Abstract
Background
Emerging evidence has shown that SARS-CoV-2 may affect the circulatory system in addition to the human respiratory system. However, no study has indicated whether patients with leukemia have a greater likelihood of SARS-CoV-2 infection or have poor treatment outcomes.
Objective
The study aimed to demonstrate the relationship between essential blood proteins and the major SARS-CoV-2 proteins by network pharmacology bioinformatics analysis.
Methods
Bioinformatics analysis was used to establish eight differentially expressed gene hubs in leukemia through differential gene screening, protein–protein interaction network analysis, and gene enrichment analysis. Molecular docking analysis was also conducted to dock the two up-regulated proteins with the spike glycoprotein in leukemia and the critical protease enzyme (Mpro) of SARS-CoV-2.
Results
We identified two up-regulated genes (PTPRC and BCL6) among the eight differentially expressed genes. The PTPRC and BCL6 also docked perfectly with the main SARS-CoV-2 structural proteins.
Conclusion and Recommendation
This study indicates that SARS-CoV-2 is likely to affect with the blood in patients with chronic leukemia. Therefore, patients with chronic leukemia require greater medical attention and precautions during the COVID-19 pandemic.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Coronaviruses (family Coronaviridae) are viruses whose genomes comprise single-stranded positive-sense RNA ranging from 27 to 34 kb in size [1]. Coronaviruses gained substantial scientific attention in early 2000 after the severe acute respiratory syndrome (SARS-CoV) and Middle East respiratory syndrome (MERS-CoV) epidemics, which caused approximately 700 and 400 deaths, respectively [2]. In early December, the reporting of SARS-CoV-2 in and around Wuhan, China, alarmed scientific communities about a disease known as COVID-19 [3,4,5]. Patients infected with SARS-CoV-2 show severe respiratory abnormalities and difficulty breathing, which may eventually result in death [6]. A highly contagious mode of transmission and the prolonged stability of the virus in the air and inert surfaces such as steel are major reasons for its spread worldwide [7]. Thus, in the present global emergency of the COVID-19 pandemic, an urgent need exists to develop an efficient treatment against SARS-CoV-2 infection. With its single-strand positive-sense RNA genome and limited structural and functional protein resources, SARS-CoV-2 can infect host cells and proliferate within them [8]. SARS-CoV-2 hijacks the host machinery at the molecular level to complete its life cycle and produce functional virion progeny [9]. Awareness of the process of COVID-19 is scarce but rapidly growing among patients with cancer, particularly hematologic malignancies. The infection rate in patients with cancer may be higher than that in the general population [10, 11]. In two studies in China, only 10 out of 1099 and 18 out of 1590 patients with COVID-19 were diagnosed with cancer [12, 13]. In one study, 60% of patients with COVID-19 with blood cancer recovered from COVID-19 within a 14-day observation period [14].
The scientific community has published findings on COVID-19 in patients with cancer worldwide [15,16,17,18]. Patients with leukemia are frequently myelosuppressed, immunosuppressed, and possibly immunoglobulin deficient, thus making them potentially highly vulnerable to COVID-19 [19]. Because of the disease biology of leukemia subtypes, associated therapy, underlying comorbidities, patient-specific aspects, and specific COVID-19-related risk factors, patients with leukemia may be at a significantly greater threat of developing SARS-CoV-2 infection [19]. Unfortunately, because of minimal reports related to leukemia, the implications are poorly understood in this particular population. Because COVID-19 is a new human virus, whether variations exist relative to other blood cancers and how the virus affects people with leukemia remain unknown. Patients with blood cancer, because of their immunocompromise due to both cancer and the effects of cancer treatment, are likely to be particularly prone to SARS-CoV-2 infection.
Our current study aimed to measure the likelihood of patients with leukemia acquiring SARS-CoV-2 infection, on the basis of a theoretical network biology approach. We studied the interaction of target genes/proteins between leukemic chronic lymphocytic and chronic myelogenous tissue and the SARS-CoV-2 virus by using computational techniques such as screening differentially expressed genes (DEGs), gene enrichment analysis (GEA), construction, protein–protein interaction network analysis (PPINA), and molecular docking analysis (MDA).
2 Materials and Methods
The detailed study workflow is depicted in Fig. 1.
2.1 Microarray Dataset Collection and Preprocessing
Three microarray datasets comprising mRNA expression profiles for leukemia and healthy groups were downloaded from NCBI’s Genome Expression Omnibus (GEO) [20]. The blood datasets included GSE8835 [21], GSE24739 [22], and GSE39411 [23]. These datasets met the following conditions: (1) samples from chronic lymphocytic leukemia (CLL) and chronic myelogenous leukemia (CML) in Homo sapiens, (2) presence of control groups, (3) expression profiling by array category, and (4) inclusion of five samples or more. The overall sample size reliably indicates DEGs or non-coding RNAs; therefore, GEO datasets encompassing at least ten samples were selected for further examination. Background data correction/normalization were performed by multiarray average (RMA) in the R affy and Lumi packages to ensure unbiased and dysregulated gene expression data. The RMA approach, including quantile normalization, was used to eliminate variations attributable to the individual Affymetrix GSE series standardization. In the fold-change calculation to identify DEGs, the sensitivity and specificity of the RMA technique were acceptable. We additionally used the Bioconductor Package (Lumi pipeline) designed to study Illumina data (BeadChip). The results verified the consistency, normalization, and stable variance of the data.
