomato
In previous experimentation, we identified the coexpression of SlUDPGT52 and a B-box protein-encoding gene under drought conditions, which prompted us to perform functional analysis. After target site sequencing, we selected three mutation sites in SlUDPGT52 for further functional analysis (Fig. 7a). To determine whether the knockout of SlUDPGT52 affects tomato responses to drought stress, seedlings of three knockout lines (CR-1, CR-2, CR-3) and WT were sown on MS medium (control) and MS supplemented with 200 mM mannitol (drought stress group), respectively (Fig. 7b). There were no significant differences between SlUDPGT52 knockout plants and WT plants in terms of plant height, root length and seedling weight compared to the control under normal condition. However, the overall growth trend of SlUDPGT52 knockout plants, including root length and seedling height, was significantly higher than that of WT plants under 200 mM mannitol treatment (Fig. 7c-e). This finding suggests that SlUDPGT52 knockout enhances drought tolerance in tomato compared to WT plants.
Analysis of SlUDPGT52 gene structure and drought tolerance of CR-SlUDPGT52 lines and WT plants. (a) SlUDPGT52 CRISPR/Cas9 knockout transgenic tomato plants target site mutation. (b) Phenotypic differences between SlUDPGT52 knockout tomato plants and WT tomatoes cultured on MS medium with or without mannitol (concentration: 200 mmol/L). Three biological replicates were performed for each line. WT: AC; CR-1, CR-2 and CR-3: SlUDPGT52 knockout lines. (c) Plant height. (d) Root length. (e) Seedlings weight of SlUDPGT52 knockout and WT tomato plants in mannitol-simulated drought treatment (*P < 0.05, **P < 0.01, **P < 0.001 and ****P < 0.0001)
The SlUDPGT52 knockout and WT tomato plants were subjected to drought treatment further to elucidate the role of SlUDPGT52 under drought stress. The WT plants started wilting on the fifth day of drought treatment, while the knockout plants maintained robust growth. On the tenth day of drought stress, the wild-type plants became severely wilted, while the SlUDPGT52 knockout plants displayed a less pronounced wilting (Fig. 8a). These results demonstrate that the knockout of SlUDPGT52 enhances drought tolerance compared to WT plants. The MDA content and electrolytic leakage of SlUDPGT52 knockout plants showed a decreasing trend compared to WT. In contrast, peroxidase (POD), superoxide dismutase (SOD) and catalase (CAT) activities and Pro content showed an increasing trend after drought treatment. Notably, the higher POD, SOD and CAT activities in the SlUDPGT52 knockout plants compared with the WT plants indicated that the knockout plants exhibited higher antioxidant enzyme activities (Fig. 8b-g). Furthermore, there was a significant increase in the internode number in the knockout plants compared to WT (Fig. 9a). After DAB and NBT staining, the leaves of SlUDPGT52 knockout lines and WT plants were lightly colored and not significantly different under normal growth conditions. However, after drought stress, the leaves of the SlUDPGT52 knockout plants were darker and more extensively colored than those of the WT plants (Fig. 9b-e). Therefore, SlUDPGT52 knockout plants exhibited enhanced tolerance to oxidative stress compared to WT plants under drought conditions. In summary, our findings demonstrate that SlUDPGT52 knockout enhances drought tolerance of tomato plants by enhancing ROS scavenging.
Phenotypic analysis and drought tolerance assay of CR-SlUDPGT52 lines and WT tomato plants. (a) The left panel shows the phenotype of a single plant, and the right panel shows the phenotypes of three biological replicates from each line. Different physiological indices were determined in Solanum lycopersicum uridine diphosphate glycosyltransferase 52 (SlUDPGT52) knockout and WT plants. (b) Malondialdehyde (MDA) content. (c) Electrolyte leakage. (d) Pro content. (e)-(g) Activities of peroxidase (POD) (e), superoxide dismutase (SOD) (f), and catalase (CAT) (g)
Knockout of Solanum lycopersicum uridine diphosphate glycosyltransferase 52 (SlUDPGT52) alters the growth and reactive oxygen species (ROS) levels in tomato plants. (a) Number of internodes in knockout and WT plants. (b), (c) Diaminobenzidine (DAB) staining between SlUDPGT52 knockout and WT plants before (b) and after drought treatment (c). (d), (e) Nitroblue tetrazolium (NBT) staining differences between SlUDPGT52 knockout plants and WT plants under normal growth conditions (d) and after drought treatment (e)
Discussion
Evolution of the UDPGT gene family in tomato
Overall, the UDPGT gene family is highly conserved among species (Ross et al. 2001). From the point of view of protein primary structure, the PSPG motif has been identified in all plant UDPGTs (Mackenzie et al. 1997). Meanwhile, in higher structural dimensions, all protein tertiary structures of SlUDPGTs predicted in this study contain two or more β/α/β Rossmann-like domains, or in other words, GT-B folds. This conclusion is consistent with previous studies (Yonekura-Sakakibara and Hanada 2011), which further confirm the high degree of conservation of the UDPGT gene family. However, certain UDPGT gene family members are less conserved among plant species. Many receptor molecules that are substrates to UDPGTs greatly differ among plant species (Lairson et al. 2008; Osmani et al. 2009). Therefore, the structure of UDPGTs must be highly diversified to adapt to different receptor molecules.
