Abstract
The early detection of colorectal polyps is crucial for the reduction of mortality rates. However, manually identifying polyps is time-consuming and expensive, increasing the risk of missing them. Our paper aims to address this issue by presenting an automated segmentation approach for colorectal polyps. This paper proposes a method that combines a skip connection with hybrid attention guidance (AG) using attention guidance (AG) and residual path frameworks to identify salient features. Furthermore, we augment test samples using original, horizontal flip, and vertical flip transformations to enhance model robustness through Test Time Augmentation (TTA). The model was trained with Kvasir-seg samples and evaluated on Kvasir-seg and CVC-ClinicDB datasets to gauge generalizability. A significant accuracy (0.9546), a Dice Similarity Coefficient (DSC) of 0.8557, a Cross-section over Union (IoU) of 0.8824, a Recall (0.8221), a Precision (0.8922), an area under Receiver Operating Characteristics (ROC-AUC) of 0.9454, and an area under Precision-Recall (AUC-PR) of 0.8717 were achieved without TTA. Through TTA integration, accuracy (0.9993), DSC (0.8663), IoU (0.8277), Recall (0.8060), Precision (0.9364), and ROC-AUC (0.9587) have been improved. A comparison of our framework with state-of-the-art models demonstrated its effectiveness and segmentation capabilities. Additionally, the proposed model contains only 0.47 million parameters and a weight size of 6.71 MB, illustrating its potential for clinical diagnostics. A computer-aided diagnosis (CAD) system improves patient outcomes by detecting colorectal polyps early and improving segmentation accuracy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
There was an increase in colorectal cancer mortality rates in 2020, with 0.93 million deaths reported out of 1.9 million new cases. Across the globe, this is a major issue that both affects men and women [1]. The abnormal proliferation of glandular tissue within the colon mucosa is the cause of this type of cancer [2]. These excessive tissue formations, known as polyps, can cause physiological complications. A minimally invasive technique called virtual colonoscopy can be used to detect and treat colon lesions accurately in response to this problem [3]. It is possible to remove polyps promptly using this method, therefore helping to prevent colon cancer. It is often necessary to hire a highly skilled endoscopist to perform precise colonoscopy examinations, which are both time-consuming and resource-intensive. Approximately 25% of polyp areas are missed during colonoscopies, increasing cancer risk [4]. This alarming statistic is highlighted in a recent global study. This problem is prompting endoscopists to use CAD systems to assist them in identifying polyps during endoscopic procedures. Polyps are less likely to be overlooked when CAD is used [5].
The term polyp refers to an abnormal protrusion of tissue on the surface of the body, which commonly occurs on mucous membranes [6]. In most cases, they appear as compact clusters or minuscule formations that resemble mushrooms. Although polyps in the colon are most common, they can also appear in the cervix, uterus, ear canal, stomach, nose, or throat. Initially regarded as benign or innocuous, these growths have the potential to develop into invasive cancer over time, leading to potentially lethal outcomes if not detected in time. In the beginning, they may appear as benign adenomas on the gastrointestinal tract or rectum, but they gradually develop into cancer over time. Additionally, feces can blend seamlessly with the surrounding mucosa, enhancing the complexity of the background. Medical professionals remain optimistic and progressive about the development of such technologies despite these challenges.
Routine colon and rectal screening and examination is the most effective method of preventing colorectal cancer (CRC) because polyps may remain asymptomatic for a long time. Therefore, colonoscopies are routinely used to identify precancerous polyps. It is difficult to distinguish between adenomatous and hyperplastic polyps with colonoscopy alone, so urologists must examine colon biopsies under the microscope. Colonoscopy, a popular variant of endoscopy, provides the most comprehensive view of the entire GI tract, including the rectum, esophagus, stomach, and duodenum. There are two types of endoscopy: Gastroscopy, upper endoscopy, colonoscopy, or lower endoscopy. A flexible, thin tube called an endoscope is inserted into the colon during a non-invasive colonoscopy. An endoscope allows you to examine inside organs without making large incisions due to its small size and powerful light source. In this procedure, the colon, large intestine, and rectum are evaluated via the rectum.
Computer-aided diagnosis (CAD) systems for medical applications are challenging. The difficulty of acquiring well-labeled, precise, and accurate medical datasets to support training and testing is one of these challenges. In addition, polyps are complex in nature, making the task even more difficult. Based on its color, size, shape, and type, a polyp can be classified as an adenoma, hyperplastic, serrated, or mixed type. There are significant limitations associated with CAD systems for colonoscopy, despite advancements. Various polyp appearances can make existing systems difficult to diagnose, which can lead to incorrect diagnoses. Additionally, the effectiveness of these systems is heavily dependent on the quality and diversity of the datasets used for training, which are often limited due to privacy concerns and the rarity of certain polyp types. High-resolution endoscopic images also pose a computational challenge, requiring a balance between accuracy and efficiency. In response to these challenges, this research aims to develop a CAD system that overcomes these limitations. Our research focuses on enhancing polyp detection accuracy using improved deep learning models, incorporating novel data augmentation techniques, and optimizing computational efficiency for practical use in real-time colonoscopy. Ultimately, the goal is to reduce the polyp miss-rate and contribute to early and accurate diagnosis of colorectal cancer, improving patient outcomes and reducing mortality.
In the medical imaging domain, deep learning techniques have largely contributed to recent advances in computer vision. There has been a rise in the number of deep learning models specialized for semantic segmentation [7,8,9,10,11]. These developments lead to notable architectures like UNet [12], Attention-UNet [13], ResUNet [14], and MultiResUNet [15]. Several downsampling phases are used in these networks in order to produce maps of essential features.
This study integrates attention guidance (AG) and residual connections within the encoder-decoder layer. Using the innovative method, vital characteristics can be extracted without the need for external supervision. In addition, it prevents redundant feature maps from being generated at different scales, resulting in an accurate representation of features.
The following contributions were made to this study:
-
1.
In this study, we propose a hybrid residual skip network, MultiResUNet + AG, that extracts relevant features from input samples using an attention mechanism. The proposed method maintains a compact size and requires fewer parameters.
-
2.
The Kvasir-SEG dataset is preprocessed using a hybrid of twenty-six different preprocessing methods to improve the model's training capabilities. The generalizability of the model was tested using the CVC-ClinicDB dataset.
-
3.
A post-processing technique is incorporated into the model to enhance segmentation capabilities and predictive power. Based on the original, horizontal, and vertical flip methodologies, we averaged our TTA operation.
-
4.
Various performance metrics were evaluated for both datasets, including DSC, IoU, precision, recall, ROC-AUC, and AUC-PR. The proposed model performed better than other existing methods.