2.2 Identification of DEGs
To examine DEGs in every GEO dataset, we used the linear model for microarray analysis (limma) package in R. By using the empirical Bayes method and decreasing the standard errors, it calculates simple t test, moderate t test, and f test results, and offers reproducible results. The limma package [24] was used to determine the DEGs between healthy and leukemia groups. DEGs were characterized as genes with p < 0.05, and logFC (0.2 ≥ logFC ≤ − 0.2) for up- and down-regulated DEGs, respectively.
2.3 Meta-Analysis of DEGs in the Gene Expression Dataset
Using the MetaMa package [25] and limma [26] in R, we performed meta-analysis of the normalized gene expression datasets by using Fisher's combined probability test technique [27]. False discovery rate adjustment was performed with Benjamini–Hochberg correction [28] by combining the p-values and fold-change values of the shared genes. Meta-analysis of datasets was conducted through generic methods of combining information by vote-counting (Table 1). BRCW (http:/jura.wi.mit.edu/bioc/tools/compare.php) was used to identify the mutual DEGs in at least two gene expression profile datasets, thus increasing the accuracy of DEG selection; the chances of a biased data compilation thus became nil. The probe numbers in the expression profile were translated to gene symbols with the Synergizer database, on the basis of the equivalent similarity between the probe and the gene in the data [29].
2.4 Functional and Pathway Enrichment Analyses
We classified DEGs by biological process (BP), molecular function (MF), cellular component, and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways to understand the significance of the listed DEGs, on the basis of Database for Annotation Visualization and Integrated Discovery (DAVID) v.6.7 [30]. The Gene Ontology (GO) and KEGG databases are used by DAVID for gene enrichment study. Pathways and roles with a p value < 0.05 were considered significant. Subsequently, ggplot2 [31] was used to construct an enrichment plot of critical biological processes, molecular, cellular components, and function pathways.
2.5 Construction of a PPI Network
For PPI network construction and analysis, we obtained specific DEGs through enrichment analysis. We used the simple concept of the correspondence of one gene to one protein and developed the DEGs' primary leukemia PPI network. The network was built with the STRING v.10.5 database [32], and the Cytoscape [33] file has been uploaded for further literature verification.
2.6 Overlap Between Leukemia and SARS-CoV-2-Associated Proteins
We identified reported COVID-19 associated genes, which were searched with GenCLiP3 [34]. GenCLiP 2.0 (http://ci.smu.edu.cn/genclip3/) is an online tool used to analyze human genes for literature mining. The literature mining gene retrieval of COVID-19-associated genes was based on user-defined query keywords. The keywords, grouped by a fuzzy algorithm, can be input by users or generated for the relevant gene established on accessible terms in the prior art. Associated Medline abstracts were linked by using the associations between genes and keywords. The co-occurrence of genes and keywords was highlighted in our literature mining.
2.7 Module Analysis
We used the molecular complex detection (MCODE) app in Cytoscape [35] to perform module analysis with the degree cutoff criteria = 2, node density cutoff = 0.1, node score cutoff = 0.2, k-core = 2, and maximum depth = 100. We also studied GO and KEGG pathway enrichment for important genes in hub modules (p value < 0.05).
2.8 Correlation of Gene Expression with Tumor-Infiltrating Immune Cells
Given the importance of immune dysregulation in leukemia, we explored the correlations between MTG1, PPP2R5B, and ANXA5 mRNA expression and tumor-infiltrating immune cells. The TIMER tool (https://cistrome.shinyapps.io/timer/) [36] was used to analyze the association between gene expression with tumor-infiltrating cells. Six tumor-infiltrating immune subsets, including B cells, CD8+ T cells, CD4+ T-cells, macrophages, neutrophils, and dendritic cells, were analyzed.
2.9 Protein Preparation
The BCL6 (PDB ID: 6XWF), KIT (PDB ID: 2EC8), and PTPRC (PDB ID: 5FMV) crystal structures were from the RCSB Protein Data Bank (PDB). Using RCSB PDB, we also downloaded the recently submitted crystal structures of COVID-19 spike glycoprotein with a single receptor-binding domain and the main protease (Mpro) of COVID-19 in complex with an inhibitor N3. PyMOL was used to optimize the structures, mainly through removal of ligands and water molecules.
2.9.1 Molecular Docking
Rigid molecular docking of proteins was performed with the Cluspro 2.0 [37] server. The files were downloaded from the top ten predictions from the Cluspro web server. Prodigy (https://bianca.science.uu.nl/prodigy/) was used to evaluate the effects of protein docking interactions and provide the binding affinity (ΔG). The ΔG specifies the solvation free energy (kcal/M) expansion after the formation of the interface. The ΔG value is computed as the difference in total solvation energies of isolated and interfacing structures. On the basis of the anticipated ΔG (Eq. 1), the dissociation constant (Kd) was determined,
where R is the ideal gas constant (in kcal K−1 mol−1), T is the temperature (in K), K is the equilibrium constant, and ΔG is the anticipated free energy 298.15 K (25 °C). Negative ΔG is associated with hydrophobic interfaces or positive protein affinity. PRODIGY [38] (PROtein binDIng enerGY prediction) is a web server for calculating binding affinity in biological complexes and determining biological interfaces from crystallographic ones. PyMOL [39] was used for the visualization of the docked structure.