Gene duplication drives biological evolution to a certain extent (Moore and Purugganan 2003), which may contribute to the diversity of SlUDPGTs. In this study, 28 tandemly duplicate gene pairs were identified, and their Ka/Ks values were calculated. The results showed that only 4 genes were not significantly affected by selection pressure, 2 genes were under strong positive selection, and 20 genes were under negative selection pressure (Fig. 2b). This result indicated that the SlUDPGT gene family is in a stage of rapid evolution. This conclusion is consistent with the low conservation between the SlUDPGT gene family members. In evolution, most of the duplicated SlUDPGT genes are adjacent to their parental genes (Fig. 3). Only one cross-chromosome duplication event occurred between chromosomes 04 and 12. The probability of duplication for each SlUDPGT gene is not equal. Genes usually duplicate more than once, and this event may be related to the degree of activity of different genes. SlUDPGTs are primarily distributed at the ends of chromosomes, influenced by duplication events and other reasons.
SlUDPGTs are involved in biotic and abiotic stress responses.
The optimal plant growth status cannot always be maintained under natural environmental conditions, necessitating various plant adaptations. Plants are exposed to various biotic and abiotic stresses, which significantly impact the yield and quality of tomatoes (Pervez et al. 2009; Zhang et al. 2017). Plant hormones serve as regulatory factors in response to stress. ABA is responsible for plant adaptation against abiotic stress, while hormones such as SA, MeJA, and ETH play a pivotal role in biotic stress responses (Verma et al. 2016). Previous studies have demonstrated the association between SlUDPGT gene family members and stress tolerance (Li et al. 2018b). Our research revealed a tight relationship between the SlUDPGT gene family regulation and the responses to a diverse range of biotic and abiotic stresses.
As observed in this study, SlUDPGT067 potentially increases the level of ABA. This was consistent with a previous study showing that SlUDPGT75C1 could mediate ABA glycosylation (Sun et al. 2017). Our results revealed that SlUDPGT067 was highly expressed in mature green, breaker stage and 10 days past breaker stage fruits (Fig. S4). Notably, the expression of SlUDPGT067 also increased significantly after the leaves were infected with TSWV, suggesting that SlUDPGT067 might also be involved in resistance to biotic stress (Fig. 4e). Moneymaker is an excellent tomato experimental variety that is not resistant to stress, insects or diseases. Furthermore, PI365967 is a variety with greater salt tolerance compared to Moneymaker (Sun et al. 2010). In this study, the expression profiles of the SlUDPGT gene family were explored in these two varieties under salt stress. Most SlUDPGT genes in Moneymaker were highly expressed under salt stress, while the expression of a few genes was reduced (Fig. 4b). The expression of SlUDPGTs in PI365967 showed a similarity to that in Moneymaker; however, the observed changes in the expression of specific genes was very high, implying a potentially pivotal role for SlUDPGTs in conferring salt stress resistance. For example, the expression of UDPGT054 increased by 30-fold after 0.5 h of salt stress and did not decrease until 2 h. The expression of UDPGT077 increased sharply after 12 h of salt treatment. There was also a significant increase in UDPGT054 and UDPGT077 expression after ABA treatment, which could regulate abiotic stress tolerance. In contrast, UDPGT050 expression decreased within 24 h after salt stress, and its expression also significantly decreased after ABA treatment (Figs. 5 and 6). The results suggest a potential association between UDPGT054 and UDPGT077 with the positive regulation of salt stress tolerance. At the same time, UDPGT050 appears to be involved in the negative regulation of salt stress adaptation and tolerance.