This research work describes the methodology, including the dataset and preprocessing procedures in Sect. 2. The proposed model is examined experimentally and quantified on both datasets in Sects. 3 and 4. In Sects. 5 and 6, a predictive analysis and a comparative analysis will be presented. The conclusion of the research is presented in Sect. 7.
2 Related Work
CAD systems have been used by many researchers to develop an automated procedure for identifying polyps. Many researchers trained their models using the Kvasir-SEG dataset, while others worked on post-processing techniques to make the models robust [2, 16,17,18,19,20,21,22,23,24,25,26]. The segmentation of endoscopic images based on semantic information has been extensively studied in medical imaging [27]. Previous research used manual descriptions (handcrafted features) to describe learning features. According to the samples' color and shape, a machine learning (ML) classifier was used to detect the lesion area [28,29,30,31,32,33,34]. However, those methods did not produce good results when it came to extracting the features. As a result, convolutional techniques were applied to improve it. To advance automatic detection, Wang et al. [34] used edge cross-section methods, while Hwang et al. [35] used elliptical shape boundary extraction techniques. Through the inclusion of global geometric constraints in the model, Tajbakhsh et al. [36] were able to increase the model's detection rate by focusing on local intensity characteristics. Their efforts, however, were limited by poor predictive accuracy and limited presentation capabilities. Moreover, both low-level and high-level polyps had difficulty being identified by the model.
Jha et al. [37] developed NanoNet for segmenting video capsule colonoscopy samples using an encoder-decoder architecture. In addition to video capsule endoscopy and conventional colonoscopy images with polyps, NanoNet was tested on a variety of datasets. Model evaluation results showed that parameters count, and performance metrics were effectively balanced. Thus, this approach achieved high levels of accuracy (0.9456), DSC (0.8227), IoU (0.7282), recall (0.8588), and precision (0.8367). A Multi-Layer Perceptron (MLP) stage is incorporated in the latent domain of the UNeXt architecture by Valanarasu et al. [38]. The MLPs preserved the essence of the representation, utilizing the tokenized MLP blocks as the main innovation. Convolutional features were tokenized and projected delicately in this block. A channel-shifting approach was also proposed to fine-tune input channel designs for MLPs, a significant advance that placed greater emphasis on local dependency learning. This method has therefore increased its overall precision to 0.9043. Based on a CNN, Bernal et al. [39] proposed a method for segmenting polyps. However, this strategy relied heavily on handcrafted methodologies, which ultimately yielded suboptimal results. As a result of difficulties acquiring spatial information from the provided data, the model was limited in its ability to identify patterns. A new convolutional architecture called HarDNet-MSEG was developed by Huang et al. [24] specifically for segmenting polyps. The model achieved a Dice score of 0.904 when tested on the Kvasir-SEG dataset.
Data augmentation was initially designed to improve the training of the model [40,41,42,43]. A relatively modest dataset is expanded by applying transformations to original data, resulting in the generation of new instances that improve performance [41]. A variety of augmentation techniques, such as flipping, shifting, and resizing, are used in image processing to enhance the training process. Multiple types of research have demonstrated that combining predictions from modified versions of test samples improves performance. Matsunaga et al. [42] used computational methods to alter test samples to ensure accurate skin lesion classification. Jin et al. [43] used rotation and translation techniques to detect nodules in pulmonary tissues. Although these efforts included TTA as a practical approach, no comprehensive formulation was provided. In addition, this method could only be used for segmentation tasks requiring uncertainty estimation.
3 Methodology
The framework presented in Fig. 1 outlines the proposed approach. Initially, the Kvasir-SEG dataset is divided into training (80%), validation (10%), and testing (10%) subsets. Afterward, the training data is processed using different data preprocessing techniques, thereby expanding the training samples. In the following steps, the model is trained using the proposed MultiResUNet + AG architecture. After training, the model's weights are leveraged to generate segmented masks with the TTA and without the TTA approach.
3.1 Dataset and Pre-processing
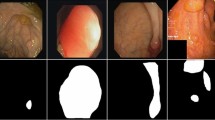
The model proposed in this study was trained with data from Kvasir-SEG [44]. The data set contains 1000 high-resolution endoscopic images, ranging from 332 × 487 to 1920 × 1072 pixels each. A physician meticulously annotated these images, which were then reviewed by an experienced gastroenterologist. Segmentation masks generated in this process are binary masks that indicate which pixels comprise the polyp. In the training strategy, 80% of images were chosen (800 images), 10% for model validation (100 images), and 10% for model assessment (100 images). This test set contains 100 images with broader, more compact, and flat polyp categories, which make up 10% of the dataset. Incorporating polyps enhances the model's learning and segmentation capabilities. Further testing was conducted using the CVC-ClinicDB dataset [45]. This dataset includes 612 images with a resolution of 384 × 288 pixels, extracted from 31 colonoscopy sequences. It features multiple examples of lesions. Similarly, a 10% test set (62 images) was chosen from this sample to test the proposed system. Figure 2 presents images of both datasets.
The pre-processing of images plays an important role in optimizing model training. Pre-processing ensures that models are not affected by variations in image quality by normalizing and reducing noise. All the images were resized to 256 × 256 pixels. Images are normalized concurrently. The training samples were increased from 800 to 20,800 using hybrid pre-processing with twenty-six augmentation techniques. In Fig. 3, the images of the overall processing are shown. Mask operations are essential for polyp segmentation, since they isolate polyps from surrounding tissue in colonoscopy images, providing a clear target for segmentation models. The precise demarcation of polyps is critical for accurate polyp detection, which can greatly influence automated diagnostic systems. By employing masks, we can simulate various polyp appearances, which is crucial for building robust models that can handle different clinical scenarios. In addition, masks reduce input dimensionality, preventing overfitting by reducing the amount of irrelevant data that passes through models. In medical imaging, masks are also effective at protecting patient privacy by obscuring features outside the area of interest. In addition to preprocessing, these operations enhance the overall effectiveness of diagnostic techniques by allowing for more precise analysis.