3 Results
The flowchart of our bioinformatics analysis of network pharmacology is shown in Fig. 1.
3.1 Extraction and Preprocessing of Microarray Data
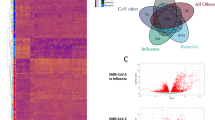
Built on the exclusion/inclusion criteria described in the methods, the microarray gene expression profiles with accession numbers GSE8835, GSE24739, and GSE39411 contain expression data from tissues from patients with CLL and CML, and healthy controls. Information associated with these datasets, such as GEO accession number, platform type, number of samples, type of study, and species, is shown in Table 1. The heatmap visualization of expression profiles for tissue samples from patients with CLL and CML and controls is shown in Fig. 2A.
A Expression heatmap showing expression levels of significant DEGs for the GSE8835, GSE24739, GSE39411 datasets. Color indicates high expression (red) and low expression (green). B Venn diagram showing the DEGs in GSE8835, GSE24739, and GSE39411 after overlap analysis. C Venn diagram showing the common genes after mapping of DEGs in leukemia and SARS-CoV-2-associated proteins
3.2 Meta-analysis and Identification of DEGs in Patients with Leukemia
The three described datasets were used to identify DEGs between patients with leukemia and healthy participants, and to perform meta-analyses for identifying mutually expressed genes across these datasets. A total of 556 DEGs were obtained after meta-analysis, 261 of which were up-regulated and 295 of which were down-regulated in patients with leukemia vs. controls on the basis of the statistical threshold of log2 (fold change) and Benjamini–Hochberg p-value.
3.3 Functional and Pathway Enrichment Analysis
We conducted GO enrichment analysis to illustrate the possible biological functions of DEGs in leukemia. GO enrichment analysis and KEGG pathway analysis for the up-and down-regulated DEGs (Supplementary Files 3 and 4) were performed. The significant enrichment of up- and down-regulated DEGs in CLL and CML is shown in Figs. 3 and 4, respectively. The common up-regulated DEGs were enriched in BPs including B-cell receptor signaling pathway, immune response, positive control of transcription, DNA-templated, innate immune response, control of cell adhesion, control of immune response, positive control of B-cell activation, phagocytosis, DNA-templated response to estradiol, T-cell distinction, antigen processing and presentation of peptide antigen via MHC class I, and negative regulation of cellular senescence (Supplementary File 3 and Fig. 3A). Down-regulated DEGs were significantly enriched in BPs including the Wnt signaling pathway, the planar cell polarity pathway, the T-cell receptor signaling pathway, mitochondrial respiratory chain complex I assembly, antigen processing and presentation of exogenous peptide antigen via MHC class II, intracellular protein transport, control of the cellular amino acid metabolic process, negative control of gene expression, negative control of cell migration, protein heteromerization, B-cell lineage commitment, immunoglobulin V(D)J recombination, etc. (Supplementary File 4 and Fig. 4A). Additionally, the MFs of the up-regulated DEGs in blood were protein binding, antigen binding, immunoglobulin receptor binding, protease binding, R-SMAD binding, etc. (Supplementary File 3 and Fig. 3B). Furthermore, the MFs of down-regulated DEGs in the blood were poly(A) RNA binding, protein binding, cadherin binding implicated in cell–cell adhesion, myosin V binding, NAD binding, catalytic activity, etc. (Supplementary File 4 and Fig. 4B). The significant cellular components of up-regulated DEGs included nucleoplasm, extracellular exosome, mast cell granule, spliceosomal complex, MHC class I protein complex, phagocytic vesicle, immunoglobulin complex, circulating, viral nucleocapsid, Golgi apparatus, crucial elements of the membrane, etc. (Supplementary File 3, Fig. 3C). The significant cellular components of down-regulated DEGs included proteasome accessory complex, respiratory chain, mitochondrial respiratory chain complex I, endosome membrane, extracellular exosome, etc. (Supplementary File 4, Fig. 4C). The significant KEGG pathways in up-regulated DEGs included salivary secretion, pathways in cancer (i.e., Wnt-pathways, GSK3 pathways, and FGF signaling pathways), signaling pathways controlling pluripotency of stem cells, cell adhesion molecules, the Jak-STAT signaling pathway, HTLV-I infection, Influenza A, the FoxO signaling pathway, the HIF-1 signaling pathway, the Ras signaling pathway, melanogenesis, endocytosis, etc. (Supplementary File 3, Fig. 3D). Most down-regulated DEGs were involved in KEGG pathways involving the blood, including biosynthesis of antibiotics, Fc gamma R-mediated phagocytosis, metabolic pathways, the Wnt signaling pathway, the Hippo signaling pathway, proteoglycans in cancer, etc., as shown in Fig. 4D (Supplementary File 4).
3.4 PPINA
Using the STRING v.10.5 databases, we built PPI networks and visualized and analyzed them in Cytoscape software v.3.4.0. The PPI evidence acquired from STRING's online DEGs and the PPI network is shown in in Fig. 5A. Network Analyzer, a Cytoscape plugin, was used to analyze the network topological properties (Supplementary File 6: Table S10) of DEGs. The 500 genes showed a considerable degree distribution, with the highest degree of 41 and lowest degree of 1.0. The average degree value was 7.029. The PPI network consisted of 385 nodes and 1353 interactions (Fig. 5A). Finally, the top ten high-grade hub genes, including CDC42, EP300, KIT, PTPRC, DVL2, BCL6, GART, HNRNPA1, HNRNPC, and SMAD3, were selected.