Biotic stress may lead to the death of large parts of plant tissues and organs, seriously affecting crop yield (Herbert 2002). Many abiotic stress conditions have been shown to weaken the defense mechanisms of plants and increase susceptibility to pathogen infection (Atkinson and Urwin 2012; Goel et al. 2008; Amtmann et al. 2008; Suzuki et al. 2014). Our results showed that the SlUDPGT gene family is also involved in the responses to a diverse range of biotic stresses. B. cinerea is an airborne plant pathogen that can cause plant tissue necrosis (Williamson et al, 2007). Several studies have been conducted on B. cinerea in tomato plants, which demonstrated its detrimental impact on tomato yield (Rguez et al. 2018). In this study, the expression patterns of the SlUDPGT gene family were investigated following B. cinerea infection. The majority of SlUDPGTs exhibited a slight upregulation in the mature green fruit stage, while they were downregulated in the ripened red fruit stage(Fig. 4b). These findings suggested that SlUDPGT expression in the fruits is potentially linked to the defense response against B. cinerea. Moreover, they might play different roles in these two different fruit development stages. TSWV causes more than $1 billion yearly loss to crops grown in fields and greenhouses (Karavina and Gubba 2017). A previous study found that members of the SlUDPGT gene family play an important role in the processes leading to plant resistance to TSWV (Campos et al. 2019). In our study, SlUDPGT006, SlUDPGT067 and SlUDPGT105 were highly expressed in roots, and many genes were highly expressed in the leaves infected with TSWV, which was in agreement with the findings of Campos et al. (Campos et al. 2019). These results indicate that SlUDPGTs may participate in biotic and abiotic stress responses. In the future, further research should be carried out to explore the roles of SlUDPGTs in the adaptation and resistance to stress conditions.
Putative functions of SlUDPGTs
UDPGTs are enzymes that catalyze glycosylation reactions. UDPGTs modify various receptor molecules through glycosylation, consequently affecting downstream biological processes, such as plant growth, flowering and fruiting (Ross et al. 2001). Many studies have also indicated that UDPGTs are involved in various biological pathways (Zhang et al. 2021). From the perspective of gene structure, SlUDPGTs exclusively possess conserved domains that are responsible for their glycosylation catalytic activity, thereby indicating a relatively unified mechanism. Several SlUDPGTs exhibit transmembrane structures and signal peptides, implying their involvement in material transportation through membranes. The analysis of cis-acting elements revealed that the promoter region of SlUDPGT genes in tomato harbored numerous stress- and hormone-related elements, including elements associated with light responses, injury responses, ABA responses, MeJA responses, GA responses, and low-temperature responses. More accurate and sensitive qRT-PCR experiments demonstrated that these conditions have a significant inducing effect on SlUDPGTs. Published studies also revealed that SlUDPGTs are associated with ABA (Dong et al. 2014; Sun et al. 2017), MeJA (Guo et al. 2016), Asian soybean rust (Langenbach et al. 2013), salt stress, and oxidative stress responses (Ahrazem et al. 2015). In addition, the transcriptomic analysis of UDPGT expression further confirmed these findings (Mamoon Rehman et al. 2016) and is consistent with the conclusions drawn from our study in tomato.
With regard to the biosynthesis of secondary metabolites, several studies have shown that UDPGTs are related to the accumulation of flavonoids in plants. MeJA is a plant hormone, and many studies have demonstrated that it can induce flavonoid biosynthesis (Guo et al. 2016; Chen et al. 2020; Premathilake et al. 2020). UDPGTs are also induced by MeJA, which in turn affects flavonoid accumulation (Guo et al. 2016). SlUDPGTs were highly expressed under MeJA treatment. This finding provides strong evidence that SlUDPGT can promote flavonoid accumulation. In addition, UDPGTs catalyze the synthesis of triterpenes (Rahimi et al. 2019) and participate in cell wall lignification in Arabidopsis (Lin et al. 2016). Although these functions could not be demonstrated in our study, UDPGT069 in tomato is highly homologous to UDPGT72B1, which regulates the Arabidopsis cell wall lignification, indicating that members of SlUDPGTs may have a similar function (Lin et al. 2016).
SlUDPGT52 was selected to determine whether UDPGTs are involved in the responses of tomato to drought stress. The results showed that SlUDPGT52 knockout plants exhibited less wilting after drought treatment compared to WT tomato plants, suggesting that SlUDPGT52 regulates drought tolerance in tomato. When grown normally, there was no significant difference in physiological parameters between CR-SlUDPGT52 and WT plants. Under drought treatment, we found that electrolytic leakage and MDA content of SlUDPGT52 knockout lines were lower compared to WT, while the Pro content and POD, SOD and CAT enzyme activities were higher. In addition, the leaves of SlUDPGT52 knockout plants showed a lighter coloration than those of WT plants after DAB and NBT staining. The above results indicate that the knockout of SlUDPGT52 has a strong capacity to induce ROS scavenging. Similarly, the downregulation of a UDP-glycosyltransferase gene in tea plants (CsUGT91Q2) decreased their ROS scavenging ability, increasing their sensitivity to low-temperature stress (Zhao et al. 2020). In contrast, overexpression of rice UGT85E1 enhanced tolerance to drought stress, resulting in an increase in Pro content and ROS scavenging capacity, and ugt85e1 tice mutants of rice exhibited greater sensitivity to drought (Liu et al. 2021). Furthermore, we found that the root growth of SlUDPGT52 knockout lines increased compared to WT when grown in a medium containing mannitol, indicating increased drought tolerance. The above finding was consistent with a previous study showing that a UDP-glycosyltransferase gene in Arabidopsis (AtUGT76E11) overexpression lines had enhanced root growth when treated with NaCl, mannitol and H2O2 treatments compared to the WT (Li et al. 2018b). In summary, SlUDPGT52 knockout can increase antioxidant enzyme activity and reduce oxidative damage and osmotic stress in tomato plants under drought stress, leading to enhanced drought tolerance.