Pre-processed images include a center crop, b crop, c random crop, d random 90-degree rotation, e transpose, f elastic transformation, g grid distortion, h optical distortion, i vertical flip, j horizontal flip, k grayscale conversion, l grayscale vertical flip, m grayscale horizontal flip, n grayscale center crop, o random brightness contrast, p random gamma, q hue saturation, r RGB shifting, s random brightness, t random contrast, u motion blur, v median blur, w gaussian blur, x gaussian noise, y channel shuffle, and z coarse dropout
3.2 MultiRes Block
Analyzing building features in aerial images from different sources requires consideration of different scenarios and contextual sizes. It is necessary to analyze multiple images in order to extract the most relevant features. The initial design of the UNet model included a 3 × 3 kernel with a pair of convolutional layers (CLs) followed by a max-pooling layer (MP) operation [46]. To enhance the multiresolution computation, different kernel sizes were used (3 × 3, 5 × 5, and 7 × 7) as parallel CLs (see Fig. 4a). A method inspired by the inception technique was created as a result. As a result of these modifications, the UNet model was able to rapidly recognize visual characteristics and understand a wide range of contextual factors. To accomplish this, the convolutional process was replaced with inception blocks. By using extra CLs, performance was improved, but memory requirements increased. To overcome this limitation, an innovative method was employed to cluster lightweight 3 × 3 CLs and deconstruct larger kernels of 5 × 5 and 7 × 7 CLs (as seen in Fig. 4b). The combination of these results resulted in scale-dependent spatial characteristics. The 5 × 5 and 7 × 7 CLs were estimated using 3 × 3 convolutional techniques. The filters were not constant throughout each CL in the first three iterations to prevent earlier layers from impacting later ones in terms of memory requirements. Moreover, a 1 × 1 kernel was used as a residual link to extract spatial details [47]. The complete design MultiRes-block is illustrated in Fig. 4c.
3.3 Residual Path
The UNet architecture includes a skip link between the encoder and decoder layers to prevent spatial features from being lost during pooling. There are, however, obstacles to this skip link. Encoder layers compute the most essential characteristics, while decoder layers add more complex characteristics. Consequently, the combined features lose some of their semantic meaning, affecting forecast accuracy. A number of convolutional operations were performed along the skip connections and an integrated residual link was used instead of a standard convolutional procedure to resolve this issue. The difference in meaning between encoder and decoder characteristics was reduced by this modification. The "Res path" skips the convolutional stages and uses a 3 × 3 filter and 1 × 1 filter with residual connections as input. The residual path connection between the encoder and decoder block is shown in Fig. 5.
3.4 Attention Guidance (AG)
The use of attention mechanisms (AG) helps the model capture complex patterns, nuanced details and attributes specific to the context while learning features. By filtering out distractions, the model can focus on the most important areas for completing the task successfully. A diagram of the overall attention-gate mechanism is shown in Fig. 6 [13].
Proposed attention gate mechanism where decoder takes the input features from skip \({\mathrm{^{\prime}}{\text{x}}}^{{\text{l}}}{\prime}\), and this value is modified by utilizing the coefficients of \({\prime}\mathrm{\alpha {\prime}}\). By combining visual context \(\mathrm{^{\prime}}\mathrm{g{\prime}}\) and activation functions extracted the relevant spatial information
In this approach, a transformation is carried out by utilizing a 1 × 1 × 1 convolution, denoted as \({\prime}{W}_{g}{\prime}\). The convolution process modifies the gating signal from the up-sample layer \({\prime}g{\prime}\). This convolution aims to pinpoint the exact region of interest and furnish a broader array of contextually relevant details. For the precise representation of the spatial attributes within the skip-connection layer \({\prime}{x}_{l}{\prime}\) originating from the encoder, executing this procedure in tandem with the linear transformation denoted as \({\prime}{W}_{x}{\prime}\) assumes significance. This combined operation yields two intermediate vectors: \({\prime}{F}_{l}{\prime}\) and \({\prime}{F}_{g}{\prime}\)emerging from a distinct linear transformation \({\prime}{W}_{x}{\prime}\) and \({\prime}{W}_{g}{\prime}\). Furthermore, the addition is performed on both vectors \({\prime}{F}_{l}{\prime}\) and \({\prime}{F}_{g}{\prime}\), simultaneously. This additive result is passed through the non-linear activation function (ReLU) and 1 × 1 × 1 convolution to attain the attention coefficients. This coefficients value is varied from \(-\infty\) to \(+\infty\). Sigmoid activation function is used to range this score in between 0 to 1. A bilinear up-sampling operation is used to resample the coefficients to perfectly match the original input size \({\prime}{x}_{l}{\prime}\) and elementwise multiplication is performed to get the final output \(\widehat{{x}^{l}}\). The coefficients ‘α’ are determined channel-by-channel, and their values are adjusted during the reverse propagation phase of the network. This fine-tuning is crucial for determining which signals can be ignored and which ones need special focus.
3.5 MultiResUNet Architecture
Within the MultiResUNet architecture, a series of MultiRes blocks were integrated, each consisting of a linear sequence of three connected CLs (see Fig. 4). The count of filter sizes within the MultiRes block is meticulously regulated. This quantification, labelled as \({\prime}W{\prime}\), is determined through a specific formula:
The term \({\prime}num\_filters{\prime}\) refers to the sizes of the filters assigned to each CL, and \({\prime}alpha{\prime}\) is the coefficient that governs these values. The crucial aspect here is regulated filter sizing, a technique used to compare and control filters within the basic UNet framework. The filter weights were distributed in standard sizes, which include the conventional stages of UNet, such as 32, 64, 128, 256, and 512. By assigning an \(alpha\) value of 1.67, \(W\) was computed based on trial and error [15]. As per Eq. 1, it is apparent that the resulting filter size will be smaller compared to the UNet model. Additionally, although maintaining uniformity, the filter size for each CL within the block must undergo variations. Subsequently, the values for the three consecutive CLs are adjusted as \(\frac{W}{6}\), \(\frac{W}{3}\), and \(\frac{W}{2}\), respectively. Section 3.2 provides a detailed explanation of the MultiRes block's intricacies. Following this, the conventional skip connection is replaced with a Residual path accompanied by an attention mechanism to achieve a more precise fusion of encoder and decoder characteristics. Sections 3.3 and 3.4 elaborate on the specifics of the residual and AG paths, respectively. This strategic approach mitigated the semantic disparity between the encoder and decoder feature maps. CLs within this res path decreased to 4, 3, 2, and 1. In addition, the residual blocks included filter sizes of 32, 64, 128, and 256. ReLU and Batch Normalization (BN) were integrated to improve the suggested block architecture [48]. Figure 7 depicts a visualization of the proposed architecture, replete with appropriate annotations.
3.6 Post-Processing Technique
Colorectal polyp segmentation requires post-processing techniques to improve accuracy and robustness. One of these approaches is TTA, which has gained prominence in recent years [49]. Multiple augmented variants of the input test images are produced by TTA by applying multiple augmentations to the input test images. Based on these augmented images and the original image, the segmentation model is applied to generate predictions. The horizontal and vertical flips are commonly used in colorectal polyp segmentation to provide variations in orientation which enhance the model's ability to handle diverse polyp shapes and directions in real-world scenarios. As a result of combining predictions from original and augmented images, the TTA method provides a more comprehensive and robust evaluation of the model's segmentation capabilities. The strategy significantly improves the model's generalization capability and overall segmentation performance, making it a valuable tool for accurate and reliable colorectal polyp segmentation. Figure 8 shows the design of the TTA approach with the proposed model.