Protein–protein interaction (PPI) network and hub clustering modules. A The PPI network of overlapping DEGs. Green nodes represent up-regulated DEGs, and cyan nodes represented down-regulated DEGs. B Module 1 (MCODE score = 7.917), Module 2 (MCODE score 7.818), Module 3 (MCODE score = 5.2). Module 5 (MCODE score = 3.2). Module 6 (MCODE score = 3.2) and Module 13 (MCODE score = 2.667)
3.5 Overlap Between Leukemia and SARS-CoV-2-Associated Proteins
The overlap between the 500 leukemia DEGs and SARS-CoV-2-associated proteins (560 proteins) was analyzed with Venn diagrams (Fig. 2C). A total of 27 leukemia-regulated SARS-CoV-2-associated proteins were identified, including ANXA1, CD38, HLA-E, BCL2, CXCL8, CDK6, FURIN, KIT, BCL6, TMEM41B, CR1, PARP1, PTPRC, BST2, CTLA4, ITPA, ABL1, CYBB, PMAIP1, HIF1A, IL7R, LY6E, CD47, NDUFA10, KPNB1, ATG16L1, and IFI44. Of the ten hub genes, three hubs, KIT, PTPRC, and BCL6, were common between leukemia and SARS-CoV-2-associated proteins. PPI network constructs with 27 genes commonly involved in leukemia and COVID-19 are shown in Fig. 5C. According to GO and KEGG pathway enrichment examination, most of the genes were implicated in multiple responses, including response to interferon, cellular response to type I interferon, viral response, control of viral genome replication, control of immune effector process, T-cell differentiation, B-cell activation, and immune response-controlling cell surface receptor signaling pathways.
3.6 Potential Mechanisms of the Critical Genes
The TIMER web tool [36] indicated that expression of the KIT, PTPRC, and BCL6 genes was meaningfully associated with one or more blood cancer (lymphoma)-infiltrating immune cell subsets. For B-cells, the expression of PTPRC and BCL6 displayed the most meaningful connection, and the expression of CDC42 was the most important relationship. CD4+ T cells have been demonstrated to support B cells to produce antibodies and help CD8+ T cells eradicate cells infected with SARS-CoV-2 viruses. Interferon-gamma, the leading player governing viral infection, is a major cytokine made by T cells [40].
3.7 Network module analysis
We imported the PPI network into Cytoscape to detect significant clustering modules. Module analysis and modules with top high scores were screened out (Fig. 6B). Eight hub nodes were present in the six modules (Table 2). According to GO and KEGG pathway enrichment analysis (p < 0.05), BCL6 in module 5 (MCODE score = 3.2) was closely associated with negative control of transcription from RNA polymerase II promoter, negative management of immune response, and negative regulation of the Notch signaling pathway. KIT in module 6 (MCODE score = 3.2) was involved in T-cell differentiation, mast cell degranulation, the RAS signaling pathway, MAPK cascade regulation of cell proliferation protease, and positive regulation of GTPase activity. PTPRC in module 3 (MCODE score = 5.2) was closely associated with antigen binding. EP300 was associated with the Jak-STAT signaling pathway, HTLV-I infection, transcriptional dysregulation in cancer, Influenza A, the FoxO signaling pathway, and melanogenesis.
Correlation analysis of MTG1, PPP2R5B, and ANXA5 mRNA expression with blood cancer (lymphoma)-infiltrating immune cells. The data are from the TIMER database (https://cistrome.shinyapps.io/timer/). A Correlation of KIT mRNA with blood cancer-infiltrating immune cells. B Correlation of BCL6 mRNA with blood cancer-infiltrating immune cells. C Correlation of PTPRC mRNA with blood cancer-infiltrating immune cells. D Correlation of CDC42 mRNA with blood cancer-infiltrating immune cells
3.8 Molecular docking
Only BCL6, KIT, and PTPRC were identified as up-regulated proteins common among the ten hub genes and 27 genes common in leukemia and SARS-CoV-2-associated proteins. The molecular docking of BCL6, KIT, and PTPRC with SARS-CoV-2 spike glycoprotein and Mpro is shown in Fig. 7. The ΔG of BCL6 and the spike protein of SARS-CoV-2 was − 56.7 kcal/mol, and the ΔG of BCL6 and the Mpro of the virus was − 6.8 kcal/mol. The \(\mathrm{\Delta G}\) of KIT and the spike glycoprotein was − 52.5 kcal/mol, and the ΔG of KIT and the Mpro of the virus was − 12.3 kcal/mol. Finally, the ΔG of PTPRC and the spike glycoprotein was − 52.5 kcal/mol, and the ΔG of PTPRC and the Mpro of the virus was − 16.1 kcal/mol. The reported ΔG indicates the solvation free energy gain after interface formation; a negative ΔG indicates hydrophobic interfaces or positive protein affinity. The dissociation constant (Kd) for each docking is given in Table 3. The three up-regulated proteins in the CLL and CML exhibited good interaction with the SARS-CoV-2 spike glycoprotein and Mpro, thus indicating that the blood tissues of patients with leukemia are vulnerable to SARS-CoV-2. The molecular docking for CDC42 (PDBID 1AJE) and SARS-CoV-2 spike glycoprotein, and CDC42 and Mpro is shown in Fig. 8. The ΔG of CDC42 and the spike glycoprotein was − 5.6 kcal/mol, and the ΔG of KIT and the Mpro of the virus was − 43.6 kcal/mol (Table 3).