Conclusions
In this study, a total of 106 UDPGT gene family members were identified and analyzed in tomato. The physical and chemical properties of the proteins, their evolutionary relationships, subcellular localization, selection pressure, cis-acting elements, gene structure, tissue expression patterns, and hormone-induced expression of these members were investigated using microarray or RNA-seq transcriptome data. qRT-PCR was conducted to more precisely assess the expression patterns of representative genes under different hormonal treatments and stresses. Most SlUDPGTs were responsive to hormone and stress treatments, indicating their crucial role in plant growth processes, especially abiotic and biotic stress responses. Furthermore, CRISPR/Cas9-mediated knockout of SlUDPGT52 enhanced drought stress tolerance in tomato, indicating that it is a negative regulator of tomato drought stress tolerance. These findings provide a foundation for the functional analysis of genes and are essential for advancing research on stress resistance mechanisms in tomatoes.
Availability of data and materials
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
Change history
13 May 2024
A Correction to this paper has been published: https://doi.org/10.1007/s44281-024-00047-2
References
Ahrazem O, Rubio-Moraga A, Trapero-Mozos A, Climentc MFL, Gomez-Cadenas A, Gomez-Gomeza L. Ectopic expression of a stress-inducible glycosyltransferase from saffron enhances salt and oxidative stress tolerance in Arabidopsis while alters anchor root formation. Plant Sci. 2015;234:60–73. https://doi.org/10.1016/j.plantsci.2015.02.004.
Amtmann A, Troufflard S, Armengaud P. The effect of potassium nutrition on pest and disease resistance in plants. Physiol Plant. 2008;133:682–91. https://doi.org/10.1111/j.1399-3054.2008.01075.x.
Artimo P, Jonnalagedda M, Arnold K, Baratin D, Csardi G, de Castro E, et al. ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res. 2012;40:W597–603.
Atkinson NJ, Urwin PE. The interaction of plant biotic and abiotic stresses: from genes to the field. J Exp Bot. 2012;63:3523–43. https://doi.org/10.1093/jxb/ers100.
Betran E, Thornton K, Long M. Retroposed new genes out of the X in Drosophila. Genome Res. 2002;12:1854–9. https://doi.org/10.1101/gr.6049.
Buchfink B, Xie C, Huson DH. Fast and sensitive protein alignment using DIAMOND. Nat Methods. 2015;12:59–60. https://doi.org/10.1038/nmeth.3176.
Campos L, Lopez-Gresa MP, Fuertes D, Belles JM, Rodrigo I, Lison P. Tomato glycosyltransferase Twi1 plays a role in flavonoid glycosylation and defence against virus. BMC Plant Biol. 2019;19:450. https://doi.org/10.1186/s12870-019-2063-9.
Cantarel BL, Coutinho PM, Rancurel C, Bernard T, Lombard V, Henrissat B. The Carbohydrate-Active EnZymes database (CAZy): an expert resource for Glycogenomics. Nucleic Acids Res. 2009;37(Database issue):D233–8. https://doi.org/10.1093/nar/gkn663.
Cantu D, Blanco-Ulate B, Yang L, Labavitch JM, Bennett AB, Powell ALT. Ripening-regulated susceptibility of tomato fruit to Botrytis cinerea requires NOR but not RIN or ethylene. Plant Physiol. 2009;150:1434–49. https://doi.org/10.1104/pp.109.138701.
Catoni M, Miozzi L, Fiorilli V, Lanfranco L, Accotto GP. Comparative analysis of expression profiles in shoots and roots of tomato systemically infected by tomato spotted wilt virus reveals organ-specific transcriptional responses. Mol Plant Microbe Interact. 2009;22:1504–13. https://doi.org/10.1094/Mpmi-22-12-1504.
Chao JT, Kong YZ, Wang Q, Sun YH, Gong DP, Lv J, et al. MapGene2Chrom, a tool to draw gene physical map based on Perl and SVG languages. Yi Chuan. 2015;37:91–7.