4 Experimental Settings and Performance Metrics
The results of our study are presented in this section, which is divided into two categories: quantitative and predictive. The proposed methodology was applied to the Kvasir-Seg dataset for training. During the training process, we used the Adam optimizer with 0.0001 learning rate and 32 batches with 50 epochs. As a result of the heuristic approach, a threshold level of 0.5 was chosen for prediction. The program was implemented using Python and the TensorFlow framework. We used Windows 10 Pro, 32 GB of RAM, and NVIDIA GeForce RTX 2080 Ti 11 GB GPUs for our experiment. In the end, a compact model of 6.71 MB was produced after the model was trained with 0.47 million parameters. A MultiResUNet + AG model processed each sample between 200 and 1000 ms during the testing phase. A MultiResUNet + AG + TTA instance takes around 200–1200 ms.
There are a variety of evaluation aspects included in the performance metrics [50].
A Dice Similarity Coefficient (DSC) measures the similarity of actual segments to predicted segments. It is calculated by dividing twice the number of true positives by the sum of twice the number of true positives, false positives, and false negatives. The metric combines precision and recall (see Eq. 5) into one. The intersection over union (IoU) measures the concordance between predicted and actual truth masks. Essentially, it is the overlap between the predicted and ground truth areas divided by the union between them. IoU does not directly use True Negatives (TN) since it focuses on the positive class—i.e. the object being detected. Typically, "negative" spaces are not taken into account in object detection since they are not bounded (see Eq. 4). The precision is determined by the proportion of accurate positive predictions compared to the total number of positive predictions. It is calculated by dividing the number of true positives by the total number of positive predictions made by the model (the sum of true positives and false positives). Last but not least, recall measures whether a model captures relevant characteristics by dividing true positive predictions by actual positive observations. In Eq. 3, the number of true positives is divided by the total number of positives (the sum of true positives and false negatives). In addition,
-
True Positive (TP): It is an outcome where the model correctly predicts a positive class. As a result, the model identifies positive instances, and they are positive.
-
True Negative (TN): This is an outcome in which the model correctly predicts the negative class. The model identifies an instance as negative, and it is indeed negative.
-
False Positive (FP): In this case, the model incorrectly predicts the positive class. Models identify instances as positive when they are negative. The error is also known as a "Type I error."
-
False Negative (FN): In this case, the model incorrectly predicts the negative class. Models identify instances as negative, but, they are positive. The error is also referred to as a Type II error.
Using Binary Cross-Entropy as the loss function, the proposed model [51] was developed. A single observation's BCE loss is calculated by a negative average of two terms: the first term is the actual label times the logarithm of the predicted probability, and the second term is one minus the actual label times the logarithm of one minus the predicted probability (see Eq. 6).
Based on these metrics, the formulas are as follows:
where, positive class defined as \(p\) and \(y\) denotes the original label.
5 Quantitative Results
Figures 9 and 10 show the performance curves for the corresponding model training times. Both curves were generated from training samples and validation samples. At 50 epochs, the loss score was 0.1319 on the training set and 0.1426 on the validation set. Moreover, the dice performance score at the model was 0.9354 on training and 0.8939 on validation. We saved the trained model's weight for testing purposes.
5.1 Testing on Kvasir-SEG Data
Multiple performance metrics are compared against various segmentation methods on the Kvasir-SEG dataset in Table 1. With respect to the UNet model, it had the lowest accuracy (0.9382) and precision (0.8756), whereas its segmentation metrics, specifically DSC and IoU, were relatively modest (0.7721 and 0.7792). There was a significant improvement in the Attention-UNet and ResUNet models, particularly in terms of DSC, IoU, and recall. A DSC score of 0.8009, an IoU score of 0.7938, a recall score of 0.7758, and a precision score of 0.8956 were achieved for the Attention-UNet model. Based on this trend, MultiResUNet achieved higher DSC, IoU, and precision than its predecessors. In addition, the MultiResUNet + AG model effectively leveraged Attention-Guided features to achieve remarkable recall (0.8221, the highest) and DSC (0.8557, the second best), with a slight precision dip (0.8922, the third best). MultiResUNet + AG + TTA, a combination of attention-guided architecture and TTA, secured peak scores across all metrics, including accuracy (0.9593), DSC (0.8663), IoU (0.8277), and precision (0.9364). Thus, architectural advancements have a cumulative effect on image segmentation's accuracy, enhancing its utility in real-world situations. In comparison with the UNet baseline, the MultiResUNet + AG + TTA model offers significant technical advances. As a result of architectural enhancements such as Attention Gates (AG) and Test-Time Augmentation (TTA), training time is reduced by approximately 64.96 percent. As a result of this efficiency, model development and fine-tuning can be accelerated. Furthermore, MultiResUNet + AG + TTA achieves a remarkable improvement in frames per second (FPS) when compared to UNet, making it the ideal solution for real-time navigation and object tracking. The model's architectural innovations and optimization strategies enable precision and speed, resulting in these improvements. In Fig. 11, various model performances are illustrated in a bar chart.
5.2 Cross Testing Performance
Second, the proposed model was tested on the CVC-ClinicDB test set to check its versatility. Table 2 provides insights into the performance of various methods based on diverse metrics. Starting with the UNet model, it showed the lowest accuracy, precision, DSC, and IoU scores. There are noticeable improvements in the DSC (0.5340 and 0.5058), IoU (0.6373 and 0.6173), and precision (0.6575 and 0.5744) of the Attention-UNet and ResUNet models, which indicate improvements in segmentation quality. As a result of further refinement of the results, the MultiResUNet model achieves competitive DSC, IoU, and precision values. The proposed MultiResUNet + AG model has demonstrated an improvement in DSC, IoU, and Recall scores of 6.13%, 1.54%, and 20.48%.
Lastly, the MultiResUNet + AG + TTA model emerged as the most robust contender, achieving the highest accuracy of 0.931 and precision of 0.7712, while significantly elevating DSC of 0.6119 and IoU of 0.6691. As a result of this collective performance, the algorithm is able to achieve precise image segmentation on a variety of datasets. Compared to UNet, MultiResUNet + AG, and MultiResUNet + AG + TTA demonstrate substantial improvements in real-time processing. The UNet operates at a modest 11 frames per second, which, while reasonable, can be limiting for applications requiring real-time performance. A significant improvement is achieved with MultiResUNet + AG, which achieves a speed of 34 frames per second, whereas MultiResUNet + AG + TTA achieves a speed of 41 frames per second. The bar chart in Fig. 12 illustrates the cross-validation of the proposed models.