4 Discussion
In this study, we selected the spike glycoprotein and Mpro as the SARS-CoV-2 target proteins by using molecular docking technologies. Through the analysis of microarray datasets, we detected 385 overlapping DEGs. Enrichment analysis revealed 500 overlapping DEGs mainly associated with genes involved in the interferon response, type I interferon cellular response, virus response, viral genome replication regulation, immune effector process regulation, differentiation of T cells, activation of B cells, and signaling pathways of immune response-regulating cell surface receptor. Through molecular analysis, we selected ten hub genes from these 500 overlapping DEGs. The MAPK cascade, GTPase regulatory activity, and other factors listed herein were associated with leukemia. Subsequently, we identified only 27 leukemia-regulated SARS-CoV-2-associated proteins. Of the ten hubs, three genes (KIT, PTPRC, and BCL6) were common to both leukemia and SARS-Cov-2-associated proteins, which have been further studied in patients with COVID-19. CDC42 in human immunodeficiency virus (HIV)-1 cell entry is the most examined aspect of CDC42 function in viral cell entry processes [41]. Prior studies have demonstrated the role of CDC42 in cell entry of other RNA viruses—a critical area for further research through similar methods.
CDC42, a protein-coding gene, is involved in pathways including nerve growth factor (NGF) and the integrin pathway. Annotations associated with this gene in GO include similar protein binding and protein kinase binding. Furthermore, RAC1 is an essential paralog of this gene. In our study, CDC42 was the top hub (degree = 41). In addition, CDC42 has a crucial role in the entry process of mouse hepatitis coronavirus (MHV CoV) [42]. In the initial phases of infection, MHV infectivity and the use of actin cytoskeleton modifying agents had related restrictive events on infection, directing to GTPase, and explicitly to the participation of CDC42 in the entry process [41, 43]. Ethyl isopropyl amiloride (EIPA) is well known for its ability to inhibit macropinocytosis through inhibiting CDC42 signaling. The findings of diminished infectivity due to Arp2/3 knockdown, disruption of the actin cytoskeleton, and EIPA have indicated that CDC42 signaling is involved in the MHV cell entry process. Because the CDC42 hub gene is a crucial protein in leukemia, we performed molecular docking between CDC42 protein in leukemia (in CLL and CML) tissues and the spike glycoprotein and Mpro, the essential structural proteins of the SARS-CoV-2 virus. The hub proteins in leukemia successfully docked with the virus's essential proteins, thereby confirming our hypothesis that patients with leukemia can have a more significant threat of being attacked by SARS-CoV-2.
Clinical studies have shown that people of all ages are generally susceptible to COVID-19. By contrast, the risk of infection with the virus increases in older people and people with underlying diseases [44]. Management of patients with leukemia in the COVID-19 pandemic can be complicated. The risk of infection with SAR-Cov-2 remains low during high-risk COVID-19 periods when optimal preventive measures and mass testing are used; however, mortality may be elevated in patients with both leukemia and COVID-19. The effects of the COVID-19 pandemic on leukemia have been evaluated in recent reports, including a study of the incidence of anxiety in patients with leukemia during the COVID-19 pandemic [14, 18], a description of physical movement and quality lifestyle in patients with leukemia during the COVID-19 pandemic [14, 18], and an investigation of the outcomes of patients with leukemia affected by COVID-19 [18]. Recent research has shown that spike glycoprotein and Mpro [45] are the main structural proteins of COVID-19. Spike glycoprotein is the main target for COVID-19 vaccines, therapeutic antibodies, and diagnostics of COVID-19 [46]. Likewise, another possible target protein is Mpro (also called 3C-like protease), a key coronavirus enzyme with an essential role in facilitating viral replication and transcription, thus providing a promising drug target for COVID-19 [46].
A higher risk of infection and likelihood of severe COVID-19 was established among cancer patients as a sub-group early in the pandemic. Thus, this research provides valuable knowledge that should help physicians make informed choices in protecting and caring for patients with leukemia and COVID-19. Furthermore, this study lays a groundwork for future relevant laboratory studies, which may enable identification of novel potential molecular targets that may be exploited to inhibit viral interactions with host cellular factors and block the spread and viral replication in the body. A better structural understanding of molecular targets, virus–host interactions, and the cause of pathogenesis is required for the development of effective therapeutic/prophylactic agents for COVID-19 prevention and treatment.
5 Conclusion
The findings of this bioinformatics-based research demonstrated that patients with chronic leukemia are at higher risk of SARS-CoV-2 infection as compared to normal individuals. Accordingly, patients with chronic leukemia require better medical attention during the COVID-19 pandemic.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary materials.