Chen J, Wang J, Wang R, Xian B, Ren CX, Liu QQ, et al. Integrated metabolomics and transcriptome analysis on flavonoid biosynthesis in safflower (Carthamus tinctorius L.) under MeJA treatment. BMC Plant Biol. 2020;20:353. https://doi.org/10.1186/s12870-020-02554-6.
Curatolo W. Glycolipid function. Biochimica Et Biophysica Acta. 1987;906:137–60. https://doi.org/10.1016/0304-4157(87)90009-8.
Dong T, Xu ZY, Park Y, Kim DH, Lee Y, Hwang I. Abscisic acid uridine diphosphate glucosyltransferases play a crucial role in abscisic acid homeostasis in Arabidopsis. Plant Physiol. 2014;165:277–89. https://doi.org/10.1104/pp.114.239210.
El-Gebali S, Mistry J, Bateman A, Eddy SR, Luciani A, Potter SC, et al. The Pfam protein families database in 2019. Nucleic Acids Res. 2019;47:D427–32. https://doi.org/10.1093/nar/gky995.
Emerson JJ, Kaessmann H, Betran E, Long MY. Extensive gene traffic on the mammalian X chromosome. Sci. 2004;303:537–40. https://doi.org/10.1126/science.1090042.
Fei ZJ, Joung JG, Tang XM, Zheng Y, Huang MY, Lee JM, et al. Tomato functional genomics database: a comprehensive resource and analysis package for tomato functional genomics. Nucleic Acids Res. 2011;39(Database issue):D1156–63. https://doi.org/10.1093/nar/gkq991.
Finn RD, Clements J, Eddy SR. HMMER web server: interactive sequence similarity searching. Nucleic Acids Res. 2011;39(Web Server issue):W29–37.
Fujiwara R, Yokoi T, Nakajima M. Structure and protein-protein interactions of human UDP-glucuronosyltransferases. Front Pharmacol. 2016;7:388. https://doi.org/10.3389/fphar.2016.00388.
Goel AK, Lundberg D, Torres MA, Matthews R, Akimoto-Tomiyama C, Farmer L, et al. The Pseudomonas syringae type III effector HopAM1 enhances virulence on water-stressed plants. Mol Plant Microbe Interact. 2008;21:361–70. https://doi.org/10.1094/Mpmi-21-3-0361.
Gong PJ, Zhang JH, Li HX, Yang CX, Zhang CJ, Zhang XH, et al. Transcriptional profiles of drought-responsive genes in modulating transcription signal transduction, and biochemical pathways in tomato. J Exp Bot. 2010;61:3563–75. https://doi.org/10.1093/jxb/erq167.
Guo A-Y. GSDS: a gene structure display server. Hereditas. 2007;29:1023–6. https://doi.org/10.1360/yc-007-1023.
Guo DD, Liu F, Tu YH, He BX, Gao Y, Guo ML. Expression patterns of three UGT genes in different chemotype safflower lines and under MeJA stimulus revealed their potential role in flavonoid biosynthesis. PLoS One. 2016;11:e0158159. https://doi.org/10.1371/journal.pone.0158159.
Herbert DA. Biotic Stress and Yield Loss. Crop Sci. 2002;42:656. https://doi.org/10.2135/cropsci2002.0656.
Horton P, Park KJ, Obayashi T, Fujita N, Harada H, Adams-Collier CJ, et al. WoLF PSORT: protein localization predictor. Nucleic Acids Res. 2007;35(1Web Server issue):585–7. https://doi.org/10.1093/nar/gkm259.
Hu YQ, Zhang MT, Lu MQ, Wu Y, Jing TT, Zhao MY, et al. Salicylic acid carboxyl glucosyltransferase UGT87E7 regulates disease resistance in Camellia sinensis. Plant Physiol. 2022;189:1507–20. https://doi.org/10.1093/plphys/kiac090.
Johnson M, Zaretskaya I, Raytselis Y, Merezhuk Y, McGinnis S, Madden TL. NCBI BLAST: a better web interface. Nucleic Acids Res. 2008;36(Web Server):W5–9.
Jumper J, Evans R, Pritzel A, Green T, Figurnov M, Ronneberger O, et al. Highly accurate protein structure prediction with AlphaFold. Nature. 2021;596:583–9. https://doi.org/10.1038/s41586-021-03819-2.
Karavina C, Gubba A. Detection and characterization of tomato spotted wilt virus infecting field and greenhouse-grown crops in Zimbabwe. Eur J Plant Pathol. 2017;149:933–44. https://doi.org/10.1007/s10658-017-1243-4.