6 Prediction
Based on the Kvasir-seg test samples, Fig. 13 presents the models' predictions. Testing samples were selected based on their size and shape—tiny, large, and flat polyps. This is significant since it validates the proposed model's detection behavior under different worst-case scenarios. The proposed MultiResUNet + AG (with and without TTA) accurately identified polyp regions when a flat type of polyp was used for segmentation (as shown in the first row of Fig. 13). Additionally, the same model was able to identify larger polyps (second-row Fig. 13) more accurately than existing models. As a result of the TTA approach, the lesion area was accurately detected. In addition, all models without UNet were able to detect a comparatively medium-sized and saturated polyp (third row, Fig. 13).
The MultiResUNet + AG model's generalizability was assessed by generating predicted masks for the CVC-ClinicDB dataset (Fig. 14). In light of the inherent variability in image samples and polyps, pinpoint accuracy in delineating polyp areas is challenging for any model. The MultiResUNet + AG model showed commendable performance, with predicted regions closely matching ground truth masks. In addition, TTA enhanced the model's performance beyond the baseline due to the collaborative effects of the augmentation ensemble.
6.1 ROC-AUC Performances
To evaluate the performance of different segmentation models, the Kvasir-SEG dataset was used to construct the Receiver Operating Characteristic Area Under the Curve (ROC-AUC). The area scores of each model provide insight into the model's ability to differentiate positive instances from negative ones. Specifically, the UNet model scored 0.9270, while the Attention-UNet model scored 0.9355, and the ResUNet model scored 0.9274. In the MultiResUNet model, the area score was 0.9381, indicating a slight improvement in segmentation. A remarkable highlight of the proposed MultiResUNet + AG model was its area score of 0.9454. The achievement exemplifies the enhanced discriminative ability enabled by Attention-Guided (AG) features. MultiResUNet + AG + TTA displayed an exceptional area score of 0.9587, which was the model with the best performance. With this remarkable result, the model has the potential to significantly improve image segmentation accuracy. It is a compelling solution for precise image analysis tasks within the Kvasir-SEG dataset due to the implementation of Attention-Guided features and Test-Time Augmentation.
As seen in Fig. 15b, the ROC-AUC results of cross-testing on the CVC-ClinicDB dataset show a reasonable fit. With an area coverage of 86.86%, MultiResUNet + AG and TTA performed exceptionally well in predicting unseen data. In the CVC-ClinicDB dataset, the area scores of each model demonstrated their ability to distinguish between positive and negative examples. A UNet model scored 0.6649, while an Attention-UNet model scored 0.8056, a significant boost. ResUNet and MultiResUNet also showed improvements in segmentation quality, with area scores of 0.7624 and 0.7946, respectively. By incorporating Attention-Guided features, MultiResUNet + AG was able to achieve an area score of 0.7590, indicating that it has the potential to enhance segmentation accuracy. Nevertheless, MultiResUNet + AG + TTA achieved an outstanding area score of 0.8686, demonstrating their synergistic effect.
6.2 AUC-PR Performances
Figure 16a presents the area under cover on precision recall (AUC-PR) scores attained by the trained models for Kvasir-SEG. The UNet model scored 0.8282, while the Attention-UNet and ResUNet achieved 0.8540 and 0.8452, respectively. However, the MultiResUNet model performed even better, scoring 0.8678, indicating improved segmentation performance. The most significant improvement was observed in the MultiResUNet + AG model, which incorporated Attention-Guided features and achieved an impressive score of 0.8717. This highlights the substantial progress that can be made by integrating attention mechanisms. With the integration of TTA, the MultiResUNet + AG + TTA model reached the highest level of performance with an outstanding score of 0.8871. This combination of AG + TTA significantly refines image segmentation outcomes. Figure 16b displays the AUC-PR performance curve for the CVC-ClinicDB set. Once again, the proposed model outperformed in accurately segmenting images. With post-processing, the model scored 0.6656, while without TTA, it scored 0.5724. This indicates the proposed model is a promising solution for robust and accurate image analysis, showing potential in real-world applications. The model has demonstrated its capacity to excel, especially in complex segmentation tasks in the colorectal area.
7 Comparison with Others Model
In this section, different methodologies for segmenting colorectal polyps on the Kvasir-SEG dataset are examined. Each row presents the performance of a different method across additional evaluation metrics. The MultiResUNet + AG model, along with its variant with TTA, is highlighted in the table, indicating their superior performance. Table 3 compares the effectiveness of several state-of-the-art (SOTA) models. In comparison, the MultiResUNet + AG model (with and without TTA) discovered only 0.47 million parameters, significantly less than other models. Although NanoNet-A identified fewer parameters, our model with TTA was more effective and improved DSC (5.03%), accuracy (1.42%), IoU (12.02%), and precision (10.03%) [37]. As a result, despite having fewer parameters, the proposed system is still able to maintain high performance. The DSC score of FCN + CNN was 0.8922, but the IoU score was 0.8022; the proposed model received 0.8277 IoU, an improvement of 3.02% [19]. The DDANet had the highest recall score, but the IoU, DSC, and precision metrics needed improvement [21]. UACANet had 69.16 million parameters, while ColonSegNet had 5.01 million; however, both models performed poorly in quantitative scoring, including accuracy, DSC, IoU, recall, and precision [18, 25]. According to the analysis presented above, the proposed model outperformed other models and contained fewer parameters, thereby reducing non-essential features during training and making it more easily implementable in practice. High FPS, like UNet + + (81.34) and ColonSegNet (73.95), often sacrifices segmentation quality. In contrast, proposed models, MultiResUNet + AG (31) and MultiResUNet + AG + TTA (38), strike a balance. They maintain competitive segmentation performance (DSC, IoU, Recall, Precision) while achieving respectable FPS values. This balance is essential in tasks where both speed and accuracy matter. Models with higher FPS may show poorer segmentation results, highlighting the effectiveness of the proposed models for real-time image segmentation applications.
Initial resizing by downsampling technique decreased resolution from a maximum of 1920 × 1072 to 256 × 256 pixels, which is thought to be one of the causes of poor results on DSC and recall scores. This enlarging process may obscure the diagnosis of various minor lesions. Loss of fine features and subtle contrasts within the Region of Interest (ROI) are possible outcomes of the downsampling process. The loss of detail or merging of fine details during downsampling reduces the ability of AI systems and human beings to detect subtle aberrations in visuals. In our future studies, we will concentrate on developing a cropping strategy in which locally defined areas provide input to the model. To ensure accurate and particular insights are acquired, this method is focused on facilitating the acquisition of exact contextual elements.