References
Wu A, Peng Y, Huang B, Ding X, Wang X, Niu P, Meng J, Zhu Z, Zhang Z, Wang J, Sheng J, Quan L, Xia Z, Tan W, Cheng G, Jiang T. Genome composition and divergence of the novel coronavirus (2019-nCoV) originating in China. Cell Host Microbe. 2020;27(3):325–8. https://doi.org/10.1016/j.chom.2020.02.001.
de Wit E, van Doremalen N, Falzarano D, Munster VJ. SARS and MERS: recent insights into emerging coronaviruses. Nat Rev Microbiol. 2016;14(8):523–34. https://doi.org/10.1038/nrmicro.2016.81.
Golamari R, Kapoor N, Devaraj T, Sahu N, Kramer C, Jain R. Experimental therapies under investigation for COVID-19. J Community Hosp Intern Med Perspect. 2021;11(2):187–93. https://doi.org/10.1080/20009666.2021.1874093.
Kumar A, Singh R, Kaur J, Pandey S, Sharma V, Thakur L, Sati S, Mani S, Asthana S, Sharma TK, Chaudhuri S, Bhattacharyya S, Kumar N. Wuhan to world: the COVID-19 pandemic. Front Cell Infect Microbiol. 2021;11:596201. https://doi.org/10.3389/fcimb.2021.596201.
Dong X, Cao YY, Lu XX, Zhang JJ, Du H, Yan YQ, Akdis CA, Gao YD. Eleven faces of coronavirus disease 2019. Allergy. 2020;75(7):1699–709. https://doi.org/10.1111/all.14289.
Rothan HA, Byrareddy SN. The epidemiology and pathogenesis of coronavirus disease (COVID-19) outbreak. J Autoimmun. 2020;109:102433. https://doi.org/10.1016/j.jaut.2020.102433.
van Doremalen N, Bushmaker T, Morris DH, Holbrook MG, Gamble A, Williamson BN, Tamin A, Harcourt JL, Thornburg NJ, Gerber SI, Lloyd-Smith JO, de Wit E, Munster VJ. Aerosol and surface stability of SARS-CoV-2 as compared with SARS-CoV-1. N Engl J Med. 2020;382(16):1564–7. https://doi.org/10.1056/NEJMc2004973.
Chen S, Luo H, Chen L, Chen J, Shen J, Zhu W, Chen K, Shen X, Jiang H. An overall picture of SARS coronavirus (SARS-CoV) genome-encoded major proteins: structures, functions and drug development. Curr Pharm Des. 2006;12(35):4539–53. https://doi.org/10.2174/138161206779010459.
de Wilde AH, Snijder EJ, Kikkert M, van Hemert MJ. Host factors in coronavirus replication. Curr Top Microbiol Immunol. 2018;419:1–42. https://doi.org/10.1007/82_2017_25.
Yu J, Ouyang W, Chua MLK, Xie C. SARS-CoV-2 Transmission in patients with cancer at a tertiary care hospital in Wuhan, China. JAMA Oncol. 2020;6(7):1108–10. https://doi.org/10.1001/jamaoncol.2020.0980.
Robilotti EV, Babady NE, Mead PA, Rolling T, Perez-Johnston R, Bernardes M, Bogler Y, Caldararo M, Figueroa CJ, Glickman MS, Joanow A, Kaltsas A, Lee YJ, Lucca A, Mariano A, Morjaria S, Nawar T, Papanicolaou GA, Predmore J, Redelman-Sidi G, Schmidt E, Seo SK, Sepkowitz K, Shah MK, Wolchok JD, Hohl TM, Taur Y, Kamboj M. Determinants of COVID-19 disease severity in patients with cancer. Nat Med. 2020;26(8):1218–23. https://doi.org/10.1038/s41591-020-0979-0.
Guan WJ, Ni ZY, Hu Y, Liang WH, Ou CQ, He JX, Liu L, Shan H, Lei CL, Hui DSC, Du B, Li LJ, Zeng G, Yuen KY, Chen RC, Tang CL, Wang T, Chen PY, Xiang J, Li SY, Wang JL, Liang ZJ, Peng YX, Wei L, Liu Y, Hu YH, Peng P, Wang JM, Liu JY, Chen Z, Li G, Zheng ZJ, Qiu SQ, Luo J, Ye CJ, Zhu SY, Zhong NS, China Medical Treatment Expert Group for Covid-19. Clinical characteristics of coronavirus disease 2019 in China. N Engl J Med. 2020;382(18):1708–20. https://doi.org/10.1056/NEJMoa2002032.
Liang W, Guan W, Chen R, Wang W, Li J, Xu K, Li C, Ai Q, Lu W, Liang H, Li S, He J. Cancer patients in SARS-CoV-2 infection: a nationwide analysis in China. Lancet Oncol. 2020;21(3):335–7. https://doi.org/10.1016/S1470-2045(20)30096-6.
Aries JA, Davies JK, Auer RL, Hallam SL, Montoto S, Smith M, Sevillano B, Foggo V, Wrench B, Zegocki K, Agrawal S, Le Dieu R, Truelove E, Erblich T, Araf S, Okosun J, Oakervee H, Cavenagh JD, Gribben JG, Riches JC. Clinical outcome of coronavirus disease 2019 in haemato-oncology patients. Br J Haematol. 2020;190(2):e64–7. https://doi.org/10.1111/bjh.16852.