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, et al. Circos: an information aesthetic for comparative genomics. Genome Res. 2009;19:1639–45. https://doi.org/10.1101/gr.092759.109.
Kumar S, Stecher G, Tamura K. MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol. 2016;33:1870–4. https://doi.org/10.1093/molbev/msw054.
Lairson LL, Henrissat B, Davies GJ, Withers SG. Glycosyltransferases: Structures, functions, and mechanisms. Annu Rev Biochem. 2008;77:521–55. https://doi.org/10.1146/annurev.biochem.76.061005.092322.
Langenbach C, Campe R, Schaffrath U, Goellner K, Conrath U. UDP-glucosyltransferase UGT84A2/BRT1 is required for Arabidopsis nonhost resistance to the Asian soybean rust pathogen Phakopsora pachyrhizi. New Phytol. 2013;198:536–45. https://doi.org/10.1111/nph.12155.
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, et al. Clustal W and clustal X version 2.0. Bioinformatics. 2007;23:2947–8. https://doi.org/10.1093/bioinformatics/btm404.
Letunic I, Bork P. 20 years of the SMART protein domain annotation resource. Nucleic Acids Res. 2018;46(D1):D493–D496. https://doi.org/10.1093/nar/gkx922.
Li P, Li YJ, Zhang FJ, Zhang GZ, Jiang XY, Yu HM, et al. The Arabidopsis UDP-glycosyltransferases UGT79B2 and UGT79B3, contribute to cold, salt and drought stress tolerance via modulating anthocyanin accumulation. Plant J. 2017b;89:85–103.
Li Q, Yu HM, Meng XF, Lin JS, Li YJ, Hou BK. Ectopic expression of glycosyltransferase UGT76E11 increases flavonoid accumulation and enhances abiotic stress tolerance in Arabidopsis. Plant Biol. 2018b;20:10–19.
Li JH, Su XX, Wang YL, Yang W, Pan Y, Su CG, et al. Genome-wide identification and expression analysis of the BTB domain-containing protein gene family in tomato. Genes Genomics. 2018a;40:1–15.
Li P, Li YJ, Wang B, Yu HM, Li Q, Hou BK. The Arabidopsis UGT87A2, a stress-inducible family 1 glycosyltransferase, is involved in the plant adaptation to abiotic stresses. Physiol Plant. 2017a;159:416–32.
Lin JS, Huang XX, Li Q, Cao YP, Bao Y, Meng XF, et al. UDP-glycosyltransferase 72B1 catalyzes the glucose conjugation of monolignols and is essential for the normal cell wall lignification in Arabidopsis thaliana. Plant J. 2016;88:26–42. https://doi.org/10.1111/tpj.13229.
Lis H, Sharon N. Protein glycosylation structural and functional aspects. Eur J Biochem. 1993;218:1–27. https://doi.org/10.1111/j.1432-1033.1993.tb18347.x.
Liu Q, Dong GR, Ma YQ, Zhao SM, Liu X, Li XK, et al. Rice glycosyltransferase gene UGT85E1 is involved in drought stress tolerance through enhancing abscisic acid response. Front Plant Sci. 2021;12:790195.
Louveau T, Leitao C, Green S, Hamiaux C, van der Rest B, Dechy-Cabaret O, et al. Predicting the substrate specificity of a glycosyltransferase implicated in the production of phenolic volatiles in tomato fruit. FEBS J. 2011;278:390–400. https://doi.org/10.1111/j.1742-4658.2010.07962.x.
Mackenzie PI, Owens IS, Burchell B, Bock KW, Bairoch A, Belanger A, et al. The UDP glycosyltransferase gene superfamily: Recommended nomenclature update based on evolutionary divergence. Pharmacogenetics. 1997;7:255–69. https://doi.org/10.1097/00008571-199708000-00001.
MamoonRehman H, Amjad Nawaz M, Bao L, Hussain Shah Z, Lee J-M, Ahmad MQ, et al. Genome-wide analysis of family-1 UDP-glycosyltransferases in soybean confirms their abundance and varied expression during seed development. J Plant Physiol. 2016;206:87–97. https://doi.org/10.1016/j.jplph.2016.08.017.
Mishra U, Rai A, Kumar R, Singh M, Pandey HP. Gene expression analysis of Solanum lycopersicum and Solanum habrochaites under drought conditions. Genom Data. 2016;9:40–1. https://doi.org/10.1016/j.gdata.2016.04.001.
Modolo LV, Li LN, Pan HY, Blount JW, Dixon RA, Wang XQ. Crystal structures of glycosyltransferase UGT78G1 reveal the molecular basis for glycosylation and deglycosylation of (iso) flavonoids. J Mol Biol. 2009;392:1292–302. https://doi.org/10.1016/j.jmb.2009.08.017.