8 Conclusions
This study has the significant advantage of minimizing missed detections, including tiny, oversaturated, flat polyps, which can improve clinical outcomes in colorectal cancer screening. This is a crucial step toward developing an automated polyp detection system for clinical use. The proposed approach combines the MultiResUNet model with a hybrid skip connection (Res + AG), utilizing the MultiRes block's robust feature extraction capabilities. A TTA technique is used to enhance boundary delineation and capture multi-scale image information, which includes original, horizontal, and vertical flip augmentations. In addition, this model is relatively lightweight, weighing only 6.71 MB and having only 0.47 million parameters. In addition, testing each sample takes 200 ms—1200 ms. A comparison between MultiResUNet + AG + TTA and SOTA proves MultiResUNet + AG + TTA to be superior for managing unseen data during real-time applications. As a result, the model performs admirably across a variety of criteria, such as precision, recall, DSC, IoU, and accuracy. The proposed model could enhance the accuracy and performance of clinical colonoscopy procedures by leveraging the capabilities of automated deep learning-based polyp segmentation. Based on the promising results obtained from this study, it is likely that MultiResUNet + AG + TTA will be used in real-world clinical settings in the future.
Data Availability
The data used in this study is available online as open source. The interested researchers could get it by contacting the authors [45].
Abbreviations
- TTA:
-
Test time augmentation
- DSC:
-
Dice similarity coefficient
- IoU:
-
Intersection over union
- ROC:
-
Receiver operating characteristics
- AUC:
-
Area under curve
- CAD:
-
Computer-aided diagnosis
- ML:
-
Machine learning
- MLP:
-
Multi-layer perceptron
- CNN:
-
Convolutional neural network
- AG:
-
Attention guidance
- CLs:
-
Convolutional layers
- MP:
-
Max-pooling
- BN:
-
Batch normalization
- HF:
-
Horizontal flip
- VF:
-
Vertical flip
- BCELOSS :
-
Binary cross-entropy loss
- PR:
-
Precision-recall
- AI:
-
Artificial intelligence
- Res:
-
Residual
- SOTA:
-
State of the art
References
Colorectal cancer, (n.d.). https://www.who.int/news-room/fact-sheets/detail/colorectal-cancer Accessed 26 Aug 2023.
Ahamed MF, Syfullah MK, Sarkar O, Islam MT, Nahiduzzaman M, Islam MR, Khandakar A, Ayari MA, Chowdhury MEH. IRv2-net: a deep learning framework for enhanced polyp segmentation performance integrating inceptionresnetV2 and UNet architecture with test time augmentation techniques. Sensors. 2023. https://doi.org/10.3390/s23187724.
Chan SCH, Liang JQ. Advances in tests for colorectal cancer screening and diagnosis, expert rev. Mol Diagn. 2022;22:449–60. https://doi.org/10.1080/14737159.2022.2065197.
Mármol I, Sánchez-de-Diego C, Pradilla Dieste A, Cerrada E, Rodriguez Yoldi MJ. Colorectal carcinoma: a general overview and future perspectives in colorectal cancer. Int J Mol Sci. 2017. https://doi.org/10.3390/ijms18010197.
Leufkens AM, Van Oijen MG, Vleggaar FP, Siersema PD. Factors influencing the miss rate of polyps in a back-to-back colonoscopy study. Endoscopy. 2012;44:470–5. https://doi.org/10.1055/s-0031-1291666.
Younas F, Usman M, Yan WQ. A deep ensemble learning method for colorectal polyp classification with optimized network parameters. Appl Intell. 2023;53:2410–33. https://doi.org/10.1007/s10489-022-03689-9.
Khan MM, Chowdhury MEH, Arefin ASMS, Podder KK, Hossain MS, Alqahtani A, Murugappan M, Khandakar A, Mushtak A, Nahiduzzaman M. A deep learning-based automatic segmentation and 3D visualization technique for intracranial hemorrhage detection using computed tomography images. Diagnostics. 2023. https://doi.org/10.3390/diagnostics13152537.
Zhuang Z, Raj AN, Jain A, Nersisson R, Chaurasia S, Li N, Lakshmanan M, Murugappan M. Nipple segmentation and localization using modified U-net on breast ultrasound images. J Med Imaging Health Inform. 2019;9:1827–37.
Ullah F, Ansari SU, Hanif M, Ayari MA, Chowdhury ME, Khandakar AA, Khan MS. Brain MR image enhancement for tumor segmentation using 3D U-Net. Sensors. 2021. https://doi.org/10.3390/s21227528.
Gul S, Khan MS, Bibi A, Khandakar A, Ayari MA, Chowdhury MEH. Deep learning techniques for liver and liver tumor segmentation: areview. Comput Biol Med. 2022;147:105620. https://doi.org/10.1016/j.compbiomed.2022.105620.
Rahman T, Khandakar A, Kadir MA, Islam KR, Islam KF, Mazhar R, Hamid T, Islam MT, Kashem S, Mahbub ZB, Ayari MA, Chowdhury MEH. Reliable tuberculosis detection using chest X-Ray with deep learning, segmentation and visualization. IEEE Access. 2020;8:191586–601. https://doi.org/10.1109/ACCESS.2020.3031384.
Negi A, Raj ANJ, Nersisson R, et al. RDA-UNET-WGAN: an accurate breast ultrasound lesion segmentation using wasserstein generative adversarial networks. Arab J Sci Eng. 2020;45:6399–410. https://doi.org/10.1007/s13369-020-04480-z.
Oktay O, Schlemper J, Folgoc LLe, Lee M, Heinrich M, Misawa K, Mori K, McDonagh S, Hammerla NY, Kainz B, Glocker B, Rueckert D. Attention U-Net: learning where to look for the pancreas; 2018. https://doi.org/10.48550/arxiv.1804.03999.
Diakogiannis FI, Waldner F, Caccetta P, Wu C. ResUNet-a: a deep learning framework for semantic segmentation of remotely sensed data. ISPRS J Photogramm Remote Sens. 2020;162:94–114. https://doi.org/10.1016/J.ISPRSJPRS.2020.01.013.
Ibtehaz N, Rahman MS. MultiResUNet : Rethinking the U-Net architecture for multimodal biomedical image segmentation. Neural Netw. 2020;121:74–87. https://doi.org/10.1016/j.neunet.2019.08.025.