Cavanna L, Citterio C, Di Nunzio C, Biasini C, Palladino MA, Ambroggi M, Madaro S, Bidin L, Porzio R, Proietto M. Prevalence of COVID-19 infection in asymptomatic cancer patients in a district with high prevalence of SARS-CoV-2 in Italy. Cureus. 2021;13(3):e13774. https://doi.org/10.7759/cureus.13774.
Miyashita H, Mikami T, Chopra N, Yamada T, Chernyavsky S, Rizk D, Cruz C. Do patients with cancer have a poorer prognosis of COVID-19? An experience in New York City. Ann Oncol. 2020;31(8):1088–9. https://doi.org/10.1016/j.annonc.2020.04.006.
Jhawar SR, Palmer JD, Wang SJ, Bitterman D, Klamer B, Huynh-Le M, Chung C, Ohri N, Stover DG, Lustberg MB, Mishra S, Warner J, Jabbour S, Goyal S, CCC19 Radiation Oncology Group. The COVID-19 & Cancer Consortium (CCC19) and opportunities for radiation oncology. Adv Radiat Oncol. 2021;6(1):100614. https://doi.org/10.1016/j.adro.2020.10.026.
Lee LYW, Cazier JB, Starkey T, et al. COVID-19 prevalence and mortality in patients with cancer and the effect of primary tumour subtype and patient demographics: a prospective cohort study. Lancet Oncol. 2020;21(10):1309–16. https://doi.org/10.1016/S1470-2045(20)30442-3.
Paul S, Rausch CR, Jain N, Kadia T, Ravandi F, DiNardo CD, Welch MA, Dabaja BS, Daver N, Garcia-Manero G, Wierda W, Pemmaraju N, Montalban Bravo G, Thompson P, Verstovsek S, Konopleva M, Kantarjian H, Jabbour E. Treating leukemia in the time of COVID-19. Acta Haematol. 2021;144(2):132–45. https://doi.org/10.1159/000508199.
Barrett T, Troup DB, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, Marshall KA, Phillippy KH, Sherman PM, Muertter RN, Holko M, Ayanbule O, Yefanov A, Soboleva A. NCBI GEO: archive for functional genomics data sets—10 years on. Nucleic Acids Res. 2011;39(Database issue):D1005–10. https://doi.org/10.1093/nar/gkq1184.
Görgün G, Holderried TA, Zahrieh D, Neuberg D, Gribben JG. Chronic lymphocytic leukemia cells induce changes in gene expression of CD4 and CD8 T cells. J Clin Investig. 2005;115(7):1797–805. https://doi.org/10.1172/JCI24176.
Affer M, Dao S, Liu C, Olshen AB, Mo Q, Viale A, Lambek CL, Marr TG, Clarkson BD. Gene expression differences between enriched normal and chronic myelogenous leukemia quiescent stem/progenitor cells and correlations with biological abnormalities. J Oncol. 2011;2011:798592. https://doi.org/10.1155/2011/798592.
Vallat L, Kemper CA, Jung N, Maumy-Bertrand M, Bertrand F, Meyer N, Pocheville A, Fisher JW 3rd, Gribben JG, Bahram S. Reverse-engineering the genetic circuitry of a cancer cell with predicted intervention in chronic lymphocytic leukemia. Proc Natl Acad Sci USA. 2013;110(2):459–64. https://doi.org/10.1073/pnas.1211130110.
Smyth GK. limma: linear models for microarray data. In: Gentleman R, Carey VJ, Huber W, Irizarry RA, Dudoit S, editors. Bioinformatics and computational biology solutions using R and bioconductor. Statistics for biology and health. New York: Springer; 2005. https://doi.org/10.1007/0-387-29362-0_23.
Marot G, Foulley JL, Mayer CD, Jaffrézic F. Moderated effect size and P-value combinations for microarray meta-analyses. Bioinformatics. 2009;25(20):2692–9. https://doi.org/10.1093/bioinformatics/btp444.
Ritchie ME, Phipson B, Wu D, Hu Y, Law CW, Shi W, Smyth GK. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015;43(7): e47. https://doi.org/10.1093/nar/gkv007.
Fisher RA. Statistical methods for research workers. In: Kotz S, Johnson NL, editors. Breakthroughs in statistics. Springer series in statistics (perspectives in statistics). New York: Springer; 1992. https://doi.org/10.1007/978-1-4612-4380-9_6.
Benjamini Y. Discovering the false discovery rate. J R Stat Soc Ser B (Stat Method). 2010;72:405–16. https://doi.org/10.1111/j.1467-9868.2010.00746.x.
Berriz GF, Roth FP. The Synergizer service for translating gene, protein and other biological identifiers. Bioinformatics. 2008;24(19):2272–3. https://doi.org/10.1093/bioinformatics/btn424.
da Huang W, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4(1):44–57. https://doi.org/10.1038/nprot.2008.211.
Wickham H. ggplot2. WIREs Comp Stat. 2011;3:180–5. https://doi.org/10.1002/wics.147.
von Mering C, Huynen M, Jaeggi D, Schmidt S, Bork P, Snel B. STRING: a database of predicted functional associations between proteins. Nucleic Acids Res. 2003;31(1):258–61. https://doi.org/10.1093/nar/gkg034.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B, Ideker T. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13(11):2498–504. https://doi.org/10.1101/gr.1239303.