Moore RC, Purugganan MD. The early stages of duplicate gene evolution. Proc Natl Acad Sci USA. 2003;100:15682–7. https://doi.org/10.1073/pnas.2535513100.
Mueller LA, Solow TH, Taylor N, Skwarecki B, Buels R, Binns J, et al. The SOL Genomics Network. a comparative resource for solanaceae biology and beyond. Plant Physiol. 2005;138:1310–7. https://doi.org/10.1104/pp.105.060707.
Osmani SA, Bak S, Moeller BL. ChemInform abstract: substrate specificity of plant UDP-dependent glycosyltransferases predicted from crystal structures and homology modeling. Chem Inform. 2009;40:325–47. https://doi.org/10.1002/chin.200928273.
Pervez MA, Ayub CM, Khan HA, Shahid MA, Ashraf I. Effect of drought stress on growth, yield and seed quality of tomato (Lycopersicon esculentum L.). Pak J Agric Sci. 2009;46:174–8.
Premathilake AT, Ni JB, Shen JQ, Bai SL, Teng YW. Transcriptome analysis provides new insights into the transcriptional regulation of methyl jasmonate-induced flavonoid biosynthesis in pear calli. BMC Plant Biol. 2020;20:388. https://doi.org/10.1186/s12870-020-02606-x.
Rahimi S, Kim J, Mijakovic I, Jung KH, Choi G, Kim SC, et al. Triterpenoid-biosynthetic UDP-glycosyltransferases from plants. Biotechnol Adv. 2019;37:107394. https://doi.org/10.1016/j.biotechadv.2019.04.016.
Rguez S, Djebali N, Ben Slimene I, Abid G, Hammemi M, Chenenaoui S, et al. Cupressus sempervirens essential oils and their major compounds successfully control postharvest grey mould disease of tomato. Ind Crop Prod. 2018;123:135–41. https://doi.org/10.1016/j.indcrop.2018.06.060.
Rombauts S, Dehais P, Van Montagu M, Rouze P. PlantCARE, a plant cis-acting regulatory element database. Nucleic Acids Res. 1999;27:295–6. https://doi.org/10.1093/nar/27.1.295.
Ross J, Li Y, Lim EK, Bowles DJ. Higher plant glycosyltransferases. Genome Biol. 2001;2:REVIEWS3004. https://doi.org/10.1186/gb-2001-2-2-reviews3004.
Shao H, He XZ, Achnine L, Blount JW, Dixon RA, Wang XQ. Crystal structures of a multifunctional triterpene/flavonoid glycosyltransferase from Medicago truncatula. Plant Cell. 2005;17:3141–54. https://doi.org/10.1105/tpc.105.035055.
Song JW, Shang LL, Li CX, Wang WQ, Wang X, Zhang CL, et al. Variation in the fruit development gene POINTED TIP regulates protuberance of tomato fruit tip. Nat Commun. 2022;13:5940. https://doi.org/10.1038/s41467-022-33648-4.
Sun W, Xu XN, Zhu HS, Liu AH, Liu L, Li JM, et al. Comparative transcriptomic profiling of a salt-tolerant wild tomato species and a salt-sensitive tomato cultivar. Plant Cell Physiol. 2010;51:997–1006. https://doi.org/10.1093/pcp/pcq056.
Sun YF, Ji K, Liang B, Du YW, Jiang L, Wang J, et al. Suppressing ABA uridine diphosphate glucosyltransferase (SlUGT75C1) alters fruit ripening and the stress response in tomato. Plant J. 2017;91:574–89. https://doi.org/10.1111/tpj.13588.
Sussman JL, Lin DW, Jiang JS, Manning NO, Prilusky J, Ritter O, et al. Protein Data Bank (PDB): database of three-dimensional structural information of biological macromolecules. Acta Crystallogr D. 1998;54:1078–84. https://doi.org/10.1107/S0907444998009378.
Suzuki N, Rivero RM, Shulaev V, Blumwald E, Mittler R. Abiotic and biotic stress combinations. New Phytol. 2014;203:32–43. https://doi.org/10.1111/nph.12797.
Verma V, Ravindran P, Kumar PP. Plant hormone-mediated regulation of stress responses. BMC Plant Biol. 2016;16:86.
Vogt T, Jones P. Glycosyltransferases in plant natural product synthesis: characterization of a supergene family. Trends Plant Sci. 2000;5:380–6. https://doi.org/10.1016/S1360-1385(00)01720-9.
Wang XQ. Structure, mechanism and engineering of plant natural product glycosyltransferases. FEBS Lett. 2009;583:3303–9. https://doi.org/10.1016/j.febslet.2009.09.042.