Jha D, Smedsrud PH, Riegler MA, Johansen D, De Lange T, Halvorsen P, Johansen HD. ResUNet++: an advanced architecture for medical image segmentation. Proc. - 2019 IEEE Int. Symp. Multimedia, ISM 2019. (2019) 225–230. https://doi.org/10.1109/ISM46123.2019.00049.
Hong LTT, Thanh NC, Long TQ. CRF-EfficientUNet: an improved UNet framework for polyp segmentation in colonoscopy images with combined asymmetric loss function and CRF-RNN Layer. IEEE Access. 2021;9:156987–7001. https://doi.org/10.1109/ACCESS.2021.3129480.
Jha D, Ali S, Tomar NK, Johansen HD, Johansen D, Rittscher J, Riegler MA, Halvorsen P. Real-time polyp detection, localization and segmentation in colonoscopy using deep learning. IEEE Access. 2021;9:40496–510. https://doi.org/10.1109/ACCESS.2021.3063716.
Wen Y, Zhang L, Meng X, Ye X. Rethinking the transfer learning for FCN based polyp segmentation in colonoscopy. IEEE Access. 2023;11:16183–93. https://doi.org/10.1109/ACCESS.2023.3245519.
Jha D, Tomar NK, Sharma V, Bagci U. TransNetR: transformer-based residual network for polyp segmentation with multi-center out-of-distribution testing; 2023. https://arxiv.org/abs/2303.07428v1.. Accessed 6 July 2023.
Tomar NK, Jha D, Ali S, Johansen HD, Johansen D, Riegler MA, Halvorsen P. DDANet: dual decoder attention network for automatic polyp segmentation. Lect. Notes Comput. Sci. (Including Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinformatics). 12668 LNCS (2021) 307–314. https://doi.org/10.1007/978-3-030-68793-9_23/COVER.
Lou A, Guan S, Loew MH. CaraNet: context axial reverse attention network for segmentation of small medical objects. J Med Imaging. 2023;10:014005. https://doi.org/10.1117/1.JMI.10.1.014005.
Duc NT, Oanh NT, Thuy NT, Triet TM, Dinh VS. ColonFormer: an efficient transformer based method for colon polyp segmentation. IEEE Access. 2022;10:80575–86. https://doi.org/10.1109/ACCESS.2022.3195241.
Huang C-H, Wu H-Y, Lin Y-L HarDNet-MSEG: a simple encoder-decoder polyp segmentation neural network that achieves over 0.9 mean dice and 86 FPS; 2021. https://arxiv.org/abs/2101.07172v2. Accessed 6 July 2023.
Kim T, Lee H, Kim D. UACANet: uncertainty augmented context attention for polyp segmentation, MM 2021 - Proc. 29th ACM Int. Conf. Multimed. (2021) 2167–75. https://doi.org/10.1145/3474085.3475375.
Dumitru R-G, Peteleaza D, Craciun C. Using DUCK-Net for polyp image segmentation. Sci Rep. 2023;13:9803. https://doi.org/10.1038/s41598-023-36940-5.
Ameling S, Wirth S, Paulus D, Lacey G, Vilarino F. Texture-Based Polyp Detection in Colonoscopy BT—Bildverarbeitung für die Medizin. In: Meinzer H-P, Deserno TM, Handels H, Tolxdorff T, editors. Springer. Berlin, Heidelberg: Berlin Heidelberg; 2009. p. 346–50.
Grosu S, Wesp P, Graser A, Maurus S, Schulz C, Knösel T, Cyran CC, Ricke J, Ingrisch M, Kazmierczak PM. Machine learning–based differentiation of benign and premalignant colorectal polyps detected with CT colonography in an asymptomatic screening population: a proof-of-concept study. Radiology. 2021;299:326–35. https://doi.org/10.1148/radiol.2021202363.
Ding J, Yue C, Wang C, Liu W, Zhang L, Chen B, Shen S, Piao Y, Zhang L. Machine learning method for the cellular phenotyping of nasal polyps from multicentre tissue scans, expert rev. Clin Immunol. 2023;19:1023–8. https://doi.org/10.1080/1744666X.2023.2207824.
Mori Y, Kudo SE. Detecting colorectal polyps via machine learning. Nat Biomed Eng. 2018. https://doi.org/10.1038/s41551-018-0308-9.
Prasath VBS. Polyp detection and segmentation from video capsule endoscopy: a review. J Imaging. 2017. https://doi.org/10.3390/jimaging3010001.
Iakovidis DK, Maroulis DE, Karkanis SA, Brokos A. A comparative study of texture features for the discrimination of gastric polyps in endoscopic video. In: 18th IEEE Symp. Comput. Med. Syst., 2005: pp. 575–580. https://doi.org/10.1109/CBMS.2005.6.
Karkanis SA, Iakovidis DK, Maroulis DE, Karras DA, Tzivras M. Computer-aided tumor detection in endoscopic video using color wavelet features. IEEE Trans. Inf. Technol. Biomed. a Publ. IEEE Eng. Med. Biol. Soc. 7 (2003) 141–52. https://doi.org/10.1109/titb.2003.813794.
Wang Y, Tavanapong W, Wong J, Oh J, de Groen PC. Part-based multiderivative edge cross-sectional profiles for polyp detection in colonoscopy. IEEE J Biomed Heal Informatics. 2014;18:1379–89. https://doi.org/10.1109/JBHI.2013.2285230.
Hwang S, Oh J, Tavanapong W, Wong J, de Groen PC. Polyp detection in colonoscopy video using elliptical shape feature. In: 2007 IEEE Int. Conf. Image Process., 2007: pp. II-465–68. https://doi.org/10.1109/ICIP.2007.4379193.
Tajbakhsh N, Gurudu SR, Liang J. Automated polyp detection in colonoscopy videos using shape and context information. IEEE Trans Med Imaging. 2016;35:630–44. https://doi.org/10.1109/TMI.2015.2487997.
Jha D, Tomar NK, Ali S, Riegler MA, Johansen HD, Johansen D, De Lange T, Halvorsen P. NanoNet: real-time polyp segmentation in video capsule endoscopy and colonoscopy. Proc IEEE Symp Comput Med Syst. 2021; 37–43. https://doi.org/10.1109/CBMS52027.2021.00014.
Valanarasu JMJ, Patel VM. UNeXt: MLP-based rapid medical image segmentation network. Lect. Notes Comput. Sci. (Including Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinformatics). 13435 LNCS (2022) 23–33. https://doi.org/10.1007/978-3-031-16443-9_3/COVER.
Bernal J, Sánchez J, Vilariño F. Towards automatic polyp detection with a polyp appearance model. Pattern Recognit. 2012;45:3166–82. https://doi.org/10.1016/j.patcog.2012.03.002.