Wang JH, Zhao LF, Wang HF, Wen YT, Jiang KK, Mao XM, Zhou ZY, Yao KT, Geng QS, Guo D, Huang ZX. GenCLiP 3: mining human genes’ functions and regulatory networks from PubMed based on co-occurrences and natural language processing. Bioinformatics. 2019. https://doi.org/10.1093/bioinformatics/btz807.
Bader GD, Hogue CW. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 2003;4:2. https://doi.org/10.1186/1471-2105-4-2.
Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, Li B, Liu XS. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 2017;77(21):e108–10. https://doi.org/10.1158/0008-5472.CAN-17-0307.
Kozakov D, Hall DR, Xia B, Porter KA, Padhorny D, Yueh C, Beglov D, Vajda S. The ClusPro web server for protein-protein docking. Nat Protoc. 2017;12(2):255–78. https://doi.org/10.1038/nprot.2016.169.
Jiménez-García B, Elez K, Koukos PI, Bonvin AM, Vangone A. PRODIGY-crystal: a web-tool for classification of biological interfaces in protein complexes. Bioinformatics. 2019;35(22):4821–3. https://doi.org/10.1093/bioinformatics/btz437.
Seeliger D, de Groot BL. Ligand docking and binding site analysis with PyMOL and Autodock/Vina. J Comput Aided Mol Des. 2010;24(5):417–22. https://doi.org/10.1007/s10822-010-9352-6.
Aronson JK, DeVito N, Plüddeman A, Ferner RE. Drug vignettes: interferons. https://www.cebm.net/covid-19/drug-vignettes-interferons/. Accessed 3 Feb 2021
Swaine T, Dittmar MT. CDC42 use in viral cell entry processes by RNA viruses. Viruses. 2015;7(12):6526–36. https://doi.org/10.3390/v7122955.
Nomura R, Kiyota A, Suzaki E, Kataoka K, Ohe Y, Miyamoto K, Senda T, Fujimoto T. Human coronavirus 229E binds to CD13 in rafts and enters the cell through caveolae. J Virol. 2004;78(16):8701–8. https://doi.org/10.1128/JVI.78.16.8701-8708.2004.
Koivusalo M, Welch C, Hayashi H, Scott CC, Kim M, Alexander T, Touret N, Hahn KM, Grinstein S. Amiloride inhibits macropinocytosis by lowering submembranous pH and preventing Rac1 and Cdc42 signaling. J Cell Biol. 2010;188(4):547–63. https://doi.org/10.1083/jcb.200908086.
Fulzele S, Sahay B, Yusufu I, Lee TJ, Sharma A, Kolhe R, Isales CM. COVID-19 virulence in aged patients might be impacted by the host cellular microRNAs abundance/profile. Aging Dis. 2020;11(3):509–22. https://doi.org/10.14336/AD.2020.0428.
Gordon DE, Jang GM, Bouhaddou M, Xu J, et al. A SARS-CoV-2 protein interaction map reveals targets for drug repurposing. Nature. 2020;583(7816):459–68. https://doi.org/10.1038/s41586-020-2286-9.
Wrapp D, Wang N, Corbett KS, Goldsmith JA, Hsieh CL, Abiona O, Graham BS, McLellan JS. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367(6483):1260–3. https://doi.org/10.1126/science.abb2507.
Funding
This work was supported by grant COVID-19 RC01 from the Dr. Sulaiman Al Habib Medical Journal Initiative to support COVID-19 research.
Author information
Authors and Affiliations
Contributions
AJA and MAM contributed to the conceptualization of the study. AJA, MAM, FAA, NAA, MZM, and MI contributed to the experimental design of the study. AJA, MAM, MZM, and NAA performed literature searches. AJA, MAM, MZM, and MI were involved in laboratory experiments and data collection. AJA, MAM, MZM, and FAA contributed to the analyses of data generated in the study. The manuscript was written by AJA, MAM, MZM, and MI, and was reviewed and approved by all authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
44229_2022_5_MOESM1_ESM.xlsx
Supplementary file1 Information on the up-regulated DEGs identified by GEO2R analysis of three GEO datasets (p-value < 0.05, |logFC|≥ 0.2 DEGs from GSE8835, GSE24739, and GSE39411. (XLSX 29 KB)
44229_2022_5_MOESM2_ESM.xlsx
Supplementary file2 Information on for the downUP-regulated DEGs identified by the GEO2R analysis of three GEO datasets (p-value < 0.05, |logFC|≤ − 0.2 DEGs from GSE8835, GSE24739, and GSE39411. (XLSX 39 KB)
44229_2022_5_MOESM3_ESM.xlsx
Supplementary file3 Gene enrichment analysis of up-regulated DEGs from chronic lymphocytic and chronic myelogenous samples of leukemia (XLSX 29 KB)
44229_2022_5_MOESM4_ESM.xlsx
Supplementary file4 Gene enrichment analysis of down-regulated DEGs from chronic lymphocytic and chronic myelogenous samples of leukemia (XLSX 31 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visithttp://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alsalman, A.J., Al Mohaini, M., Malik, M.Z. et al. Elevated Vulnerability of Chronic Leukemia Patients to COVID-19 Infection: A Systems Biology Approach. Dr. Sulaiman Al Habib Med J 4, 32–45 (2022). https://doi.org/10.1007/s44229-022-00005-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s44229-022-00005-y