Wang D, Zhang Y, Zhang Z, Zhu J, Yu J. KaKs_Calculator 2.0: a toolkit incorporating gamma-series methods and sliding window strategies. Genomics Proteomics Bioinformatics. 2010;8:77–80. https://doi.org/10.1016/s1672-0229(10)60008-3.
Wang L, Zhou Y, Ding Y, Chen C, Chen X, Su N, et al. Novel flavin-containing monooxygenase protein FMO1 interacts with CAT2 to negatively regulate drought tolerance through ROS homeostasis and ABA signaling pathway in tomato. Hort Res. 2023;10:uhad037. https://doi.org/10.1093/hr/uhad037.
Williamson B, Tudzynsk B, Tudzynski P, van Kan JAL. Botrytis cinerea: the cause of grey mould disease. Mol Plant Pathol. 2007;8:561–80. https://doi.org/10.1111/J.1364-3703.2007.00417.X.
Yonekura-Sakakibara K, Hanada K. An evolutionary view of functional diversity in family 1 glycosyltransferases. Plant J. 2011;66:182–93. https://doi.org/10.1111/j.1365-313X.2011.04493.x.
Zhang PF, Senge M, Dai YY. Effects of salinity stress at different growth stages on tomato growth, yield, and water-use efficiency. Commun Soil Sci Plant Anal. 2017;48:624–34. https://doi.org/10.1080/00103624.2016.1269803.
Zhang K, Sun Y, Li MN, Long RC. CrUGT87A1, a UDP-sugar glycosyltransferases (UGTs) gene from Carex rigescens, increases salt tolerance by accumulating flavonoids for antioxidation in Arabidopsis thaliana. Plant Physiol Bioch. 2021;159:28–36. https://doi.org/10.1016/j.plaphy.2020.12.006.
Zhao MY, Zhang N, Gao T, Jin JY, Jing TT, Wang JM, et al. Sesquiterpene glucosylation mediated by glucosyltransferase UGT91Q2 is involved in the modulation of cold stress tolerance in tea plants. New Phytol. 2020;226:362–72. https://doi.org/10.1111/nph.16364.
Acknowledgments
Not applicable.
Funding
This work was supported by the National Natural Science Foundation of China (No. 31872123), Graduate Student Innovation Program of Southwest University (SWUS23103), the Fundamental Research Funds for the Central Universities (XDJK2020B060) and the Special Key Project of Technological Innovation and Application Development of Chongqing (No. CSTB2023TIAD-KPX0026).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation and experiments were performed by LQ, XC1, YF, LW, HZ, XC2 and YD. Data collection and analysis were performed by LQ, XC1, HH, XH, QY, HM and JL. The first draft of the manuscript was written by JL, LQ, XC1, HH and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
Authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: the funding section is corrected.
Supplementary Information
Additional file 1:
Supplementary Figure S1. Solanum lycopersicum uridine diphosphate glycosyltransferases (SlUDPGTs) gene structure.
Additional file 2:
Supplementary Figure S2. Protein tertiary structures of partial members of uridine diphosphate glycosyltransferase (UDPGT) gene family in tomato.
Additional file 3:
Supplementary Figure S3. Location of the cis-acting element in each Solanum lycopersicum uridine diphosphate glycosyltransferase (SlUDPGTs) gene promoter.
Additional file 4
: Supplementary Figure S4. Expression patterns of Solanum lycopersicum uridine diphosphate glycosyltransferases (SlUDPGTs) in 10 tissues/stages.
Additional file 5:
Supplementary Table S1. List of physicochemical properties and subcellular localization of Solanum lycopersicum uridine diphosphate glycosyltransferases (SlUDPGTs) identified in tomato. Supplementary Table S2. List of primers used for qRT-PCR and knockout target site of Solanum lycopersicum uridine diphosphate glycosyltransferase 52 (SlUDPGT52). Supplementary Table S3. KaKs of all Solanum lycopersicum uridine diphosphate glycosyltransferases (SlUDPGTs) tandem repeat gene pairs. Supplementary Table S4. Cis-elements identified in the promoters of Solanum lycopersicum uridine diphosphate glycosyltransferases (SlUDPGTs). Supplementary Table S5. Cis-elements identified in the promoters of more than 20 Solanum lycopersicum uridine diphosphate glycosyltransferases (SlUDPGTs). Supplementary Table S6. Information of probe sets used for microarray expression analysis.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qiu, L., Chen, X., Hou, H. et al. Genome-wide characterization of the tomato UDP-glycosyltransferase gene family and functional identification of SlUDPGT52 in drought tolerance. HORTIC. ADV. 1, 14 (2023). https://doi.org/10.1007/s44281-023-00016-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44281-023-00016-1