Krizhevsky A, Sutskever I, Hinton GE. Imagenet classification with deep convolutional neural networks. In: Pereira F, Burges CJ, Bottou L, Weinberger KQ, editors. Curran Associates Inc Proceedings. USA; 2012. p. 1097–106.
Wang G, Li W, Aertsen M, Deprest J, Ourselin S, Vercauteren T. Aleatoric uncertainty estimation with test-time augmentation for medical image segmentation with convolutional neural networks. Neurocomputing. 2019;338:34–45. https://doi.org/10.1016/j.neucom.2019.01.103.
Matsunaga K, Hamada A, Minagawa A, Koga H. Image classification of Melanoma, Nevus and Seborrheic Keratosis by deep neural network ensemble. CoRR. abs/1703.0; 2017. http://arxiv.org/abs/1703.03108.
Jin H, Li Z, Tong R, Lin L. A deep 3D residual CNN for false-positive reduction in pulmonary nodule detection. Med Phys. 2018;45:2097–107. https://doi.org/10.1002/mp.12846.
Jha D, Smedsrud PH, Riegler MA, Halvorsen P, de Lange T, Johansen D, Johansen HD. Kvasir-SEG: a segmented polyp dataset. Lect. Notes Comput. Sci. (Including Subser. Lect. Notes Artif. Intell. Lect. Notes Bioinformatics). 11962 LNCS (2020) 451–462. https://doi.org/10.1007/978-3-030-37734-2_37/COVER.
Bernal J, Sánchez FJ, Fernández-Esparrach G, Gil D, Rodríguez C, Vilariño F. WM-DOVA maps for accurate polyp highlighting in colonoscopy: validation vs saliency maps from physicians. Comput Med Imaging Graph. 2015;43:99–111. https://doi.org/10.1016/J.COMPMEDIMAG.2015.02.007.
Ronneberger O, Fischer P, Brox T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In: Navab N, Hornegger J, Wells WM, Frangi AF, editors. BT - Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. Cham: Springer International Publishing; 2015. p. 234–41.
Drozdzal M, Vorontsov E, Chartrand G, Kadoury S, Pal C. The importance of skip connections in biomedical image segmentation. In: Gustavo C, Diana M, Loïc P, Andrew B, João Manuel RST, Vasileios B, João PP, Jacinto CN, Marco L, Zhi L, Jaime SC, Julien C, editors. International Workshop on Deep Learning in Medical Image Analysis. Greece: Springer; 2016. p. 179–87. https://doi.org/10.1007/978-3-319-46976-8_19.
Abdollahi A, Pradhan B. Integrated technique of segmentation and classification methods with connected components analysis for road extraction from orthophoto images. Expert Syst Appl. 2021;176:114908. https://doi.org/10.1016/j.eswa.2021.114908.
Jha D, Smedsrud PH, Johansen D, de Lange T, Johansen HD, Halvorsen P, Riegler MA. A comprehensive study on colorectal polyp segmentation with ResUNet++, conditional random field and test-time augmentation. IEEE J Biomed Heal Informat. 2021;25:2029–40. https://doi.org/10.1109/JBHI.2021.3049304.
Lian L, Zhu T, Zhu F, Zhu H. Deep Learning for Caries Detection and Classification. Diagnostics. 2021. https://doi.org/10.3390/diagnostics11091672.
Bai Z, Wang J, Zhang X-L, Chen J. End-to-end speaker verification via curriculum bipartite ranking weighted binary cross-entropy. IEEE/ACM Trans Audio Speech Lang Process. 2022;30:1330–44. https://doi.org/10.1109/TASLP.2022.3161155.
Jha D, Riegler MA, Johansen D, Halvorsen P, Johansen HD. DoubleU-Net: a deep convolutional neural network for medical image segmentation. Proc IEEE Symp Comput Med Syst. 2020; 558–564. https://doi.org/10.1109/CBMS49503.2020.00111.
Zhou Z, Siddiquee MMR, Tajbakhsh N, Liang J. UNet++: a nested U-Net architecture for medical image segmentation. Deep Learn. Med. Image Anal. Multimodal Learn. Clin. Decis. Support 4th Int. Work. DLMIA 2018, 8th Int. Work. ML-CDS 2018, Held Conjunction with MICCAI 2018, Granada, Spain, 11045; 2018. pp. 3–11. https://doi.org/10.1007/978-3-030-00889-5_1.
Funding
This research is supported via funding from Prince Sattam Bin Abdulaziz University project number (PSAU/2023/R/1444). The statements made herein are solely the responsibility of the authors. The open access publication cost is covered by Qatar National Library.
Author information
Authors and Affiliations
Contributions
Md. Faysal Ahamed and Md. Rabiul Islam; Conceptualization, Md. Faysal Ahamed, Md. Rabiul Islam, and Md. Nahiduzzaman; methodology, Md. Faysal Ahamed, Md. Rabiul Islam, Muhammad E. H. Chowdhury, and M. Murugappan; validation, Md. Faysal Ahamed, Md. Rabiul Islam, Md. Nahiduzzaman, Muhammad E. H. Chowdhury, and M. Murugappan; formal analysis, Md. Faysal Ahamed, Md. Rabiul Islam, and Md. Nahiduzzaman; investigation, Muhammad E. H. Chowdhury and Abdulrahman Alqahtani; resources, Muhammad E. H. Chowdhury, Abdulrahman Alqahtani, and M. Murugappan; data curation, Md. Faysal Ahamed; writing—original draft, Md. Faysal Ahamed, M. Murugappan and Md. Nahiduzzaman; writing—review and editing, Md. Rabiul Islam, Muhammad E. H. Chowdhury, and M. Murugappan; visualization, Md. Rabiul Islam, Muhammad E. H. Chowdhury, and M. Murugappan; supervision, Muhammad E. H. Chowdhury, Abdulrahman Alqahtani, and M. Murugappan; project administration, Muhammad E. H. Chowdhury, Abdulrahman Alqahtani, and M. Murugappan; funding acquisition, Md. Rabiul Islam, Muhammad E. H. Chowdhury, Abdulrahman Alqahtani, and M. Murugappan;; All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Conflicts of Interest
The authors declare no conflict of interest.
Ethical approval
Not Applicable.
Consent for publication
All the author(s) in the paper have given their consent to submit and publish the paper in HCIS.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahamed, M.F., Islam, M.R., Nahiduzzaman, M. et al. Automated Colorectal Polyps Detection from Endoscopic Images using MultiResUNet Framework with Attention Guided Segmentation. Hum-Cent Intell Syst 4, 299–315 (2024). https://doi.org/10.1007/s44230-024-00067-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s44230-024-00067-1